Abstract
Key message
In barley (Hordeum vulgare), MTOPVIB is critical for meiotic DSB and accompanied SC and CO formation while dispensable for meiotic bipolar spindle formation.
Abstract
Homologous recombination during meiosis assures genetic variation in offspring. Programmed meiotic DNA double-strand breaks (DSBs) are repaired as crossover (CO) or non-crossover (NCO) during meiotic recombination. The meiotic topoisomerase VI (TopoVI) B subunit (MTOPVIB) plays an essential role in meiotic DSB formation critical for CO-recombination. More recently MTOPVIB has been also shown to play a role in meiotic bipolar spindle formation in rice and maize. Here, we describe a meiotic DSB-defective mutant in barley (Hordeum vulgare L.). CRISPR-associated 9 (Cas9) endonuclease-generated mtopVIB plants show complete sterility due to the absence of meiotic DSB, synaptonemal complex (SC), and CO formation leading to the occurrence of univalents and their unbalanced segregation into aneuploid gametes. In HvmtopVIB plants, we also frequently found the bi-orientation of sister kinetochores in univalents during metaphase I and the precocious separation of sister chromatids during anaphase I. Moreover, the near absence of polyads after meiosis II, suggests that despite being critical for meiotic DSB formation in barley, MTOPVIB seems not to be strictly required for meiotic bipolar spindle formation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Meiosis in sexually reproducing organisms assures genetic variation through homologous recombination (HR) (Hunter, 2015; Mercier et al., 2015). Meiotic HR starts with DNA double-strand break (DSB) formation catalyzed by the topoisomerase-like protein SPO11 (Bergerat et al., 1997; Keeney et al., 1997). Meiotic DSBs repaired using the homologous chromosome as template result either in crossovers (COs; reciprocal genetic exchange between homologous chromosomes) or non-crossovers (NCOs; only rather short stretches of DNA are copied to the broken chromosome) (Hunter, 2015; Mercier et al., 2015; Wang and Copenhaver, 2018). Typically, only a minor fraction of meiotic DSBs becomes repaired into COs in plants. Meiotic DSBs are generated in excess with more than 90% of plant DSBs being resolved as NCO or being repaired by using the sister chromatid as a template (Mercier et al., 2015). The remaining DSBs can mature into two types of COs in plants: Interference-sensitive class I COs that are farther apart along the chromosome than expected by chance (CO interference; (Berchowitz and Copenhaver, 2010)) and depend on the activity of a group of proteins called ZMM proteins (named after the budding yeast proteins Zip1-4, Mer3, Msh4 and Msh5 (Pyatnitskaya et al., 2019)) across organisms including different plants (e.g., Chelysheva et al., 2012; Shen et al., 2012; Desjardins et al., 2020) as well as ZMM-independent COs that are interference-insensitive (class II) and depend in part on MUS81 or FANCD2 (Berchowitz et al., 2007; Higgins et al., 2008; Kurzbauer et al., 2018). In plants, the majority of CO are ZMM-dependent class I COs (Mercier et al., 2015; Wang and Copenhaver, 2018).
While in many species a single SPO11 gene is present (e.g., mammals and fungi) with two functionally distinct splice variants as in mice and humans (Bellani et al., 2010; Kauppi et al., 2011), three SPO11 genes are found in A. thaliana. Two of them, SPO11-1 and SPO11-2, become differentially spliced (Sprink and Hartung, 2014, 2021) and are required for meiotic DSB formation (Grelon et al., 2001; Stacey et al., 2006; Hartung et al., 2007), while the third, SPO11-3, is critical for somatic development (Hartung et al., 2002; Sugimoto-Shirasu et al., 2002; Yin et al., 2002) but not required for meiotic DSB formation. In rice, among five SPO11 genes at least SPO11-1 and SPO11-2 are involved in meiotic DSB formation (Yu et al., 2010; An et al., 2011; Fayos et al., 2020). In wheat, SPO11-1 and SPO11-2 (Benyahya et al., 2020; Da Ines et al., 2020), and in maize, SPO11-1 (Ku et al., 2020) are critical for meiotic DSB formation. In A. thaliana (L.) Heynh., SPO11-1 and SPO11-2 form a heterotetramer with two copies of the distant archaeal topoisomerase VI (TopoVI) B subunit homolog MTOPVIB (Vrielynck et al., 2016) constituting the meiotic TopoVI-like complex (core catalytic meiotic DSB induction complex). MTOPVIB contains four protein motifs that are conserved among flowering plants, named B1, B2, B3, and B4 from N- to C-terminus. The B1 and B2 motifs are part of the GHKL domain, necessary for ATP binding. The function of the B3 motif is currently unclear. The B4 motif is part of the Transducer domain mediating the interaction with SPO11-1 and SPO11-2 (Vrielynck et al., 2016). MTOPVIB is critical for meiotic DSB induction in A. thaliana, rice, maize, and mice (Fu et al., 2016; Robert et al., 2016a; Vrielynck et al., 2016; Xue et al., 2016; Jing et al., 2020). Further accessory proteins are also required for DSB formation in plants (e.g., PRD1-3 or DFO in A. thaliana; (De Muyt et al., 2007, De Muyt et al., 2009, Zhang et al., 2012, Vrielynck et al., 2021)). Functional impairment of any of the core DSB induction complex components or any accessory factor results in abolition of meiotic DSB formation, leading to a severe fertility reduction due to the absence of meiotic HR and accompanied unbalanced gamete formation.
In addition to its role in meiotic DSB formation, MTOPVIB, initially in rice (Xue et al., 2019) but also in maize (Jing et al., 2020), has been implicated in being critical for bipolar spindle formation at metaphase I, which is independent of its role in DSB formation. In mtopVIB, a high frequency of polyads (up to octads) is found in rice (~ 86% Xue et al., 2019), maize (~ 63% Jing et al., 2020), and in A. thaliana (~ 70% Tang et al., 2017). Moreover, at metaphase I, sister kinetochores in rice mtopVIB univalents are typically mono-oriented while in univalents in other rice mutants defective for meiotic DSB formation, frequent bi-orientation of sister kinetochores is found (Xue et al., 2019). Hence, likely the bi-orientation of kinetochores in (at least some) univalents at metaphase I is critical for the bipolar spindle organization in case of the absence of meiotic DSBs in an MTOPVIB-dependent manner (Xue et al., 2019).
Here we describe the function of MTOPVIB in barley. Cas9 endonuclease-generated mtopVIB plants are entirely sterile due to defective meiosis. In HvmtopVIB, no meiotic DSBs form, synaptonemal complex (SC) formation is abolished, and the absence of CO formation leads to univalents that undergo unbalanced segregation into aneuploid gametes. Moreover, frequent bi-orientation of sister kinetochores during metaphase I, precocious separation of sister chromatids during anaphase I, and rare polyad formation were found in mtopVIB lines. Hence, barley MTOPVIB seems not to be required for bipolar spindle formation despite its role in meiotic DSB formation.
Materials and methods
Plant material, growing conditions, and crossing
Grains of the wild-type (WT) barley (Hordeum vulgare L.) cultivar Golden Promise and mtopVIB mutants were germinated on wet filter paper in a Petri dish. After seven days, plantlets were transferred to soil pots and grown in a greenhouse at 18/15 °C and under 16 h light/8 h darkness conditions with a relative humidity of 60–70%.
Plant crossings were done as described (Ahn et al., 2020) by using five-to-seven-week-old plants. Spikes for emasculation were selected based on the size and appearance of emerging awns. All three immature anthers from each spikelet along the whole spike were removed. Two days after emasculation, 2–3 freshly collected mature anthers were introduced into each of the emasculated spikelets. The developing caryopses became visible after 10 to 12 days in cases of a successful pollination. Mature spikes were collected to estimate the number of generated grains per cross.
Resequencing of HvMTOPVIB and expression analysis
Total RNA was isolated from 100 mg of barley anthers using Trizol (Invitrogen). One µg of RNA was employed for cDNA synthesis using the Invitrogen FirstStrand cDNA Synthesis Using SuperScript II RT kit following manufacturer’s instructions. cDNA samples were used for PCR amplification using the NEB Phusion High Fidelity PCR Kit and Sanger resequencing of the coding sequence of barley MTOPVIB. For primer sequences see Online resource 1.
Selection of Cas9/gRNA target motifs and Cas9-mediated in vitro digestion
The coding sequence of HvMTOPVIB was cloned using the CloneJET PCR cloning kit following the manufacturer's instructions (ThermoFisher). The resulting plasmid DNA was linearized with XbaI. Cas9/gRNA target motifs were selected within the coding sequence of HvMTOPVIB according to criteria previously described (Kumlehn et al., 2018). Three accordingly designed HvMTOPVIB crRNAs (single-strand RNAs that contain the gRNA target sequence) for exon 1 (gRNA#1: GAGCTTCCGGTGGGGGGAGG), exon 2 (gRNA#2: GGATGTCGGAGTCGCAGTGC), and exon 6 (gRNA#3: GACTTCATATTATGGCTGGT) were hybridized to the tracrRNA by dissolving 2 nmol each in Nuclease-Free Duplex Buffer (Integrated DNA Technologies, IDT). To generate the ribonucleoprotein (RNP) complex (Cas9 endonuclease combined with the hybridized gRNAs), 1 µl of the RNA mix was combined with 1 µl of Cas9 solution (1 µM Cas9 endonuclease purchased from IDT, 1 × PBS, and 50% glycerol) in 1 × PBS. The digestion was performed by mixing the RNP complex solution with 200 ng of linearized plasmid DNA followed by incubation at 37 °C for 1 h and analyzed by running the samples in an agarose gel.
Construction of transformation vectors
Using the CasCADE vector system (Hoffie et al., 2022), target-specific sequences (see Online resource 1) were integrated downstream of a wheat TaU6 promoter into generic gRNA modules which were then used to create guide RNA assembly vector pGH577. Then, the final assembly vector pGH584 was produced including a maize codon-optimized xcas9 (Hu et al., 2018) driven by the maize Polyubiquitin 1 promoter along with the 5’-UTR including intron 1. From pGH584, the SfiI-fragment containing gRNA and xcas9 expression units were transferred to the compatible generic binary vector 271p6i-2 × 35 s-TE9 (DNA Cloning Service, Hamburg, Germany) to create pGH615.
Plant transformation
The generation of transgenic barley plants was done as previously described (Hensel et al., 2008; Marthe et al., 2015). In brief, immature embryos were dissected from surface-sterilized caryopses, co-cultivated upon inoculation with the hypervirulent Agrobacterium tumefaciens strain AGL1 (Lazo et al., 1991) harboring plasmid pGH615, which was followed by callus induction and plant regeneration under selective conditions using Timentin to remove Agrobacterium and hygromycin to ensure preferential development of transgenic cells and tissue. When the plantlets had developed roots, they were transferred into soil.
Identification of mutations within HvMTOPVIB
The genomic regions addressed for targeted mutagenesis (a fragment of 628 bp spanning the target motifs of gRNA#1 and gRNA#2 as well as a fragment of 411 bp harboring the cognate motif of gRNA#3) were PCR-amplified from flag leaf genomic DNA of primary transformants (T0) and Sanger-sequenced. For primer sequences see Online resource 1. In segregating T1 families (35 siblings per T0 parent), the target regions were PCR-amplified and resulting agarose gel-purified amplicons were sent to Sanger-sequencing. T-DNA-free individuals were PCR-selected using primers for cas9 and hpt. For primer sequences see Online resource 1.
Plant genotyping
HvmtopVIB is associated with a disruption of a BtsI restriction site within HvMTOPVIB, which facilitates genotyping. After PCR amplification of the previously described 628-bp PCR fragment spanning the target motifs of gRNA#1 and gRNA#2 (for primer sequences see Online resource 1), resulting amplicons were purified and used in a BtsI restriction digest. PCR amplicons from the HvMTOPVIB WT allele are digested by Btsl (287, 252, and 90 bp fragments) while PCR amplicons from plants carrying a mutant mtopVIB allele are only partially digested (343 and 287 bp fragments) enabling plant genotyping.
Male meiotic chromosome preparations
Spikes were fixed in an ice-cold 3:1 (75% ethanol, 25% acetic acid) solution for at least 24 h. One anther per spikelet was used to determine the meiotic stage by squashing it on a microscopic slide in a drop of acetocarmine (MORPHISTO) and evaluation under a light microscope. The remaining two anthers of selected spikelets were either used immediately or stored in freshly-prepared 3:1 solution. Male meiotic chromosomes were prepared by squashing as described (Li et al., 2018). Anthers were disrupted with the help of forceps in a drop of acetocarmine solution, the preparation was heated by passing the slide 2–3 times above a flame (the acetocarmine should not boil), and a glass coverslip was placed on top pressing gently and firmly with a thumb between filter paper. At this stage, male meiotic chromosomes can be visualized under a light microscope. Staining can be enhanced by adding additional acetocarmine solution at the edges of the coverslip, allowing it to penetrate by diffusion, and by incubating slides in a moist chamber at 4 °C overnight. This also enables staining of the meiotic spindle. To counterstain chromosomes with DAPI, slides were frozen in liquid nitrogen, the coverslips removed with the help of a razor blade, and immediately transferred to an ethanol series (70, 85, 100%) 2 min each for dehydration. After airdrying, slides were counterstained with DAPI in Vectashield mounting media (1.5 µg/ml, Vector Laboratories).
Immunohistochemistry
One fresh anther per spikelet was used to determine meiotic stages as indicated above. The remaining two anthers of selected spikelets were used for chromosome spreading as described (Armstrong et al., 2009; Cuacos et al., 2021) with minor modifications. Two anthers per slide were digested in 25 µl of enzyme mix for 8 min at 37 °C in a moist chamber, disrupting the material with a brass rod after the first 4 min. Spreading was done with 17 µl of 1.5% Lipsol solution and fixation with 17 µl of 4% paraformaldehyde. Primary antibodies were incubated overnight at 4 °C, and secondary antibodies for 1 h at 37 °C. The following primary antibodies and dilutions were used: anti-HvHEI10 (guinea pig (Desjardins et al., 2020), 1:200), anti-OsγH2AX (rabbit (Miao et al., 2013), 1:200), anti-HvASY1 (guinea pig, 1:500), anti-HvZYP1 (rabbit, 1:500), and anti-grass CENH3 (rabbit (Sanei et al., 2011), 1:300). Secondary antibodies and dilutions used were: goat anti-guinea pig Alexa488 (Invitrogen), goat anti-guinea pig Alexa594 (Invitrogen), donkey anti-rabbit Alexa488 (Jackson Immunology Research), and donkey anti-rabbit Alexa594 (Invitrogen), all at 1:500.
Fluorescence in situ hybridization
All steps were performed at room temperature unless otherwise indicated. Slides with DAPI-stained male meiotic chromosomes prepared as described above were rinsed in 2xSSC to let coverslips fall, followed by an alcohol series (70, 85, 100%) and air-dried. Next, slides were washed twice in 2xSSC for 5 min, treated with 45% acetic acid for 10 min, washed in 2xSSC for 10 min, treated with 0.1% pepsin in 0.01 N HCl for 10 min at 37 °C, rinsed twice with 2xSSC for 5 min, fixed in 4% formaldehyde (5 ml 37% formaldehyde + 42 ml 2xSSC) for 10 min followed by three washes in 2xSSC of 5 min each. Afterward, slides were dehydrated in ethanol (70, 85, 100%, 2 min each) and air-dried for 1 h. 20 µl hybridization mix (10 µl deionized formamide, 5 µl 4 × buffer (4 × buffer: 80 µl 20xSSC, 8 µl 1 M Tris–HCL pH 8.0, 1.6 µl 0.5 M EDTA, 99.2 µl double-distilled water (ddH2O)), 2 µl ddH2O and 3 µl probe) was added to each slide under a 22 × 22 mm coverslip, slides were sealed with rubber cement (Fixogum, Marabu), denatured at 80 °C for 2 min on a hot plate and immediately transferred to a moist chamber for incubation overnight at 37 °C. On the next day, coverslips were carefully removed, slides were washed in 2xSSC for 20 min at 58 °C, transferred to 2xSSC at room temperature, and dehydrated in 70, 85, 100% ethanol, 2 min each. After airdrying, slides were counterstained with DAPI in Vectashield mounting media and evaluated under a fluorescence microscope.
Probes utilized were: 5S (pCT4.2, Campell et al., 1992) and 45S (pTa71, Gerlach and Bedbrook, 1979) rDNA as well as HvT01 (Rey et al., 2018). FISH probes were labeled by nick translation with Texas Red and Atto488 (NT labeling kits, Jena Biosciences).
Microscopy
Images were acquired with a Nikon Eclipse Ni-E fluorescence microscope equipped with a Nikon DS-Qi2 camera and NIS-Elements-AR version 4.60 software (Nikon, Tokyo, Japan). Images were processed with GIMP 2.10 (www.gimp.org). To detect the ultrastructural chromatin organization of meiocytes at a resolution of ~ 120 nm (super-resolution achieved with a 488 nm laser excitation) spatial structured illumination microscopy (3D-SIM) was performed with a 63 × /1.4 Oil Plan-Apochromat objective of an ElyraPS.1 microscope system and the software ZENBlack (Carl Zeiss GmbH). Images were captured separately for each fluorochrome using the 561, 488, and 405 nm laser lines for excitation and appropriate emission filters (Weisshart et al., 2016; Kubalová et al., 2021). Maximum intensity projections of whole cells were calculated via the ZENBlack software.
Yeast two-hybrid assays
To fuse HvSPO11-1, HvSPO11-2, HvMTOPVIB or HvmtopVIB to the Gal4activation domain (AD, prey vector) or the Gal4DNA-binding domain (BD, bait vector), each CDS was cloned into pGADT7 and pGBKT7 vectors. For primer sequences see Online resource 1. Different combinations of prey and bait plasmids were co-transformed into the yeast strain Y2HGold. Handling of yeast transformation and selection assays were performed according to the manufacturer’s instructions (Takara, #630,489). Transformed yeast cells were grown on different selective media: TDO (Minimal Media Triple Dropouts, SD/–His/–Leu/–Trp, #630,419), QDO (Minimal Media Quadruple Dropouts, SD/–Ade/–His/–Leu/–Trp, #630,428) and DDO (Minimal Media Double Dropouts, SD/–Leu/–Trp, #630,417).
Antibody production
To produce anti-HvZYP1 antibodies, a synthetic peptide (HPANIGELFSEGSLNPYADD) corresponding to aa 839–858 of H. vulgare ZYP1 was used for immunization of rabbits by LifeTein. Rabbit anti-HvZYP1 was affinity-purified against the synthetic peptide by LifeTein.
To produce anti-HvASY1, the complete HvASY1 CDS was used to produce recombinant HvASY1 proteins by Biomatik. Recombinant HvASY1 proteins were used for the immunization of guinea pigs by Davids Biotechnologie GmbH.
Results
Identification of barley MTOPVIB
To identify potential candidates involved in meiotic DSB induction in the cereal crop barley, the barley reference genome was initially queried for putative SPO11 and MTOPVIB homologs (The International Barley Genome Sequencing Consortium 2012). In the latest MorexV3 genome reference (Mascher et al., 2021) three putative SPO11 candidates (SPO11-1—HORVU.MOREX.r3.5HG0511100.1, SPO11-2—HORVU.MOREX.r3.7HG0699350.1, SPO11-3—HORVU.MOREX.r3.4HG0385890.1) and a single MTOPVIB candidate (HORVU.MOREX.r3.7HG0726050) were found. We focused on the single copy MTOPVIB candidate. The MTOPVIB candidate locus represents a putative ORF of 7477 bp predicted to encode for twelve exons and two different CDS variants of 1449 and 1452 bp (The International Barley Genome Sequencing Consortium 2012) (Fig. 1a). While the first predicted CDS variant results in a 484 aa protein containing all four conserved MTOPVIB domains (B1, B2, B3, and B4), the second predicted variant results in a 354 aa protein lacking part of the Transducer domain. In cDNA samples from anthers of cv. Golden Promise, only the first but not the second CDS variant was found at detectable levels, suggesting abundant expression of the first variant in reproductive tissues.
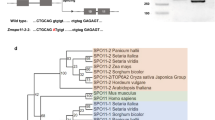
Identification of the HvMTOPVIB gene and isolation of a HvmtopVIB mutant. a HvMTOPVIB schematic gene structure depicting exons and introns. a, b Three selected gRNAs targeting exon one (gRNA#1), two (gRNA#2), or six (gRNA#3) of HvMTOPVIB are active in vitro. HvmtopVIB mutation (T insertion present in HvmtopVIB as compared with the HvMTOPVIB CDS) recovered in independent transgenic families and plants. c Schematic protein model of HvMTOPVIB depicting HvmtopVIB and all four conserved MTOPVIB domains (B1, B2, B3, and B4). d HvMTOPVIB while not HvmtopVIB interacts with HvSPO11-1 and HvSPO11-2 in “yeast two-hybrid assays”. No interaction was found between HvSPO11-1 and HvSPO11-2. Interactions were verified by the growth of yeast cells on different selective media: TDO (Minimal Media Triple Dropouts, SD/–His/–Leu/–Trp), QDO (Minimal Media Quadruple Dropouts, SD/–Ade/–His/–Leu/–Trp) and DDO (Minimal Media Double Dropouts, SD/–Leu/–Trp). BD, bait vector; AD, prey vector
Generation of HvmtopVIB plants by Cas9-triggered mutagenesis
To functionally dissect the role of meiotic DSB induction for CO formation in barley, the single copy candidate MTOPVIB was selected as the target for RNA-guided Cas9 to isolate DSB-defective plants. MTOPVIB was initially identified in the Morex reference genome. Typically, cv. Golden Promise is employed for stable genetic transformation including the application of Cas endonuclease technology (Koeppel et al., 2019). The conservation of the MTOPVIB coding sequence between cvs. Morex and Golden Promise was confirmed based on sequencing of the MTOPVIB coding sequence from respective anther cDNA samples. Three Cas9-compatible gRNAs addressing either exon one (gRNA#1), two (gRNA#2), or six (gRNA#3) of HvMTOPVIB were selected and their target-specific activity was confirmed in vitro (Fig. 1a, b). These three gRNAs were independently placed under the control of a wheat U6 promoter in a binary vector carrying cas9 under control of the maize Ubiquitin 1 promoter and hpt as plant selection marker, and used for stable genetic transformation of cv. Golden Promise via Agrobacterium-mediated DNA transfer to immature embryo explants. Among 18 regenerated T0 plants, Sanger sequencing of Cas9/gRNA target regions revealed potential mutations in four independent plants (named E1, E7, E9, and E12) at the target motifs in exon one and exon two. In 35 progeny of each E1, E7, E9, and E12 T0 plants, heritable edits were only recovered in the gRNA#2-addressed target motif (in exon two) within the T1 families of E7 and E9. Among these, three cas9-free T1 plants with a heterozygous one-base pair thymidine insertion 189 bp downstream of the ATG of HvMTOPVIB (termed HvmtopVIB) were identified (Fig. 1b, c). The presence of this mutation (at position + 103 relative to the ATG of the CDS; exon 2) was confirmed based on sequencing of anther cDNA samples. HvmtopVIB leads to a predicted frameshift in the HvMTOPVIB reading frame from the middle of the B1 protein motif, which results in a nonsense aa sequence and a premature STOP codon. The predicted aberrant truncated protein is 77 aa long and lacks all conserved functional domains of MTOPVIB. To confirm whether the insertion had an impact on HvMTOPVIB function, the HvMTOPVIB cDNA of HvmtopVIB and WT plants was used in Y2H experiments together with the CDS of HvSPO11-1 and HvSPO11-2 (Fig. 1d). Only yeast cells transformed with HvMTOPVIB and HvSPO11-1 or HvSPO11-2 grew on selective media suggesting an interaction of HvMTOPVIB with HvSPO11-1 and HvSPO11-2. Notably, the one-base pair insertion in HvmtopVIB disrupts the interaction of HvMTOPVIB with HvSPO11-1 and HvSPO11-2, suggesting that the mutation recovered in independent transgenic families and plants results in an aberrant truncated protein disrupting the function of HvMTOPVIB.
HvmtopVIB plants are sterile despite normal vegetative growth
Vegetative growth and development of homo- and heterozygous HvmtopVIB plants were similar to WT plants. However, homozygous HvmtopVIB plants were completely sterile, i.e., spikes were devoid of grains, while WT and heterozygous HvmtopVIB plants were fertile (Fig. 2a, b). To address whether also female fertility was disturbed, HvmtopVIB plants as well as segregating WT plants were pollinated with WT pollen. While pollinated WT plants formed grains, in the case of HvmtopVIB plants, no grains were recovered whatsoever, suggesting that both male and female fertility were severely impaired.
Phenotype of HvmtopVIB barley plants: Vegetative growth and development similar to WT, but complete sterility due to meiotic defects. a, b Six-weeks old plants and mature spikes from WT (Golden Promise) (a) and HvmtopVIB (b). No grains formed in HvmtopVIB. c Male meiotic chromosome analysis in WT, HvmtopVIB, and HvmtopVIB + / + . Compared with WT and segregating HvmtopVIB + / + plants, in HvmtopVIB the absence of thick chromosomes indicates no synapsis during pachytene and the presence of 14 univalents indicates the absence of COs at metaphase I. During anaphase I, random segregation of univalents to opposite poles or separation of sister chromatids (arrowheads) finally leading to the formation of unbalanced tetrads and micronuclei in HvmtopVIB. DNA stained with DAPI and shown in grey. d Quantification of bivalent/univalent formation in WT, HvmtopVIB + / + and HvmtopVIB. Left: among all bivalents, percentage of ring and rod bivalents (in WT and HvmtopVIB + / +) and bivalent-like (“biv-like”) structures and “unpaired” bivalents (i.e., pairs of univalents) in HvmtopVIB. Right: absolute values. Bars represent 10 µm
Absence of bivalent and CO formation as well as unbalanced chromosome segregation and gametes in HvmtopVIB mutants
Male meiotic chromosome analysis was performed to assess meiotic chromosome behavior in HvmtopVIB mutants. During WT meiosis (cv. Golden Promise) and in segregating WT plants of HvmtopVIB mutants (Fig. 2c), chromosomes appeared as thin threads during leptotene, gradually thickening as homologous chromosomes get physically connected, being completely synapsed during pachytene. Chromosome condensation led to seven bivalents connected via chiasmata (cytological visualization of COs), typically forming rings (representing at least one chiasma at each chromosome arm) and occasionally rods (at least one chiasma in one chromosome arm). During metaphase I, bivalents align at the equatorial plate, and homologous chromosomes segregate to opposite poles during anaphase I. Four haploid cells are found upon separation of chromatids during meiosis II, forming a tetrad that will give rise to male gametes. In HvmtopVIB (Fig. 2c), meiotic prophase I initially appeared similar to the WT. However, in HvmtopVIB, no cells showing thick chromosome threads representing pachytene were found. Moreover, during metaphase I, chromosomes appeared as 14 univalents, suggesting a failure to form chiasmata, which resulted in a lack of bivalent formation. During anaphase I, univalents segregated randomly to opposite poles resulting in unbalanced dyads and subsequently also in the formation of unbalanced tetrads and micronuclei. Notably, commonly also the precocious separation of sister chromatids was found during meiosis I. While in the WT, seven bivalents were invariably found, in HvmtopVIB, no bivalent formation was seen. However, structures reminiscent of a rod-like bivalent (1–3 per cell) were frequently observed in HvmtopVIB (Fig. 2d). To resolve whether infrequent bivalent formation occurred involving homologous chromosomes, FISH using HvT01 as well as 5S and 45S ribosomal DNA ribosomal DNA probes was performed (Fig. 3a–c). Among 38 studied bivalent-like structures, only one was formed by homologous chromosomes, a frequency which is even lower than the number expected by random associations. Notably, in 32 out of 38 cases, one of the two, or both, 45S rDNA-containing chromosomes participated in such bivalent-like structures, as previously observed even in the absence of recombination (Stronghill et al., 2010; Da Ines et al., 2012). However, not necessarily the 45S rDNA-containing chromosome arm was always involved. We also immunolocalized the class I CO marker HEI10 (Chelysheva et al., 2012; Wang et al., 2012) (Fig. 3d). While in the WT, 16.5 ± 1.3 HEI10 foci (range 14–20, n = 66) were found during diakinesis and metaphase I, in HvmtopVIB no HEI10 foci (n = 100) were found, including a complete lack of HEI10 foci at bivalent-like structures, suggesting the complete absence of ZMM-dependent class I CO. Hence, these bivalent-like structures likely represent random achiasmatic associations of chromosomes or chromosomes being nearby by chance.
Bivalent-like structures in HvmtopVIB mutants are achiasmatic. a, b FISH with 45S (green) and 5S rDNA as well as HvT01 (purple) probes facilitate the identification of all homologous chromosome pairs. Among 38 bivalent-like structures, bivalent-like associations primarily occur between non-homologous chromosomes (a) and only in one case between homologous chromosomes (b). c Proportion of each type of bivalent-like structure (homologous, green; or non-homologous, red range) considering chromosomes involved (chromosome with large 45S rDNA signal (“45Sb”), small signal (“45Ss”) or no 45S signal (“X”)). d Immunolocalization of HEI10 in WT reveals numerous foci on diakinesis-metaphase I chromosomes. No HEI10 foci were detected on HvmtopVIB bivalent-like structures (asterisks) or univalents. DNA stained with DAPI and shown in blue. Bars represent 10 µm
Absence of meiotic DSB and SC formation in HvmtopVIB mutants
Rapid phosphorylation of histone H2AX around DSB sites is found and in diverse species including plants an antibody to detect the phosphorylated form of H2AX (γH2AX) is used to detect DSB sites (Rogaku et al., 1998, Paull et al., 2000; Mahadevaiah et al., 2001; Sanchez-Moran et al., 2007; Wu et al., 2015). Hence, we performed immunolocalization against γH2AX as a possible proxy for DSB formation in barley together with the axis-associated protein ASY1 (Caryl et al., 2000; Armstrong et al., 2002; Sanchez-Moran et al., 2007; Higgins et al., 2012) (Fig. 4a). In the WT, abundant γH2AX foci were detected during early zygotene (average 466.5 ± 113.2, range 352–675, n = 10), which is consistent with previous reports in barley (Higgins et al., 2012). In contrast, in HvmtopVIB, a strong decrease in γH2AX foci or complete absence was found during early prophase I (average 2.3 ± 2.8, range 0–8, n = 11), suggesting (near) absence of meiotic DSB formation. Thus, the absence of typical bivalent structures or ZMM-dependent CO formation is due to an abolished HR induction.
Absence of DSB and SC formation in HvmtopVIB mutants. a Immunolocalization of ASY1 and γH2AX visualized by 3D-SIM reveals numerous axis-associated γH2AX foci in WT but none in HvmtopVIB. b Immunolocalization of ASY1 and ZYP1 indicates initially polarized synapsis in WT spreading across the whole nucleus until all chromosome pairs are synapsed with ASY1 getting gradually depleted from ZYP1-positive synapsed regions (from top to bottom). In HvmtopVIB, only ZYP1 foci varying in number and size are present. No linear ZYP1 signals are found, indicating the absence of synapsis. Bars represent 5 µm
Simultaneous immunolocalization of ASY1 and ZYP1 (transverse filament protein of the SC; Higgins et al., 2005) was performed to explore whether SC formation occurs in HvmtopVIB (Fig. 4b). In the WT, ASY1 localizes to the meiotic chromosome axis during leptotene, and it becomes depleted from synapsed regions upon loading of ZYP1 (Wojtasz et al., 2009; Lambing et al., 2015). Hence, during zygotene-pachytene, the presence of ASY1 combined with the absence of ZYP1 marks un-synapsed chromosome regions, while the almost absence of ASY1 combined with the presence of ZYP1 marks synapsed chromosome regions. Moreover, a spatiotemporal asymmetry in meiotic progression was found, i.e., chromosome axis and SC formation are initially polarized to one area of the nucleus before gradually extending across the whole nucleus as previously described (Higgins et al., 2012). In HvmtopVIB, ASY1 localization was similar to WT during leptotene and zygotene including initial polarization before spreading throughout the nucleus. In the case of ZYP1, in HvmtopVIB, no typical linear signals were found. ZYP1 formed varying numbers of foci that did not elongate. Also, larger aggregates or polycomplexes that varied in number and size appeared. Moreover, in HvmtopVIB (n = 51) no sign of polarized localization of these ZYP1 foci or aggregates/polycomplexes was found. Thus, the absence of ZYP1 installation is consistent with abolished synapsis due to the lack of meiotic DSBs and consequently HR.
Limited polyad formation and frequent bi-orientation of kinetochores in HvmtopVIB univalents during meiosis I
MTOPVIB is critical for bipolar spindle formation during meiosis I in maize and rice (Xue et al., 2019; Jing et al., 2020). In mtopVIB, independent of meiotic DSB formation, multipolar spindles form, resulting in a high frequency of polyads instead of tetrads. Also, univalents preferentially have mono-oriented sister kinetochores during metaphase I. We asked whether sister kinetochore mono-orientation is prevalent also in HvmtopVIB, and polyads occur in high frequency. Surprisingly, in HvmtopVIB similar to the WT, we found only dyads after the first meiotic division, and predominantly tetrads (unbalanced and showing micronuclei formation) after the second meiotic division (Fig. 5a). Only one out of 179 spores in HvmtopVIB was a polyad, specifically a pentad. In accordance with this, spindles were consistently arranged in a bipolar fashion during the two meiotic divisions (83 cells) (Fig. 5b).
In HvmtopVIB mutants bipolar spindle formation including split sister centromeres in univalents and a limited frequency of polyads appears. a In HvmtopVIB, tetrads form predominantly; occasionally dyads similar to WT, sometimes triads, and only once a pentad. Acetocarmine-staining examples of a dyad (II), two triads (III), two tetrads (IV), and a pentad (V). b Acetocarmine-stained male meiotic cells reveal exclusively bipolar spindle formation in HvmtopVIB. From left to right: metaphase I, anaphase I, telophase I, metaphase II, anaphase II. c Immunolocalization of CENH3 (red) reveals split sister centromeres in univalents during diakinesis-metaphase I (two CENH3 signals per univalent, asterisks, vs. one signal that indicates fused sister centromeres). Chromosomes stained with DAPI and shown in blue. Bars represent 10 µm
Both in WT and HvmtopVIB, dyads were found together with tetrads in the same preparations (in both backgrounds ~ 10% of all scored cells), likely representing asynchrony in meiotic progression among meiocytes within the same anthers as described for cv. Golden Promise grown at 20 °C (Schindfessel et al., 2021), which is thus unrelated to the MTOPVIB mutation.
Notably, in HvmtopVIB in addition to dyads and tetrads also triads were found (~ 23%), but never tripolar spindles or frequent polyads (Fig. 5a, b). A similar situation was found in other meiotic mutants forming two (dyads) or three (triads) nuclei unrelated to spindle alterations (e.g., dmc1 in barley, where also pentads were found (Szurman-Zubrzycka et al., 2019), pair2 in rice (Nonomura et al., 2004) or mms21 in Arabidopsis (Liu et al., 2014)). Likely, due to defects in chromosome segregation in HvmtopVIB caused by the lack of recombination leading to unbalanced tetrads, cytokinesis that produces three or four (or very rarely five) microspores (often with micronuclei) occurs.
Using antibodies against the centromere-specific histone H3 variant CENH3, we quantified the number of split and fused CENH3 signals in HvmtopVIB univalents to quantify bi- and mono-orientation, respectively, of centromeres/kinetochores during meiosis I. All metaphase I cells contained at least 1 and up to 7 univalents with split centromeres (average 3 univalents/cell with split centromeres, N = 27 cells) (Fig. 5c), suggesting that bi-orientation of sister centromeres is frequent in univalents, and this could lead to the frequent observation of precocious sister separation found during anaphase I.
Discussion
In this work, we have characterized MTOPVIB in barley. Mutant plants display an absence of DSB, SC, and CO formation resulting in the occurrence of univalents at metaphase I, unbalanced gametes, and sterility. Hence, HvMTOPVIB-dependent meiotic DSB formation is essential for meiotic recombination. Unlike other plants, MTOPVIB seems not to be required for bipolar meiotic spindle formation in barley.
HvMTOPVIB is critical for meiotic DSB induction in barley
Disruption of HvMTOPVIB results in the absence of meiotic DSB formation and complete sterility, while vegetative growth and development are similar to WT. Reciprocal crosses suggest that both male and female meiosis are defective. No Y2H interaction was found between HvSPO11-1 and HvSPO11-2 while HvMTOPVIB interacts in yeast with both HvSPO11-1 and HvSPO11-2 similar to Arabidopsis and rice (Vrielynck et al., 2016; Xue et al., 2016). The interaction of HvSPO11-1 or HvSPO11-2 with HvMTOPVIB was disrupted in the case of HvmtopVIB, which lacks all four HvMTOPVIB domains conserved among flowering plants including the B4 motif part of the Transducer domain that mediates the interaction with SPO11-1 and SPO11-2 in plants (Vrielynck et al., 2016; Xue et al., 2016). Hence, similar to other plant species, also in barley, MTOPVIB may form a heterotetrameric complex with SPO11-1 and SPO11-2 constituting the meiotic TopoVI-like complex (core catalytic meiotic DSB induction complex) (Robert et al., 2016b). However, whether both HvSPO11-1 and HvSPO11-2 are indeed required for meiotic DSB formation and whether MTOPVIB indeed forms a heterotetrameric complex with SPO11-1 and SPO11-2 is unclear and needs further studies. Future studies are also needed to identify whether HvSPO11-3 is required for meiotic DSB formation or whether HvSPO11-3 is critical for somatic development and not required for meiotic DSB formation similar to AtSPO11-3 (Hartung et al., 2002; Sugimoto-Shirasu et al., 2002; Yin et al., 2002).
Absence of SC and CO formation in HvmtopVIB mutants
In HvmtopVIB, the absence of meiotic DSB formation results in severe meiotic defects that entail complete sterility. Meiotic chromosome axis formation during leptotene and zygotene, including an initial polarization of ASY1 before spreading throughout the nucleus was similar to WT in HvmtopVIB. However, synapsis was abolished in HvmtopVIB. Instead, ZYP1 formed varying numbers of foci that did not elongate. Hence, similar to most species including plants, also synapsis is meiotic DSB-dependent in barley.
Consistent with the absence of meiotic DSBs and accompanied meiotic homologous repair processes, we found no signs of meiotic CO formation. While in WT, seven bivalents are invariably found during metaphase I, in HvmtopVIB, only univalents were evident. The absence of bivalent formation resulted in unbalanced segregation of univalents into aneuploid gametes. Besides, precocious sister chromatid separation during meiosis I (see below) occurred. Notably, in some cells, bivalent-like structures were found at diakinesis/metaphase I. These structures lacked HEI10 foci and were randomly composed of either homologous or mostly non-homologous chromosomes. Due to this, and given the (near) absence of DSB (virtually no γH2AX foci) and SC formation (no ZYP1 polymerization), we consider these bivalent-like structures to be achiasmatic chromosome associations. Taken together, in barley, SC and CO formation requires meiotic DSB formation in an MTOPVIB-dependent manner.
Bipolar spindle and limited polyad formation in HvmtopVIB mutants
Independent of its role in meiotic DSB formation, MTOPVIB is required for bipolar spindle formation during meiosis I in maize and rice (Xue et al., 2019; Jing et al., 2020). Moreover, regular spore formation is impaired in mtopVIB leading to primary polyads (up to octads) with a frequency of ~ 86% in rice (Xue et al., 2019), ~ 63% in maize (Jing et al., 2020), and ~ 70% in A. thaliana (Tang et al., 2017). In rice, at metaphase I, sister kinetochores of univalents were mono-oriented in mtopVIB but not in other DSB-defective mutant plants, where frequent bi-orientation of sister kinetochores was found. Hence, it was suggested that kinetochore bi-orientation in (at least some) univalents at metaphase I is essential for bipolar spindle organization and depends on MTOPVIB (Xue et al., 2019).
In HvmtopVIB meiotic bipolar spindles were found, and similar to the WT, the majority of tetrads were composed of four spores, that however, showed micronuclei and were genetically unbalanced. Thus, MTOPVIB is unlikely to be required for bipolar spindle formation in barley when univalents exist due to the absence of meiotic DSB formation. Furthermore, the frequent occurrence of bi-orientation of sister kinetochores in univalents during metaphase I in HvmtopVIB mutants, when compared with rice or maize mtopVIB univalents, where centromere mono-orientation was prevalent (Xue et al., 2019; Jing et al., 2020), suggests that bipolar spindle formation may indeed depend on sister kinetochore bi-orientation as proposed (Xue et al., 2019).
In rice, MTOPVIB interacts via its C-terminus with PRD2 (Xue et al., 2019) and in Arabidopsis prd2 unequal bipolar or multipolar spindles form (Jiang et al., 2009), suggesting a possible functional interaction of MTOPVIB and PRD2. In HvmtopVIB, despite showing a DSB null phenotype and being the interactions with SPO11-1 and SPO11-2 disrupted in yeast, we cannot exclude that a truncated protein with residual function during metaphase I/kinetochore organization is formed. However, this seems rather unlikely since the predicted truncated HvmtopVIB lacks all conserved domains, including the domain required in rice for the interaction with PRD2 (Xue et al., 2019).
In any case, future studies of further DSB-defective barley mutants including additional mutant alleles of mtopVIB are required to address whether bi-orientation of sister kinetochores in univalents and meiotic bipolar spindle formation are indeed meiotic DSB-independent in barley.
Together, we showed that HvMTOPVIB is critical for meiotic DSB induction and that all accompanied downstream DSB-dependent processes including SC and CO formation are impaired in HvmtopVIB mutants. We assume that this mutant presents a valuable resource for future research on barley meiosis. Moreover, different from other species, MTOPVIB seems not to be required for meiotic bipolar spindle formation in barley.
Author contribution statement
SS, MC, MAA, and CF performed experiments. VS performed super-resolution microscopy. IH, GH and JK designed and cloned transformation vectors and performed genetic engineering. SS, MC, and SH analyzed the data. SH acquired funding. SS, MC, and SH drafted the manuscript. All authors read and approved the final text.
References
An XJ, Deng ZY, Wang T (2011) OsSpo11-4, a rice homologue of the archaeal TopVIA protein, mediates double-strand DNA cleavage and interacts with OsTopVIB. PLoS ONE 6:e20327
Armstrong SJ, Caryl AP, Jones GH, Franklin FCH (2002) Asy1, a protein required for meiotic chromosome synapsis, localizes to axis-associated chromatin in arabidopsis and brassica. J Cell Sci 115:3645–3655
Armstrong SJ, Sanchez-Moran E, Franklin FCH (2009) Cytological analysis of arabidopsis thaliana meiotic chromosomes. In: Keeney S (ed) Meiosis: vol 2, cytological methods. Humana Press, Totowa, pp 131–145
Bellani MA, Boateng KA, McLeod D, Camerini-Otero RD (2010) The expression profile of the major mouse SPO11 isoforms indicates that SPO11β introduces double strand breaks and suggests that SPO11α has an additional role in prophase in both spermatocytes and oocytes. Mol Cell Biol 30:4391–4403
Benyahya F, Nadaud I, Da Ines O, Rimbert H, White C, Sourdille P (2020) SPO11.2 is essential for programmed double-strand break formation during meiosis in bread wheat (Triticum aestivum L.). Plant J 104:30–43
Berchowitz LE, Copenhaver GP (2010) Genetic interference: don’t stand so close to me. Curr Genomics 11:91–102
Berchowitz LE, Francis KE, Bey AL, Copenhaver GP (2007) The role of AtMUS81 in interference-insensitive crossovers in a thaliana. PLOS Genetics 3:e132
Bergerat A, de Massy B, Gadelle D, Varoutas P-C, Nicolas A, Forterre P (1997) An atypical topoisomerase II from archaea with implications for meiotic recombination. Nature 386:414–417
Campell BR, Song Y, Posch TE, Cullis CA, Town CD (1992) Sequence and organization of 5S ribosomal RNA-encoding genes of Arabidopsis thaliana. Gene 112:225–228
Caryl AP, Armstrong SJ, Jones GH, Franklin FCH (2000) A homologue of the yeast HOP1 gene is inactivated in the Arabidopsis meiotic mutant asy1. Chromosoma 109:62–71
Chelysheva L, Vezon D, Chambon A, Gendrot G, Pereira L, Lemhemdi A, Vrielynck N, Le Guin S, Novatchkova M, Grelon M (2012) The Arabidopsis HEI10 Is a New ZMM Protein Related to Zip3. PLOS Genet 8:e1002799
Desjardins SD, Ogle DE, Ayoub MA, Heckmann S, Henderson IR, Edwards KJ, Higgins JD (2020) MutS homologue 4 and MutS homologue 5 maintain the obligate crossover in wheat despite stepwise gene Loss following polyploidization. Plant Physiol 183:1545–1558
Cuacos M, Lambing C, Pachon-Penalba M, Osman K, Armstrong SJ, Henderson IR, Sanchez-Moran E, Franklin FCH, Heckmann S (2021) Meiotic chromosome axis remodelling is critical for meiotic recombination in brassica rapa. J Exp Bot 72:3012–3027
Da Ines O, Abe K, Goubely C, Gallego ME, White CI (2012) Differing requirements for RAD51 and DMC1 in meiotic pairing of centromeres and chromosome arms in Arabidopsis thaliana. PLoS Genet 8:e1002636
Da Ines O, Michard R, Fayos I, Bastianelli G, Nicolas A, Guiderdoni E, White C, Sourdille P (2020) Bread wheat TaSPO11-1 exhibits evolutionarily conserved function in meiotic recombination across distant plant species. Plant J 103:2052–2068
De Muyt A, Pereira L, Vezon D, Chelysheva L, Gendrot G, Chambon A, Lainé-Choinard S, Pelletier G, Mercier R, Nogué F, Grelon M (2009) A high throughput genetic screen identifies new early meiotic recombination functions in arabidopsis thaliana. PLoS Genet 5:e1000654
De Muyt A, Vezon D, Gendrot G, Gallois J-L, Stevens R, Grelon M (2007) AtPRD1 is required for meiotic double strand break formation in Arabidopsis thaliana. EMBO J 26:4126–4137
Fayos I, Meunier AC, Vernet A, Navarro-Sanz S, Portefaix M, Lartaud M, Bastianelli G, Périn C, Nicolas A, Guiderdoni E (2020) Assessment of the roles of SPO11–2 and SPO11–4 in meiosis in rice using CRISPR/Cas9 mutagenesis. J Experimen Botany. 71(22):7046–7058
Fu M, Wang C, Xue F, Higgins J, Chen M, Zhang D, Liang W (2016) The DNA topoisomerase VI-B Subunit OsMTOPVIB Is essential for meiotic recombination initiation in rice. Mol Plant 9:1539–1541
Gerlach WL, Bedbrook JR (1979) Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acids Res 7:1869–1885
Grelon M, Vezon D, Gendrot G, Pelletier G (2001) AtSPO11-1 is necessary for efficient meiotic recombination in plants. EMBO J 20:589–600
Hartung F, Angelis KJ, Meister A, Schubert I, Melzer M, Puchta H (2002) An archaebacterial topoisomerase homolog not present in other eukaryotes is indispensable for cell proliferation of plants. Curr Biol 12:1787–1791
Hartung F, Wurz-Wildersinn R, Fuchs Jr, Schubert I, Suer S, Puchta H (2007) The catalytically active tyrosine residues of both SPO11-1 and SPO11-2 are required for meiotic double-strand break induction in arabidopsis. Plant Cell 19:3090–3099
Hensel G, Valkov V, Middlefell-Williams J, Kumlehn J (2008) Efficient generation of transgenic barley: the way forward to modulate plant–microbe interactions. J Plant Physiol 165:71–82
Higgins JD, Buckling EF, Franklin FCH, Jones GH (2008) Expression and functional analysis of AtMUS81 in arabidopsis meiosis reveals a role in the second pathway of crossing-over. Plant J 54:152–162
Higgins JD, Perry RM, Barakate A, Ramsay L, Waugh R, Halpin C, Armstrong SJ, Franklin FCH (2012) Spatiotemporal asymmetry of the meiotic program underlies the predominantly distal distribution of meiotic crossovers in barley. Plant Cell 24:4096–4109
Higgins JD, Sanchez-Moran E, Armstrong SJ, Jones GH, Franklin FC (2005) The arabidopsis synaptonemal complex protein ZYP1 is required for chromosome synapsis and normal fidelity of crossing over. Genes Dev 19:2488–2500
Hoffie, I, Daghma, D, Mirzakhmedov, M, Chamas, S, Egorova, A, Fontana, I, Hoffie, R, Ehrhardt, M, Marthe, C, Büchner, H, Hiekel, S, Kumlehn, J (2022) CasCADE: a modular and versatile vector system for Cas endonuclease-mediated genome modifications validated in mono- and dicotyledonous plants in preparation
Hu JH, Miller SM, Geurts MH, Tang W, Chen L, Sun N, Zeina CM, Gao X, Rees HA, Lin Z, Liu DR (2018) Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556:57–63
Hunter N (2015) Meiotic Recombination: The Essence of Heredity. Cold Spring Harb Perspect Biol 7:a016618
Jiang H, Wang F-F, Wu Y-T, Zhou X, Huang X-Y, Zhu J, Gao J-F, Dong R-B, Cao K-M, Yang Z-N (2009) MULTIPOLAR SPINDLE 1 (MPS1), a novel coiled-coil protein of Arabidopsis thaliana, is required for meiotic spindle organization. Plant J 59:1001–1010
Jing JL, Zhang T, Kao YH, Huang TH, Wang CJR, He Y (2020) ZmMTOPVIB enables DNA double-strand break formation and bipolar spindle assembly during maize meiosis. Plant Physiol 4:1811–1822
Kauppi L, Barchi M, Baudat F, Romanienko PJ, Keeney S, Jasin M (2011) Distinct properties of the XY pseudoautosomal region crucial for male meiosis. Science 331:916–920
Keeney S, Giroux CN, Kleckner N (1997) Meiosis-specific dna double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell 88:375–384
Koeppel I, Hertig C, Hoffie R, Kumlehn J (2019) Cas endonuclease technology—a quantum leap in the advancement of barley and wheat genetic engineering. Int J Mol Sci 20:2647
Ku J-C, Ronceret A, Golubovskaya I, Lee DH, Wang C, Timofejeva L, Kao Y-H, Gomez Angoa AK, Kremling K, Williams-Carrier R, Meeley R, Barkan A, Cande WZ, Wang C-JR (2020) Dynamic localization of SPO11-1 and conformational changes of meiotic axial elements during recombination initiation of maize meiosis. PLoS Genet 16:e1007881
Kubalová I, Němečková A, Weisshart K, Hřibová E, Schubert V (2021) Comparing Super-Resolution Microscopy Techniques to Analyze Chromosomes. Int J Mol Sci 22:1903
Kumlehn J, Pietralla J, Hensel G, Pacher M, Puchta H (2018) The CRISPR/Cas revolution continues: from efficient gene editing for crop breeding to plant synthetic biology. J Integr Plant Biol 60:1127–1153
Kurzbauer M-T, Pradillo M, Kerzendorfer C, Sims J, Ladurner R, Oliver C, Janisiw MP, Mosiolek M, Schweizer D, Copenhaver GP, Schlögelhofer P (2018) Arabidopsis thaliana FANCD2 Promotes Meiotic Crossover Formation. Plant Cell 30:415–428
Lambing C, Osman K, Nuntasoontorn K, West A, Higgins JD, Copenhaver GP, Yang J, Armstrong SJ, Mechtler K, Roitinger E, Franklin FCH (2015) Arabidopsis PCH2 mediates meiotic chromosome remodeling and maturation of crossovers. PLoS Genet 11:e1005372
Lazo GR, Stein PA, Ludwig RA (1991) A DNA transformation-competent arabidopsis genomic library in agrobacterium. Bio/technology 9:963–967
Li Y, Qin B, Shen Y, Zhang F, Liu C, You H, Du G, Tang D, Cheng Z (2018) HEIP1 regulates crossover formation during meiosis in rice. Proc Natl Acad Sci U S A 115:10810–10815
Liu M, Shi S, Zhang S, Xu P, Lai J, Liu Y, Yuan D, Wang Y, Du J, Yang C (2014) SUMO E3 ligase AtMMS21 is required for normal meiosis and gametophyte development in arabidopsis. BMC Plant Biol 14:153
Mahadevaiah SK, Turner JMA, Baudat F, Rogakou EP, de Boer P, Blanco-Rodríguez J, Jasin M, Keeney S, Bonner WM, Burgoyne PS (2001) Recombinational DNA double-strand breaks in mice precede synapsis. Nat Genet 27:271–276
Marthe C, Kumlehn J, Hensel G (2015) Barley (Hordeum vulgare L) Transformation Using Immature Embryos. Agrobacterium Protocols: Volume 1 Wang, K. Springer, New York, pp 71–83
Mascher M, Wicker T, Jenkins J, Plott C, Lux T, Koh CS, Ens J, Gundlach H, Boston LB, Tulpová Z, Holden S, Hernández-Pinzón I, Scholz U, Mayer KFX, Spannagl M, Pozniak CJ, Sharpe AG, Šimková H, Moscou MJ, Grimwood J, Schmutz J, Stein N (2021) Long-read sequence assembly: a technical evaluation in barley. Plant Cell 33:1888–1906
Mercier R, Mézard C, Jenczewski E, Macaisne N, Grelon M (2015) The Molecular Biology of Meiosis in Plants. Annu Rev Plant Biol 66:297–327
Miao C, Tang D, Zhang H, Wang M, Li Y, Tang S, Yu H, Gu M, Cheng Z (2013) Central region component1, a novel synaptonemal complex component, is essential for meiotic recombination initiation in rice. Plant Cell 25:2998–3009
Nonomura KI, Nakano M, Murata K, Miyoshi K, Eiguchi M, Miyao A, Hirochika H, Kurata N (2004) An insertional mutation in the rice PAIR2 gene, the ortholog of Arabidopsis ASY1, results in a defect in homologous chromosome pairing during meiosis. Mol Genet Genomics 271:121–129
Paull TT, Rogakou EP, Yamazaki V, Kirchgessner CU, Gellert M, Bonner WM (2000) A critical role for histone H2AX in recruitment of repair factors to nuclear foci after DNA damage. Curr Biol 10:886–895
Pyatnitskaya A, Borde V, De Muyt A (2019) Crossing and zipping: molecular duties of the ZMM proteins in meiosis. Chromosoma 128:181–198
Rey MD, Moore G, Martín AC (2018) Identification and comparison of individual chromosomes of three accessions of hordeum chilense, hordeum vulgare, and triticum aestivum by FISH. Genome 61:387–396
Robert T, Nore A, Brun C, Maffre C, Crimi B, Guichard V, Bourbon H-M, de Massy B (2016a) The TopoVIB-like protein family is required for meiotic DNA double-strand break formation. Science 351:943–949
Robert T, Vrielynck N, Mézard C, de Massy B, Grelon M (2016b) A new light on the meiotic DSB catalytic complex. Semin Cell Dev Biol 54:165–176
Rogakou EP, Pilch DR, Orr AH, Ivanova VS, Bonner WM (1998) DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J Biol Chem 273:5858–5868
Sanchez-Moran E, Santos J-L, Jones GH, Franklin FCH (2007) ASY1 mediates AtDMC1-dependent interhomolog recombination during meiosis in arabidopsis. Genes Dev 21:2220–2233
Sanei M, Pickering R, Kumke K, Nasuda S, Houben A (2011) Loss of centromeric histone H3 (CENH3) from centromeres precedes uniparental chromosome elimination in interspecific barley hybrids. Proc Natl Acad Sci U S A 108:E498-505
Schindfessel C, Drozdowska Z, De Mooij L, Geelen D (2021) Loss of obligate crossovers, defective cytokinesis and male sterility in barley caused by short-term heat stress. Plant Reprod 34:243–253
Shen Y, Tang D, Wang K, Wang M, Huang J, Luo W, Luo Q, Hong L, Li M, Cheng Z (2012) ZIP4 in homologous chromosome synapsis and crossover formation in rice meiosis. J Cell Sci 125:2581–2591
Sprink T, Hartung F (2014) The splicing fate of plant SPO11 genes. Front Plant Sci 5:214
Sprink T, Hartung F (2021) Heterologous complementation of SPO11-1 and -2 depends on the splicing pattern. Int J Mol Sci 22:9346
Stacey NJ, Kuromori T, Azumi Y, Roberts G, Breuer C, Wada T, Maxwell A, Roberts K, Sugimoto-Shirasu K (2006) Arabidopsis SPO11-2 functions with SPO11-1 in meiotic recombination. Plant J 48:206–216
Stronghill P, Pathan N, Ha H, Supijono E, Hasenkampf C (2010) Ahp2 (Hop2) function in arabidopsis thaliana (Ler) is required for stabilization of close alignment and synaptonemal complex formation except for the two short arms that contain nucleolus organizer regions. Chromosoma 119:443–458
Sugimoto-Shirasu K, Stacey NJ, Corsar J, Roberts K, McCann MC (2002) DNA topoisomerase VI Is essential for endoreduplication in arabidopsis. Curr Biol 12:1782–1786
Szurman-Zubrzycka M, Baran B, Stolarek-Januszkiewicz M, Kwaśniewska J, Szarejko I, Gruszka D (2019) The dmc1 mutant allows an insight into the DNA double-strand break repair during meiosis in barley (Hordeum vulgare L.). Front Plant Sci 10:761
Tang Y, Yin Z, Zeng Y, Zhang Q, Chen L, He Y, Lu P, Ye D, Zhang X (2017) MTOPVIB interacts with AtPRD1 and plays important roles in formation of meiotic DNA double-strand breaks in Arabidopsis. Sci Rep 7:10007
The International Barley Genome Sequencing Consortium (2012) A physical, genetic and functional sequence assembly of the barley genome. Nature 491:711–716
Vrielynck N, Chambon A, Vezon D, Pereira L, Chelysheva L, De Muyt A, Mézard C, Mayer C, Grelon M (2016) A DNA topoisomerase VI–like complex initiates meiotic recombination. Science 351:939–943
Vrielynck N, Schneider K, Rodriguez M, Sims J, Chambon A, Hurel A, De Muyt A, Ronceret A, Krsicka O, Mézard C, Schlögelhofer P, Grelon M (2021) Conservation and divergence of meiotic DNA double strand break forming mechanisms in Arabidopsis thaliana. Nucleic Acids Res 49:9821–9835
Wang K, Wang M, Tang D, Shen Y, Miao C, Hu Q, Lu T, Cheng Z (2012) The role of rice HEI10 in the formation of meiotic crossovers. PLoS Genet 8:e1002809
Wang Y, Copenhaver GP (2018) Meiotic recombination: mixing it up in plants. Annu Rev Plant Biol 69:577–609
Weisshart K, Fuchs J, Schubert V (2016) Structured illumination microscopy (SIM) and photoactivated localization microscopy (PALM) to analyze the abundance and distribution of RNA polymerase ii molecules on flow-sorted arabidopsis nuclei. Bio-Protoc 6:e1725
Wojtasz L, Daniel K, Roig I, Bolcun-Filas E, Xu H, Boonsanay V, Eckmann CR, Cooke HJ, Jasin M, Keeney S, McKay MJ, Toth A (2009) Mouse HORMAD1 and HORMAD2, two conserved meiotic chromosomal proteins, are depleted from synapsed chromosome axes with the help of TRIP13 AAA-ATPase. PLoS Genet 5:e1000702
Wu Z, Ji J, Tang D, Wang H, Shen Y, Shi W, Li Y, Tan X, Cheng Z, Luo Q (2015) OsSDS is essential for DSB formation in rice meiosis. Front Plant Sci. https://doi.org/10.3389/fpls.2015.00021
Xue Z, Li Y, Zhang L, Shi W, Zhang C, Feng M, Zhang F, Tang D, Yu H, Gu M, Cheng Z (2016) OsMTOPVIB promotes meiotic DNA double-strand break formation in rice. Mol Plant 9:1535–1538
Xue Z, Liu C, Shi W, Miao Y, Shen Y, Tang D, Li Y, You A, Xu Y, Chong K, Cheng Z (2019) OsMTOPVIB is required for meiotic bipolar spindle assembly. Proc Natl Aca Sci 116:201821315
Yin Y, Cheong H, Friedrichsen D, Zhao Y, Hu J, Mora-Garcia S, Chory J (2002) A crucial role for the putative arabidopsis topoisomerase VI in plant growth and development. Proc Natl Acad Sci 99:10191–10196
Yu H, Wang M, Tang D, Wang K, Chen F, Gong Z, Gu M, Cheng Z (2010) OsSPO11-1 is essential for both homologous chromosome pairing and crossover formation in rice. Chromosoma 119:625–636
Ahn Yun-Jae, Cuacos Maria, Ayoub Mohammad A, Kappermann Judith, Houben Andreas, Heckmann Stefan (2020) In planta delivery of chemical compounds into barley meiocytes: EdU as compound example. In: Pradillo Mónica, Heckmann Stefan (eds) plant meiosis: methods and protocols. Springer New York, New York, NY, pp 381–402. https://doi.org/10.1007/978-1-4939-9818-0_27
Zhang C, Song Y, Cheng Z-H, Wang Y-X, Zhu J, Ma H, Xu L, Yang Z-N (2012) The arabidopsis thaliana DSB formation (AtDFO) gene is required for meiotic double-strand break formation. Plant J 72:271–281
Acknowledgements
We are grateful to Jana Lorenz, Franziska Hartmann, Sabine Sommerfeld and Marius Doelling for excellent technical assistance, to the IPK gardeners for horticultural help, to Zhukuan Cheng for kindly providing the γH2AX antibodies, to Andreas Houben for kindly providing the CENH3 antibodies, and to all lab members for fruitful discussions. We would also like to thank the German Federal Ministry of Education and Research (BMBF) for financial support in the frame of the grant HERBY (FKZ-031B0188). Costs for open access publishing were partially funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation, grant 491250510).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Author notes
Stefan Steckenborn and Maria Cuacos have contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Communicated by Raphael Mercier.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Steckenborn, S., Cuacos, M., Ayoub, M.A. et al. The meiotic topoisomerase VI B subunit (MTOPVIB) is essential for meiotic DNA double-strand break formation in barley (Hordeum vulgare L.). Plant Reprod 36, 1–15 (2023). https://doi.org/10.1007/s00497-022-00444-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00497-022-00444-5