Abstract
Purpose
Hyaluronic acid plays an essential role in water retention of the intervertebral disc (IVD) and thus provides flexibility and shock absorbance in the spine. Hyaluronic acid gets degraded by hyaluronidases (HYALs), and some of the resulting fragments were previously shown to induce an inflammatory and catabolic response in human IVD cells. However, no data currently exist on the expression and activity of HYALs in IVD health and disease.
Methods
Gene expression, protein expression and activity of HYALs were determined in human IVD biopsies with different degrees of degeneration (n = 50 total). Furthermore, freshly isolated human IVD cells (n = 23 total) were stimulated with IL-1β, TNF-α or H2O2, followed by analysis of HYAL-1, HYAL-2 and HYAL-3 gene expression.
Results
Gene expression of HYAL-1 and protein expression of HYAL-2 significantly increased in moderate/severe disc samples when compared to samples with no or low IVD degeneration. HYAL activity was not significantly increased due to high donor–donor variation, but seemed overall higher in the moderate/severe group. An inflammatory environment, as seen during IVD disease, did not affect HYAL-1, HYAL-2 or HYAL-3 expression, whereas exposure to oxidative stress (100 µM H2O2) upregulated HYAL-2 expression relative to untreated controls.
Conclusion
Although HYAL-1, HYAL-2 and HYAL-3 are all expressed in the IVD, HYAL-2 seems to have the highest pathophysiological relevance. Nonetheless, further studies will be needed to comprehensively elucidate its significance and to determine its potential as a therapeutic target.
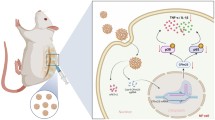
Graphic abstract
These slides can be retrieved under Electronic Supplementary Material.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glycosaminoglycans (GAGs), which are major components of the extracellular matrix (ECM) of numerous connective tissues, including the intervertebral disc (IVD), are a family of negatively charged heteropolysaccharides. Chondroitin sulfate and keratan sulfate as the IVD-typical GAGs bind to an extended protein core, resulting in the brush-like proteoglycan (PG) named aggrecan. The G1 domain at the N-terminal end of aggrecan interacts with hyaluronic acid (HA) and link protein and thus provides the hydrated gel structure seen in healthy IVDs, specifically in the nucleus pulposus (NP) [1, 2]. Hyaluronic acid (HA) itself is composed of repeated disaccharide units of d-glucuronic acid and N-acetyl-d-glucosamine with a molecular weight of 103–104 kDa (high molecular weight, HMW-HA) [3, 4].
Ample evidence has been provided over the past decades for the degradation of aggrecan with IVD degeneration. In fact, enzymatic action by matrix metalloproteinases (MMPs) and aggrecanases can target the interglobular domain (between the G1 and G2 domains), with MMP degradation typically happening earlier than aggrecanase degradation [5, 6]. Interestingly, mechanical overloading as a contributor to degeneration was shown to enhance MMP-associated cleavage of aggrecan [7]. Furthermore, changes in spatial distribution and disaccharide sulphation patterns of GAGs are commonly observed during IVD aging and degeneration [8,9,10].
HA degradation can occur via prokaryotic enzymes, including bacterial β-endoglycosidases and bacterial β-exoglycosidases [11]. However, in the context of ECM maintenance and degeneration in human health and disease, the eukaryotic HA-degrading endo-β-n-acetylhexosaminidases—also termed hyaluronidases (HYALs)—are of highest relevance, albeit other eukaryotic enzymes (e.g., β-glucuronidase) also possess HA degrading activity [11]. HYALs preferentially degrade HA through cleavage at the β-(1,4)-linkage [12], but can also affect the integrity of chondroitin sulfate at reduced activity [13].
Six HYAL-like gene sequences have been identified in humans so far, sharing about 40% of their identity with one another: HYAL-1, HYAL-2, HYAL-3, HYAL-4, PH20 (= SPAM1) as well as a pseudo gene HYAL-Phyal1 that is transcribed in humans, but is not translated [13]. Of these, HYAL-1 and HYAL-2 have been investigated in most detail. While both of these enzymes are located on chromosome 3p21.3 [14], they show differences with regard to substrates, location within the cellular compartment and cleavage products [11, 15].
The process of HA cleavage (Fig. 1) is initiated by HYAL-2, a cell surface glycosylphosphatidylinositol-anchored protein that acts in cooperation with CD44 [16, 17], by degrading HMW-HA into intermediate sized fragments of approximately 10–20 kDa [14]. These fragments are partially taken up by the cell upon interaction with surface HA receptors (endocytosis) [14, 15] and partially released into the extracellular space [17]. Upon endocytosis, HYAL-1 further degrades these fragments into oligosaccharides (predominantly tetrasaccharides) within endosomal-lysosomal structures in the cell. Fragments are then released by exocytosis or further cleaved by the lysosomal enzymes β-d-glucuronidase and β-N-acetyl-d-hexosaminidase into individual sugars [13, 15, 18]. Aside from enzymatic reactions, in vivo degradation of HA can also occur through reactive oxygen species (ROS), as described in detail by Stern et al. [11].
Mechanism of HA degradation. a HA is cleaved by HYALs at hexosaminidic β (1–4) linkages between glucuronic acid and N-acetylglucosamine. b Membrane-anchored HYAL-2 cleaves HMW HA to small MW HA, which is then internalized into lysozomes and further cleaved to oligosaccharides by HYAL-1. In the IVD, oligosaccharides have been shown to exhibit biological (e.g., pro-inflammatory) activity. In the IVD, the expression of HYAL-2 is increased with oxidative stress as well as with degeneration (but in a subset of samples), yet not in a simulated inflammatory environment
Larger HYAL-2-induced fragments (oligomers) and smaller HYAL-1-induced fragments (e.g., tetramers) have both been described to possess biological activity, thereby affecting a variety of cellular processes in a cell-type and size-specific manner [19,20,21,22,23]. More specifically, various types of HA fragments were shown to be able to promote inflammation in, e.g., immune cells (10–400 kDa) [19] (35kDA) [24], chondrocytes (6-mer oligosaccharides) [25] and also IVD cells (6–12 disaccharides) [26] (Fig. 1), although not all studies show an effect (4 mer as well as range between 4 and 200 kDa) [27, 28]. Interestingly, inflammatory conditions have been described to further induce the expression and activity of HYALs in certain cell and tissue types [29, 30], although it is currently unclear whether cells in degenerated IVDs (which are characterized by inflammation [31, 32]) retain a similar feedback mechanism.
Aside from inflammation, the degenerating IVD possesses various other microenvironmental characteristics that might affect HYAL metabolism, including reduced pH [33, 34] and enhanced oxidative stress [35]. It has previously been shown that the activity of HYALs is pH-dependent. While human HYAL-1 as a lysosomal enzyme is most active around pH 4.0 [36], the pH optimum for the membrane-associated HYAL-2 is around 6.0 [17]. Such a reduced pH is typically achieved through the activation of the Na+/H+ exchanger 1 (NHE1) upon interaction with CD44 [16]. As degeneration-associated accumulation of lactic acid in IVDs leads to a drop in pH as low as 5.7 in severely affected tissues [33, 34], the pH microenvironment of the IVD may further modulate HYAL-2 activity. Concomitant with a degeneration-associated reduction in pH, enhanced generation of ROS has been described in the IVD [35, 37]. While ROS have previously been shown to induce posttranslational oxidative modification of collagens such as crosslinking and unfolding [38, 39], their role in modulating the HYAL metabolism have not yet been investigated. However, bronchial epithelial cells demonstrated increased HYAL-2 expression and activity when exposed to ROS [40] and similar response patterns may also exist for IVD cells.
Despite the functional importance of HA (and PGs) in the IVD, the role and regulation of HYALs in IVD health and disease are currently unknown. Therefore, the aim of this study was to identify the expression and activity of HYAL-1, HYAL-2 and HYAL-3 in human IVD tissue with different degrees of degeneration and to test whether their expression is regulated under inflammatory or oxidative stress conditions in vitro.
Materials and methods
Human IVD tissue
The collection of human IVD tissue from patients undergoing spinal surgery was approved by the Cantonal ethics Committee Zurich, Switzerland (#EK-16/05) as well as by the Ethics Committee of the Charité Berlin, Germany (#EA2/087/11). Informed consent was obtained from all patients.
Samples used for direct analysis of HYAL expression/activity (n = 34, mixture of NP and AF) were immediately cooled after intraoperative excision and thereafter shock frozen at − 80 °C. Using preoperative MRIs, the degree of IVD degeneration in these samples was determined according to Pfirrmann et al. [41], using an adopted 4-grade classification scale as previously described [42, 43]. Specifically, discs were graded as non-degenerated (grade 1), mildly degenerated (grade 2), moderately degenerated (grade 3) and severely degenerated (grade 4) (Table 1).
Samples used for cell isolation (Pfirmann grade 3–4, disc herniation or DDD) were transferred into DMEM/F12 (D8437, Sigma-Aldrich, USA) with 3% anti–anti (15240062, Gibco, USA) upon intraoperative excision and immediately transported to the laboratory for further processing (n = 15). Due to the degeneration status of the specimens as well as the posterior surgical approach used in most cases, no separation of nucleus pulposus and annulus fibrosus was performed.
Primary cell culture
Tissue samples of mixed degeneration grades were cut into pieces and incubated in a sterile solution of 0.2% collagenase NB4 (17454, Serva, Heidelberg, Germany) and 0.3% dispase II (04942078001, Roche, Basel, Switzerland) in phosphate-buffered saline (PBS) for 4–8 h. Then, the cell suspension was filtered using a 70-μm cell strainer (352350, BD Bioscience, Switzerland) and expanded up to passage 3 in a 2D monolayer culture containing DMEM/F12 media with 10% FCS (F7524, Sigma-Aldrich, USA) and 1% anti–anti, with medium changes twice a week.
Cell stimulation with IL-1β or TNF-α
For cytokine stimulation experiments, IVD cells (n = 5) were seeded into 6-well plates at a density of 33,000 cells/cm2. After 24 h, cells were rendered serum free (DMEM/F12 + 1% anti–anti) for 2 h. To determine the best cytokine stimulatory response, cells were incubated with 0.1, 1, 5 or 10 ng/mL recombinant IL-1β (211-11, Peprotech/LuBioScience, Switzerland) or 0.1, 1, 10 or 100 ng/mL recombinant TNF-α (315-01A, Peprotech/ LuBioScience, Switzerland) in serum-free medium for 18 h. All concentrations were controlled for cytotoxicity (data not shown). In addition, a time course experiment with stimulation times of 2, 6 and 18 h for 5 ng/ml IL-1β or 10 ng/ml TNF-α was conducted.
Cell stimulation with H2O2
For H2O2 stimulation experiments, IVD cells (n = 5) were seeded at a density of only 11,500 cells/cm2 into T25 flasks or 6-well plates and allowed to settle and adapt to culturing conditions for further 48 h prior to the oxidant treatment. Thereafter, cells were rendered serum free for 2 h and consequently exposed for an additional 2 h to H2O2 (H1009, Sigma-Aldrich, Switzerland) at concentrations of 25, 50 and 100 μM to ascertain the strongest physiological ROS effect on the cells. After the treatment, the oxidative medium was immediately replaced by fresh DMEM/F12 supplemented with 10% FCS and 1% anti–anti and cells allowed to recover from oxidative stress for 24 h [44]. All concentrations were controlled for cytotoxicity (data not shown).
HYAL gene expression in IVD tissue and treated IVD cells
To isolate RNA from IVD biopsies with normal/mild (n = 8) and moderate/severe (n = 16) degeneration, shock-frozen tissues were pulverized in liquid nitrogen using a custom-made device, immersed in Trizol (15596-018, Invitrogen, USA), exposed to phase separation with chloroform and subsequently subjected to column-based purification (12183025, PureLink RNA Mini Kit, ThermoFisher, Switzerland) as previously described [45]. To isolate RNA from cultured cells (n = 5/treatment), the PureLink RNA Mini Kit or the Trizol/choloroform method was used.
RNA from tissues or cells was reverse transcribed to cDNA using the Reverse Transcription Reagents (4374966, ThermoFisher, Switzerland), and real-time PCR analysis was performed by TaqMan Gene Expression assays (ThermoFisher, Switzerland), using primers/probes for HYAL-1, HYAL-2 and HYAL-3 (Table 2). Additionally, HO-1 mRNA expression was measured to verify induction of oxidative stress in H2O2 treated cells. TBP was used as the housekeeping gene for all experiments except for H2O2 treatments, in which GAPDH was more stably expressed than TBP. Gene expression in the tissue was normalized to the housekeeping gene and shown as 2−dCt values. Gene expression of stimulated cells was normalized to the housekeeping gene and to the untreated control and shown as 2−ddCt.
HYAL protein expression
To isolate protein from IVD biopsies with normal/mild (n = 5) and moderate/severe (n = 5) degeneration, samples were pulverized as described above and then immersed in PBS supplemented with 100x protease inhibitors (78425, Pierce, USA) with three freeze-thaw cycles at − 72 °C on dry ice, as recommended by the ELISA’s manufacturer.
Total protein concentration was determined by Bradford assay as described by the manufacturer (500-0006, Bio-Rad, Switzerland). Protein expression of HYAL-1 (tissue/cells), HYAL-2 (tissue/cells) and HYAL-3 (tissue) was detected on total protein samples (20–40 μg for tissue, 10–20 μg for cells, 100 μl for supernatants) by ELISA (MBS703230, My BioSource, USA; E1126 h, Lubio Science, Switzerland). Results are expressed as ng HYAL per mg of total protein in the IVD tissue or as the ng per ml in cultured cells, based on the sum of cell lysates and supernatants.
Total HYAL activity
Total HYAL activity in IVD biopsies with normal/mild (n = 5) and moderate/severe (n = 5) degeneration was analyzed with a commercial HYAL activity kit (Ra003-01-HAK, Amsbio, UK), according to the manufacturers instruction. Briefly, defrosted and washed tissue biopsies were immersed in base buffer defined by the manufacturer (10–50 mg tissue/ml) and then homogenized by Polytron mixer. The homogenized sample were centrifuged, the supernatant collected and total protein assessed by Bradford assay. The specific HYAL activity was determined on 50 μl supernatant and expressed as ng HA removed per minute and mg total protein applied.
Statistical analysis
Comparisons between treatment groups and control group were conducted using unpaired t test if data were normally distributed, or Mann–Whitney test if data were not normally distributed. Comparison between different groups was analyzed using one-way ANOVA for normally distributed data, with the Tukey’s multiple comparisons test. All analyses were done using the Graph Pad Prism statistical program, with a significance level of p < 0.05.
Results
The expression of hyaluronidases in normal and degenerated discs
Gene expression of HYAL-1, HYAL-2 and HYAL-3 in human disc tissue was analyzed in relation to the degeneration grade (grades 1 and 2 = normal/mild, grade 3–5 = moderate/severe, as defined by Pfirrmann et al. [41] (Fig. 2). The expression of HYAL-1 significantly increased in moderate/severe disc samples when compared to the normal/mild group (p = 0.0098). The expression of HYAL-2 was not significantly affected by degeneration (p = 0.4167), although selected donors showed exceptionally high expression levels. Gene expression of HYAL-3 was not significantly different between the grades (p = 0.834). Protein expression of HYAL-1 and HYAL-2 in human disc tissue was analyzed as well (Fig. 3a, b). HYAL-2 protein significantly increased in moderate/severe disc samples, when compared to the normal/mild samples (p = 0.0412), while HYAL-1 protein was unchanged (p = 0.769). Overall HYAL activity tended to increase in the moderate/severe group (p = 0.1886) (Fig. 3c).
Gene expression of HYALs in IVD tissue. Disc tissues were collected from patients undergoing elective spinal surgeries. a Gene expression of HYAL-1 was upregulated in moderate/severe disc degeneration group, when compared with the normal/mild group. b Gene expression of HYAL-2 was not changed; however, some degenerated samples showed high gene expression. c Gene expression of HYAL-3 was unchanged. Results were calculated by 2−ΔCt method relative to TBP and analyzed by Mann–Whitney U test due to non-normality of data (n = 8 in normal/mild group, n = 16 in moderate/severe group)
Protein expression and activity of HYALs in IVD tissue. Disc tissues were collected from patients undergoing elective spinal surgeries. a Protein expression of HYAL-1 was not significantly different between the normal/mild disc degeneration group and the moderate/severe group. b Protein expression of HYAL-2 was upregulated in moderate/severe samples. c Total activity of HYALs tended to increase in the moderate/severe group. Results were calculated as ng of HYAL present (a, b) or removed (c) per mg of total protein and analyzed by unpaired t test due to normality of data (n = 5 in each group)
The effects of inflammation and oxidative stress on the expression of hyaluronidases in disc cells
As shown above, HYAL expression/activity was partially upregulated in discs with higher degeneration grade. Therefore, the influence of main hallmarks of DDD (inflammation and oxidative stress) on the expression of HYALs was tested next. Gene expression of HYAL-1, HYAL-2 and HYAL-3 was analyzed in primary human disc cells treated with increasing concentrations of pro-inflammatory cytokines IL-1β (0.1–10 ng/mL) (Fig. 4a–c) and TNF-α (0.1–100 ng/mL) (Fig. 4d–f). Pro-inflammatory cytokines did not influence gene expression of HYAL-1, HYAL-2 and HYAL-3. Gene expression of HYAL-1, HYAL-2 and HYAL-3 was also tested in primary human disc cells treated with increasing concentrations of oxidative stress inducer H2O2 (25–100 µM) (Fig. 5a–c). H2O2 at 100 µM dose-dependently upregulated gene expression of HYAL-2 (p = 0.0445 to control) and HYAL-3 (p < 0.05–25 and 50 µM), while the expression of HYAL-1 only tended to increase (p = 0.1276).
Gene expression of HYALs in primary IVD cells treated with pro-inflammatory cytokines. Primary cell cultures were prepared from disc tissues collected during spinal surgeries and treated with increasing concentration of IL-1β and TNF-α for 18 h. The effects of IL-1β (0.1–10 ng/mL) on gene expression of a HYAL1, b HYAL2 and c HYAL3. The effects of TNF-α (0.1–100 ng/mL) on gene expression of d HYAL1, e HYAL2 and f HYAL3. Results were calculated by 2−ΔΔCt method relative to the untreated control, with TBP as housekeeping gene and analyzed by one-way ANOVA with Tukey post hoc test due to normality of data (n = 5)
Gene expression of HYALs in primary disc cells treated with reactive oxygen species. Primary cell cultures were prepared from disc tissues collected during spinal surgeries and treated with increasing concentration of H2O2 for 2 h. The effects of H2O2 (25–100 µM) on gene expression of a HYAL-1, b HYAL-2 and c HYAL-3. Results were calculated by 2−ΔΔCt method relative to the untreated control, with GAPDH as housekeeping gene and analyzed by one-way ANOVA with Tukey post hoc test due to normality of data (n = 5)
Discussion
This study showed for the first time that HYAL-1, HYAL-2 and HYAL-3, the three major HA-degrading enzymes, are expressed in IVD issue and isolated cells. Interestingly, HYAL-1 expression was increased with increasing degeneration, although this was only observed on the mRNA level, but not on the protein level. This may possibly be explained by negative feedback loops related to HA metabolism and/or alternative splicing. It has been shown that under certain conditions (e.g., hypoxia), the HYAL-1 mRNA can undergo alternative splicing (previously identified, e.g., in kidney cells and tumors), generating multiple mRNA species, only one of which is translated into protein [13, 46, 47]. While HYAL-2 gene expression was not significantly increased with degeneration, several severely degenerated samples demonstrated explicitly high HYAL-2 mRNA levels that may have contributed to the significant increase in the protein level with progressing degeneration, as shown in Fig. 3. However, the hyaluronidase activity of HYAL-2 remains controversial and has been described as limited, especially compared to HYAL-1 [18], possibly explaining why its significant protein enhancement with degeneration did not coincide with a significant (but merely tendential) increase in HYAL activity. The majority of our control samples came from patients with traumatic IVD injuries or vertebral fractures. Although our tissue donors are usually operated soon after the traumatic incident, we cannot exclude the possibility that the expression and/or activity of HYALs were altered in some control samples. Nevertheless, our data grouped as traumatic IVDs vs. other samples did not show any pattern that would indicate an effect of the trauma itself.
It should furthermore be noted that the high data variability within the two degeneration groups for all targets indicates that HYAL expression and activity is not only affected by the grade of degeneration, but also by other factors that were not controlled for due to the small sample size. Possible examples include, but are not limited to, diabetic conditions [48], P. acnes infection [49] and tissue loading patterns [50, 51]. Higher number of samples will be needed to better identify trends and statistically significant differences in future studies. Furthermore, future studies would ideally investigate NP and AF cells separately, although our own preliminary data on bovine NP and AF cells indicate marginal zonal differences in HYAL expression.
Increased HYAL expression and/or activity have been described in numerous pathologies, such as during asthma (within the epithelium) [30], cartilage hypertrophy [52] or osteoarthritis and rheumatoid arthritis (within the knee synovium/synoviocytes) [53, 54]. HYAL deficiency or knockdown/knockout have also been described to promote certain pathologies, leading, e.g., to swelling of the periarticular masses [55] as well as to progression of osteoarthritis [56, 57]. These studies demonstrate that HYALs seem to play a crucial role in cartilage homeostasis, which not only shows similar composition [58], but also comparable degenerative processes and pathophysiological conditions [59] as the IVD. As in other tissues, HA in the IVD is cleaved by HYALs at hexosaminidic β (1–4) linkages between glucuronic acid and N-acetylglucosamine. Membrane-anchored HYAL-2 cleaves HMW HA to small MW HA, which is then internalized into lysozomes and further cleaved by HYAL-1 to oligosaccharides, which possess biological (e.g., pro-inflammatory) activity (Fig. 1).
Importantly, pathological degeneration of the IVD and cartilage are both characterized by increased level of inflammatory mediators [31, 32, 60,61,62] and reactive oxygen species (ROS) [35, 63, 64] within the tissue. Our goal was thus to analyze whether HYAL expression is regulated by exposure to pro-inflammatory cytokines or ROS, as would occur during DDD. However, we could not observe any significant changes in HYAL-1, HYAL-2 or HYAL-3 expression upon stimulation with IL-1β or TNF-α, independent of the concentration (Fig. 4) or the analysis time point (data not shown). In contrast, articular cartilage chondrocytes were previously shown to respond to IL-1β and TNF-α, treatment with an upregulation of HYAL-1, HYAL-2 and HYAL-3 gene expression [29] and similar response patterns were observed in airway epithelial cells [30].
Although we did not observe an upregulation of HYALs upon IL-1β or TNF-α stimulation, IVD cells responded to higher concentrations of H2O2 (100 μM) with increased expression of HYAL-2 and HYAL-3 (Figs. 1, 5). Feng et al. [65] summarized that various disc cells derived from different species show mitochondrion-dependent ROS production and that ROS can induce p38, ERKs, JNKs, p65 and Akt in IVD cells, thus leading to increased expression of matrix degrading enzymes and pro-inflammatory cytokines. In this study, we used H2O2 for cell stimulation as H2O2 has previously been identified in human NP tissues, hence representing a physiological type of ROS [66]. Similar to IVD cells, exposure to ROS (by xanthine oxidase) increased HYAL-2 expression and activity via the p38MAPK signaling pathway in human bronchial epithelial cells [40].
Overall, our and other studies demonstrate that ROS is not only involved in HA metabolism via direct cleavage of HA [11], but also via induction of HYAL expression, which in turn will then contribute to HA degradation. This notion is supported by the fact that exposure of airway cells to ROS leads to sustained HA degradation that lasts for at least 24 h [40]. Furthermore, GAG degradation by HYALs is also associated with increased expression of various MMPs [67], a group of enzymes that is known for its crucial role in ECM degradation in various tissues, including the IVD [68, 69].
In conclusion, our data not only show expression of HYAL-1, HYAL-2 and HYAL-3 in the IVD, but furthermore points to HYAL-2 as the most relevant HYAL as it increased during degeneration and further upregulated by ROS, a hallmark of DDD. Further studies will be needed to confirm the relevance of HYAL-2 in IVD health and disease and to determine its potential as a therapeutic target.
References
Sivan SS, Wachtel E, Roughley P (2014) Structure, function, aging and turnover of aggrecan in the intervertebral disc. Biochim Biophys Acta 1840:3181–3189. https://doi.org/10.1016/j.bbagen.2014.07.013
Roughley PJ (2004) Biology of intervertebral disc aging and degeneration: involvement of the extracellular matrix. Spine (Phila Pa 1976) 29:2691–2699
Lipson SJ, Muir H (1981) Proteoglycans in experimental intervertebral-disk degeneration. Spine 6:194–210. https://doi.org/10.1097/00007632-198105000-00002
Bushell GR, Ghosh P, Taylor TF, Akeson WH (1977) Proteoglycan chemistry of the intervertebral disks. Clin Orthop Relat Res 129:115–123
Sztrolovics R, Alini M, Roughley PJ, Mort JS (1997) Aggrecan degradation in human intervertebral disc and articular cartilage. Biochem J 326(Pt 1):235–241. https://doi.org/10.1042/bj3260235
Patel KP, Sandy JD, Akeda K, Miyamoto K, Chujo T, An HS, Masuda K (2007) Aggrecanases and aggrecanase-generated fragments in the human intervertebral disc at early and advanced stages of disc degeneration. Spine 32:2596–2603. https://doi.org/10.1097/BRS.0b013e318158cb85
Iatridis JC, Godburn K, Wuertz K, Alini M, Roughley PJ (2011) Region-dependent aggrecan degradation patterns in the rat intervertebral disc are affected by mechanical loading in vivo. Spine 36:203–209. https://doi.org/10.1097/BRS.0b013e3181cec247
Iatridis JC, MacLean JJ, O’Brien M, Stokes IA (2007) Measurements of proteoglycan and water content distribution in human lumbar intervertebral discs. Spine (Phila Pa 1976) 32:1493–1497. https://doi.org/10.1097/brs.0b013e318067dd3f
Antoniou J, Steffen T, Nelson F, Winterbottom N, Hollander AP, Poole RA, Aebi M, Alini M (1996) The human lumbar intervertebral disc: evidence for changes in the biosynthesis and denaturation of the extracellular matrix with growth, maturation, ageing, and degeneration. J Clin Invest 98:996–1003. https://doi.org/10.1172/JCI118884
Liu X, Krishnamoorthy D, Lin L, Xue P, Zhang F, Chi L, Linhardt RJ, Iatridis JC (2018) A method for characterising human intervertebral disc glycosaminoglycan disaccharides using liquid chromatography-mass spectrometry with multiple reaction monitoring. Eur Cells Mater 35:117–131. https://doi.org/10.22203/eCM.v035a09
Stern R, Kogan G, Jedrzejas MJ, Soltes L (2007) The many ways to cleave hyaluronan. Biotechnol Adv 25:537–557. https://doi.org/10.1016/j.biotechadv.2007.07.001
Wang W, Wang J, Li F (2017) Hyaluronidase and chondroitinase. Adv Exp Med Biol 925:75–87. https://doi.org/10.1007/5584_2016_54
Stern R, Jedrzejas MJ (2006) Hyaluronidases: their genomics, structures, and mechanisms of action. Chem Rev 106:818–839. https://doi.org/10.1021/cr050247k
Girish KS, Kemparaju K (2007) The magic glue hyaluronan and its eraser hyaluronidase: a biological overview. Life Sci 80:1921–1943. https://doi.org/10.1016/j.lfs.2007.02.037
Racine R, Mummert ME (2012) Hyaluronan endocytosis: mechanisms of uptake and biological functions. Mol Regul Endocytosis 1:377–390. https://doi.org/10.5772/45976
Bourguignon LY, Singleton PA, Diedrich F, Stern R, Gilad E (2004) CD44 interaction with Na+–H+ exchanger (NHE1) creates acidic microenvironments leading to hyaluronidase-2 and cathepsin B activation and breast tumor cell invasion. J Biol Chem 279:26991–27007. https://doi.org/10.1074/jbc.M311838200
Harada H, Takahashi M (2007) CD44-dependent intracellular and extracellular catabolism of hyaluronic acid by hyaluronidase-1 and -2. J Biol Chem 282:5597–5607. https://doi.org/10.1074/jbc.M608358200
Stern R (2003) Devising a pathway for hyaluronan catabolism: are we there yet? Glycobiology 13:105r–115r. https://doi.org/10.1093/glycob/cwg112
de la Motte C, Nigro J, Vasanji A, Rho H, Kessler S, Bandyopadhyay S, Danese S, Fiocchi C, Stern R (2009) Platelet-derived hyaluronidase 2 cleaves hyaluronan into fragments that trigger monocyte-mediated production of proinflammatory cytokines. Am J Pathol 174:2254–2264. https://doi.org/10.2353/ajpath.2009.080831
West DC, Hampson IN, Arnold F, Kumar S (1985) Angiogenesis induced by degradation products of hyaluronic-acid. Science 228:1324–1326. https://doi.org/10.1126/science.2408340
West DC, Kumar S (1989) The effect of hyaluronate and its oligosaccharides on endothelial-cell proliferation and monolayer integrity. Exp Cell Res 183:179–196. https://doi.org/10.1016/0014-4827(89)90428-X
Taylor KR, Trowbridge JM, Rudisill JA, Termeer CC, Simon JC, Gallo RL (2004) Hyaluronan fragments stimulate endothelial recognition of injury through TLR4. J Biol Chem 279:17079–17084. https://doi.org/10.1074/jbc.M310859200
Stern R, Asari AA, Sugahara KN (2006) Hyaluronan fragments: an information-rich system. Eur J Cell Biol 85:699–715. https://doi.org/10.1016/j.ejcb.2006.05.009
McKee CM, Penno MB, Cowman M, Burdick MD, Strieter RM, Bao C, Noble PW (1996) Hyaluronan (HA) fragments induce chemokine gene expression in alveolar macrophages: the role of HA size and CD44. J Clin Investig 98:2403–2413. https://doi.org/10.1172/Jci119054
Campo GM, Avenoso A, Campo S, D’Ascola A, Nastasi G, Calatroni A (2010) Small hyaluronan oligosaccharides induce inflammation by engaging both toll-like-4 and CD44 receptors in human chondrocytes. Biochem Pharmacol 80:480–490. https://doi.org/10.1016/j.bcp.2010.04.024
Quero L, Klawitter M, Schmaus A, Rothley M, Sleeman J, Tiaden AN, Klasen J, Boos N, Hottiger MO, Wuertz K, Richards PJ (2013) Hyaluronic acid fragments enhance the inflammatory and catabolic response in human intervertebral disc cells through modulation of toll-like receptor 2 signalling pathways. Arthritis Res Ther 15:R94. https://doi.org/10.1186/ar4274
Cowman MK, Shortt C, Arora S, Fu Y, Villavieja J, Rathore J, Huang X, Rakshit T, Jung GI, Kirsch T (2019) Role of hyaluronan in inflammatory effects on human articular chondrocytes. Inflammation. https://doi.org/10.1007/s10753-019-01043-9
Olsson M, Bremer L, Aulin C, Harris HE (2018) Fragmented hyaluronan has no alarmin function assessed in arthritis synovial fibroblast and chondrocyte cultures. Innate Immun Lond 24:131–141. https://doi.org/10.1177/1753425918759590
Flannery CR, Little CB, Hughes CE, Caterson B (1998) Expression and activity of articular cartilage hyaluronidases. Biochem Biophys Res Commun 251:824–829. https://doi.org/10.1006/bbrc.1998.9561
Monzon ME, Manzanares D, Schmid N, Casalino-Matsuda SM, Forteza RM (2008) Hyaluronidase expression and activity is regulated by pro-inflammatory cytokines in human airway epithelial cells. Am J Resp Cell Mol 39:289–295. https://doi.org/10.1165/rcmb.2007-0361OC
Johnson ZI, Schoepflin ZR, Choi H, Shapiro IM, Risbud MV (2015) Disc in flames: roles of TNF-alpha and IL-1beta in intervertebral disc degeneration. Eur Cell Mater 30:104–116 discussion 107–116
Wuertz K, Haglund L (2013) Inflammatory mediators in intervertebral disk degeneration and discogenic pain. Global Spine J 3:175–184. https://doi.org/10.1055/s-0033-1347299
Diamant B, Karlsson J, Nachemson A (1968) Correlation between lactate levels and ph in discs of patients with lumbar rhizopathies. Experientia 24:1195. https://doi.org/10.1007/bf02146615
Kitano T, Zerwekh JE, Usui Y, Edwards ML, Flicker PL, Mooney V (1993) Biochemical-changes associated with the symptomatic human intervertebral-disk. Clin Orthop Relat R 293:372–377
Feng C, Yang M, Lan M, Liu C, Zhang Y, Huang B, Liu H, Zhou Y (2017) ROS: crucial intermediators in the pathogenesis of intervertebral disc degeneration. Oxid Med Cell Longev 2017:5601593. https://doi.org/10.1155/2017/5601593
Zhang L, Bharadwaj AG, Casper A, Barkley J, Barycki JJ, Simpson MA (2009) Hyaluronidase activity of human hyal1 requires active site acidic and tyrosine residues. J Biol Chem 284:9433–9442. https://doi.org/10.1074/jbc.M900210200
Poveda L, Hottiger M, Boos N, Wuertz K (2009) Peroxynitrite induces gene expression in intervertebral disc cells. Spine 34:1127–1133. https://doi.org/10.1097/BRS.0b013e31819f2330
Elgawish A, Glomb M, Friedlander M, Monnier VM (1996) Involvement of hydrogen peroxide in collagen cross-linking by high glucose in vitro and in vivo. J Biol Chem 271:12964–12971
Scharf B, Clement CC, Yodmuang S, Urbanska AM, Suadicani SO, Aphkhazava D, Thi MM, Perino G, Hardin JA, Cobelli N, Vunjak-Novakovic G, Santambrogio L (2013) Age-related carbonylation of fibrocartilage structural proteins drives tissue degenerative modification. Chem Biol 20:922–934. https://doi.org/10.1016/j.chembiol.2013.06.006
Monzon ME, Fregien N, Schmid N, Falcon NS, Campos M, Casalino-Matsuda SM, Forteza RM (2010) Reactive oxygen species and hyaluronidase 2 regulate airway epithelial hyaluronan fragmentation. J Biol Chem 285:26126–26134. https://doi.org/10.1074/jbc.M110.135194
Pfirrmann CW, Metzdorf A, Zanetti M, Hodler J, Boos N (2001) Magnetic resonance classification of lumbar intervertebral disc degeneration. Spine (Phila Pa 1976) 26:1873–1878
Tiaden AN, Klawitter M, Lux V, Mirsaidi A, Bahrenberg G, Glanz S, Quero L, Liebscher T, Wuertz K, Ehrmann M, Richards PJ (2012) Detrimental role for human high temperature requirement serine protease A1 (HTRA1) in the pathogenesis of intervertebral disc (IVD) degeneration. J Biol Chem 287:21335–21345. https://doi.org/10.1074/jbc.M112.341032
Klawitter M, Hakozaki M, Kobayashi H, Krupkova O, Quero L, Ospelt C, Gay S, Hausmann O, Liebscher T, Meier U, Sekiguchi M, Konno S, Boos N, Ferguson SJ, Wuertz K (2014) Expression and regulation of toll-like receptors (TLRs) in human intervertebral disc cells. Eur Spine J 23:1878–1891. https://doi.org/10.1007/s00586-014-3442-4
Krupkova O, Handa J, Hlavna M, Klasen J, Ospelt C, Ferguson SJ, Wuertz-Kozak K (2016) The natural polyphenol epigallocatechin gallate protects intervertebral disc cells from oxidative stress. Oxid Med Cell Longev 2016:7031397. https://doi.org/10.1155/2016/7031397
Wuertz K, Godburn K, MacLean JJ, Barbir A, Donnelly JS, Roughley PJ, Alini M, Iatridis JC (2009) In vivo remodeling of intervertebral discs in response to short- and long-term dynamic compression. J Orthop Res 27:1235–1242. https://doi.org/10.1002/jor.20867
Chao KL, Muthukumar L, Herzberg O (2007) Structure of human hyaluronidase-1, a hyaluronan hydrolyzing enzyme involved in tumor growth and angiogenesis. Biochemistry 46:6911–6920. https://doi.org/10.1021/bi700382g
Junker N, Latini S, Petersen LN, Kristjansen PE (2003) Expression and regulation patterns of hyaluronidases in small cell lung cancer and glioma lines. Oncol Rep 10:609–616
Dogne S, Rath G, Jouret F, Caron N, Dessy C, Flamion B (2016) Hyaluronidase 1 deficiency preserves endothelial function and glycocalyx integrity in early streptozotocin-induced diabetes. Diabetes 65:2742–2753. https://doi.org/10.2337/db15-1662
Slaby O, McDowell A, Bruggemann H, Raz A, Demir-Deviren S, Freemont T, Lambert P, Capoor MN (2018) Is IL-1beta further evidence for the role of propionibacterium acnes in degenerative disc disease? Lessons from the study of the inflammatory skin condition acne vulgaris. Front Cell Infect Microbiol 8:272. https://doi.org/10.3389/fcimb.2018.00272
Kitamura R, Tanimoto K, Tanne Y, Kamiya T, Huang YC, Tanaka N, Tanaka E, Tanne K (2010) Effects of mechanical load on the expression and activity of hyaluronidase in cultured synovial membrane cells. J Biomed Mater Res A 92a:87–93. https://doi.org/10.1002/jbm.a.32345
Tanimoto K, Kitamura R, Tanne Y, Kamiya T, Kunimatsu R, Yoshioka M, Tanaka N, Tanaka E, Tanne K (2010) Modulation of hyaluronan catabolism in chondrocytes by mechanical stimuli. J Biomed Mater Res A 93:373–380. https://doi.org/10.1002/jbm.a.32540
Tanimoto K, Suzuki A, Ohno S, Honda K, Tanaka N, Doi T, Nakahara-Ohno M, Yoneno K, Nakatani Y, Ueki M, Yanagida T, Kitamura R, Tanne K (2004) Hyaluronidase expression in cultured growth plate chondrocytes during differentiation. Cell Tissue Res 318:335–342. https://doi.org/10.1007/s00441-004-0966-7
Yoshida M, Sai S, Marumo K, Tanaka T, Itano N, Kimata K, Fujii K (2004) Expression analysis of three isoforms of hyaluronan synthase and hyaluronidase in the synovium of knees in osteoarthritis and rheumatoid arthritis by quantitative real-time reverse transcriptase polymerase chain reaction. Arthritis Res Ther 6:R514–R520. https://doi.org/10.1186/ar1223
Uzuki M, Sasaki Y, Tokunaga S, Kamataki A, Nomi K, Kitagawa H, Kaiyama J, Sawai T (2008) Activity and expression of hyaluronidases associated with hyaluronan synthases expression and change of molecular weight of hyaluronan in the joint fluid. In: OARSI world congress on osteoarthritis, osteoathritis and cartilage
Natowicz MR, Short MP, Wang Y, Dickersin GR, Gebhardt MC, Rosenthal DI, Sims KB, Rosenberg AE (1996) Clinical and biochemical manifestations of hyaluronidase deficiency. New Engl J Med 335:1029–1033. https://doi.org/10.1056/Nejm199610033351405
Martin DC, Atmuri V, Hemming RJ, Farley J, Mort JS, Byers S, Hombach-Klonisch S, Stern R, Triggs-Raine BL (2008) A mouse model of human mucopolysaccharidosis IX exhibits osteoarthritis. Hum Mol Genet 17:1904–1915. https://doi.org/10.1093/hmg/ddn088
Higuchi Y, Nishida Y, Kozawa E, Zhuo LS, Arai E, Hamada S, Morita D, Ikuta K, Kimata K, Ushida T, Ishiguro N (2017) Conditional knockdown of hyaluronidase 2 in articular cartilage stimulates osteoarthritic progression in a mice model. Sci Rep Uk 7:7028. https://doi.org/10.1038/s41598-017-07376-5
Lee CR, Sakai D, Nakai T, Toyama K, Mochida J, Alini M, Grad S (2007) A phenotypic comparison of intervertebral disc and articular cartilage cells in the rat. Eur Spine J 16:2174–2185. https://doi.org/10.1007/s00586-007-0475-y
Rustenburg CME, Emanuel KS, Peeters M, Lems WF, Vergroesen PPA, Smit TH (2018) Osteoarthritis and intervertebral disc degeneration: quite different, quite similar. JOR Spine 1:e1033
Loeser RF (2006) Molecular mechanisms of cartilage destruction: mechanics, inflammatory mediators, and aging collide. Arthritis Rheum 54:1357–1360. https://doi.org/10.1002/art.21813
Kapoor M, Martel-Pelletier J, Lajeunesse D, Pelletier JP, Fahmi H (2011) Role of proinflammatory cytokines in the pathophysiology of osteoarthritis. Nat Rev Rheumatol 7:33–42. https://doi.org/10.1038/nrrheum.2010.196
Fernandes JC, Martel-Pelletier J, Pelletier JP (2002) The role of cytokines in osteoarthritis pathophysiology. Biorheology 39:237–246
Suzuki S, Fujita N, Hosogane N, Watanabe K, Ishii K, Toyama Y, Takubo K, Horiuchi K, Miyamoto T, Nakamura M, Matsumoto M (2015) Excessive reactive oxygen species are therapeutic targets for intervertebral disc degeneration. Arthritis Res Ther 17:316. https://doi.org/10.1186/s13075-015-0834-8
Lepetsos P, Papavassiliou AG (2016) ROS/oxidative stress signaling in osteoarthritis. Biochim Biophys Acta 1862:576–591. https://doi.org/10.1016/j.bbadis.2016.01.003
Feng CC, Yang MH, Lan MH, Liu C, Zhang Y, Huang B, Liu H, Zhou Y (2017) ROS: crucial intermediators in the pathogenesis of intervertebral disc degeneration. Oxidative Medicine and Cellular Longevity. https://doi.org/10.1155/2017/5601593
Kim KW, Chung HN, Ha KY, Lee JS, Kim YY (2009) Senescence mechanisms of nucleus pulposus chondrocytes in human intervertebral discs. Spine J 9:658–666. https://doi.org/10.1016/j.spinee.2009.04.018
Ohno-Nakahara M, Honda K, Tanimoto K, Tanaka N, Doi T, Suzuki A, Yoneno K, Nakatani Y, Ueki M, Ohno S, Knudson W, Knudson CB, Tanne K (2004) Induction of CD44 and MMP expression by hyaluronidase treatment of articular chondrocytes. J Biochem 135:567–575. https://doi.org/10.1093/jb/mvh069
Bachmeier BE, Nerlich A, Mittermaier N, Weiler C, Lumenta C, Wuertz K, Boos N (2009) Matrix metalloproteinase expression levels suggest distinct enzyme roles during lumbar disc herniation and degeneration. Eur Spine J 18:1573–1586. https://doi.org/10.1007/s00586-009-1031-8
Weiler C, Nerlich AG, Zipperer J, Bachmeier BE, Boos N (2002) 2002 SSE Award Competition in Basic Science: expression of major matrix metalloproteinases is associated with intervertebral disc degradation and resorption. Eur Spine J 11:308. https://doi.org/10.1007/s00586-002-0472-0
Acknowledgements
Open access funding provided by Paracelsus Medical University. This study was supported by a start-up grant from the Center for Applied Biotechnology and Molecular Medicine (CABMM).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest. The authors have full control of all primary data.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Krupkova, O., Greutert, H., Boos, N. et al. Expression and activity of hyaluronidases HYAL-1, HYAL-2 and HYAL-3 in the human intervertebral disc. Eur Spine J 29, 605–615 (2020). https://doi.org/10.1007/s00586-019-06227-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00586-019-06227-3