Abstract
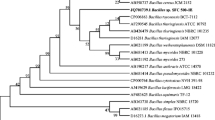
Pentachlorophenol (PCP) is a recalcitrant biocide that bioaccumulates in the environment due to its persistent nature and has been listed as a priority pollutant due to its toxicological and health effects. In this study, a novel PCP-degrading Bacillus cereus strain AOA-CPS1 (BcAOA) was isolated from wastewater and characterized for PCP biotransformation in a batch reactor. The degradation kinetics were elucidated via substrate inhibition models, while PCP biotransformation was established by spectrophotometric and GC–MS analysis. BcAOA shared 95% sequence homology with Bacillus cereus strain XS2 and is closely related to some B. cereus strains which are previously reported to degrade PCP and other related pollutants. BcAOA degraded 74% of 350 mg l−1 of PCP within 9 days in a batch culture. The biotransformation of PCP by BcAOA followed the first and zero-order kinetics at low and high PCP concentration, respectively, with biokinetic constants: maximum biotransformation rate (0.0996 mg l−1 h−1); substrate inhibition constant (723.75 mg l−1); half-saturation constant (171.198 mg l−1) and R2 (0.98). The genes (pcpABCDE, cytochrome P450) encoding the enzymes involved in the biodegradation of PCP were amplified from the genomic DNA of BcAOA. Further, depending upon the genes amplified and identified metabolites using GC–MS, two different PCP biotransformation pathways were proposed in this study. Cloning and expression of the catabolic genes are underway to map out the concise pathway for PCP biotransformation by BcAOA.






Similar content being viewed by others
References
Ammeri RW, Mehri I, Badi S, Hassen W, Hassen A (2017) Pentachlorophenol degradation by Pseudomonas fluorescens. Water Qual Res J Canada 52:99–108. https://doi.org/10.2166/wqrj.2017.003
Andrews JF (1968) A Mathematical model for the continuous culture of microorganisms utilizing inhibitory substrates. Biotechnol Bioeng X:707–723. https://doi.org/10.1002/bit.260100602
Apajalahti JH, Salkinoja-Salonen MS (1987) Complete dechlorination of tetrachlorohydroquinone by cell extracts of pentachlorophenol-induced Rhodococcus chlorophenolicus. J Bacteriol 169:5125–5130. https://doi.org/10.1128/jb.169.11.5125-5130.1987
Arnaldos M, Amerlinck Y, Rehman U, Maere T, Van Hoey S, Naessens W, Nopens I (2015) From the affinity constant to the half-saturation index: understanding conventional modeling concepts in novel wastewater treatment processes. Water Res 70:458–470. https://doi.org/10.1016/j.watres.2014.11.046
ATSDR (2017) Agency for toxic substances and disease registry. substance priority list (candidates for toxicological profiles). 190. https://www.atsdr.cdc.gov/. Accessed 30 Sept 2020
Bhowmik A (2014) Development of a cost effective medium for enhanced production of Bacillus thuringiensis δ-endotoxin in partial fulfilment of the requirments for the MS degree in biotechnology. BRAC University, Dhaka. https://core.ac.uk/download/pdf/61805939.pdf. Accessed 30 Sept 2020
Cai M, Xun L (2002) Organization and regulation of pentachlorophenol-degrading genes in Sphingobium chlorophenolicum ATCC 39723. Society 184:4672–4680. https://doi.org/10.1128/JB.184.17.4672
Cassidy MB, Lee H, Trevors JT, Zablotowicz RB (1999) Chlorophenol and nitrophenol metabolism by Sphingomonas sp UG30. J Ind Microbiol Biotechnol 23:232–241. https://doi.org/10.1038/sj/jim/2900749
CDC (2018) Fourth National Report on Human Exposure to Environmental Chemicals Updated Table. URL: https://www.cdc.gov/exposurereport/pdf/FourthReport_UpdatedTables_Jul2014.pdf. Accessed 30 Sept 2020
Chanama M, Chanana S (2011) Expression of pentachlorophenol degradative genes of Sphingobium chlorophenolicum ATCC39723 in Escherichia coli. Asian J Public Heal 2:78–83
Chanama S, Crawford RL (1997) Mutational analysis of pcpA and its role in pentachlorophenol degradation by Sphingomonas (Flavobacterium) chlorophenolica ATCC 39723. Appl Environ Microbiol 63:4833–4838
Chandra R, Raj A, Yadav S, Patel DK (2009) Reduction of pollutants in pulp paper mill effluent treated by PCP-degrading bacterial strains. Environ Monit Assess 155:1–11. https://doi.org/10.1007/s10661-008-0413-4
Chen L, Yang J (2008) Biochemical characterization of the tetrachlorobenzoquinone reductase involved in the biodegradation of pentachlorophenol. Int J Mol Sci 9:198–212. https://doi.org/10.3390/ijms9030198
Chen L, Maloney K, Krol E, Zhu B, Yang J (2009) Cloning, overexpression, purification, and characterization of the maleylacetate reductase from Sphingobium chlorophenolicum strain ATCC 53874. Curr Microbiol 58:599–603. https://doi.org/10.1007/s00284-009-9377-z
Dai M, Rogers BJ, Warner JR, Copley SD (2003) A previously unrecognized step in pentachlorophenol degradation in Sphingobium chlorophenolicum is catalyzed by tetrachlorobenzoquinone reductase (PcpD). J Bacteriol 185:302–310. https://doi.org/10.1128/JB.185.1.302-310.2003
Darko G, Akoto O, Oppong C (2008) Persistent organochlorine pesticide residues in fish, sediment and water from Lake Bosomtwi, Ghana. Chemosphere 72(1):21–24. https://doi.org/10.1016/j.chemosphere.2008.02.052
DOH (2005) Pesticides registered in South Africa, including information on chemical classification, application use and relevant crops as indicated by South African MRL levels (South African Department of Health, 2005)
Durruty I, Okada E, González JF, Murialdo SE (2011a) Multisubstrate monod kinetic model for simultaneous degradation of chlorophenol mixtures. Biotechnol Bioprocess Eng 16:908–915. https://doi.org/10.1007/s12257-010-0418-z
Durruty I, Okada E, González JF, Murialdo SE (2011b) Degradation of chlorophenol mixtures in a fed-batch system by two soil bacteria. Water SA 37:547–552. https://doi.org/10.4314/wsa.v37i4.13
El-Bialy HA, Khalil OAA, Gomaa OM (2018) Bacterial-mediated biodegradation of pentachlorophenol via electron shuttling. Environ Technol 40(18):2416–2424. https://doi.org/10.1080/09593330.2018.1442501
Engst R, Macholz RM, Kujawa M, Jochen HL, Plass R (1976) The metabolism of lindane and its metabolites gamma-2,3,4,5,6-pentachlorocyclohexene, pentachlorobenzene, and pentachlorophenol in rats and the pathways of lindane metabolism. J Environ Sci Heal Part B 11:95–117. https://doi.org/10.1080/03601237609372028
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
Gakuba E, Moodley B, Ndungu P, Birungi G (2018) Partition distribution of selected organochlorine pesticides in water, sediment pore water and surface sediment from Umgeni river, Kwazulu-Natal, South Africa. Water SA 44:232–249. https://doi.org/10.4314/wsa.v44i2.09
Gong X, Qi S, Wang Y, Julia EB, Lv C (2007) Historical contamination and sources of organochlorine pesticides in sediment cores from Quanzhou Bay, Southeast China. Mar Pollut Bull 54:1434–1440. https://doi.org/10.1016/j.marpolbul.2007.05.006
IARC (2019a) Pentachlorophenol and some related compounds. IARC monographs on the evaluation of carcinogenic risks to humans. URL: https://publications.iarc.fr/Book-And-Report-Series/Iarc-Monographs-On-The-Identification-Of-Carcinogenic-Hazards-To-Humans/Pentachlorophenol-And-Some-Related-Compounds-2019. Accessed 30 Sept 2020
IARC (2019b) International Agency for Research on Cancer monographs on the identification of carcinogenic hazards to humans. In: Agents Classified by the IARC Monographs, Volumes 1–127. URL: https://monographs.iarc.fr/agents-classified-by-the-iarc/. Accessed 30 Sept 2020
IARC (2016) International Agency for Research on Cancer (IARC) Monographs evaluate pentachlorophenol and some related compounds. Lyon, France. URL: https://www.iarc.fr/wp-content/uploads/2018/07/Volume-117_news-item.pdf. Accessed 30 Sept 2020
Icgen Y, Icgen B, Ozcengiz G (2002) Regulation of crystal protein biosynthesis by Bacillus thuringiensis: I. Effects of mineral elements and pH. Res Microbiol 153:599–604. https://doi.org/10.1016/s0923-2508(02)01366-9
Igbinosa EO, Odjadjare EE, Chigor VN, Igbinosa IH, Emoghene AO, Ekhaise FO, Igiehon NO, Idemudia OG (2013) Toxicological profile of chlorophenols and their derivatives in the environment: the public health perspective. Sci World J 2013:460215. https://doi.org/10.1155/2013/460215
IPEN POPRC (2013) Pentachlorophenol is one of the world’s worst chemicals—Alaska Community Action On Toxics. https://www.akaction.org/pentachlorophenol_2013_10_18/. Accessed 30 Sept 2020
Jenkins D, Wanner J (2014) Activated sludge—100 years and counting. IWA Publishing, London
Joshi VV, Prewitt ML, Ma D-P, Borazjani H (2015) Enhanced remediation of pentachlorophenol (PCP)-contaminated groundwater by bioaugmentation with known PCP-degrading bacteria. Bioremediat J 19:160–170. https://doi.org/10.1080/10889868.2014.995369
Karn SK, Pan X (2017) Diversity of arsenite-oxidizing gene ( aioA gene ) from arsenic-rich gold mine dump of Xinjiang, China. Bioinfolet 14:149–151
Karn SK, Chakrabarty SK, Reddy MS (2010) Pentachlorophenol degradation by Pseudomonas stutzeri CL7 in the secondary sludge of pulp and paper mill. J Environ Sc 22:1608–1612. https://doi.org/10.1016/s0923-2508(02)01366-9
Khessairi A, Fhoula I, Jaouani A, Turki Y, Cherif A, Boudabous A, Hassen A, Ouzari H (2014) Pentachlorophenol degradation by Janibacter sp., a new Actinobacterium isolated from saline sediment of arid land. Biomed Res Int. https://doi.org/10.1155/2014/296472
Kim S, Chen J, Cheng T, Gindulyte A, He J, He S, Li Q, Shoemaker BA, Thiessen PA, Yu B, Zaslavsky L, Zhang J, Bolton EE (2019) PubChem 2019 update: improved access to chemical data. Nucleic Acids Res 47(D1):D1102–D1109. https://doi.org/10.1093/nar/gky1033
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Bio Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Li D, Park J, Oh JR (2001) Silyl derivatization of alkylphenols, chlorophenols, and bisphenol A for simultaneous GC/MS determination. Anal Chem 73:3089–3095. https://doi.org/10.1021/ac001494l
Lin LD, Chen YF, Wang SY, MingJer T (2009) Leachability, metal corrosion, and termite resistance of wood treated with copper-based preservative. Int Biodeterior Biodegrad 63:533–538. https://doi.org/10.1016/j.ibiod.2008.07.012
Lopez-Echartea E, Macek T, Demnerova K, Uhlik O (2016) Bacterial biotransformation of pentachlorophenol and micropollutants formed during its production process. J Environ Res Public Health Int. https://doi.org/10.3390/ijerph13111146
Marchesi JR, Sato T, Weightman AJ, Martin TA, Fry JC, Hiom SJ, Wade WG (1998) Design and evaluation of useful bacterium-specific PCR primers that amplify genes coding for bacterial 16S rRNA. Appl Environ Microbiol 64:795–799
McAllister KA, Lee H, Trevors JT (1996) Microbial degradation of pentachlorophenol. Biodegradation 7:1–40
Mikesell MD, Boyd SA (1986) Complete reductive dechlorination and mineralization of pentachlorophenol by anaerobic microorganisms. Appl Env Microbiol 52:861–865
Monod J (1949) The Growth of bacterial cultures. Annu Rev Microbiol 3:371–394. https://doi.org/10.1146/annurev.mi.03.100149.002103
Nthunya LN, Khumalo NP, Verliefde AR, Mamba BB, Mhlanga SD (2019) Quantitative analysis of phenols and PAHs in the Nandoni Dam in Limpopo Province, South Africa: A preliminary study for dam water quality management. Phys Chem Earth 112:228–236. https://doi.org/10.1016/j.pce.2019.02.003
Olaniran AO, Igbinosa EO (2011) Chlorophenols and other related derivatives of environmental concern: properties, distribution and microbial degradation processes. Chemosphere 83:1297–1306. https://doi.org/10.1016/j.chemosphere.2011.04.009
Olatunji OS (2019) Evaluation of selected polychlorinated biphenyls (PCBs) congeners and dichlorodiphenyltrichloroethane (DDT) in fresh root and leafy vegetables using GC-MS. Sci Rep 9:538. https://doi.org/10.1038/s41598-018-36996-8
Orser CS, Dutton J, Lange C, Jablonski P, Xun L, Hargis M (1993) Characterization of a Flavobacterium glutathione S-transferase gene involved in reductive dechlorination. J Bacteriol 175:2640–2644
Orser CS, Lange CC, Xun L, Zahrt TC, Schneider BJ (1993) Cloning, sequence analysis, and expression of the Flavobacterium pentachlorophenol-4-monooxygenase gene in Escherichia coli. J Bacteriol 175:411–416. https://doi.org/10.1128/jb.175.2.411-416.1993
Osibanjo O, Bouwman H, Bashir NHH, Okond’Ahoka J, Choong Kwet Yve R, Onyoyo HA (2003) Regionally based assessment of persistent toxic substances. Sub- Saharan Africa regional report. UNEP Chemicals, Geneva. http://ee-net.ne.jp/mec2004/UNEP-PTS/RBA%20PTS%20Global%20Report.pdf. Accessed 30 Sept 2020
Patachia S, Croitoru C (2016) Biopolymers for wood preservation. In: Jonkers HM (ed) Biopolymers and biotech admixtures for eco-efficient construction materials. Elsevier, New York, pp 305–332. https://doi.org/10.1016/B978-0-08-100214-8.00014-2
Patel BP, Kumar A (2016a) Multi-substrate biodegradation of chlorophenols by defined microbial consortium. 3 Biotech 6(2):191. https://doi.org/10.1007/s13205-016-0511-x
Patel BP, Kumar A (2016b) Biodegradation of 2,4-dichlorophenol by Bacillus endophyticus strain: optimization of experimental parameters using response surface methodology and kinetic study. Desalin Water Treat 57:15932–15940. https://doi.org/10.1080/19443994.2015.1076351
Patel BP, Kumar A (2016c) Optimization study for maximizing 2,4-dichlorophenol degradation by Kocuria rhizophila strain using response surface methodology and kinetic study. Desalin Water Treat 57:18314–18325. https://doi.org/10.1080/19443994.2015.1091988
Quinn LP, de Vos BJ, Fernandes-Whaley M, Roos C, Bouwman H, Kylin H, Pieters R, van den Berg J (2011) Pesticide use in South Africa: One of the largest importers of pesticides in Africa, Pesticides in the modern world - pesticides use and management, Stoytcheva M (ed) InTech. http://www.intechopen.com/articles/show/title/pesticide-use-in-south-africa-one-of-the-largest-importers-of-pesticides-in-africa. Accessed 30 Sept 2020
Ren H, Li Q, Zhan Y, Fang X, Yu D (2016) 2,4-Dichlorophenol hydroxylase for chlorophenol removal: substrate specificity and catalytic activity. Enzyme Microb Technol 82:74–81. https://doi.org/10.1016/j.enzmictec.2015.08.008
Saber DL, Crawford RL (1985) Isolation and characterization of Flavobacterium strains that degrade pentachlorophenol. Appl Environ Microbiol 50:1512–1518
Sharma A, Thakur IS, Dureja P (2009) Enrichment, isolation and characterization of pentachlorophenol degrading bacterium Acinetobacter sp. ISTPCP-3 from effluent discharge site. Biodegradation 20:643–650. https://doi.org/10.1007/s10532-009-9251-5
Silva V, Mol HGJ, Zomer P, Tienstra M, Ritsema CJ, Geissen V (2019) Pesticide residues in European agricultural soils—a hidden reality unfolded. Sci Total Environ 653:1532–1545. https://doi.org/10.1016/j.scitotenv.2018.10.441
Singh S, Singh ABB, Chandra AR, Patel ADK, Rai AV (2009) Synergistic biodegradation of pentachlorophenol by Bacillus cereus (DQ002384), Serratia marcescens (AY927692) and Serratia marcescens (DQ002385). World J Microbiol Biotechnol. https://doi.org/10.1007/s11274-009-0083-6
Smith RM (2003) Before the injection–modern methods of sample preparation for separation techniques. J Chromatogr A 1000:3–27. https://doi.org/10.1016/s0021-9673(03)00511-9
Stockholm convention (2019) Stockholm convention on persistent organic pollutants. (UNEP/POPS/COP.9/INF/16). Geneva, Switzerland. http://chm.pops.int/TheConvention/ConferenceoftheParties/Meetings/COP9/tabid/7521/ctl/Download/mid/20310/Default.aspx?id=5&ObjID=27106. Accessed 30 Sept 2020
Takeuchi M, Hamana K, Hiraishi A (2001) Proposal of the genus Sphingomonas sensu stricto and three new genera, Sphingobium, Novosphingobium and Sphingopyxis, on the basis of phylogenetic and chemotaxonomic analyses. Int J Syst Evol Microbiol 51:1405–1417. https://doi.org/10.1099/00207713-51-4-1405
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci USA 101:11030–11035. https://doi.org/10.1073/pnas.0404206101
Thompson LA, Darwish WS, Ikenaka Y, Nakayama MMS, Mizukawa H, Ishizuka M (2017) Organochlorine pesticide contamination of foods in Africa: incidence and public health significance. J Vet Med Sci. https://doi.org/10.1292/jvms.16-0214
Tripathi M, Garg SK (2010) Studies on selection of efficient bacterial strain simultaneously tolerant to hexavalent chromium and pentachlorophenol isolated from treated tannery effluent. Res J Microbiol 5:707–716. https://doi.org/10.3923/jm.2010.707.716
Uotila JS, Kitunen VH, Saastamoinen T, Coote T, Haggblom MM, Salkinoja-Salonen MS (1992) Characterization of aromatic dehalogenases of Mycobacterium fortuitum CG-2. J Bacteriol 174:5669–5675
Villemur R (2013) The pentachlorophenol-dehalogenating Desulfitobacterium hafniense strain PCP-1. Philos Trans R Soc Lond B Biol Sci 368:20120319. https://doi.org/10.1098/rstb.2012.0319
WRC (2015) Persistent organic pollutants threats: threats posed by the organic pollutants to a tourist destination. http://www.wrc.org.za/wp-content/uploads/mdocs/PB_1977_PoPs%20in%20DBN%20estuaries.pdf. Accessed 30 Sept 2020
WRC (2018) Water research commission: Annual report 2017-2018. http://www.wrc.org.za/about-us/annual-reports/. Accessed 30 Sept 2020
Xu L, Resing K, Lawson SL, Babbitt PC, Copley SD (1999) Evidence That pcpA encodes 2,6-Dichlorohydroquinone dioxygenase, the ring cleavage enzyme required for pentachlorophenol degradation in Sphingomonas chlorophenolica Strain ATCC 39723. Biochemistry 38:7659. https://doi.org/10.1021/bi990103y
Yahaya A, Okoh OO, Agunbiade FO, Okoh AI (2019) Occurrence of phenolic derivatives in Buffalo River of Eastern Cape South Africa: exposure risk evaluation. Ecotoxicol Environ Saf 171:887–893. https://doi.org/10.1016/j.ecoenv.2019.01.037
Yang CF, Lee CM, Wang C (2006) Isolation and physiological characterization of the pentachlorophenol degrading bacterium Sphingomonas chlorophenolica. Chemosphere 62:709–714. https://doi.org/10.1016/j.chemosphere.2005.05.012
Yuancai L, Chen Y, Song W, Hu Y (2014) Enhanced selection of micro-aerobic pentachlorophenol degrading granular sludge. J Hazard Mater 280:134–142. https://doi.org/10.1016/j.jhazmat.2014.07.067
Zhang W, Sun Z (2008) Random local neighbor joining: A new method for reconstructing phylogenetic trees. Mol Phylogenet Evol 47(1):117–128. https://doi.org/10.1016/j.ympev.2008.01.019
Zhang J, Liu X, Xu Z, Chen H, Yang Y (2008) Degradation of chlorophenols catalyzed by laccase. Int Biodeterior Biodegrad 61:351–356. https://doi.org/10.1016/J.IBIOD.2007.06.015
Acknowledgments
O.A. thank University of KwaZulu Natal for awarding fellowship during PH.D. studies.
Funding
National Research Foundation, South Africa (Grant No: 94036 and 92803).
Author information
Authors and Affiliations
Contributions
O.A. and A.O conceived and designed the project. O.A. performed the experiments. M.P.M. contributed reagents and materials. O.A., A.K., M.P.M. and A.O. wrote the manuscript. All the authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare he has no conflict of interest.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Aregbesola, O.A., Mokoena, M.P. & Olaniran, A.O. Biotransformation of pentachlorophenol by an indigenous Bacillus cereus AOA-CPS1 isolated from wastewater effluent in Durban, South Africa. Biodegradation 31, 369–383 (2020). https://doi.org/10.1007/s10532-020-09915-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-020-09915-w