Abstract
Cultivated meat, also known as cultured or cell-based meat, is meat produced directly from cultured animal cells rather than from a whole animal. Cultivated meat and seafood have been proposed as a means of mitigating the substantial harms associated with current production methods, including damage to the environment, antibiotic resistance, food security challenges, poor animal welfare, and—in the case of seafood—overfishing and ecological damage associated with fishing and aquaculture. Because biomedical tissue engineering research, from which cultivated meat draws a great deal of inspiration, has thus far been conducted almost exclusively in mammals, cultivated seafood suffers from a lack of established protocols for producing complex tissues in vitro. At the same time, fish such as the zebrafish Danio rerio have been widely used as model organisms in developmental biology. Therefore, many of the mechanisms and signaling pathways involved in the formation of muscle, fat, and other relevant tissue are relatively well understood for this species. The same processes are understood to a lesser degree in aquatic invertebrates. This review discusses the differentiation and maturation of meat-relevant cell types in aquatic species and makes recommendations for future research aimed at recapitulating these processes to produce cultivated fish and shellfish.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
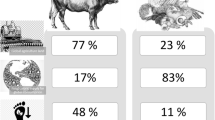
Large-scale industrial meat production causes negative externalities related to the environment, food security, antibiotic resistance, and animal welfare (Godfray et al. 2018; Hilborn et al. 2018). The idea that meat might be cultivated from isolated stem cells has been proposed as a solution to these challenges (Datar and Betti 2010). The concept of cultivated meat (CM) production has been reported in manuscripts published in as early as the 1930s (Birkenhead 1930), while the first patent describing a process to produce meat from cells at large scale was granted in 1999 to Dutch researcher Willem van Eelen (van Eelen et al. 1999). By 2013, the first CM prototype—a beef hamburger—was eaten in a live tasting, and the process of obtaining muscle fibers that composed the cultivated hamburger was further described in a publication the following year (Post 2014). Over the past decade, CM has grown from an idea into a nascent field consisting of various academic labs and for-profit companies (Choudhury et al. 2020; Nyika et al. 2021).
While the primary focus has been on farmed terrestrial animals, companies and researchers are also attempting to cultivate seafood (Rubio et al. 2019a; Tsuruwaka and Shimada 2022; Goswami et al. 2022a). Indeed, one of the first academic publications on CM described an attempt to expand a filet of goldfish meat in vitro as a protein source for long-term space travel (Benjaminson et al. 2002). Although seafood’s impacts vary widely across species, regions, and production practices, both wild-caught fish and aquaculture may be associated with significant challenges (Pauly and Zeller 2016; Boone Kauffman et al. 2017; Watts et al. 2017; Lima et al. 2018; Parker et al. 2018). Cultivated seafood (CS), obtained following cellular agriculture approaches, has the potential to ameliorate many of the negative externalities associated with seafood production (Reis et al. 2021). However, realizing these benefits depends on several discrete, intermediate successes, including the need for CS products to be both cost-competitive and viewed as acceptable substitutes by consumers (Halpern et al. 2021). As of the end of 2021, twenty companies globally were working on CS, with nine of them being established earlier in that year (Azoff 2022).
The relationship between the biology of tissue and the organoleptic and nutritional properties of meat is complex (Listrat et al. 2016). To produce CM, stem cells must be differentiated into mature myofibers, adipocytes, and other meat-relevant cell types (Lee et al. 2022). Therefore, methods for inducing differentiation and maturation that are efficient, cost-effective, and food-safe must be identified. Existing knowledge of cell differentiation pathways should be used to generate hypotheses and identify candidate strategies to promote cells’ differentiation, which may then be empirically tested and optimized. While this knowledge is limited for many seafood species, cellular differentiation pathways are reasonably well understood in zebrafish (Guyon et al. 2007; Salmerón 2018; Keenan and Currie 2019). Indeed, it has been suggested that the wealth of biological information available for zebrafish makes it a good candidate for early studies and perhaps even product development for CS (Potter et al. 2020).
This review discusses molecular signals involved in differentiating muscle, fat, and other cell types necessary for producing high-quality CS. Where possible, data from popular seafood species are discussed. While the primary focus is on fish, which have been better studied, relevant differentiation pathways in aquatic invertebrates are briefly discussed. Where possible, recommendations are provided as to how existing data may be used to inform future efforts to improve differentiation protocols for CS.
Cell Sources for CS
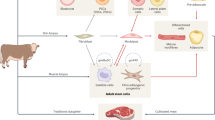
Various pluripotent and adult stem cell types have been investigated or proposed as sources for CM and CS production (Reiss et al. 2021). Most of the mature cell types likely to be necessary for CM and CS belong to the mesodermal lineage (Fig. 1 and Table 1). Because the most critical cells for the final product are muscle and fat cells, the precursors to these cell types—mesenchymal stem cells, satellite cells, fibro-adipogenic progenitors, and preadipocytes—are the most likely candidates among the adult stem cells to be used for CM and CS production (Reiss et al. 2021). It was recently demonstrated that fibroblast-like cells isolated from filefish fins could be easily differentiated into several cell types, including skeletal muscle–like cells and adipocytes (Tsuruwaka and Shimada 2022). If similar results are found for other fish species and under serum-free conditions, fish fin–derived fibroblasts could serve as an alternative cell source with the potential to make cell line development and culture substantially easier. Cell types can be identified experimentally by the expression of marker genes, but these markers are somewhat less well characterized in seafood species than amniotes. Methods for isolation, identification, culture, and enhancement of proliferation for some key cell types are discussed below, emphasizing data from fish where possible.
Lineage relationships between cell types that may be used as starting materials for CM and CS and desired cell types likely to be present in the final product (gray background). CS products will be composed primarily of fast myofibers and adipocytes, likely with smaller quantities of some of the other listed cell types. Dashed lines indicate cell type transitions that are not canonically understood to be part of normal developmental or regenerative processes, but that have been observed experimentally (Blagden et al. 1997; Du et al. 1997; Asakura et al. 2001; Potthoff et al. 2007; Tsuruwaka and Shimada 2022) and may be useful for producing the cell types desired in CS products. Some examples of genes/proteins that serve as useful markers are listed next to the associated cell type (Devoto et al. 1996; Todorcević et al. 2010; Bricard et al. 2014; Landemaine et al. 2014; Ma et al. 2018; Peng et al. 2019; Li et al. 2020; Reiss et al. 2021)
ESC-Like Cells and iPSCs
Embryonic stem cell (ESC)–like lines have been established from several fish species, including sea bass (Chen et al. 2003a; Buonocore et al. 2006; Parameswaran et al. 2007), catfish (Dash et al. 2010; Barman et al. 2014), sea bream (Béjar et al. 1999; Chen et al. 2003b), tilapia (Fan et al. 2017), turbot (Chen et al. 2005), cod (Holen et al. 2010), rohu carp (Goswami et al. 2012), zebrafish (Collodi et al. 1992; Driever and Rangini 1993; He et al. 2006; Ho et al. 2014), and medaka (Wakamatsu et al. 1994; Hong et al. 1996; Yuan and Hong 2017). In contrast to true ESCs from other species such as mouse (West et al. 2006), it is not clear that any of the described ESC-like lines from fish can give rise to germ cells. However, given the cell types involved, existing ESC-like cells are expected to be sufficient as a cell source for most CS products, perhaps with the exception of roe. Many of these studies reported culturing fish ES-like cells in Dulbecco’s Modified Eagle’s Medium (DMEM) with β-mercaptoethanol, selenium, glutamine, pyruvate, non-essential amino acids, fibroblast growth factor 2 (FGF2), leukemia inhibitory factor (LIF), fetal bovine serum (FBS), fish serum, and fish embryo extract (Hong et al. 1996). Under these conditions, fish ES-like cells generally maintain markers of pluripotency. DMEM supplemented with insulin-like growth factor 2 (IGF-2) maintained pluripotent medaka ES-like cells, though with a reduced growth rate relative to FBS-containing media (Yuan and Hong 2017). Leibovitz’s L-15 with FBS has also been shown to support the growth of ES-like cells from some fish species (Bryson et al. 2006; Parameswaran et al. 2007). ESC-like cells from shrimp have been successfully cultured for up to ten passages, though contamination was a substantial problem and ultimately a continuous line was not successfully developed (Fan and Wang 2002).
Zebrafish fibroblasts have been successfully reprogrammed to induced pluripotent stem cells (iPSCs) using the Yamanaka reprogramming factors Oct3/4, Sox2, Klf4, and c-Myc (Rosselló et al. 2013; Peng et al. 2019). More recently, iPSC-like cells were generated from koi fibroblasts using a chemical reprogramming method (Xu et al. 2022). The fact that fish cells can be reprogrammed has the potential to be used for CS, especially for species from which it is difficult to obtain tissues or embryos. Beyond the simple translation of iPSC reprogramming methods to common seafood species, it will be critical to transition to methods not reliant on antibiotic-inducible systems and, ideally, footprint-free methods (Rao and Malik 2012) to alleviate potential concerns related to genetic modification.
MSCs
Mesenchymal stromal cells (MSCs) are a potential cell source for CS due to their high self-renewal capacity, high proliferation rate, and multipotency. Isolation of MSCs from a variety of fish tissues, including visceral adipose tissue (gilthead sea bream (Salmerón et al. 2016), rainbow trout (Bou et al. 2017), and Atlantic salmon (Ytteborg et al. 2015)), vertebra bone (gilthead sea bream (Salmerón et al. 2016; Riera-Heredia et al. 2019)), heart (zebrafish (Fathi et al. 2019)), and liver (zebrafish (Fathi et al. 2019)) have been reported. Typically in such studies, tissues are mechanically disrupted and digested using collagenase, and MSCs are cultured in DMEM supplemented with 10% FBS.
Adipose-derived MSCs (AMSCs) have been isolated from human adipose tissue obtained by liposuction and processing of the raw aspirate to obtain the heterogeneous cell population known as the stromal vascular fraction (SVF) (Zuk et al. 2001). AMSCs may be also found within adipose tissue of bovines and other mammals (Mehta et al. 2019). Centrifugation of the collagenase-digested tissue results in three-layered fractions: a top fraction containing floating lipids and mature and lysed adipocytes, followed by an aqueous fraction composed of enzymes and medium, and the SVF, where MSCs and adipogenic progenitors can be found (Mehta et al. 2019). Because the SVF is heterogeneous, fluorescence- or magnetic-activated cell sorting (FACS or MACS) may be used to select a more defined population, as demonstrated by Ishimura et al. (2008) in mouse cells. However, the utility of this method depends on the availability of suitable antibodies, which may be a limitation for many aquatic species.
In culture, fish MSCs can be identified by ability to adhere to tissue culture polystyrene plates, specific cell surface markers, lack of hematopoietic and endothelial markers, morphology, and multi-lineage differentiation capacity. Fathi et al. (2019) showed that MSCs from zebrafish heart and liver presented a fibroblast-like morphology and expressed Nanog, Oct4, and Sox2, common pluripotency markers. These cells were positive for CD44 and CD90 and negative for CD31 and CD34. Moreover, zebrafish MSCs could differentiate into osteocyte, adipocyte, and chondrocyte lineages, representing a distinctive characteristic of MSCs (Dominici et al. 2006; Fathi et al. 2019).
For CS applications, the multi-lineage potential of MSCs could be evaluated to grow different relevant tissues, such as muscle and adipose tissue. Adipogenic differentiation of gilthead sea bream, rainbow trout, and Atlantic salmon MSCs can be induced using a differentiation media containing insulin, 3-isobutyl-1-methylxanthine, dexamethasone, and a lipid mixture (Ytteborg et al. 2015; Salmerón et al. 2016; Bou et al. 2017). Myogenic differentiation has not yet been reported in the literature for fish MSCs. However, appropriate physical, mechanical, and bio/chemical conditions (through culture medium supplementation) could induce myoblast formation from mammalian MSCs (Wang et al. 2012a; Xu et al. 2015; Witt et al. 2017), including AMSCs (Zuk et al. 2001, 2002; Zheng et al. 2006).
Satellite Cells and Myoblasts
Satellite cells (SCs, also called myosatellite cells) (Mauro 1961) are another likely cell source for CM (Hanga et al. 2020). While their canonical role is in muscle regeneration, mouse SCs can be differentiated into myogenic, osteogenic, and adipogenic lineages (Asakura et al. 2001). Hollway et al. (2007) identified a population of presumptive SCs in adult zebrafish originating from the anterior somite. This population contributes to the repair of injured muscle (Seger et al. 2011; Knappe et al. 2015) via asymmetric cell division (Gurevich et al. 2016). SCs in adult zebrafish are concentrated primarily in slow muscle, express the transcription factor paired box protein 7 (Pax7), are characterized by dense heterochromatin, and are located between the muscle cell membrane and the basal lamina (Berberoglu et al. 2017). Pax3 and Pax7 are consistent markers of adult muscle SCs and embryonic myogenic progenitors in species including mice (Asakura et al. 2001; Relaix et al. 2005), chicken, quail (Gros et al. 2005), zebrafish (Yin et al. 2018; Ganassi et al. 2018), pearlfish (Marschallinger et al. 2009), and rainbow trout (Bricard et al. 2014; Villasante et al. 2016), although in fish they may remain briefly expressed during early stages of differentiation (Devoto et al. 2006; Marschallinger et al. 2009; Seger et al. 2011). Other commonly used SC markers were reviewed by Siegel et al. (2013).
Montserrat et al. (2007, 2012) described methods for primary SC culture from gilthead seabream, and similar protocols have been employed for rainbow trout (Fauconneau and Paboeuf 2000; Castillo et al. 2002, 2004). Proportions of SCs in carp decrease with age (Koumans et al. 1991), suggesting that isolation from younger fish is preferable. More recently, a spontaneously immortalized myogenic mackerel cell line has been described (Saad et al. 2022). Culture of myogenic cells from aquatic invertebrates has been even less thoroughly studied, though a recent description of methods for primary culture of lobster myogenic cells (Jang et al. 2022) may pave the way for future studies.
Myostatin, a transforming growth factor β (TGF-β) superfamily member, is well-characterized as an inhibitor of proliferation and differentiation in mammalian muscle. Loss of function of myostatin also increases muscle growth in many fish species (Lee et al. 2009; Chisada et al. 2011; Gao et al. 2016; Torres-Velarde et al. 2020), though not in all fish species tested (Terova et al. 2013). Evidence from mammals (Morissette et al. 2009) and fish (Liu et al. 2020) indicates that myostatin inhibits Akt, a positive regulator of muscle hypertrophy. In partial contrast to mammalian systems, myostatin inhibited proliferation but not differentiation in rainbow trout SCs (Seiliez et al. 2012; Garikipati and Rodgers 2012a, b). Also unlike in mammals, it has been suggested that the effects of fish myostatin are not specific to muscle (Gabillard et al. 2013). Myostatin loss of function, therefore, offers a possible strategy for improving proliferation rates as part of the CS bioprocess, but this strategy will need to be tested and may be effective in only some species.
Whereas IGFs in mammalian muscle tend to promote differentiation (Retamales et al. 2015; Pourquié et al. 2018), their effects in fish may be more complex. IGF-1 and IGF-2 stimulated proliferation in primary cultures of myoblasts or SCs from rainbow trout (Castillo et al. 2004; Gabillard et al. 2010; Garikipati and Rodgers 2012a, b). In cultured gilthead seabream SCs, treatment with IGF-2 and IGF-1 induced markers of early and later stages of differentiation, respectively (Jiménez-Amilburu et al. 2013). Consistent with this, IGF-2 was more effective than IGF-1 in stimulating myoblast proliferation (Rius-Francino et al. 2011). Together, these results suggest that IGF-2 and IGF-1 may tend to promote proliferation and differentiation, respectively, but that their effects depend on species and culture conditions. IGF-1’s effects on rat myoblast differentiation are partially mediated through myostatin (Retamales et al. 2015), which lacks its canonical function as an inhibitor of differentiation in at least some fish species. Treatment with specific IGFs—or IGF mimics—during the proliferation or differentiation stages may be helpful for CS, but predicting their effects is not entirely straightforward.
The adenylate cyclase activator forskolin promoted proliferation without effects on differentiation in both zebrafish and mouse SCs (Xu et al. 2013). Anthocyanidin treatment of rainbow trout primary myogenic cells increased expression of Pax7 and non-significantly reduced expression of differentiation markers (Villasante et al. 2016), suggesting that these compounds might help maintain myogenic stem cells in a proliferative state.
Fibro-/Adipogenic Progenitors (FAPs)
Fibro-/adipogenic progenitors (FAPs) are multipotent non-myogenic MSCs that can be isolated from the muscle’s SVF (Joe et al. 2010; Low et al. 2017). FAPs can be found in the interstitial space of skeletal muscle and support myogenic development and regeneration following muscle injury (Joe et al. 2010). Low et al. described protocols to isolate FAPs from the SVF of murine skeletal muscle using antibodies to cell-specific surface antigens and FACS, resulting in adhered cells with a spindle shape with short projections (Low et al. 2017).
FAPs express the platelet-derived growth factor receptor-α (PDGFRα or CD140a) but can be also recognized by the expression of vimentin, delta like non-canonical Notch ligand 1 (Dlk1) or preadipocyte factor 1 (Pref1), and stem cells antigen 1 (Sca1) (Li et al. 2020). Additionally, in mice, such cells are identified by the absence of CD31, CD45, and integrin-α7 (Uezumi et al. 2011; Judson et al. 2017).
Different studies using mammalian cells, including bovines, had shown the potential of FAPs to differentiate into fibroblasts (Joe et al. 2010) and adipocytes (Arrighi et al. 2015; Uezumi et al. 2010). Therefore, FAPs have been proposed as a cell source for CM (Melzener et al. 2021; Dohmen et al. 2022), namely, for production of connective and fat tissues of meat (Reiss et al. 2021). It remains to be determined whether an equivalent population exists in fish.
FAP proliferation and determination are highly dependent on the niche environment, regulated by crosstalk with SCs, myotubes, and immune cells (Biferali et al. 2019). While proliferation is positively regulated with interleukin-4 (IL-4), interleukin-15 (IL-15), TGF-β1, and myostatin, the commitment to either adipogenesis or fibrogenesis is more complex (Li et al. 2020). Further studies are required for their elucidation and to understand similarities and differences between mammals and fish. The control of those pathways will be crucial to improve CM and CS production efficiency.
Preadipocytes
Adipogenesis is characterized by two phases—determination and differentiation—requiring progressive induction of genes responsible for functions such as lipid uptake and the secretion of adipokines. Cells that have undergone determination—and thus are committed to the adipocyte lineage, but are not yet differentiated—are often called preadipocytes (Salmerón 2018).
Studies in mice indicate that adipogenic precursor cells express mesenchymal markers such as SCA1, CD34, and CD29 but do not express mice hematopoietic (CD45) and endothelial (CD31) markers. As mouse adipogenic precursors become further committed to the adipocyte lineage, they lose their expression of CD24 (Berry and Rodeheffer 2013; Hepler et al. 2017). Mouse preadipocytes also express the zinc finger protein ZFP423 (Gupta et al. 2010).
Preadipocytes from Atlantic salmon have a fibroblast-like morphology and do not contain lipid droplets (Vegusdal et al. 2003). Identified preadipocyte markers may differ somewhat between fish species but include peroxisome proliferator-activated receptor gamma (pparγ); CCAAT/enhancer-binding protein (c/ebp), namely, c/ebpα and c/ebpβ, transgelin, and fatty acid synthase (fas) in Atlantic salmon (Todorcević et al. 2010); and glucose-6-phosphate dehydrogenase (g6pdh) in sea bream (Salmerón et al. 2016; Salmerón 2018).
Like MSCs and FAPs, fish preadipocytes can be isolated directly from the SVF (Todorcević et al. 2010; Wang et al. 2012b; Liu et al. 2018; Salmerón 2018). In zebrafish, pancreatic white adipose tissue starts to develop 12 days after fertilization, followed by an increase in visceral, subcutaneous, and cranial depots (Imrie and Sadler 2010). Gilthead seabream preadipocytes obtained from fish specimens weighing 50 g have improved proliferative capacity compared with the ones sourced from 500 g fish specimens (Salmerón et al. 2013), suggesting that the age of the donor animal is an important factor.
The induction of the Wnt/β-catenin pathway has been shown to maintain preadipocytes in an undifferentiated and proliferative state in mammals (Ross et al. 2000) and in carp (Liu et al. 2018). Supplementing the culture medium with IGF-1, insulin, and the growth hormone somatotropin has been shown to improve the proliferation capacity of preadipocytes from gilthead seabream (Salmerón et al. 2013). Insulin promoted, while tumor necrosis factor alpha (TNFɑ) and docosahexaenoic acid (DHA) inhibited, the proliferation of preadipocytes from large yellow croaker (Wang et al. 2012b). Some of these molecules may offer opportunities for optimization of preadipocyte proliferation media, thereby enabling greater efficiency in CS.
Myogenesis in Fish
The main transcription factors, structural proteins, morphogenetic processes, and other critical elements of muscle fiber formation and function are conserved across the vertebrate lineage, including in non-teleost fish species such as sturgeon (Steinbacher et al. 2006). However, both Xu et al. (2000) and Costa et al. (2002) found that zebrafish, chick, and mouse begin to express myogenesis-related genes in different orders, suggesting differences in the underlying regulatory network. Differences in myosin composition during the early stages of regeneration suggest that different pools of cells might be involved in zebrafish cells’ myogenesis compared to seabream (Rowlerson et al. 1997). Thus, caution is warranted in generalizing results from amniotes to fish or even between different fish species. Conservation of signaling pathways across species means that findings from one species offer a promising hypothesis for another species, but such hypotheses must be carefully tested and not simply assumed to be true.
Markers of Myogenic Differentiation
Most of the genes used to mark various stages of myogenesis in fish (Fig. 2) are the same as those used in other vertebrates, though some species differences exist in the expression pattern across cell types or stages.
Genes involved in fish muscle differentiation and maturation. a Steps involved in the differentiation and maturation process. Line graph schematic shows the approximate timing of activation and downregulation of Pax7 and the four MRFs relative to these steps (Weinberg et al. 1996; Hinits et al. 2007; Schnapp et al. 2009; Chen and Galloway 2014; Farnsworth et al. 2020). b UMAP plot of gene expression data (Farnsworth et al. 2020) from developing zebrafish reveals several clusters corresponding to developing and mature muscle. Manually drawn borders correspond to higher level categories, e.g., slow muscle, some of which contain several distinct, numbered clusters. Gene expression data in this panel is from myod, shown for reference. Legend applies to panels b and c. Created using the UCSC Cell Browser (https://zebrafish-dev.cells.ucsc.edu) (Speir et al. 2021). c Expression of several genes relevant to muscle development or function is shown, with key steps in the development process outlined (Farnsworth et al. 2020; Speir et al. 2021). Mrf4 (not shown) shows a similar temporal pattern to that of myog (Schnapp et al. 2009)
Myoblast determination protein (MyoD) and myogenic factor 5 (Myf5) mark cells committed to the myogenic lineage in most vertebrate species, including fish (Weinberg et al. 1996), and together with muscle regulatory factor 4 (Mrf4) and myogenin (Myog) are referred to as the muscle regulatory factors (MRFs). In rainbow trout SC cultures, myoD expression appears 4–48 h after seeding and remains following differentiation (Rescan et al. 1994). Subsequent studies in zebrafish (Weinberg et al. 1996; Coutelle et al. 2001), pacu (de Almeida et al. 2008), and flounder (Zhang et al. 2006) found high myoD transcript levels in somites and juvenile muscle tissue and lower levels in adult muscle. Expression of myf5 follows a similar pattern but shows a slightly earlier onset of expression and a sharper decline between differentiating and mature muscle tissue (Chen et al. 2001; Coutelle et al. 2001; Farnsworth et al. 2020).
Expression of myog in zebrafish cells follows that of myoD by several hours and is found in a subset of the myoD-expressing cells (Weinberg et al. 1996). Myog expression is commonly used as a marker for the transition from proliferation to differentiation in fish (Millan-Cubillo et al. 2019) and rodents (Asakura et al. 2001).
Xu et al. (2000) found that genes for muscle-specific sarcomeric proteins become expressed at different times throughout zebrafish somitogenesis (see Fig. 2), making them valuable markers of differentiation and maturation stages. Myosin heavy and light chain isoform composition changes as fish mature (Martinez et al. 1991; Veggetti et al. 1993; Johnston et al. 1998; Cole et al. 2004), suggesting that particular myosin isoforms may serve as markers of immature or mature fibers. Proteins such as alpha-actinin, titin, F-actin, and desmin are found in a striated pattern, which may be used to assess the extent of maturation and confirm the proper organization of muscle fibers (Costa et al. 2002; Ganassi et al. 2018).
A variety of markers are commonly used to identify slow muscle fibers and their precursors, including slow myosin heavy chain (Roy et al. 2001; Wolff et al. 2003; Baxendale et al. 2004; Hinits et al. 2009; Yao et al. 2013; Ganassi et al. 2018) and the transcription factors PR domain containing 1 (Prdm1/U-boot) (Hinits et al. 2009; Yin et al. 2018), prospero homeobox 1 (Prox1) (Roy et al. 2001; Wolff et al. 2003; Baxendale et al. 2004; Seger et al. 2011), and myocyte enhancer factor 2ca (Mef2ca) (Hinits et al. 2009). The Sonic hedgehog (Shh) target Patched1 (Ptc1) marks slow muscle precursors (Hinits et al. 2009). Markers of fish fast muscle include fast myosin heavy and light chains (Xu et al. 2000; Hinits et al. 2009; Yao et al. 2013; Ganassi et al. 2018), although myosin light chain 2 (mylz2) transcript is also weakly expressed in immature slow muscle (Hinits et al. 2009). Other markers include muscle α-actin (acta1), α-tropomyosin (tpma), troponin C (tnnc), troponin T (tnnt), and parvalbumin (pvalb) (Xu et al. 2000). In gilthead seabream, myoD is found in two isoforms, which may be differentially expressed between slow and fast muscle (Tan and Du 2002).
Signals Initiating Myogenic Commitment, Fusion, and Maturation
Various cues control the transition from stem cell to mature myofiber, some of which are outlined in Fig. 3. In cases where differences exist, the differentiation process will be discussed primarily in the context of fast muscle fibers; signals specific to slow fibers are discussed in the “Signals Regulating the Decision to Become Fast or Slow Fibers” section.
Signaling pathways involved in myogenic commitment, differentiation, fusion, and maturation in fish (Weintraub 1993; Maroto et al. 1997; Hamade et al. 2006; Feng et al. 2006; Hinits et al. 2009, 2011; Schnapp et al. 2009; Xu et al. 2013; Windner et al. 2015; Abraham 2016; Ferrari et al. 2019; Osborn et al. 2020). Gold indicates proteins primarily associated with maintenance of quiescent or proliferative states, teal indicates those primarily associated with differentiation, and gray indicates a mixed role. Dashed lines indicate interactions based on evidence in mammalian systems that have not been directly observed in fish (Kim et al. 2008; Dey et al. 2011)
MRFs
In the absence of both MyoD and Myf5, no muscle is formed in zebrafish embryos (Schnapp et al. 2009; Hinits et al. 2011). Although zebrafish Mrf4 does not usually participate in myogenic commitment (Hinits et al. 2009), ectopic mrf4 expression in myoD/myf5 morphant embryos induced expression of MyoD and myogenic commitment (Schnapp et al. 2009). For cultivated fish, inducing expression of mrf4 alone may therefore be a viable means of inducing both myogenic commitment and differentiation. Besides its role in promoting differentiation, Mrf4 is necessary for proper myofiber alignment in vivo (Wang et al. 2008). Zebrafish MyoD has been suggested to promote myogenesis by inducing the expression of myog (Hinits et al. 2009) and miR-206 (Hinits et al. 2011).
Fgf
Fgf signaling induces myod expression (Groves et al. 2005) via T-box16 (Tbx16) (Osborn et al. 2020). Consistent with this, overactivation of Fgf signaling causes zebrafish embryonic Pax7-expressing cells to differentiate into fast muscle, while inhibition leads to an overabundance of Pax7-expressing cells (Yin et al. 2018). Retinoic acid has been shown to bi-directionally modulate myoD and myog expression via Fgf8, preferentially in what will become fast fibers (Hamade et al. 2006). While treatment of zebrafish embryonic cells with FGF2 induces myf5 and mylz2 expression, expression of those genes was blocked by phosphoinositide 3-kinase (PI3K) or mammalian target of rapamycin (mTOR) inhibitors (Xu et al. 2013). However, FGF treatment also slightly impaired fusion into myotubes (Xu et al. 2013). Fgf promotes dedifferentiation in regenerating zebrafish extraocular muscles (Saera-Vila et al. 2016), and can stimulate the proliferation of committed myoblasts (Gabillard et al. 2010). These latter observations suggest that Fgf in fish may promote myogenic commitment and early differentiation while limiting later stages of differentiation, fusion, and maturation, consistent with findings in other model systems (Moore et al. 1991; Olwin and Rapraeger 1992; Hutson et al. 2010).
Wnt Pathway
It has been demonstrated in mouse C2C12 cells that Wnt signaling promotes myogenic differentiation (Abraham 2016), likely via β-catenin’s ability to interact with and enhance the activity of MyoD (Kim et al. 2008). The role of the Wnt pathway in fish myogenesis has been less well studied, though it has been shown that histone deacetylase 8 (Hdac8) positively regulates differentiation via the Wnt pathway (Ferrari et al. 2019).
Myomixer and Myomaker
Although the activation of muscle-associated genes and the fusion of myoblasts into multinucleated myofibers occur concurrently in vivo, they can be decoupled experimentally. The membrane-associated peptide Myomixer acts in concert with the transmembrane protein Myomaker to trigger the fusion of mammalian myoblasts, but does not affect myosin expression (Bi et al. 2017). The function of this peptide appears to be conserved across mammalian and fish lineages (Bi et al. 2017). Consistent with the idea that Myomixer and Myomaker specifically affect fusion and not differentiation generally, zebrafish embryos with a Myomaker loss of function show impaired fusion without losing expression of muscle-specific markers (Landemaine et al. 2014; Zhang and Roy 2017). Fusion primarily depends on activation of Myomaker and Myomixer by Myog, though some cells in the medial myotome remain fusion-competent due to activation of Myomaker by notochord-derived Hedgehog (Ganassi et al. 2018). However, it has also been demonstrated that Hedgehog overexpression inhibits expression of Myomixer (Wu et al. 2022) and Myomaker (Shi et al. 2018), leading to defects in fusion. This seeming discrepancy might be explained by the characteristics of different cell populations or by different responses to moderate versus high levels of Hedgehog.
Mesp-b, Tbx6, and Ripply1
In zebrafish, mesoderm posterior homolog B (Mesp-b) maintains embryonic muscle progenitors in an undifferentiated, proliferative state by inducing mesenchyme homeobox (meox1) expression and inhibiting myoD and myf5 expression (Windner et al. 2015). Ripply1 causes cells to differentiate by degrading Tbx6, an upstream regulator of mesp-b and ripply1 itself (Kinoshita et al. 2018). Thus, in the context of a cultivated fish bioprocess, manipulating the balance between Mesp-b/Tbx6 and Ripply1 could help maintain myogenic stem cells in an undifferentiated state or induce their differentiation.
Alignment, Stiffness, and Other Physical Cues
Myogenic commitment can be induced and maturation enhanced in vitro by various physical cues, including alignment, matrix stiffness, stretch/strain, and electrical stimulation (Lee et al. 2022). Human MSCs could be steered toward the myogenic lineage when grown on micropatterned fibronectin stripes (Yu et al. 2013). Increasing the alignment of myoblasts or MSCs by growing them on decellularized plants (Campuzano et al. 2020; Allan et al. 2021) or curved substrates (Wang et al. 2012a; Connon and Gouveia 2021) also enhances differentiation. Commitment and differentiation can also be influenced by the stiffness of the substrate on which the cells are grown (Engler et al. 2004, 2006; Freeman and Kelly 2017). To our knowledge, the effects of physical cues on myogenic differentiation have not been investigated in fish cells. Determining whether these effects exist in fish—and, if so, what specific values for variables such as stiffness and groove width most efficiently induce myogenesis and how these variables act in combination—may substantially improve product quality and scalability for CS.
Inducing Myogenesis in Culture
Ultimately, a successful CS bioprocess will require a reliable and efficient means of inducing myogenic commitment, differentiation, and maturation at the desired time. Table 2 summarizes several protocols demonstrated to induce myogenesis in cultured fish cells. Future work should use these results as a starting point, together with a detailed understanding of molecular pathways involved in myogenic differentiation that may be insufficiently activated by the published protocols—as well as other strategies such as the manipulation of cellular alignment—to guide the development of sufficiently robust methods.
Other methods for inducing myogenic differentiation have been demonstrated in cultured mammalian cells (Zhu et al. 2014) but not reported in fish. The reported strategies rely on myogenic inducers such as a specific skeletal muscle cell growth medium (SkGM™)-2 BulletKit™ (Lonza) containing human epidermal growth factor (hEGF), fetuin, FBS, dexamethasone, and insulin, supplemented with 5-azacytidine (Stern-Straeter et al. 2014; Okamura et al. 2018). Messmer et al. (2022) found that transferrin, insulin, lysophosphatidic acid, and glucagon increased differentiation of bovine SCs grown in serum-free conditions and that acetylcholine enhanced maturation and fusion. Chal et al. (2016) developed a protocol for in vitro generation of myofibers and SCs from human PSCs that relied on the sequential application of optimized culture media formulations. Based on the zebrafish blastomere screen described above, Xu et al. (2013) designed a differentiation cocktail containing FGF2, the adenylate cyclase activator forskolin, and the GSK3beta inhibitor BIO, which differentiated human iPSCs into multinucleated myotubes. Tanaka et al. (2013) demonstrated the direct conversion of human iPSCs to myocytes by expression of MYOD1, and Watanabe et al. (2011) noted that mouse myoblast-derived iPSCs may have improved myogenic differentiation potential compared to those derived from other cell types. Several studies have reported myogenic differentiation of MSCs in response to coculture with, or treatment with conditioned media from, myogenic cell lines (Stern-Straeter et al. 2014; Patruno et al. 2017; Korovina 2019). Because the overall process of myogenesis shares many molecular details between species, efforts to produce better methods for differentiating fish muscle should also explore the hypothesis that these same protocols might be successful in fish.
Signals Regulating the Decision to Become Fast or Slow Fibers
Fish white muscle, made up of fast fibers, is generally perceived most positively in a food context (Listrat et al. 2016). However, red muscle in the correct ratio and geometry may be required for specific products such as hamachi or to produce desirable flavor compounds during cooking. Because several transcription factors have dual roles in promoting cell differentiation and maturation and influencing muscle fiber type specification (Hinits and Hughes 2007; Potthoff and Olson 2007), monitoring fast and slow muscle marker expression during bioprocesses development will be necessary to avoid unwanted organoleptic effects. Proteins thought to be involved in fiber type specification are outlined in Fig. 4.
Proteins involved in specifying slow versus fast muscle fate in fish (Du et al. 1997; Barresi et al. 2000; Du and Dienhart 2001; Roy et al. 2001; Hinits and Hughes 2007; Potthoff et al. 2007; Potthoff and Olson 2007; Yao et al. 2013). Proteins shown in blue primarily promote development of slow muscle, and those shown in yellow primarily promote a fast muscle fate. Those that play an important role in both fiber types are shown in gray, with blue outlines indicating those playing a substantially greater role in promoting a slow muscle fate. Dashed lines indicate mechanisms that have been demonstrated in other model systems but not in fish
Fast and slow muscle progenitors can be distinguished within the zebrafish presomitic mesoderm (Devoto et al. 1996; Coutelle et al. 2001; Jackson and Ingham 2013). However, in some instances, cells’ commitment toward a specific fiber type could be reprogrammed with relative ease (Potthoff et al. 2007). Slow fibers develop from the notochord-adjacent adaxial cells, while the paraxial cells become fast muscle (Hatta et al. 1991; Devoto et al. 1996). Adult fish slow fibers are mononucleated, most of which are found in a thin layer below the skin as the “superficial slow fibers,” while those that form the horizontal myoseptum are derived from precursors known as the “muscle pioneers” (Hatta et al. 1991; Roy et al. 2001). The spatial separation of slow and fast muscle means that slow- and fast-fated myoblast cultures can be initiated simply by careful dissection of the muscle (Duran et al. 2020).
Many studies have implicated the Hh pathway, especially notochord-derived Shh, in steering zebrafish muscle cells toward maturation into slow fibers (Weinberg et al. 1996; Barresi et al. 2000; Du and Dienhart 2001; Coutelle et al. 2001; Osborn et al. 2011). Slow muscle development in vivo can be prevented by blocking Shh signaling (Blagden et al. 1997; Du et al. 1997). Together with the fact that zebrafish adaxial cells express fast muscle-specific genes prior to somite formation (Xu et al. 2000; Hinits and Hughes 2007), this may imply that myocytes become fast fibers by default unless exposed to sufficient Shh levels. Cell differentiation into slow fibers failed when gilthead seabream embryos were treated with forskolin, an adenylate cyclase activator, presumably due to Hh pathway inhibition (Tan and Du 2002). However, the effects of Hh signaling depend on dosage and timing, and in some cases, Hh may also promote differentiation into fast muscle fibers (Wolff et al. 2003; Feng et al. 2006).
Downstream signals mediating the effects of Hh signaling on zebrafish slow muscle include smoothened (Smo) (Barresi et al. 2000), glioma-associated oncogene (Gli2) (Du and Dienhart 2001), cyclin-dependent kinase inhibitor 1C (Cdkn1c) (Osborn et al. 2011), Prdm1a, and Prox1 (Roy et al. 2001). Prdm1a is antagonized by the transcription factor Pre-B-cell leukemia transcription factor (Pbx) (Yao et al. 2013). Pbx2 and Pbx4 cooperate with MyoD to promote differentiation into fast fibers (Maves et al. 2007).
Class II HDACs promote cells’ differentiation into fast muscle fibers in mice by repressing the transcription factor myocyte enhancer factor 2 (MEF2) (Potthoff et al. 2007). It is unknown whether this phenomenon exists in fish and, if so, which Mef2 and Hdac isoforms are involved. MEF2 regulates the differentiation of multiple cell types (Potthoff and Olson 2007) and zebrafish Mef2 plays a role in both fast and slow fiber maturation (Hinits and Hughes 2007). Therefore, it might be possible to control fiber type specification by manipulating this pathway in committed cell types such as SCs, though the effects may be sensitive to expression levels, isoforms (Ticho et al. 1996), splice variants, transcriptional states, or other factors.
Adipogenesis in Fish
Skeletal muscle-associated adipocytes are present in teleost species from multiple orders. In rainbow trout, red seabream, and pacific herring, adipocytes can be found in white muscle, myosepta, slow muscle, and in gaps between muscle bundles (Kaneko et al. 2016). In pelagic fish such as salmon, Pacific herring, and Pacific saury, intracellular lipid deposition can also be found in red muscle (Kaneko et al. 2016).
Adipocytes can differentiate from cells including MSCs (Ytteborg et al. 2015; Salmerón et al. 2016; Bou et al. 2017), FAPs (Reiss et al. 2021), preadipocytes, and—at least in mice—SCs (Asakura et al. 2001), as well as possibly from fibroblast-like cells (Tsuruwaka and Shimada 2022). The FAP differentiation process is conserved among species and potentially can be applicable for fish species. However, during fish adipogenesis, the timeline of events should be adjusted when translating such knowledge (Li et al. 2020).
Markers of Adipogenic Differentiation
Terminal differentiation of adipocytes can be observed as an increase in lipid accumulation, often visualized using stains such as oil red O (Vegusdal et al. 2003)—a standard readout of adipogenic differentiation—and characterized by increased expression of genes related to lipid metabolism (e.g., pparg, cebpb, and fas; Fig. 5).
Genes involved in fat differentiation and maturation in fish (Vegusdal et al. 2003; Oku and Umino 2008; Hesslein et al. 2009; Todorcević et al. 2010; Huang et al. 2012a; Mota de Sá et al. 2017; Liu et al. 2018; Salmerón 2018). Genes are ordered according to their approximate order of activation during adipogenic differentiation. Numbers to the left of some rows indicate the timing of activation within the zebrafish somite in hours post-fertilization (Den Broeder et al. 2015). Genes fading out to the right are downregulated in mature fat tissue relative to precursors. Red indicates genes associated with zebrafish MSCs (Fathi et al. 2019) and gold indicates those primarily associated with FAP appearance in mammalian systems (Li et al. 2020). Blue indicates the genes related to adipogenesis from the preadipocyte state until achieving a mature adipocyte
Salmon cells undergoing adipogenesis begin to express c/ebpβ, pparγ, fas, c/ebpα, bone morphogenetic protein 4 (bmp4), lipoprotein lipase (lpl), and c/ebpδ in approximately that order (Todorcević et al. 2010; Huang et al. 2010). Pparγ, C/ebP/α, and leptin expression have been previously reported in differentiated salmon adipocytes (Vegusdal et al. 2003), as have pparγ, c/ebpα, c/ebpγ, and fas in differentiating preadipocytes from grass carp (Liu et al. 2018). The most useful markers to distinguish preadipocytes from cells that have already begun adipogenesis are those whose expression can be allocated to specific stages. These include c/ebpγ, fas (Liu et al. 2018), c/ebpα (Todorcević et al. 2010; Huang et al. 2010), fatp1 (Huang et al. 2010), and bmp4 (Todorcević et al. 2010), which become expressed during intermediate stages of differentiation, and fabp11 (Huang et al. 2010) during later stages.
Signals Initiating Adipogenic Differentiation
As with muscle, various genes and signaling pathways are involved in activating or repressing adipogenesis in fish (Fig. 6).
Signaling pathways involved in adipogenic commitment, differentiation, and maturation in fish (Ross et al. 2000; Vegusdal et al. 2003; Oku and Umino 2008; Christodoulides et al. 2009; Hesslein et al. 2009; Todorcević et al. 2010; Cawthorn et al. 2012; Huang et al. 2012a; Ytteborg et al. 2015; Mota de Sá et al. 2017; Liu et al. 2018; Salmerón 2018). Gold indicates signals primarily associated with maintenance of quiescent or proliferative states and teal indicates those primarily associated with differentiation
Peroxisome Proliferator-Activated Receptors (Ppar ɑ, β, and γ)
Three Ppar isoforms (ɑ, β, and γ) are present in teleost fish, and these nuclear receptors have important roles in adipogenesis (Cruz-Garcia et al. 2009). Pparγ is the major adipogenic regulator that leads to transcriptional activation of genes that facilitate lipid storage and fatty acid metabolism (Salmerón 2018). This occurs when Pparγ attaches to the cis-retinoic acid receptor alpha (Rxrα) and binds the promoters of those genes. Pparγ is required for the in vitro differentiation of Atlantic salmon adipocytes and cooperates with other regulators such as C/Ebpα (Vegusdal et al. 2003). Oku and Umino (2008) supported that Pparγ is essential in adipocyte differentiation, although in red sea bream cultures this marker was not directly linked to adipocyte differentiation. While in humans (and mice) PPARγ was reported to have two isoforms (PPARγ1 and PPARγ2), in zebrafish there is only one (Pparγ1) (Den Broeder et al. 2015; Wafer et al. 2017).
CCAAT/Enhancer-Binding Proteins (C/Ebpα, C/Ebpβ, C/Ebpγ, and C/Ebpδ)
C/Ebps are a family of six transcription factors with several domains that allow DNA recognition followed by gene regulation, where C/Ebpα, β, γ, and δ specifically regulate genes that promote adipogenesis. The expression of each transcription factor is dependent on the species, but usually in mammals, C/Ebpβ and C/EbpPδ are induced in the early stages of adipogenesis and together induce expression of C/Ebpα (Mota de Sá et al. 2017). In fish, C/Ebpα and C/Ebpβ appear to have important roles in zebrafish embryonic development (Lyons et al. 2001), and both have significant protein homology to both human and mouse orthologs (Imrie and Sadler 2010). However, in studies performed with salmon preadipocytes, c/ebpα is expressed relatively late during adipogenesis (Todorcević et al. 2010; Huang et al. 2010). Moreover, C/Ebpα can impact its own production and the expression of Pparγ through a positive feedback mechanism (Rosen et al. 2002; Mota de Sá et al. 2017). In studies using carp preadipocytes, a high expression of c/ebpγ was also detected (Liu et al. 2018).
Zinc-Finger Protein 423 (Zfp423)
Zfp423 is a transcription regulator required in early adipogenic commitment with limited sequence variation among vertebrates (Gupta et al. 2010; Hamilton 2020). Zfp423 becomes active when two inhibitory complexes, Wnt1 inducible signaling path-way protein 2 (WISP2)–Zfp423 and ZNF52–Ebf1, disassociate, allowing Zfp423 and Ebf1 to enter the nucleus and activate Pparγ transcription (Hesslein et al. 2009; Hammarstedt et al. 2013). This process can be blocked by bta-miR-23a via direct targeting of Zfp423 (Guan et al. 2017). Therefore, Zfp423 expression is essential to initialize determination and to allow the commitment of preadipocytes. In fact, Zfp423 knockdown in mouse embryos resulted in impaired adipose tissue development (Gupta et al. 2010; Shao et al. 2017). The same knockdown in bovine adipose cells prevented their differentiation in vitro, while overexpression improved their adipogenic potential (Huang et al. 2012b). Moreover, studies using immortalized adipogenic progenitor cells from bovine muscle SVF found that Zfp423 regulation is required for intramuscular fat development and that highly adipogenic cells (cells that stain intensely with Oil-Red-O) expressed Zfp423, Pparγ, C/Ebpα, and C/Ebpβ (Huang et al. 2012b).
FGF-1 and FGF-2
Supplementation with FGF-1 and FGF-2 can positively impact adipogenic differentiation. In human AMSCs, FGF-1 supplementation before adipogenic induction with a hormone cocktail increased mRNA expression of C/EBPα in a dose-dependent manner (Kim et al. 2015). Other studies also reported that FGF-2 increases proliferation and directly affects adipogenic differentiation by inducing the expression of PPARγ (Kakudo et al. 2007). The mRNA expression of CEBPα was inversely correlated with FGF-2 concentration (Kim et al. 2015). Besides media supplementation, incorporation of FGF-2 in microspheres composed of gelatin enabled preadipocytes to differentiate into adipose tissue in a mouse model (Tabata et al. 2000; Yun et al. 2010).
TGF-β and BMP4
The TGF-β pathway, the conservation of which was investigated by Huminiecki et al. (2009), is composed of TGF-β and BMP signaling ligands, surface receptors, and SMAD signaling proteins. TGF-β ligands attach to TGF-β receptors that become phosphorylated, consequently leading to phosphorylation of SMAD3. Phosphorylated SMAD3 can cross the nuclear membrane and inhibit the expression of CEBPs and PPARγ (Li and Wu 2020). Consequently, early TGF-β signaling appears to promote adipogenic commitment but also to inhibit adipogenic differentiation. TGF-β is somewhat conserved across species, with peptide sequences from rainbow trout strongly aligned with those of striped bass (Harms et al. 2000). Carp Tgf-β2 appears to be loosely related to avian and mammalian TGF-β2 isoforms.
BMP4, a member of the TGF-β superfamily, is secreted by differentiated preadipocytes and induces the adipogenic commitment of precursor cells (Gustafson et al. 2015). When BMP4 is released, it activates a receptor that promotes the dissociation of the inhibitory WISP2–ZFP423 complex (Hammarstedt et al. 2013), thereby activating PPARγ transcription. In addition, cultures of Atlantic salmon primary adipose SVF cells demonstrated an increased expression of Bmp4 after chemical induction of adipocyte differentiation (Todorcević et al. 2010).
Wnt Pathways Regulate Adipogenesis
The Wnt family of signaling proteins has an important role in regulating tissue maintenance and remodeling. Activation of the Wnt pathway and signaling through β-catenin represses adipogenesis by inhibiting the expression of PPARγ and CEBPα (Christodoulides et al. 2009). One mechanism by which Wnt family proteins prevent activation of PPARγ is thought to be activation of WISP2, the aforementioned inhibitor of Zfp423 (Hammarstedt et al. 2013). As mentioned in the “Preadipocytes” section, the Wnt/β-catenin pathway has maintained preadipocytes in an undifferentiated and proliferative state in carp (Liu et al. 2018). In cultures of grass carp preadipocytes, this was observed after 7 days of DHA (100 µM) supplementation, where carp adipocytes had decreased expression of C/Ebpα, Pparγ, C/Ebpγ, and Fas (Liu et al. 2018). In 3T3-L1 preadipocytes, a similar treatment resulted in expression of Wnt1, thus inhibiting the activation of PPARγ gene expression (Moldes et al. 2003).
In addition to this canonical pathway, Wnt family members can also activate non-canonical signaling through Wnt5a and Wnt5b, both of whose gene expression in salmon MSCs increases throughout the adipogenic process (Ytteborg et al. 2015). In 3T3-L1 preadipocytes, Wnt5b promotes the adipogenic process by down-regulating β-catenin (Kanazawa et al. 2005). Both Wnt5a and Wnt5b have been suggested to promote early adipogenesis by enhancing the expression of PPARγ (Kanazawa et al. 2005; van Tienen et al. 2009). Besides those factors, Wnt6, Wnt10a, and Wnt10b are also early regulators of adipogenic commitment and their overexpression leads to a decrease in the mRNA expression of PPARγ and C/EBPα, mediated by β-catenin (Cawthorn et al. 2012). Moreover, Wnt10b helps maintain the preadipocyte state, and its blockage induces transdifferentiation of myoblasts into adipocytes (Ross et al. 2000). While most of these studies are on mammalian cell models, their conservation among different animals may inspire strategies to induce adipogenesis in fishes.
IGF-1 and Insulin
Insulin and insulin-like growth factor 1 (IGF-1) are important adipogenic regulators, as demonstrated in vivo by reductions in adipose tissue formation in transgenic mice lacking insulin and/or IGF-1 receptors (Boucher et al. 2016), and also influence preadipocyte proliferation (see Sect. 2.5). While both insulin and IGF-1 increased differentiation and lipid accumulation of gilthead seabream preadipocytes, a stronger effect was observed from IGF-1 (Salmerón et al. 2013).
Hedgehog (Hh) Pathway
The Hh pathway regulates cell fate determination, proliferation, migration, polarity, and gene expression. In adipogenesis, the Hh pathway is involved with expression of PPARγ, leading to possible alterations in the fates of precursor cells. In 3T3-L1, NIH-3T3 cells, and porcine AMSCs, Hh pathway activation inhibits adipogenic differentiation (Fan et al. 2018). Such inhibition was also observed during fat formation in 3T3-L1 preadipocytes and in the Drosophila fat body, suggesting a conserved role for the Hh pathway as an adipogenic regulator in vertebrates and invertebrates (Suh et al. 2006). Even with less information on the impact of Hh pathway modulation on fish fat tissue, Wynne et al. (2021) considered that this pathway is also associated with cell fate and proliferation in teleost fishes.
MicroRNAs
MicroRNAs (miRNAs) are short RNAs that regulate gene expression via multiple mechanisms and have a well-characterized role in regulating mammalian adipogenesis (Romao et al. 2011), including a potential role in bovine intramuscular and subcutaneous fat (Guo et al. 2017; Mir et al. 2020). As of March 3, 2022, miRBase—a database of published miRNA sequences (Griffiths-Jones 2004; Griffiths-Jones et al. 2006, 2008; Kozomara and Griffiths-Jones 2011, 2014; Kozomara et al. 2019)—contained sequences of 1917 precursors and 2654 mature mRNAs from human, 1064 precursors/1025 mature from bovine, 371 precursors/497 mature from salmon, and 355 precursors/373 mature from zebrafish.
Various miRNAs have been identified that promote or inhibit adipogenesis, including in fish. MiR-143 has been characterized as a marker of lipid deposition in rainbow trout, and some evidence suggests a mechanistic role in promoting adipogenesis via inhibition of the α/β hydrolase-domain 5 (abhd5) gene (Mennigen et al. 2013). Similarly, miR-150-4p expression in chickens promoted differentiation of intramuscular adipocytes by targeting retinoid X receptor gamma (Zhang et al. 2018).
Depleting miR-27b in zebrafish increased adipocyte hyperplasia, lipid accumulation, and expression of adipocyte-related genes including Pparγ and C/Ebpɑ, indicating that it serves as a negative regulator of adipogenesis (Hsu et al. 2018). This is consistent with findings in other vertebrates; for example, in human MSC-derived adipocytes, miR-27b has been shown to directly suppress PPARγ and to inhibit lipid accumulation and adipogenesis-associated marker gene expression when overexpressed (Karbiener et al. 2009). In both mouse and human cells, miR-182 inhibits adipogenesis by targeting C/EBPɑ and is downregulated temporarily during early adipogenesis (Dong et al. 2020).
Terminal Differentiation and Accumulation of Lipid Droplets (i.e., Lipogenesis)
A lipid source (e.g., oleic acid or a lipid cocktail) is routinely added to the medium for adipogenic differentiation of fish preadipocytes (Vegusdal et al. 2003; Todorcević et al. 2010; Liu et al. 2018; Salmerón 2018), though it has also been reported that long-chain omega-3 fatty acids may inhibit differentiation (Huang et al. 2010; Liu et al. 2018). Differentiation of Atlantic salmon preadipocytes was induced by a cocktail of medium supplements including insulin, dexamethasone, biotin, triiodothyronine, pantothenate, isobutylmethylxanthine (IBMX), fatty acids, and cholesterol (Todorcević et al. 2010) (Table 3).
In differentiated adipocytes from Atlantic salmon, the expression of genes for adipokines adipsin and visfatin coincides with the accumulation of lipid droplets (Todorcević et al. 2010). An increase in expression of NADPH-related genes, such as glucose-6-phosphate dehydrogenase (g6pd) or 6-phosphogluconate dehydrogenase (pgd), has been reported in terminal differentiation of salmon adipocytes, which is aligned with the need of NADPH for triacylglycerol/lipid production and accumulation (Todorcević et al. 2010).
Insulin decreased lipolysis in the mature adipocytes, whereas Tnfɑ increased this process (Wang et al. 2012b). However, while another study in adipocytes isolated from gilthead seabream found that insulin decreased lipolysis in some experiments, these effects were inconsistent (Albalat et al. 2005).
Tnf-related genes are down-regulated in salmon SVF cells upon adipogenic induction, though up-regulated at initial stages before reaching confluence (Todorcević et al. 2010). Further research in gilthead seabream mesenteric adipocytes suggests that Tnfɑ regulation of adipogenic factors varies amongst fat and lean phenotypes (Cruz-Garcia et al. 2009). In this study, the authors reported that Tnfɑ had lipolytic effects and reduced lipid accumulation characterized by Pparγ downregulation in adipocytes from lean fish. In contrast, the adipocytes from fat specimens had Pparβ-mediated lipolytic effects or no apparent changes from Tnfɑ supplementation.
Fish Connective Tissue, Vascular Tissue, and Skin
While muscle and fat cells are the main contributors to the organoleptic and nutritional properties of meat, connective tissue also plays an essential role in both the mechanical properties of the tissue and the changes to those properties during the cooking process (Listrat et al. 2016). In the context of CS, the scaffold might fully or partially substitute this role. Fibroblasts or other extracellular matrix-secreting cell types could also be incorporated. Fortunately, fibroblast-like cells and cells derived from fin tissue are abundant among fish cell lines (Thangaraj et al. 2021) and are easily isolated and cultured. Future research into the use of fibroblasts in CS should focus on identifying optimal culture conditions for serum-free co-cultures of fibroblasts with myogenic and adipogenic cells and investigating the effects of these cell types on one another’s proliferation and differentiation.
Bricard et al. (2014) identified a population of extracellular matrix-secreting cells in the myosepta of trout embryos that appeared analogous to mammalian tenocytes and expressed the tenocyte marker scleraxis. This population was later shown to be dependent on Hh signaling, and its loss was shown to lead to a muscle detachment phenotype (Ma et al. 2018). This latter observation suggests that tenocytes might substantially contribute to fish’s mechanical and organoleptic properties.
Although vascularization of tissue is not likely to contribute meaningfully to taste or texture (Listrat et al. 2016), some scaffolding strategies might require the presence of endothelial or smooth muscle cells. Adding pro-angiogenic factors (Huang et al. 2012a) at strategic points during the bioprocess might facilitate the creation of vascularized tissues from a single multi- or pluripotent cell line.
For many CS product applications, a lack of skin will be an advantage (Rubio et al. 2019a). In cases where skin is desirable, co-cultures of fibroblasts and scale-derived epithelial cells may be used (Rakers et al. 2011), likely in conjunction with methods to encourage slow muscle growth (see Section 3.4).
Myogenesis and Adipogenesis in Aquatic Invertebrates
Myogenesis and adipogenesis are even less well understood in aquatic invertebrates than in fish. Table 4 summarizes some of the molecular players and pathways thought to be involved in invertebrate myogenesis.
Crustaceans
Myogenesis
In many crustaceans, muscle fiber types appear to exist on a spectrum rather than in distinct fast and slow categories, with each type having a unique expression profile of myofibrillar proteins and isoforms (Medler and Mykles 2003). Relative composition of fast or slow types, and that of tissues undergoing protein synthesis or degradation, vary continually to accommodate the remodeling required for ongoing molt cycles (Mykles 1997). These variations may have implications for cultivated crustacean meat.
Early myogenesis appears to be similar to that of the fruit fly Drosophila melanogaster where “founder” cells migrate from the mesoderm and fuse with undifferentiated myoblasts to form muscle progenitors (Kreissl et al. 2008; Jirikowski et al. 2010; Harzsch and Kreissl 2010). Myogenesis also occurs during appendage regeneration in both groups, though in crustaceans, this process more closely replicates embryogenesis where a developing blastema forms the pool of undifferentiated cells from which myogenic precursors emerge (Hopkins et al. 1999; Hopkins and Das 2015).
Little is known about the molecular drivers of myogenesis in crustaceans; however, it is recognized that there are likely similarities with Drosophila (Mykles and Medler 2015), where the key myogenic transcription factors are Twist, Nautilus, and Mef2 (Taylor 2006). Twist, most similar in function to vertebrate MyoD, is expressed early and is important for initial determination and patterning of the mesoderm, and then for formation of early myogenic progenitors and founder cells (Taylor 2006; Bothe and Baylies 2016). Nautilus, more similar to MyoD in sequence (Taylor 2006), is expressed later and is important for founder cell patterning (Wei et al. 2007). Nautilus has also been shown to initiate the myogenic program in adult fibroblasts (Zhang et al. 1999) and cardioblasts using the GAL4-targeted system (Keller et al. 1997). Mef2 works synergistically with Twist and Nautilus, as it does with the vertebrate MRFs, and is important for cells’ terminal differentiation and fusion (Bour et al. 1995; Taylor 2006; Bryantsev et al. 2012).
In the isopod crustacean Parhyale hawaiensis, expression of Twist is also evident at the mesodermal patterning stage and induces Mef2, which is required for later muscle determination and differentiation (Price and Patel 2008). In various penaeid shrimp, Mef2 appears to be expressed earlier than Twist but is also necessary for myogenic determination and differentiation (Wei et al. 2016). Studies on crustacean Nautilus were not found; however, in the crayfish Cherax destructor, Pax3 is implicated in embryonic myogenesis as well as after molting and during appendage regeneration (White et al. 2005). A regenerative role for Pax3 has also been seen in Parhyale (Konstantinides and Averof 2014).
Myostatin in crustaceans appears to have multiple functions beyond myogenic regulation and its direct effect on myogenesis remains unclear (Mykles and Medler 2015; Yan et al. 2020). Studies on shrimp show both a positive effect (De Santis et al. 2011) and a negative effect (Yan et al. 2020; Wang et al. 2021), and its effect on different muscle types (thoracic versus claw) within the same crab species also appear to be contradictory (MacLea et al. 2012).
Molt Hormones and Growth Factors
Molting is intrinsically linked to muscle growth and development in all arthropods, driven by ecdysteroids such as ecdysone (Mykles and Medler 2015). In Drosophila, ecdysone appears to induce myogenesis through activation of Mef2 via Twist (Lovato et al. 2005). Several in vitro insect studies have shown ecdysone media supplementation induces terminal differentiation in various myogenic cells, particularly myoblasts (Baryshyan et al. 2012; Rubio et al. 2020). In crustaceans, high titers of ecdysteroids have been shown to stimulate protein synthesis in different muscle types across the molt cycle, although apparently through varying pathways such as myostatin or Rheb (Mykles 1997; Covi et al. 2010; MacLea et al. 2012). Figure 7 outlines these potential pathways in the claw muscle of premolt land crab. This hormone is readily accessible and thus an ideal candidate with which to begin crustacean muscle differentiation experiments.
Potential signaling pathway of the molt hormone ecdysone in land crab claw muscle during the premolt stage (Covi et al. 2010; MacLea et al. 2012). The active form, 20HE, binds to the EcR-RxR nuclear receptor and activates Rheb, either directly or through the repression of myostatin (possibly via a corepressor). Rheb is a major activator of mTOR, known to stimulate protein translation. Gold indicates where inactivation is required for myogenic protein synthesis, and teal indicates activation. Dashed lines indicate steps where the exact mechanism(s) involved are unclear
The effect of growth factors has also been observed on crustacean muscle growth, with IGF supplementation both in vitro and in vivo showing increases in muscle protein synthesis in crayfish (Chaulet et al. 2012; Jayesh et al. 2015). Other studies have shown varying results using different recombinant growth factors, with one review highlighting that growth factors obtained from more species-relevant serums or tissue extracts are likely to have more promising results (Ma et al. 2017). Once identified, crustacean growth factor homologs could potentially be overexpressed to induce myogenesis or made recombinantly and used in media supplementation.
Crustacean Fat Synthesis
Rather than intramuscular adipocytes as in vertebrates, crustacean muscle lipid content appears to be derived mainly from cell membrane phospholipids (PL) and sterols, with a large component of the PLs being long chain (lc) polyunsaturated fatty acids (PUFA), such as the nutritionally important omega-3 fats, eicosapentaenoic acid (EPA) and DHA (Chapelle 1977; Zhao et al. 2015; Shu-Chien et al. 2017; Lu et al. 2020). Although some lc-PUFA synthesis genes have shown expression in muscle tissue (Toledo 2019), crustacean lipid synthesis and oxidation primarily occurs in the hepatopancreas (the organ functionally equivalent to the liver and adipose tissue in vertebrates, and the fat body in insects) with lipids (predominantly phospholipids) being disseminated to other tissues, including muscle, via the hemolymph (O’Connor et al. 1968; Teshima et al. 1986; Garofalaki et al. 2006). A close examination of the hepatopancreas/hemolymph/muscle relationship could therefore inform attempts to ensure cultivated crustacean meat contains the correct lipid profiles. This might involve co-culturing cells or even a feeder cell system akin to Integriculture’s CultNet system, which allows media to be circulated between chambers containing different cell types (Hanyu and Kawashima 2017). However, because some lc-PUFAs, including EPA and DHA, are considered essential fatty acids in many crustaceans, unable to be synthesized by either organ (Zhao et al. 2015; Shu-Chien et al. 2017; Toledo 2019), such a system on its own is likely to be insufficient. As such, appropriate lipid profiles might be simply achieved with media or scaffold supplementation of just the essential—or perhaps all—required lipids that can be derived from plant sources or precision fermentation.
Alternatively, if driving endogenous adipogenesis is a consideration, then a deeper understanding of molecular mechanisms is needed. There is no known PPARγ equivalent in flies or crustaceans, although some suggest the multifunctional and molt-related nuclear receptor E75 may fill a similar role (Hong and Park 2010).
Mollusks
The culture of molluscan cells is a relatively new and under-explored area, as reviewed by Yoshino et al. (2013).
Cephalopods
Cephalopod muscle structure is substantially different from amniotes and fish, with three sets of short (typically under one millimeter), mononucleated muscle fibers oriented in roughly perpendicular directions (Gosline and DeMont 1985; Kier 2016). Cephalopod fast fibers are distinguished by shorter sarcomeres and lower paramyosin content rather than by differences in myosin isoform expression as in vertebrates (Kier and Schachat 1992). As paramyosin contributes to the gel characteristics of squid meat (Sano et al. 1986), paramyosin content—along with muscle fiber ultrastructure—may be a key variable to optimize when developing differentiation protocols for cultivated cephalopod meat. The transcription factor NK4 is thought to play a role analogous to vertebrate Pax3/7 in determining myogenic precursors, while the Hh pathway is thought to be involved in muscle precursor proliferation (Zullo et al. 2017). For example, Grimaldi et al. found that Hh and Ptc are both expressed in cuttlefish myoblasts fated to become radial fast fibers and that inhibiting the pathway induced apoptosis and reduced the muscle precursors’ proliferation rate (Grimaldi et al. 2008). Myf5 and MyoD share functions between vertebrate and cephalopod lineages, while the factors involved in myotome determination and in differentiation are largely unknown (Zullo et al. 2017).
Bivalves
Our understanding of the molecular pathways involved in bivalve myogenesis is incomplete, though the Hh pathway has been implicated in myogenesis in oysters (Li et al. 2018). The insulin, PI3K-Akt, and mTOR pathways have been shown to correlate with growth rates in clams (Nie et al. 2021), though causal relationships have not been definitively established and effects in tissues besides muscle cannot be ruled out. In oysters, these pathways seem to be similarly correlated to muscle growth (Choi et al. 2018; Kim and Choi 2019) and a number of newly characterized insulin-like peptides appear to be critically, but variously, involved in muscle growth regulation (Li et al. 2021). A temporal expression analysis in scallop muscle has highlighted twist2 as a key component for myogenic differentiation of MSCs (Sun et al. 2021). In mussels, myogenic differentiation is influenced by the presence of particular extracellular matrix molecules, with collagen substrates tending to reversibly inhibit differentiation, and fibronectin, poly-L-lysine, and carbon-coated substrates promoting differentiation (Odintsova et al. 2010; Dyachuk 2013).
Gastropods
Myogenesis in gastropods is similarly poorly understood. Downregulation of abalone myostatin led to upregulation of the insulin pathway, which was taken as a proxy for somatic growth (Carrera-Naipil et al. 2016). Numerous microRNAs and long non-coding RNAs have been identified as differentially expressed between the muscle tissue of large and small abalone, suggesting that some of these might regulate myogenesis (Huang et al. 2018a, b). An abalone homolog of the homeobox gene Mox (also known as Meox) has been identified and, based on its expression pattern with the developing somite, was hypothesized to play a similar role in the development of the early mesoderm and the muscle lineage as in vertebrates (Hinman and Degnan 2002).
Recommendations for Future Research
Controlling Proliferation and Differentiation
While culture methods for fish embryonic and adult stem cells exist, optimizing culture media and growth conditions for long-term stemness maintenance and increased proliferation rates will help facilitate the scale-up process. This may be accomplished by optimizing component concentrations in existing formulations, likely assisted by statistical methods such as design of experiments (Cosenza et al. 2021) and testing recombinant species-specific growth factors (Venkatesan et al. 2022). Compounds such as IGF-2 (Rius-Francino et al. 2011) and anthocyanidins (Villasante et al. 2016), which have been demonstrated in fish myogenic stem cells to stimulate proliferation and increase expression of stem cell markers, respectively, may also be considered as media additives.
General strategies for inducing differentiation can be broadly divided into physical methods, in which the geometry or mechanical properties of the culture environment are manipulated, and chemical or molecular methods (Fig. 8). In both cases, gene expression patterns corresponding to the desired cell type are activated. Most of the existing data on fish myogenic and adipogenic differentiation come from studies in zebrafish. This makes zebrafish an invaluable tool for CS research (Potter et al. 2020), but also means that translating this knowledge to other species—a necessity if CS is to recreate the variety of seafood products available today—will require extensive research. Differences in the biology of marine and freshwater fish may make this challenge more difficult for marine species.
Several general strategies for inducing differentiation of cultured cells have been described in the literature and may be relevant to CM and CS, though not all have been applied to fish or other aquatic species. Starting and desired cell types shown are those especially relevant to CM and CS, but are not an exhaustive list. Strategies can be broadly categorized into physical methods and chemical or molecular methods, and multiple strategies may be combined to enhance differentiation
Physical methods have shown promising results in mammalian tissue engineering studies (Lee et al. 2022). They should be investigated as one possible strategy—perhaps in combination with chemical and molecular methods—for scalably inducing myogenic and adipogenic differentiation in cells from various seafood species. However, energy needs for strategies such as cyclic stretch and electrical stimulation must be considered.
Chemical and molecular methods have been better studied in fish cells, but more work is needed to ensure that the reagents used to control cell fate are food-safe, low-cost, scalable, and environmentally sustainable. It will be essential to consider the food safety implications of such reagents at the concentrations—including those of byproducts—expected in the final product (Ong et al. 2021). This includes perceived risks; for example, some consumers may be hesitant toward products in which genetic modification is used for cellular reprogramming or to induce differentiation. Such concerns may or may not correlate with scientifically backed safety concerns and may show regional variation (Bryant et al. 2020). Many of the compounds commonly used in differentiation media for research purposes, e.g., IBMX for adipogenesis, are not food-grade (Fish et al. 2020). Knowledge of the signaling pathways mediating the effects of such reagents on cell fate may inform efforts to replace them with safe and effective alternatives.
Fish Myogenesis
Reagents previously demonstrated to induce myogenesis in fish stem cells include IGF1, IGF2, FGF2, GSK3b inhibitors, calpain inhibitors, and adenylate cyclase activators (see Table 2). Forskolin is a plant-based adenylate cyclase activator, sometimes taken as a dietary supplement, which has been shown to stimulate zebrafish SC proliferation and ESC-like cell myogenesis (Xu et al. 2013). While its safety as a supplement has not been conclusively demonstrated, existing evidence indirectly suggests the possibility of its safe use in inducing myogenesis in CS. Godard et al. (2005) used 25 mg forskolin per day to study weight loss in humans, while Xu et al. (2013) used 50 μM forskolin to induce zebrafish ESC-like cell myogenesis. Therefore, 1 l of the differentiation media would contain approximately the equivalent of one daily dose of forskolin. The amount remaining in a serving of CS—presumably considerably less since it is not known to accumulate in animal tissue—would need to be tested along with a rigorous safety profile.
Serum starvation is a reliable method for inducing myogenic differentiation in various species, including fish (Gabillard et al. 2010). While animal-derived serum is a poor choice for use in CM or CS, RNA sequencing of serum-starved cells was recently used to guide the development of a serum-free formulation for myogenesis of cultured bovine SCs (Messmer et al. 2022). Similar strategies could be employed for CS.
Muscle Maturation, Fiber Type, and the Need for Consumer and Sensory Research
While myogenic differentiation is undoubtedly necessary for CS, it will be essential to understand the relationship between the extent of maturation and organoleptic properties. Given the differences in overall toughness and typical fiber lengths (Listrat et al. 2016), it would be reasonable to hypothesize that extensive fusion and maturation might be more necessary for terrestrial meat than fish.
Similarly, sensory and consumer research will determine how red muscle and skin—whether as part of the final product or removed after cooking and before eating—influence product acceptability. Where slow muscle is desirable, spatially defined cues that activate genes such as Mef2, Prdm1a, or the Hh pathway could be introduced. If cells develop into undesired slow fibers, strategies to activate class II Hdacs, Pbx, or Protein kinase A (PKA) may be helpful. However, because some evidence points to fast muscle as the “default” (Blagden et al. 1997; Du et al. 1997; Xu et al. 2000; Hinits and Hughes 2007), such cues may be unnecessary. In vivo, muscle pioneers are essential for maintaining the chevron shape of the myomeres, but make up a small percentage of slow fibers (Keenan and Currie 2019). It is conceivable that one might try to recapitulate the in vivo self-organization processes that lead to chevron formation (Rost et al. 2014; Tlili et al. 2019) using known signals for muscle pioneer or superficial slow fiber identity (Nguyen-Chi et al. 2012) as part of a CS bioprocess for whole cut filets. However, the muscle pioneers may otherwise be dispensable.
Fish Adipogenesis
Development of food-grade differentiation protocols for fat will similarly require extensive optimization based on an understanding of the molecular pathways involved, followed by empirical testing. The fact that lipids tend to promote differentiation (Vegusdal et al. 2003) is an advantage since lipid-containing media may be used to simultaneously induce differentiation and control the lipid profile of the final product. However, tradeoffs may exist between the health benefits of omega-3 fats in the final product and their sometimes detrimental effects on differentiation (Huang et al. 2010; Liu et al. 2018).
Invertebrates
Compared to fish, even less is known about differentiation into meat-relevant cell types in aquatic invertebrates. Muscle fiber structure and type composition vary considerably both from vertebrates and among invertebrates, and crustacean muscle structure varies considerably across the molt cycle (Mykles 1997), the organoleptic implications of which are not well understood. Knowledge of the molecular drivers and pathways involved in myogenesis and adipogenesis for these animals is particularly poor. However, as for fish, what is known can inform potential avenues for analysis.
Identification of a crustacean Nautilus homolog and its potential to initiate myogenesis in different stem cell types would be informative, as would investigating the effects of Twist, Mef2, and Pax3. There is still uncertainty around myostatin’s positive or negative effect on crustacean myogenesis, so this needs further elucidation. Some potential reagents to investigate are ecdysone and various growth factors, with emphasis on more species-relevant proteins.
As crustacean muscle tissue does not synthesize fat locally and lacks intramuscular adipocytes, fat supplementation via the media or scaffold may be necessary and sufficient to achieve appropriate fat profiles and distributions in cultivated crustacean meat.
Because crustaceans and insects share a close evolutionary relationship, knowledge about the mechanisms of myogenesis and adipogenesis may provide a “shortcut” to understanding these processes in crustaceans. It has also been suggested that cultured insect cells might be used directly as cell sources for cultivated CS, as the necessary culture conditions to grow such cells are both flexible and well-understood (Rubio et al. 2019b, c; Letcher et al. 2022). If the organoleptic properties of such products meet consumers’ expectations, this strategy may be effective at addressing the challenges related to the cost and scale of CS.
Research into mollusk myogenic differentiation could build on findings concerning molecular pathways such as Hh and various transcription factors. The work of Dyachuk (2013) and Odinstova et al. (2010) provide clear avenues to investigate culture substrate effects on myogenesis.
Conclusion
The complexity of differentiation is a challenge for researchers attempting to identify the most efficient means of inducing the desired cellular behavior. It is also an opportunity because it offers many possible strategies. The task of future research will be to select those methods—or combinations of methods—that achieve the desired results in a manner that is cost-effective, food-safe, and sustainable.
For the potential benefits of CM and CS to be realized, it is essential that production costs be compatible with commodity meat prices. Several techno-economic assessments have been published, many of which model a wide range of scenarios and generate a similarly wide range of modeled costs. Published estimates include costs of USD $22—51/kg (Humbird 2021), $1.95—437,000/kg (Risner et al. 2021), $6.43—22,423/kg (Vergeer et al. 2021), $13.00—$30.40/kg (Negulescu et al. 2022), and $63/kg (Garrison et al. 2022). A primary cost driver—especially in nearer-term scenarios—is the culture medium, including macronutrients (primarily amino acids and glucose), growth factors, and other recombinant proteins (primarily albumin). Therefore, development of low-cost culture media is of paramount importance for CM and CS to achieve price parity with conventional animal products and be profitable. In some modeled scenarios where the cost of media had already been substantially reduced, capital expenditures and labor also made substantial contributions to the cost of production. Improvements to bioreactor technologies will help both to reduce capital expenditures and to increase the achievable cell densities and growth rates. When considering large-scale CM or CS production in bioreactors, many factors must be optimized to improve cell proliferation, including reducing shear-stress, adequately monitoring variables such as pH and carbon dioxide (Bellani et al. 2020), and removing waste metabolites from media. Promising solutions for the latter that are being pursued in for-profit organizations include medium recycling systems based on dialysis (Nahmias 2020) or genetic engineering techniques that reduce the production of toxic products such as ammonia (Genovese et al. 2021). Large-scale commercialization will also be enabled by innovations in standardized tissue sampling, cell banking, immortalization, reprogramming, scaffolding, and end product characterization. A recent analysis of spent media from cultured mouse and chicken cells revealed substantial variation in utilization of various key nutrients, suggesting that media formulations are unlikely to be interchangeable across species (O’Neill et al. 2022). It is likely to be generally true that cells will have different metabolic needs and different needs for specific growth factors to control their proliferation and differentiation depending on their species and cell type. By better understanding the needs of individual cell lines, it will be possible to improve the efficiency of CM and CS bioprocesses, thereby reducing both the environmental impacts and the cost of production for future products.
Data Availability
Not applicable.
Code Availability
Not applicable.
References
Abraham ST (2016) A role for the Wnt3a/β-catenin signaling pathway in the myogenic program of C2C12 cells. In Vitro Cell Dev Biol Anim 52:935–941
Albalat A, Gómez-Requeni P, Rojas P et al (2005) Nutritional and hormonal control of lipolysis in isolated gilthead seabream (Sparus aurata) adipocytes. Am J Physiol Regul Integr Comp Physiol 289:R259–R265
Allan SJ, Ellis MJ, De Bank PA (2021) Decellularized grass as a sustainable scaffold for skeletal muscle tissue engineering. J Biomed Mater Res A 109:2471–2482
Arrighi N, Moratal C, Clément N et al (2015) Characterization of adipocytes derived from fibro/adipogenic progenitors resident in human skeletal muscle. Cell Death Dis 6:e1733
Asakura A, Komaki M, Rudnicki M (2001) Muscle satellite cells are multipotential stem cells that exhibit myogenic, osteogenic, and adipogenic differentiation. Differentiation 68:245–253
Azoff M (2022) 2021 Industry update: Alternative seafood. The Good Food Institute. https://gfi.org/wpcontent/uploads/2022/04/2021-Alternative-Seafood-Industry-Update.docx-2.pdf. Accessed 05 Oct 2022
Barman AS, Lal KK, Rathore G et al (2014) Derivation and characterization of a ES-like cell line from Indian catfish Heteropneustes fossilis blastulas. Sci World J 2014:427497
Barresi MJ, Stickney HL, Devoto SH (2000) The zebrafish slow-muscle-omitted gene product is required for Hedgehog signal transduction and the development of slow muscle identity. Development 127:2189–2199
Baryshyan AL, Woods W, Trimmer BA, Kaplan DL (2012) Isolation and maintenance-free culture of contractile myotubes from Manduca sexta embryos. PLoS ONE 7:e31598
Baxendale S, Davison C, Muxworthy C et al (2004) The B-cell maturation factor Blimp-1 specifies vertebrate slow-twitch muscle fiber identity in response to Hedgehog signaling. Nat Genet 36:88–93
Béjar J, Hong Y, Alvarez MC (1999) Towards obtaining ES cells in the marine fish species Sparus aurata; multipassage maintenance, characterization and transfection. Genet Anal 15:125–129
Bellani CF, Ajeian J, Duffy L et al (2020) Scale-up technologies for the manufacture of adherent cells. Front Nutr 7:575146
Benjaminson MA, Gilchriest JA, Lorenz M (2002) In vitro edible muscle protein production system (MPPS): stage 1, fish. Acta Astronaut 51:879–889
Berberoglu MA, Gallagher TL, Morrow ZT et al (2017) Satellite-like cells contribute to pax7-dependent skeletal muscle repair in adult zebrafish. Dev Biol 424:162–180
Berry R, Rodeheffer MS (2013) Characterization of the adipocyte cellular lineage in vivo. Nat Cell Biol 15:302–308
Bi P, Ramirez-Martinez A, Li H et al (2017) Control of muscle formation by the fusogenic micropeptide myomixer. Science 356:323–327
Biferali B, Proietti D, Mozzetta C, Madaro L (2019) Fibro-Adipogenic Progenitors Cross-Talk in Skeletal Muscle: The Social Network. Front Physiol 10:1074
Birkenhead FES (1930) The world in 2030 AD. Hodder and Stoughton, London
Blagden CS, Currie PD, Ingham PW, Hughes SM (1997) Notochord induction of zebrafish slow muscle mediated by Sonic hedgehog. Genes Dev 11:2163–2175
Boone Kauffman J, Arifanti VB, Hernández Trejo H et al (2017) The jumbo carbon footprint of a shrimp: carbon losses from mangrove deforestation. Front Ecol Environ 15:183–188
Bothe I, Baylies MK (2016) Drosophila myogenesis. Curr Biol 26:R786–R791
Bou M, Montfort J, Le Cam A et al (2017) Gene expression profile during proliferation and differentiation of rainbow trout adipocyte precursor cells. BMC Genomics 18:347
Boucher J, Softic S, El Ouaamari A et al (2016) Differential roles of insulin and IGF-1 receptors in adipose tissue development and function. Diabetes 65:2201–2213
Bour BA, O’Brien MA, Lockwood WL et al (1995) Drosophila MEF2, a transcription factor that is essential for myogenesis. Genes Dev 9:730–741
Bricard Y, Rallière C, Lebret V et al (2014) Early fish myoseptal cells: insights from the trout and relationships with amniote axial tenocytes. PLoS ONE 9:e91876
Bryant C, van Nek L, Rolland NCM (2020) European markets for cultured meat: a comparison of Germany and France. Foods 9. https://doi.org/10.3390/foods9091152
Bryantsev AL, Baker PW, Lovato TL et al (2012) Differential requirements for Myocyte Enhancer Factor-2 during adult myogenesis in Drosophila. Dev Biol 361:191–207
Bryson SP, Joyce EM, Martell DJ et al (2006) A cell line (HEW) from embryos of haddock (Melanogrammus aeglefinius) and its capacity to tolerate environmental extremes. Mar Biotechnol 8:641–653
Buonocore F, Libertini A, Prugnoli D et al (2006) Production and characterization of a continuous embryonic cell line from sea bass (Dicentrarchus labrax L.). Mar Biotechnol 8:80–85
Campuzano S, Mogilever NB, Pelling A (2020) Decellularized plant-based scaffolds for guided alignment of myoblast cells. bioRxiv 2020.02.23.958686. https://doi.org/10.1101/2020.02.23.958686
Carrera-Naipil C, Valenzuela-Muñoz V, Valdés JA et al (2016) RNA interference in Haliotis rufescens myostatin evidences upregulation of insulin signaling pathway. Agri Gene 1:93–99
Castillo J, Codina M, Martínez ML et al (2004) Metabolic and mitogenic effects of IGF-I and insulin on muscle cells of rainbow trout. Am J Physiol Regul Integr Comp Physiol 286:R935–R941
Castillo J, Le Bail P-Y, Paboeuf G et al (2002) IGF-I binding in primary culture of muscle cells of rainbow trout: changes during in vitro development. Am J Physiol Regul Integr Comp Physiol 283:R647–R652
Cawthorn WP, Bree AJ, Yao Y et al (2012) Wnt6, Wnt10a and Wnt10b inhibit adipogenesis and stimulate osteoblastogenesis through a β-catenin-dependent mechanism. Bone 50:477–489
Chal J, Al Tanoury Z, Hestin M et al (2016) Generation of human muscle fibers and satellite-like cells from human pluripotent stem cells in vitro. Nat Protoc 11:1833–1850
Chapelle S (1977) Lipid composition of tissues of marine crustaceans. Biochem Syst Ecol 5:241–248
Chaulet A, Medesani DA, Freitas J et al (2012) Induction of somatic growth in juvenile crayfish Cherax quadricarinatus (Decapoda, Parastacidae), by ecdysone and insulin growth factor. Aquaculture 370–371:1–6
Chen JW, Galloway JL (2014) The development of zebrafish tendon and ligament progenitors. Development 141:2035–2045
Chen S-L, Ren G-C, Sha Z-X, Hong Y (2005) Development and characterization of a continuous embryonic cell line from turbot (Scophthalmus maximus). Aquaculture 249:63–68
Chen S-L, Sha Z-X, Ye H-Q (2003) Establishment of a pluripotent embryonic cell line from sea perch (Lateolabrax japonicus) embryos. Aquaculture 218:141–151
Chen S-L, Ye H-Q, Sha Z-X, Hong Y (2003) Derivation of a pluripotent embryonic cell line from red sea bream blastulas. J Fish Biol 63:795–805
Chen YH, Lee WC, Liu CF, Tsai HJ (2001) Molecular structure, dynamic expression, and promoter analysis of zebrafish (Danio rerio) myf-5 gene. Genesis 29:22–35
Chisada S-I, Okamoto H, Taniguchi Y et al (2011) Myostatin-deficient medaka exhibit a double-muscling phenotype with hyperplasia and hypertrophy, which occur sequentially during post-hatch development. Dev Biol 359:82–94
Choi YH, Kim E-Y, Nam TJ (2018) Involvement of insulin-like growth factor in intraspecific variation in growth of Pacific oyster Crassostrea gigas during winter. Fish Sci 84:1017–1024
Choudhury D, Tseng TW, Swartz E (2020) The business of cultured meat. Trends Biotechnol 38:573–577
Christodoulides C, Lagathu C, Sethi JK, Vidal-Puig A (2009) Adipogenesis and WNT signalling. Trends Endocrinol Metab 20:16–24
Ciarlo CA, Zon LI (2016) Embryonic cell culture in zebrafish. Methods Cell Biol 133:1–10
Cole NJ, Hall TE, Martin CI et al (2004) Temperature and the expression of myogenic regulatory factors (MRFs) and myosin heavy chain isoforms during embryogenesis in the common carp Cyprinus carpio L. J Exp Biol 207:4239–4248
Collodi P, Kamei Y, Ernst T et al (1992) Culture of cells from zebrafish (Brachydanio rerio) embryo and adult tissues. Cell Biol Toxicol 8:43–61
Connon CJ, Gouveia RM (2021) Milliscale substrate curvature promotes myoblast self-organization and differentiation. Adv Biol 5:e2000280
Cosenza Z, Block DE, Baar K (2021) Optimization of muscle cell culture media using nonlinear design of experiments. Biotechnol J 16:e2100228. https://doi.org/10.1002/biot.202100228
Costa ML, Escaleira RC, Rodrigues VB et al (2002) Some distinctive features of zebrafish myogenesis based on unexpected distributions of the muscle cytoskeletal proteins actin, myosin, desmin, α-actinin, troponin and titin. Mech Dev 116:95–104
Coutelle O, Blagden CS, Hampson R et al (2001) Hedgehog signalling is required for maintenance of myf5 and myoD expression and timely terminal differentiation in zebrafish adaxial myogenesis. Dev Biol 236:136–150
Covi JA, Bader BD, Chang ES, Mykles DL (2010) Molt cycle regulation of protein synthesis in skeletal muscle of the blackback land crab, Gecarcinus lateralis, and the differential expression of a myostatin-like factor during atrophy induced by molting or unweighting. J Exp Biol 213:172–183
Cruz-Garcia L, Saera-Vila A, Navarro I et al (2009) Targets for TNFalpha-induced lipolysis in gilthead sea bream (Sparus aurata L.) adipocytes isolated from lean and fat juvenile fish. J Exp Biol 212:2254–2260
Dash C, Routray P, Tripathy S et al (2010) Derivation and characterization of embryonic stem-like cells of Indian major carp Catla catla. J Fish Biol 77:1096–1113
Datar I, Betti M (2010) Possibilities for an in vitro meat production system. Innov Food Sci Emerg Technol 11:13–22
de Almeida FLA, Carvalho RF, Pinhal D et al (2008) Differential expression of myogenic regulatory factor MyoD in pacu skeletal muscle (Piaractus mesopotamicus Holmberg 1887: Serrasalminae, Characidae, Teleostei) during juvenile and adult growth phases. Micron 39:1306–1311
Den Broeder MJ, Kopylova VA, Kamminga LM, Legler J (2015) Zebrafish as a model to study the role of peroxisome proliferating-activated receptors in adipogenesis and obesity. PPAR Res 2015:358029
De Santis C, Wade NM, Jerry DR et al (2011) Growing backwards: an inverted role for the shrimp ortholog of vertebrate myostatin and GDF11. J Exp Biol 214:2671–2677
Devoto SH, Melançon E, Eisen JS, Westerfield M (1996) Identification of separate slow and fast muscle precursor cells in vivo, prior to somite formation. Development 122:3371–3380
Devoto SH, Stoiber W, Hammond CL et al (2006) Generality of vertebrate developmental patterns: evidence for a dermomyotome in fish. Evol Dev 8:101–110
Dey BK, Gagan J, Dutta A (2011) miR-206 and -486 induce myoblast differentiation by downregulating Pax7. Mol Cell Biol 31:203–214
Dohmen RGJ, Hubalek S, Melke J et al (2022) Muscle-derived fibro-adipogenic progenitor cells for production of cultured bovine adipose tissue. NPJ Sci Food 6:6
Dominici M, Le Blanc K, Mueller I et al (2006) Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy 8:315–317
Dong M, Ye Y, Chen Z et al (2020) MicroRNA 182 is a novel negative regulator of adipogenesis by targeting CCAAT/Enhancer-binding protein α. Obesity 28:1467–1476
Driever W, Rangini Z (1993) Characterization of a cell line derived from zebrafish (Brachydanio rerio) embryos. In Vitro Cell Dev Biol Anim 29A:749–754
Du SJ, Devoto SH, Westerfield M, Moon RT (1997) Positive and negative regulation of muscle cell identity by members of the hedgehog and TGF-beta gene families. J Cell Biol 139:145–156
Du SJ, Dienhart M (2001) Gli2 mediation of Hedgehog signals in slow muscle induction in zebrafish. Differentiation 67:84–91
Duran BO da S, Dal-Pai-Silva M, Garcia de la Serrana D (2020) Rainbow trout slow myoblast cell culture as a model to study slow skeletal muscle, and the characterization of mir-133 and mir-499 families as a case study. J Exp Biol 223. https://doi.org/10.1242/jeb.216390
Dyachuk V (2013) Extracellular matrix is required for muscle differentiation in primary cell cultures of larval Mytilus trossulus (Mollusca: Bivalvia). Cytotechnology 65:725–735
Engler AJ, Griffin MA, Sen S et al (2004) Myotubes differentiate optimally on substrates with tissue-like stiffness: pathological implications for soft or stiff microenvironments. J Cell Biol 166:877–887
Engler AJ, Sen S, Sweeney HL, Discher DE (2006) Matrix elasticity directs stem cell lineage specification. Cell 126:677–689
Fan C, Zhang Y, Wang J, Cheng J (2018) Roles of Hedgehog signaling pathway in adipogenic differentiation potential of porcine adipose-derived mesenchymal stem cells. R Bras Zootec 47. https://doi.org/10.1590/rbz4720170019
Fan T-J, Wang X-F (2002) In vitro culture of embryonic cells from the shrimp, Penaeus chinensis J Exp Mar Biol Ecol 267:175–184
Fan Z, Liu L, Huang X et al (2017) Establishment and growth responses of Nile tilapia embryonic stem-like cell lines under feeder-free condition. Dev Growth Differ 59:83–93
Farnsworth DR, Saunders LM, Miller AC (2020) A single-cell transcriptome atlas for zebrafish development. Dev Biol 459:100–108
Fathi E, Farahzadi R, Sheikhzadeh N (2019) Immunophenotypic characterization, multi-lineage differentiation and aging of zebrafish heart and liver tissue-derived mesenchymal stem cells as a novel approach in stem cell-based therapy. Tissue Cell 57:15–21
Fauconneau B, Paboeuf G (2000) Effect of fasting and refeeding on in vitro muscle cell proliferation in rainbow trout (Oncorhynchus mykiss). Cell Tissue Res 301:459–463
Feng X, Adiarte EG, Devoto SH (2006) Hedgehog acts directly on the zebrafish dermomyotome to promote myogenic differentiation. Dev Biol 300:736–746
Ferrari L, Bragato C, Brioschi L et al (2019) HDAC8 regulates canonical Wnt pathway to promote differentiation in skeletal muscles. J Cell Physiol 234:6067–6076
Fish KD, Rubio NR, Stout AJ et al (2020) Prospects and challenges for cell-cultured fat as a novel food ingredient. Trends Food Sci Technol 98:53–67
Freeman FE, Kelly DJ (2017) Tuning alginate bioink stiffness and composition for controlled growth factor delivery and to spatially direct MSC fate within bioprinted tissues. Sci Rep 7:17042
Gabillard J-C, Biga PR, Rescan P-Y, Seiliez I (2013) Revisiting the paradigm of myostatin in vertebrates: insights from fishes. Gen Comp Endocrinol 194:45–54
Gabillard J-C, Sabin N, Paboeuf G (2010) In vitro characterization of proliferation and differentiation of trout satellite cells. Cell Tissue Res 342:471–477
Ganassi M, Badodi S, Ortuste Quiroga HP et al (2018) Myogenin promotes myocyte fusion to balance fibre number and size. Nat Commun 9:4232
Gao Y, Dai Z, Shi C et al (2016) Depletion of myostatin b promotes somatic growth and lipid metabolism in zebrafish. Front Endocrinol 7:88
Garikipati DK, Rodgers BD (2012) Myostatin stimulates myosatellite cell differentiation in a novel model system: evidence for gene subfunctionalization. Am J Physiol Regul Integr Comp Physiol 302:R1059–R1066
Garikipati DK, Rodgers BD (2012) Myostatin inhibits myosatellite cell proliferation and consequently activates differentiation: evidence for endocrine-regulated transcript processing. J Endocrinol 215:177–187
Garofalaki TF, Miniadis-Meimaroglou S, Sinanoglou VJ (2006) Main phospholipids and their fatty acid composition in muscle and cephalothorax of the edible Mediterranean crustacean Palinurus vulgaris (spiny lobster). Chem Phys Lipids 140:55–65
Garrison GL, Biermacher JT, Brorsen BW (2022) How much will large-scale production of cell-cultured meat cost? J Agric Food Res 10:100358
Genovese NJ, Schulze EN, Desmet DN (2021) Compositions and methods for increasing the efficiency of cell cultures used for food production (U.S. Patent No. 20210340570:A1). U.S. Patent and Trademark Office. https://patents.google.com/patent/US20210340570A1/en
Godard MP, Johnson BA, Richmond SR (2005) Body composition and hormonal adaptations associated with forskolin consumption in overweight and obese men. Obes Res 13:1335–1343
Godfray HCJ, Aveyard P, Garnett T et al (2018) Meat consumption, health, and the environment. Science 361. https://doi.org/10.1126/science.aam5324
Gosline JM, DeMont ME (1985) Jet-propelled swimming in squids. Sci Am 252:96–103
Goswami M, Belathur Shambhugowda Y, Sathiyanarayanan A et al (2022) Cellular aquaculture: prospects and challenges. Micromachines 13:828
Goswami M, Lakra WS, Yadav K, Jena JK (2012) Development of an ES-like cell culture system (RESC) from rohu, Labeo rohita (Ham.). Fish Physiol Biochem 38:1775–1783
Goswami M, Yashwanth BS, Trudeau V, Lakra WS (2022) Role and relevance of fish cell lines in advanced in vitro research. Mol Biol Rep 49:2393–2411
Griffiths-Jones S (2004) The microRNA registry. Nucleic Acids Res 32:D109–D111
Griffiths-Jones S, Grocock RJ, van Dongen S et al (2006) miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 34:D140–D144
Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ (2008) miRBase: tools for microRNA genomics. Nucleic Acids Res 36:D154–D158
Grimaldi A, Tettamanti G, Acquati F et al (2008) A hedgehog homolog is involved in muscle formation and organization of Sepia officinalis (mollusca) mantle. Dev Dyn 237:659–671
Gros J, Manceau M, Thomé V, Marcelle C (2005) A common somitic origin for embryonic muscle progenitors and satellite cells. Nature 435:954–958
Groves JA, Hammond CL, Hughes SM (2005) Fgf8 drives myogenic progression of a novel lateral fast muscle fibre population in zebrafish. Development 132:4211–4222
Guan L, Hu X, Liu L et al (2017) bta-miR-23a involves in adipogenesis of progenitor cells derived from fetal bovine skeletal muscle. Sci Rep 7:43716
Guo Y, Zhang X, Huang W, Miao X (2017) Identification and characterization of differentially expressed miRNAs in subcutaneous adipose between Wagyu and Holstein cattle. Sci Rep 7:44026
Gupta RK, Arany Z, Seale P et al (2010) Transcriptional control of preadipocyte determination by Zfp423. Nature 464:619–623
Gurevich DB, Nguyen PD, Siegel AL et al (2016) Asymmetric division of clonal muscle stem cells coordinates muscle regeneration in vivo. Science 353:aad9969
Gustafson B, Hammarstedt A, Hedjazifar S et al (2015) BMP4 and BMP antagonists regulate human white and beige adipogenesis. Diabetes 64:1670–1681
Guyon JR, Steffen LS, Howell MH et al (2007) Modeling human muscle disease in zebrafish. Biochim Biophys Acta 1772:205–215
Halpern BS, Maier J, Lahr HJ et al (2021) The long and narrow path for novel cell-based seafood to reduce fishing pressure for marine ecosystem recovery. Fish Fish 22:652–664
Hamade A, Deries M, Begemann G et al (2006) Retinoic acid activates myogenesis in vivo through Fgf8 signalling. Dev Biol 289:127–140
Hamilton BA (2020) ZNF423 orthologs are highly constrained in vertebrates but show domain-level plasticity across invertebrate lineages. bioRxiv 2020.03.09.984518. https://doi.org/10.1101/2020.03.09.984518
Hammarstedt A, Hedjazifar S, Jenndahl L et al (2013) WISP2 regulates preadipocyte commitment and PPARγ activation by BMP4. Proc Natl Acad Sci U S A 110:2563–2568
Hanga MP, Ali J, Moutsatsou P et al (2020) Bioprocess development for scalable production of cultivated meat. Biotechnol Bioeng 117:3029–3039
Hanyu Y, Kawashima I (2017) Growth induction system, growth induction control device, growth induction control method, and growth induction control program (Japanese Patent No. JP:6111510:B1). Japan Patent Office. https://patents.google.com/patent/JP6111510B1/en
Harms CA, Kennedy-Stoskopf S, Horne WA et al (2000) Cloning and sequencing hybrid striped bass (Morone saxatilis x M. chrysops) transforming growth factor-beta (TGF-beta), and development of a reverse transcription quantitative competitive polymerase chain reaction (RT-qcPCR) assay to measure TGF-beta mRNA of teleost fish. Fish Shellfish Immunol 10:61–85
Harzsch S, Kreissl S (2010) Myogenesis in the thoracic limbs of the American lobster. Arthropod Struct Dev 39:423–435
Hatta K, Bremiller R, Westerfield M, Kimmel CB (1991) Diversity of expression of engrailed-like antigens in zebrafish. Development 112:821–832
He S, Salas-Vidal E, Rueb S et al (2006) Genetic and transcriptome characterization of model zebrafish cell lines. Zebrafish 3:441–453
Hepler C, Vishvanath L, Gupta RK (2017) Sorting out adipocyte precursors and their role in physiology and disease. Genes Dev 31:127–140
Hesslein DGT, Fretz JA, Xi Y et al (2009) Ebf1-dependent control of the osteoblast and adipocyte lineages. Bone 44:537–546
Hilborn R, Banobi J, Hall SJ et al (2018) The environmental cost of animal source foods. Front Ecol Environ 16:329–335
Hinits Y, Hughes SM (2007) Mef2s are required for thick filament formation in nascent muscle fibres. Development 134:2511–2519
Hinits Y, Osborn DPS, Carvajal JJ et al (2007) Mrf4 (myf6) is dynamically expressed in differentiated zebrafish skeletal muscle. Gene Expr Patterns 7:738–745
Hinits Y, Osborn DPS, Hughes SM (2009) Differential requirements for myogenic regulatory factors distinguish medial and lateral somitic, cranial and fin muscle fibre populations. Development 136:403–414
Hinits Y, Williams VC, Sweetman D et al (2011) Defective cranial skeletal development, larval lethality and haploinsufficiency in Myod mutant zebrafish. Dev Biol 358:102–112
Hinman VF, Degnan BM (2002) Mox homeobox expression in muscle lineage of the gastropod Haliotis asinina: evidence for a conserved role in bilaterian myogenesis. Dev Genes Evol 212:141–144
Ho SY, Goh CWP, Gan JY et al (2014) Derivation and long-term culture of an embryonic stem cell-like line from zebrafish blastomeres under feeder-free condition. Zebrafish 11:407–420
Holen E, Kausland A, Skjærven K (2010) Embryonic stem cells isolated from Atlantic cod (Gadus morhua) and the developmental expression of a stage-specific transcription factor ac-Pou2. Fish Physiol Biochem 36:1029–1039
Hollway GE, Bryson-Richardson RJ, Berger S et al (2007) Whole-somite rotation generates muscle progenitor cell compartments in the developing zebrafish embryo. Dev Cell 12:207–219
Hong J-W, Park KW (2010) Further understanding of fat biology: lessons from a fat fly. Exp Mol Med 42:12–20
Hong Y, Winkler C, Schartl M (1996) Pluripotency and differentiation of embryonic stem cell lines from the medakafish (Oryzias latipes). Mech Dev 60:33–44
Hopkins PM, Chung ACK, Durica DS (1999) Limb Regeneration in the Fiddler Crab, Uca pugilator: Histological, Physiological and Molecular Considerations. Am Zool 39:513–526
Hopkins PM, Das S (2015) Regeneration in crustaceans. Nat Hist Crustacea 4:168–198
Hsu C-C, Lai C-Y, Lin C-Y et al (2018) MicroRNA-27b depletion enhances endotrophic and intravascular lipid accumulation and induces adipocyte hyperplasia in zebrafish. Int J Mol Sci 19. https://doi.org/10.3390/ijms19010093
Huang T-S, Todorcević M, Ruyter B, Torstensen BE (2010) Altered expression of CCAAT/enhancer binding protein and FABP11 genes during adipogenesis in vitro in Atlantic salmon (Salmo salar). Aquacult Nutr 16:72–80
Huang H, Lindgren A, Wu X et al (2012) High-throughput screening for bioactive molecules using primary cell culture of transgenic zebrafish embryos. Cell Rep 2:695–704
Huang Y, Das AK, Yang Q-Y et al (2012) Zfp423 promotes adipogenic differentiation of bovine stromal vascular cells. PLoS One 7:e47496
Huang J, Luo X, Huang M et al (2018) Identification and characteristics of muscle growth-related microRNA in the Pacific abalone. Haliotis Discus Hannai BMC Genomics 19:915
Huang J, Luo X, Zeng L et al (2018) Expression profiling of lncRNAs and mRNAs reveals regulation of muscle growth in the Pacific abalone. Haliotis Discus Hannai Sci Rep 8:16839
Humbird D (2021) Scale-up economics for cultured meat. Biotechnol Bioeng 118:3239–3250
Huminiecki L, Goldovsky L, Freilich S et al (2009) Emergence, development and diversification of the TGF-beta signalling pathway within the animal kingdom. BMC Evol Biol 9:28
Hutson MR, Zeng XL, Kim AJ et al (2010) Arterial pole progenitors interpret opposing FGF/BMP signals to proliferate or differentiate. Development 137:3001–3011
Imrie D, Sadler KC (2010) White adipose tissue development in zebrafish is regulated by both developmental time and fish size. Dev Dyn 239:3013–3023
Ishimura D, Yamamoto N, Tajima K et al (2008) Differentiation of adipose-derived stromal vascular fraction culture cells into chondrocytes using the method of cell sorting with a mesenchymal stem cell marker. Tohoku J Exp Med 216:149–156
Jackson HE, Ingham PW (2013) Control of muscle fibre-type diversity during embryonic development: the zebrafish paradigm. Mech Dev 130:447–457
Jang M, Scheffold J, Bruheim P (2022) Isolation and cultivation of primary muscle cells from Lobster (Homarus gammarus). In Vitro Cell Dev Biol - Animal 58:446–451
Jayesh P, Philip R, Singh ISB (2015) Multifactorial interaction of growth factors on Penaeus monodon lymphoid cells and the impact of IGFs in DNA synthesis and metabolic activity in vitro. Cytotechnology 67:559–571
Jiménez-Amilburu V, Salmerón C, Codina M et al (2013) Insulin-like growth factors effects on the expression of myogenic regulatory factors in gilthead sea bream muscle cells. Gen Comp Endocrinol 188:151–158
Jirikowski G, Kreissl S, Richter S, Wolff C (2010) Muscle development in the marbled crayfish—insights from an emerging model organism (Crustacea, Malacostraca, Decapoda). Dev Genes Evol 220:89–105
Joe AWB, Yi L, Natarajan A et al (2010) Muscle injury activates resident fibro/adipogenic progenitors that facilitate myogenesis. Nat Cell Biol 12:153–163
Johnston IA, Cole NJ, Abercromby M, Vieira VLA (1998) Embryonic temperature modulates muscle growth characteristics in larval and juvenile herring. J Exp Biol 201:623–646
Judson RN, Low M, Eisner C, Rossi FM (2017) Isolation, culture, and differentiation of Fibro/Adipogenic Progenitors (FAPs) from skeletal muscle. Methods Mol Biol 1668:93–103
Kakudo N, Shimotsuma A, Kusumoto K (2007) Fibroblast growth factor-2 stimulates adipogenic differentiation of human adipose-derived stem cells. Biochem Biophys Res Commun 359:239–244
Kanazawa A, Tsukada S, Kamiyama M et al (2005) Wnt5b partially inhibits canonical Wnt/beta-catenin signaling pathway and promotes adipogenesis in 3T3-L1 preadipocytes. Biochem Biophys Res Commun 330:505–510
Kaneko G, Shirakami H, Hirano Y et al (2016) Diversity of lipid distribution in fish skeletal muscle. Zoolog Sci 33:170–178
Karbiener M, Fischer C, Nowitsch S et al (2009) microRNA miR-27b impairs human adipocyte differentiation and targets PPARgamma. Biochem Biophys Res Commun 390:247–251
Keenan SR, Currie PD (2019) The developmental phases of zebrafish myogenesis. J Dev Biol 7. https://doi.org/10.3390/jdb7020012
Keller CA, Erickson MS, Abmayr SM (1997) Misexpression of nautilus Induces myogenesis in cardioblasts and alters the pattern of somatic muscle fibers. Dev Biol 181:197–212
Kier WM (2016) The musculature of coleoid cephalopod arms and tentacles. Front Cell Dev Biol 4:10
Kier WM, Schachat FH (1992) Biochemical comparison of fast- and slow-contracting squid muscle. J Exp Biol 168:41–56
Kim C-H, Neiswender H, Baik EJ et al (2008) β-Catenin interacts with MyoD and regulates its transcription activity. Mol Cell Biol 28:2941–2951
Kim E-Y, Choi YH (2019) Regulation of adductor muscle growth by the IGF-1/AKT pathway in the triploid Pacific oyster, Crassostrea gigas. Fish Aquat Sci 22:1–10
Kim S, Ahn C, Bong N et al (2015) Biphasic effects of FGF2 on adipogenesis. PLoS One 10:e0120073
Kinoshita H, Ohgane N, Fujino Y et al (2018) Functional roles of the Ripply-mediated suppression of segmentation gene expression at the anterior presomitic mesoderm in zebrafish. Mech Dev 152:21–31
Knappe S, Zammit PS, Knight RD (2015) A population of Pax7-expressing muscle progenitor cells show differential responses to muscle injury dependent on developmental stage and injury extent. Front Aging Neurosci 7:161
Konstantinides N, Averof M (2014) A common cellular basis for muscle regeneration in arthropods and vertebrates. Science 343:788–791
Korovina DG (2019) The use of bovine multipotent mesenchymal stem cells isolated from bone marrow and adipose tissue as sources to obtain muscle cells in vitro. IOP Conf Ser: Earth Environ Sci 315:042040
Koumans JT, Akster HA, Booms GH et al (1991) Numbers of myosatellite cells in white axial muscle of growing fish: Cyprinus carpio L. (Teleostei). Am J Anat 192:418–424
Kozomara A, Birgaoanu M, Griffiths-Jones S (2019) miRBase: from microRNA sequences to function. Nucleic Acids Res 47:D155–D162
Kozomara A, Griffiths-Jones S (2011) miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res 39:D152–D157
Kozomara A, Griffiths-Jones S (2014) miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res 42:D68-73
Kreissl S, Uber A, Harzsch S (2008) Muscle precursor cells in the developing limbs of two isopods (Crustacea, Peracarida): an immunohistochemical study using a novel monoclonal antibody against myosin heavy chain. Dev Genes Evol 218:253
Landemaine A, Rescan P-Y, Gabillard J-C (2014) Myomaker mediates fusion of fast myocytes in zebrafish embryos. Biochem Biophys Res Commun 451:480–484
Lee C-Y, Hu S-Y, Gong H-Y et al (2009) Suppression of myostatin with vector-based RNA interference causes a double-muscle effect in transgenic zebrafish. Biochem Biophys Res Commun 387:766–771
Lee K-Y, Loh H-X, Wan ACA (2022) Systems for muscle cell differentiation: from bioengineering to future food. Micromachines 13:71
Letcher SM, Rubio NR, Ashizawa RN et al (2022) In vitro insect fat cultivation for cellular agriculture applications. ACS Biomater Sci Eng. https://doi.org/10.1021/acsbiomaterials.2c00093
Li H, Li Q, Yu H (2018) Molecular characterization of the hedgehog signaling pathway and its necessary function on larval myogenesis in the pacific oyster Crassostrea gigas. Front Physiol 9:1536
Li S-N, Wu J-F (2020) TGF-β/SMAD signaling regulation of mesenchymal stem cells in adipocyte commitment. Stem Cell Res Ther 11:41
Li X, Fu X, Yang G, Du M (2020) Review: enhancing intramuscular fat development via targeting fibro-adipogenic progenitor cells in meat animals. Animal 14:312–321
Li Y, Fu H, Zhang F et al (2021) Identification, characterization, and expression profiles of insulin-like peptides suggest their critical roles in growth regulation of the Pacific oyster. Crassostrea Gigas Gene 769:145244
Lima LB, Oliveira FJM, Giacomini HC, Lima-Junior DP (2018) Expansion of aquaculture parks and the increasing risk of non-native species invasions in Brazil. Rev Aquacult 10:111–122
Listrat A, Lebret B, Louveau I et al (2016) How muscle structure and composition influence meat and flesh quality. Sci World J 2016:3182746
Liu J, Pan M, Huang D et al (2020) Myostatin-1 inhibits cell proliferation by inhibiting the mTOR signal pathway and MRFs, and activating the ubiquitin-proteasomal system in skeletal muscle cells of Japanese Flounder Paralichthys olivaceus. Cells 9:2376
Liu P, Tian J-J, Ji H et al (2018) The Wnt/β-catenin pathway contributes to the regulation of adipocyte development induced by docosahexaenoic acid in grass carp, Ctenopharyngodon idellus. Comp Biochem Physiol B Biochem Mol Biol 216:18–24
Löhle M, Hermann A, Glass H et al (2012) Differentiation efficiency of induced pluripotent stem cells depends on the number of reprogramming factors. Stem Cells 30:570–579
Lovato TL, Benjamin AR, Cripps RM (2005) Transcription of Myocyte enhancer factor-2 in adult Drosophila myoblasts is induced by the steroid hormone ecdysone. Dev Biol 288:612–621
Low M, Eisner C, Rossi F (2017) Fibro/Adipogenic Progenitors (FAPs): isolation by FACS and culture. Methods Mol Biol 1556:179–189
Lu T, Shen Y, Cui G-X et al (2020) Detailed analysis of lipids in edible viscera and muscles of cooked crabs Portunus trituberculatus and Portunus pelagicus. J Aquat Food Prod Technol 29:391–406
Lyons SE, Shue BC, Lei L et al (2001) Molecular cloning, genetic mapping, and expression analysis of four zebrafish c/ebp genes. Gene 281:43–51
MacLea KS, Abuhagr AM, Pitts NL et al (2012) Rheb, an activator of target of rapamycin, in the blackback land crab, Gecarcinus lateralis: cloning and effects of molting and unweighting on expression in skeletal muscle. J Exp Biol 215:590–604
Ma J, Zeng L, Lu Y (2017) Penaeid shrimp cell culture and its applications. Rev Aquacult 9:88–98
Ma RC, Jacobs CT, Sharma P et al (2018) Stereotypic generation of axial tenocytes from bipartite sclerotome domains in zebrafish. PLoS Genet 14:e1007775
MacLea KS, Covi JA, Kim H-W et al (2010) Myostatin from the American lobster, Homarus americanus: Cloning and effects of molting on expression in skeletal muscles. Comp Biochem Physiol A Mol Integr Physiol 157:328–337
Maroto M, Reshef R, Münsterberg AE et al (1997) Ectopic Pax-3 activates MyoD and Myf-5 expression in embryonic mesoderm and neural tissue. Cell 89:139–148
Marschallinger J, Obermayer A, Sänger AM et al (2009) Postembryonic fast muscle growth of teleost fish depends upon a nonuniformly distributed population of mitotically active Pax7+ precursor cells. Dev Dyn 238:2442–2448
Martinez I, Christiansen JS, Ofstad R, Olsen RL (1991) Comparison of myosin isoenzymes present in skeletal and cardiac muscles of the Arctic charr Salvelinus alpinus (L.). Sequential expression of different myosin heavy chains during development of the fast white skeletal muscle. Eur J Biochem 195:743–753
Mauro A (1961) Satellite cell of skeletal muscle fibers. J Biophys Biochem Cytol 9:493–495
Maves L, Waskiewicz AJ, Paul B et al (2007) Pbx homeodomain proteins direct Myod activity to promote fast-muscle differentiation. Development 134:3371–3382
Medler S, Mykles DL (2003) Analysis of myofibrillar proteins and transcripts in adult skeletal muscles of the American lobster Homarus americanus: variable expression of myosins, actin and troponins in fast, slow-twitch and slow-tonic fibres. J Exp Biol 206:3557–3567
Mehta F, Theunissen R, Post MJ (2019) Adipogenesis from Bovine Precursors. Methods Mol Biol 1889:111–125
Melzener L, Verzijden KE, Buijs AJ et al (2021) Cultured beef: from small biopsy to substantial quantity. J Sci Food Agric 101:7–14
Mennigen JA, Skiba-Cassy S, Panserat S (2013) Ontogenetic expression of metabolic genes and microRNAs in rainbow trout alevins during the transition from the endogenous to the exogenous feeding period. J Exp Biol 216:1597–1608
Messmer T, Klevernic I, Furquim C et al (2022) A serum-free media formulation for cultured meat production supports bovine satellite cell differentiation in the absence of serum starvation. Nat Food 3:74–85. https://doi.org/10.1038/s43016-021-00419-1
Millan-Cubillo AF, Martin-Perez M, Ibarz A et al (2019) Proteomic characterization of primary cultured myocytes in a fish model at different myogenesis stages. Sci Rep 9:14126
Mir BA, Reyer H, Komolka K et al (2020) Differentially expressed miRNA-Gene targets related to intramuscular fat in musculus Longissimus Dorsi of Charolais × Holstein F2-Crossbred Bulls. Genes 11. https://doi.org/10.3390/genes11060700
Moldes M, Zuo Y, Morrison RF et al (2003) Peroxisome-proliferator-activated receptor gamma suppresses Wnt/beta-catenin signalling during adipogenesis. Biochem J 376:607–613
Montserrat N, Capilla E, Navarro I, Gutiérrez J (2012) Metabolic effects of insulin and IGFs on Gilthead Sea Bream (Sparus aurata) muscle cells. Front Endocrinol 3:55
Montserrat N, Sánchez-Gurmaches J, García de la Serrana D et al (2007) IGF-I binding and receptor signal transduction in primary cell culture of muscle cells of gilthead sea bream: changes throughout in vitro development. Cell Tissue Res 330:503–513
Moore JW, Dionne C, Jaye M, Swain JL (1991) The mRNAs encoding acidic FGF, basic FGF and FGF receptor are coordinately downregulated during myogenic differentiation. Development 111:741–748
Morissette MR, Cook SA, Buranasombati C et al (2009) Myostatin inhibits IGF-I-induced myotube hypertrophy through Akt. Am J Physiol Cell Physiol 297:C1124–C1132
Mota de Sá P, Richard AJ, Hang H, Stephens JM (2017) Transcriptional regulation of adipogenesis. Compr Physiol 7:635–674
Mykles DL (1997) Crustacean muscle plasticity: molecular mechanisms determining mass and contractile properties. Comp Biochem Physiol B Biochem Mol Biol 117:367–378
Mykles DL, Medler S (2015) Skeletal muscle differentiation growth and plasticity. In: Chang E, Theil M (eds) Physiology. Oxford University Press, pp 134–167
Nahmias Y (2020) Systems and methods for growing cells in vitro (U.S. Patent No. 20200080050:A1). U.S. Patent and Trademark Office. https://patents.google.com/patent/US20200080050A1/en
Navet S, Bassaglia Y, Baratte S et al (2008) Somatic muscle development in Sepia officinalis (cephalopoda - mollusca): a new role for NK4. Dev Dyn 237:1944–1951
Negulescu PG, Risner D, Spang ES, et al (2022) Techno-Economic Modelling and Assessment of Cultivated Meat: Impact of Production Bioreactor Scale. engrXiv
Nguyen-Chi ME, Bryson-Richardson R, Sonntag C et al (2012) Morphogenesis and cell fate determination within the adaxial cell equivalence group of the zebrafish myotome. PLoS Genet 8:e1003014
Nie H, Zheng M, Wang Z et al (2021) Transcriptomic analysis provides insights into candidate genes and molecular pathways involved in growth of Manila clam Ruditapes philippinarum. Funct Integr Genomics 21:341–353
Nyika J, Mackolil J, Workie E et al (2021) Cellular agriculture research progress and prospects: insights from bibliometric analysis. Curr Res Biotechnol. https://doi.org/10.1016/j.crbiot.2021.07.001
O’Connor JD, Gilbert LI, Dr H et al (1968) Aspects of lipid metabolism in Crustaceans [with Discussion]. Am Zool 8:529–543
Odintsova NA, Dyachuk VA, Nezlin LP (2010) Muscle and neuronal differentiation in primary cell culture of larval Mytilus trossulus (Mollusca: Bivalvia). Cell Tissue Res 339:625–637
Okamura LH, Cordero P, Palomino J et al (2018) Myogenic differentiation potential of mesenchymal stem cells derived from fetal bovine bone marrow. Anim Biotechnol 29:1–11
Oku H, Umino T (2008) Molecular characterization of peroxisome proliferator-activated receptors (PPARs) and their gene expression in the differentiating adipocytes of red sea bream Pagrus major. Comp Biochem Physiol B Biochem Mol Biol 151:268–277
Olwin BB, Rapraeger A (1992) Repression of myogenic differentiation by aFGF, bFGF, and K-FGF is dependent on cellular heparan sulfate. J Cell Biol 118:631–639
O’Neill EN, Ansel JC, Kwong GA et al (2022) Spent media analysis suggests cultivated meat media will require species and cell type optimization. NPJ Sci Food 6:46
Ong KJ, Johnston J, Datar I et al (2021) Food safety considerations and research priorities for the cultured meat and seafood industry. Compr Rev Food Sci Food Saf 20:5421–5448. https://doi.org/10.1111/1541-4337.12853
Osborn DPS, Li K, Cutty SJ et al (2020) Fgf-driven Tbx protein activities directly induce myf5 and myod to initiate zebrafish myogenesis. Development 147:dev184689
Osborn DPS, Li K, Hinits Y, Hughes SM (2011) Cdkn1c drives muscle differentiation through a positive feedback loop with Myod. Dev Biol 350:464–475
Parameswaran V, Shukla R, Bhonde R, Hameed ASS (2007) Development of a pluripotent ES-like cell line from Asian sea bass (Lates calcarifer)–an oviparous stem cell line mimicking viviparous ES cells. Mar Biotechnol 9:766–775
Parker RWR, Blanchard JL, Gardner C et al (2018) Fuel use and greenhouse gas emissions of world fisheries. Nat Clim Chang 8:333–337
Patruno M, Gomiero C, Sacchetto R et al (2017) Tat-MyoD fused proteins, together with C2c12 conditioned medium, are able to induce equine adult mesenchimal stem cells towards the myogenic fate. Vet Res Commun 41:211–217
Pauly D, Zeller D (2016) Catch reconstructions reveal that global marine fisheries catches are higher than reported and declining. Nat Commun 7:10244
Peng L, Zhou Y, Xu W et al (2019) Generation of stable induced pluripotent stem-like cells from adult zebra fish fibroblasts. Int J Biol Sci 15:2340–2349
Post MJ (2014) An alternative animal protein source: cultured beef. Ann N Y Acad Sci 1328:29–33
Potter G, Smith AST, Vo NTK et al (2020) A more open approach is needed to develop cell-based fish technology: it starts with zebrafish. One Earth 3:54–64
Potthoff MJ, Olson EN (2007) MEF2: a central regulator of diverse developmental programs. Development 134:4131–4140
Potthoff MJ, Wu H, Arnold MA et al (2007) Histone deacetylase degradation and MEF2 activation promote the formation of slow-twitch myofibers. J Clin Invest 117:2459–2467
Pourquié O, Al Tanoury Z, Chal J (2018) The long road to making muscle in vitro. Curr Top Dev Biol 129:123–142
Price AL, Patel NH (2008) Investigating divergent mechanisms of mesoderm development in arthropods: the expression of Ph-twist and Ph-mef2 in Parhyale hawaiensis. J Exp Zool B Mol Dev Evol 310:24–40
Rakers S, Klinger M, Kruse C, Gebert M (2011) Pros and cons of fish skin cells in culture: long-term full skin and short-term scale cell culture from rainbow trout, Oncorhynchus mykiss. Eur J Cell Biol 90:1041–1051
Rao MS, Malik N (2012) Assessing iPSC reprogramming methods for their suitability in translational medicine. J Cell Biochem 113:3061–3068
Reis GG, Heidemann MS, Goes HAA, Molento CFM (2021) Can radical innovation mitigate environmental and animal welfare misconduct in global value chains? The case of cell-based tuna. Technol Forecast Soc Change 169:120845
Reiss J, Robertson S, Suzuki M (2021) Cell sources for cultivated meat: applications and considerations throughout the production workflow. Int J Mol Sci 22:7513
Relaix F, Rocancourt D, Mansouri A, Buckingham M (2005) A Pax3/Pax7-dependent population of skeletal muscle progenitor cells. Nature 435:948–953
Rescan PY, Gauvry L, Paboeuf G, Fauconneau B (1994) Identification of a muscle factor related to MyoD in a fish species. Biochim Biophys Acta 1218:202–204
Retamales A, Zuloaga R, Valenzuela CA et al (2015) Insulin-like growth factor-1 suppresses the Myostatin signaling pathway during myogenic differentiation. Biochem Biophys Res Commun 464:596–602
Riera-Heredia N, Lutfi E, Gutiérrez J et al (2019) Fatty acids from fish or vegetable oils promote the adipogenic fate of mesenchymal stem cells derived from gilthead sea bream bone potentially through different pathways. PLoS ONE 14:e0215926
Risner D, Li F, Fell JS et al (2021) Preliminary techno-economic assessment of animal cell-based meat. Foods 10:3
Rius-Francino M, Acerete L, Jiménez-Amilburu V et al (2011) Differential effects on proliferation of GH and IGFs in sea bream (Sparus aurata) cultured myocytes. Gen Comp Endocrinol 172:44–49
Romao JM, Jin W, Dodson MV et al (2011) MicroRNA regulation in mammalian adipogenesis. Exp Biol Med 236:997–1004
Rosen ED, Hsu C-H, Wang X et al (2002) C/EBPalpha induces adipogenesis through PPARgamma: a unified pathway. Genes Dev 16:22–26
Rosselló RA, Chen C-C, Dai R et al (2013) Mammalian genes induce partially reprogrammed pluripotent stem cells in non-mammalian vertebrate and invertebrate species. Elife 2:e00036
Ross SE, Hemati N, Longo KA et al (2000) Inhibition of adipogenesis by Wnt signaling. Science 289:950–953
Rost F, Eugster C, Schröter C et al (2014) Chevron formation of the zebrafish muscle segments. J Exp Biol 217:3870–3882
Rowlerson A, Radaelli G, Mascarello F, Veggetti A (1997) Regeneration of skeletal muscle in two teleost fish: Sparus aurata and Brachydanio rerio. Cell Tissue Res 289:311–322
Roy S, Wolff C, Ingham PW (2001) The u-boot mutation identifies a Hedgehog-regulated myogenic switch for fiber-type diversification in the zebrafish embryo. Genes Dev 15:1563–1576
Rubio N, Datar I, Stachura D et al (2019) Cell-based fish: a novel approach to seafood production and an opportunity for cellular agriculture. Front Sustain Food Syst 3:43
Rubio NR, McCartney NE, Trimmer BA, Kaplan DL (2020) Biofabrication with insect cells. Trends in Entomology 16:1–17
Rubio NR, Fish KD, Trimmer BA, Kaplan DL (2019) In vitro insect muscle for tissue engineering applications. ACS Biomater Sci Eng 5:1071–1082
Rubio NR, Fish KD, Trimmer BA, Kaplan DL (2019) Possibilities for engineered insect tissue as a food source. Frontiers in Sustainable Food Systems 3:24
Saad MK, Yuen JSK, Joyce CM et al (2022) Continuous fish muscle cell line with capacity for myogenic and adipogenic-like phenotypes. bioRxiv 2022.08.22.504874. https://doi.org/10.1101/2022.08.22.504874
Saera-Vila A, Kish PE, Kahana A (2016) Fgf regulates dedifferentiation during skeletal muscle regeneration in adult zebrafish. Cell Signal 28:1196–1204
Salmerón C (2018) Adipogenesis in fish. J Exp Biol 221:jeb161588
Salmerón C, Acerete L, Gutiérrez J et al (2013) Characterization and endocrine regulation of proliferation and differentiation of primary cultured preadipocytes from gilthead sea bream (Sparus aurata). Domest Anim Endocrinol 45:1–10
Salmerón C, Riera-Heredia N, Gutiérrez J et al (2016) Adipogenic gene expression in gilthead sea bream mesenchymal stem cells from different origin. Front Endocrinol 7:113
Sano T, Fnoguchi S, Tsuchiya T, Matsumoto JJ (1986) Contribution of paramyosin to marine meat gel characteristics. J Food Sci 51:946–950
Schnapp E, Pistocchi AS, Karampetsou E et al (2009) Induced early expression of mrf4 but not myog rescues myogenesis in the myod/myf5 double-morphant zebrafish embryo. J Cell Sci 122:481–488
Seger C, Hargrave M, Wang X et al (2011) Analysis of Pax7 expressing myogenic cells in zebrafish muscle development, injury, and models of disease. Dev Dyn 240:2440–2451
Seiliez I, Sabin N, Gabillard J-C (2012) Myostatin inhibits proliferation but not differentiation of trout myoblasts. Mol Cell Endocrinol 351:220–226
Shao M, Hepler C, Vishvanath L et al (2017) Fetal development of subcutaneous white adipose tissue is dependent on Zfp423. Mol Metab 6:111–124
Shi J, Cai M, Si Y et al (2018) Knockout of myomaker results in defective myoblast fusion, reduced muscle growth and increased adipocyte infiltration in zebrafish skeletal muscle. Hum Mol Genet 27:3542–3554
Shu-Chien AC, Han W-Y, Carter CG et al (2017) Effect of dietary lipid source on expression of lipid metabolism genes and tissue lipid profile in juvenile spiny lobster Sagmariasus verreauxi. Aquaculture 479:342–351
Siegel AL, Gurevich DB, Currie PD (2013) A myogenic precursor cell that could contribute to regeneration in zebrafish and its similarity to the satellite cell. FEBS J 280:4074–4088
Speir ML, Bhaduri A, Markov NS et al (2021) UCSC cell browser: visualize your single-cell data. Bioinformatics. https://doi.org/10.1093/bioinformatics/btab503
Steinbacher P, Haslett JR, Sänger AM, Stoiber W (2006) Evolution of myogenesis in fish: a sturgeon view of the mechanisms of muscle development. Anat Embryol 211:311–322
Stern-Straeter J, Bonaterra GA, Juritz S et al (2014) Evaluation of the effects of different culture media on the myogenic differentiation potential of adipose tissue- or bone marrow-derived human mesenchymal stem cells. Int J Mol Med 33:160–170
Suh JM, Gao X, McKay J et al (2006) Hedgehog signaling plays a conserved role in inhibiting fat formation. Cell Metab 3:25–34
Sun X, Li L, Wu B et al (2021) Cell type diversity in scallop adductor muscles revealed by single-cell RNA-Seq. Genomics 113:3582–3598
Tabata Y, Miyao M, Inamoto T et al (2000) De novo formation of adipose tissue by controlled release of basic fibroblast growth factor. Tissue Eng 6:279–289
Tan X, Du SJ (2002) Differential expression of two MyoD genes in fast and slow muscles of gilthead seabream (Sparus aurata). Dev Genes Evol 212:207–217
Tanaka A, Woltjen K, Miyake K et al (2013) Efficient and reproducible myogenic differentiation from human iPS cells: prospects for modeling Miyoshi Myopathy in vitro. PLoS ONE 8:e61540
Taylor MV (2006) Comparison of muscle development in drosophila and vertebrates. In: Sink H (ed) Muscle development in drosophila. Springer, Austin, pp 169–203
Terova G, Rimoldi S, Bernardini G, Saroglia M (2013) Inhibition of myostatin gene expression in skeletal muscle of fish by in vivo electrically mediated dsRNA and shRNAi delivery. Mol Biotechnol 54:673–684
Teshima S-I, Kanazawa A, Kakuta Y (1986) Effects of dietary phospholipids on lipid transport in the juvenile prawn. Bull Jpn Soc Sci Fish 52:159–163
Thangaraj RS, Narendrakumar L, Prasannan Geetha P et al (2021) Comprehensive update on inventory of finfish cell lines developed during the last decade (2010–2020). Rev Aquac 13:2248–2288
Ticho BS, Stainier DY, Fishman MC, Breitbart RE (1996) Three zebrafish MEF2 genes delineate somitic and cardiac muscle development in wild-type and mutant embryos. Mech Dev 59:205–218
Tlili S, Yin J, Rupprecht J-F et al (2019) Shaping the zebrafish myotome by intertissue friction and active stress. Proc Natl Acad Sci U S A 116:25430–25439
Todorcević M, Skugor S, Krasnov A, Ruyter B (2010) Gene expression profiles in Atlantic salmon adipose-derived stromo-vascular fraction during differentiation into adipocytes. BMC Genomics 11:39
Toledo TM (2019) Effect of dietary lipid on the molecular and metabolic profile of a freshwater crayfish. PhD, Queensland University of Technology. https://eprints.qut.edu.au/134470/
Torres-Velarde J, Llera-Herrera R, Ibarra-Castro L et al (2020) Post-transcriptional silencing of myostatin-1 in the spotted rose snapper (Lutjanus guttatus) promotes muscle hypertrophy. Mol Biol Rep 47:443–450
Tsuruwaka Y, Shimada E (2022) Reprocessing seafood waste: challenge to develop aquatic clean meat from fish cells. NPJ Sci Food 6:7
Uezumi A, Fukada S-I, Yamamoto N et al (2010) Mesenchymal progenitors distinct from satellite cells contribute to ectopic fat cell formation in skeletal muscle. Nat Cell Biol 12:143–152
Uezumi A, Ito T, Morikawa D et al (2011) Fibrosis and adipogenesis originate from a common mesenchymal progenitor in skeletal muscle. J Cell Sci 124:3654–3664
van Eelen WF, Van Kooten WJ, Westerhof W (1999) Industrial scale production of meat from in vitro cell cultures (World Patent No. 1999031222:A1). World Intellectual Property Organization. https://patents.google.com/patent/WO1999031222A1/en
van Tienen FHJ, Laeremans H, van der Kallen CJH, Smeets HJM (2009) Wnt5b stimulates adipogenesis by activating PPARgamma, and inhibiting the beta-catenin dependent Wnt signaling pathway together with Wnt5a. Biochem Biophys Res Commun 387:207–211
Veggetti A, Mascarello F, Scapolo PA et al (1993) Muscle growth and myosin isoform transitions during development of a small teleost fish, Poecilia reticulata (Peters) (Atheriniformes, Poeciliidae): a histochemical, immunohistochemical, ultrastructural and morphometric study. Anat Embryol 187:353–361
Vegusdal A, Sundvold H, Gjøen T, Ruyter B (2003) An in vitro method for studying the proliferation and differentiation of Atlantic salmon preadipocytes. Lipids 38:289–296
Venkatesan M, Semper C, Skrivergaard S et al (2022) Recombinant production of growth factors for application in cell culture. iScience 25:105054. https://doi.org/10.1016/j.isci.2022.105054
Vergeer R, Sinke P, Odegard I (2021) TEA of cultivated meat - Future projections of different scenarios. CE Delft. https://www.cedelft.eu/en/publications/2609/tea-of-cultivated-meat-future-projections-of-different-scenarios. Accessed 23 Apr 2021
Villasante A, Powell MS, Moutou K et al (2016) Effects of anthocyanidins on myogenic differentiation and antioxidant defense in primary myogenic cells isolated from rainbow trout (Oncorhynchus mykiss). Aquaculture 454:81–89
Wafer R, Tandon P, Minchin JEN (2017) The Role of Peroxisome Proliferator-Activated Receptor Gamma (PPARG) in Adipogenesis: Applying Knowledge from the Fish Aquaculture Industry to Biomedical Research. Front Endocrinol 8:102
Wakamatsu Y, Ozato K, Sasado T (1994) Establishment of a pluripotent cell line derived from a medaka (Oryzias latipes) blastula embryo. Mol Mar Biol Biotechnol 3:185–191
Wang J, Li J, Ge Q, Li J (2021) A potential negative regulation of myostatin in muscle growth during the intermolt stage in Exopalaemon carinicauda. Gen Comp Endocrinol 314:113902
Wang P-Y, Li W-T, Yu J, Tsai W-B (2012) Modulation of osteogenic, adipogenic and myogenic differentiation of mesenchymal stem cells by submicron grooved topography. J Mater Sci Mater Med 23:3015–3028
Wang X, Huang M, Wang Y (2012) The effect of insulin, TNFα and DHA on the proliferation, differentiation and lipolysis of preadipocytes isolated from large yellow croaker (Pseudosciaena crocea R.). PLoS One 7:e48069
Wang Y-H, Li C-K, Lee G-H et al (2008) Inactivation of zebrafish mrf4 leads to myofibril misalignment and motor axon growth disorganization. Dev Dyn 237:1043–1050
Watanabe S, Hirai H, Asakura Y et al (2011) MyoD gene suppression by Oct4 is required for reprogramming in myoblasts to produce induced pluripotent stem cells. Stem Cells 29:505–516
Watts JEM, Schreier HJ, Lanska L, Hale MS (2017) The rising tide of antimicrobial resistance in aquaculture: sources. Sinks and Solutions Mar Drugs 15:158
Wei J, Glaves RS, Sellars MJ et al (2016) Expression of the prospective mesoderm genes twist, snail, and mef2 in penaeid shrimp. Dev Genes Evol 226:317–324
Wei Q, Rong Y, Paterson BM (2007) Stereotypic founder cell patterning and embryonic muscle formation in Drosophila require nautilus (MyoD) gene function. Proc Natl Acad Sci U S A 104:5461–5466
Weinberg ES, Allende ML, Kelly CS et al (1996) Developmental regulation of zebrafish MyoD in wild-type, no tail and spadetail embryos. Development 122:271–280
Weintraub H (1993) The MyoD family and myogenesis: redundancy, networks, and thresholds. Cell 75:1241–1244
West JA, Park I-H, Daley GQ, Geijsen N (2006) In vitro generation of germ cells from murine embryonic stem cells. Nat Protoc 1:2026–2036
White RB, Lamey TM, Ziman M, Koenders A (2005) Isolation and expression analysis of a Pax group III gene from the crustacean Cherax destructor. Dev Genes Evol 215:306–312
Windner SE, Doris RA, Ferguson CM et al (2015) Tbx6, Mesp-b and Ripply1 regulate the onset of skeletal myogenesis in zebrafish. Development 142:1159–1168
Witt R, Weigand A, Boos AM et al (2017) Mesenchymal stem cells and myoblast differentiation under HGF and IGF-1 stimulation for 3D skeletal muscle tissue engineering. BMC Cell Biol 18:15
Wolff C, Roy S, Ingham PW (2003) Multiple muscle cell identities induced by distinct levels and timing of hedgehog activity in the zebrafish embryo. Curr Biol 13:1169–1181
Wu P, Yong P, Zhang Z et al (2022) Loss of myomixer results in defective myoblast fusion, impaired muscle growth, and severe myopathy in zebrafish. Mar Biotechnol. https://doi.org/10.1007/s10126-022-10159-3
Wynne R, Archer LC, Hutton SA et al (2021) Alternative migratory tactics in brown trout (Salmo trutta) are underpinned by divergent regulation of metabolic but not neurological genes. Ecol Evol 11:8347–8362
Xu C, Tabebordbar M, Iovino S et al (2013) A zebrafish embryo culture system defines factors that promote vertebrate myogenesis across species. Cell 155:909–921
Xu W, Li H, Peng L et al (2022) Fish pluripotent stem-like cell line induced by small-molecule compounds from caudal fin and its developmental potentiality. Front Cell Dev Biol 9:817779
Xu Y, He J, Wang X et al (2000) Asynchronous activation of 10 muscle-specific protein (MSP) genes during zebrafish somitogenesis. Dev Dyn 219:201–215
Xu Y, Li Z, Li X et al (2015) Regulating myogenic differentiation of mesenchymal stem cells using thermosensitive hydrogels. Acta Biomater 26:23–33
Yan Y, Lu X, Kong J et al (2020) Molecular characterization of myostatin and its inhibitory function on myogenesis and muscle growth in Chinese Shrimp, Fenneropenaeus chinensis. Gene 758:144986
Yao Z, Farr GH 3rd, Tapscott SJ, Maves L (2013) Pbx and Prdm1a transcription factors differentially regulate subsets of the fast skeletal muscle program in zebrafish. Biol Open 2:546–555
Yin J, Lee R, Ono Y et al (2018) Spatiotemporal coordination of FGF and Shh signaling underlies the specification of myoblasts in the zebrafish embryo. Dev Cell 46:735-750.e4
Yoshino TP, Bickham U, Bayne CJ (2013) Molluscan cells in culture: primary cell cultures and cell lines. Can J Zool 91.https://doi.org/10.1139/cjz-2012-0258
Ytteborg E, Todorcevic M, Krasnov A et al (2015) Precursor cells from Atlantic salmon (Salmo salar) visceral fat holds the plasticity to differentiate into the osteogenic lineage. Biol Open 4:783–791
Yu T, Chua CK, Tay CY et al (2013) A generic micropatterning platform to direct human mesenchymal stem cells from different origins towards myogenic differentiation. Macromol Biosci 13:799–807
Yuan Y, Hong Y (2017) Medaka insulin-like growth factor-2 supports self-renewal of the embryonic stem cell line and blastomeres in vitro. Sci Rep 7:78
Yun Y-R, Won JE, Jeon E et al (2010) Fibroblast growth factors: biology, function, and application for tissue regeneration. J Tissue Eng 2010:218142
Zhang JM, Chen L, Krause M et al (1999) Evolutionary conservation of MyoD function and differential utilization of E proteins. Dev Biol 208:465–472
Zhang M, Li D-H, Li F et al (2018) Integrated analysis of MiRNA and genes associated with meat quality reveals that Gga-MiR-140-5p affects intramuscular fat deposition in chickens. Cell Physiol Biochem 46:2421–2433
Zhang W, Roy S (2017) Myomaker is required for the fusion of fast-twitch myocytes in the zebrafish embryo. Dev Biol 423:24–33
Zhang Y, Tan X, Zhang P-J, Xu Y (2006) Characterization of muscle-regulatory gene, MyoD, from flounder (Paralichthys olivaceus) and analysis of its expression patterns during embryogenesis. Mar Biotechnol 8:139–148
Zhao J, Wen X, Li S et al (2015) Effects of dietary lipid levels on growth, feed utilization, body composition and antioxidants of juvenile mud crab Scylla paramamosain (Estampador). Aquaculture 435:200–206
Zheng B, Cao B, Li G, Huard J (2006) Mouse adipose-derived stem cells undergo multilineage differentiation in vitro but primarily osteogenic and chondrogenic differentiation in vivo. Tissue Eng 12:1891–1901
Zhu X, Fu L, Yi F et al (2014) Regeneration: making muscle from hPSCs. Cell Res 24:1159–1161
Zuk PA, Zhu M, Ashjian P et al (2002) Human adipose tissue is a source of multipotent stem cells. Mol Biol Cell 13:4279–4295
Zuk PA, Zhu M, Mizuno H et al (2001) Multilineage cells from human adipose tissue: implications for cell-based therapies. Tissue Eng 7:211–228
Zullo L, Fossati SM, Imperadore P, Nödl M-T (2017) Molecular determinants of cephalopod muscles and their implication in muscle regeneration. Front Cell Dev Biol 5:53
Acknowledgements
The authors would like to thank Dr. Catherine Walsh, Dr. Mahmoudreza Ovissipour, Dr. Alexandrea A. Duscher, Dr. Mukunda Goswami, Dr. Simone Costa, Ms. Subha Subramanian, Dr. Serene Chng, and Dr. Ka Yi Ling for providing helpful and constructive feedback on the manuscript. All figures were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
Conceptualization: CB, EAS. Visualization: CB, DMCM. Writing—original draft (including literature review): CB, LM, DMCM, GFF. Writing—review and editing: CB, LM, DMCM, GFF, FCF, EAS.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bomkamp, C., Musgrove, L., Marques, D.M.C. et al. Differentiation and Maturation of Muscle and Fat Cells in Cultivated Seafood: Lessons from Developmental Biology. Mar Biotechnol 25, 1–29 (2023). https://doi.org/10.1007/s10126-022-10174-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-022-10174-4