Abstract
Spo11 is a homolog of a subunit of archaebacterial topoisomerase, which catalyzes DNA double-strand breaks and initiates homologous chromosome recombination. In the present study, we silenced the SPO11-1 gene in rice (Oryza sativa) using RNAi. Rice plants with loss-of-function of OsSPO11-1 have no apparent growth defects during vegetative development, but homologous chromosome pairing and recombination are significantly obstructed. Telomeres can be assembled as bouquet during the zygotene stage of the OsSPO11-1-deficient plants, just as that in wild type. Although the two axial-associated proteins, REC8 and PAIR2, are loaded onto the chromosomes, the depletion of PAIR2 from the chromosomes is much later than in wild type. The central element of the synaptonemal complex (SC), ZEP1, does not load onto the chromosomes normally, implying that SC formation is disturbed severely. The crossover protein, MER3, isn't efficiently assembled onto chromosomes and the lack of bivalent suggests that crossovers are also affected in the absence of OsSPO11-1. Thus, OsSPO11-1 is essential for both homologous chromosomes pairing and crossover formation during meiosis in rice.







Similar content being viewed by others
References
Allers T, Lichten M (2001) Differential timing and control of noncrossover and crossover recombination during meiosis. Cell 106:47–57
Bailis JM, Roeder GS (1998) Synaptonemal complex morphogenesis and sister-chromatid cohesion require Mek1-dependent phosphorylation of a meiotic chromosomal protein. Genes Dev 12:3551–3563
Bergerat A, de Massy B, Gadelle D, Varoutas PC, Nicolas A, Forterre P (1997) An atypical topoisomerase II from Archaea with implications for meiotic recombination. Nature 386:414–417
Brar GA, Kiburz BM, Zhang Y, Kim JE, White F, Amon A (2006) Rec8 phosphorylation and recombination promote the step-wise loss of cohesins in meiosis. Nature 441:532–536
De Muyt A, Vezon D, Gendrot G, Gallois JL, Stevens R, Grelon M (2007) AtPRD1 is required for meiotic double strand break formation in Arabidopsis thaliana. EMBO J 26:4126–4137
De Muyt A, Pereira L, Vezon D, Chelysheva L, Gendrot G, Chambon A, Laine-Choinard S, Pelletier G, Mercier R, Nogue F, Grelon M (2009) A high throughput genetic screen identifies new early meiotic recombination functions in Arabidopsis thaliana. PLoS Genet 5:e1000654
Dernburg AF, McDonald K, Moulder G, Barstead R, Dresser M, Villeneuve AM (1998) Meiotic recombination in C. elegans initiates by a conserved mechanism and is dispensable for homologous chromosome synapsis. Cell 94:387–398
Grelon M, Vezon D, Gendrot G, Pelletier G (2001) AtSPO11-1 is necessary for efficient meiotic recombination in plants. EMBO J 20:589–600
Jain M, Tyagi AK, Khurana JP (2006) Overexpression of putative topoisomerase 6 genes from rice confers stress tolerance in transgenic Arabidopsis plants. FEBS J 273:5245–5260
Keeney S, Giroux CN, Kleckner N (1997) Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell 88:375–384
Kelly KO, Dernburg AF, Stanfield GM, Villeneuve AM (2000) Caenorhabditis elegans msh-5 is required for both normal and radiation-induced meiotic crossing over but not for completion of meiosis. Genetics 156:617–630
Klein F, Mahr P, Galova M, Buonomo SB, Michaelis C, Nairz K, Nasmyth K (1999) A central role for cohesins in sister chromatid cohesion, formation of axial elements, and recombination during yeast meiosis. Cell 98:91–103
Kugou K, Fukuda T, Yamada S, Ito M, Sasanuma H, Mori S, Katou Y, Itoh T, Matsumoto K, Shibata T et al (2009) Rec8 guides canonical Spo11 distribution along yeast meiotic chromosomes. Mol Biol Cell 20:3064–3076
Li W, Ma H (2006) Double-stranded DNA breaks and gene functions in recombination and meiosis. Cell Res 16:402–412
Liu QQ, Zhang JL, Wang ZY, Hong MM, Gu MH (1998) A highly efficient transformation system mediated by Agrobacterium tumefaciens in rice (Oryza sativa L.). Acta Phytophysiologica Sinica 24:259–271
Mahadevaiah SK, Turner JM, Baudat F, Rogakou EP, de Boer P, Blanco-Rodriguez J, Jasin M, Keeney S, Bonner WM, Burgoyne PS (2001) Recombinational DNA double-strand breaks in mice precede synapsis. Nat Genet 27:271–276
Maleki S, Neale MJ, Arora C, Henderson KA, Keeney S (2007) Interactions between Mei4, Rec114, and other proteins required for meiotic DNA double-strand break formation in Saccharomyces cerevisiae. Chromosoma 116:471–486
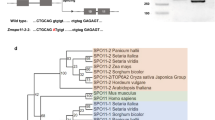
Malik SB, Ramesh MA, Hulstrand AM, Logsdon JM Jr (2007) Protist homologs of the meiotic Spo11 gene and topoisomerase VI reveal an evolutionary history of gene duplication and lineage-specific loss. Mol Biol Evol 24:2827–2841
McEachern MJ, Krauskopf A, Blackburn EH (2000) Telomeres and their control. Annu Rev Genet 34:331–358
McKim KS, Hayashi-Hagihara A (1998) mei-W68 in Drosophila melanogaster encodes a Spo11 homolog: evidence that the mechanism for initiating meiotic recombination is conserved. Genes Dev 12:2932–2942
Mercier R, Jolivet S, Vezon D, Huppe E, Chelysheva L, Giovanni M, Nogue F, Doutriaux MP, Horlow C, Grelon M, Mezard C (2005) Two meiotic crossover classes cohabit in Arabidopsis: one is dependent on MER3, whereas the other one is not. Curr Biol 15:692–701
Murakami H, Keeney S (2008) Regulating the formation of DNA double-strand breaks in meiosis. Genes Dev 22:286–292
Nakagawa T, Ogawa H (1999) The Saccharomyces cerevisiae MER3 gene, encoding a novel helicase-like protein, is required for crossover control in meiosis. EMBO J 18:5714–5723
Nichols MD, DeAngelis K, Keck JL, Berger JM (1999) Structure and function of an archaeal topoisomerase VI subunit with homology to the meiotic recombination factor Spo11. EMBO J 18:6177–6188
Nonomura K, Nakano M, Eiguchi M, Suzuki T, Kurata N (2006) PAIR2 is essential for homologous chromosome synapsis in rice meiosis I. J Cell Sci 119:217–225
Osman F, Dixon J, Doe CL, Whitby MC (2003) Generating crossovers by resolution of nicked Holliday junctions: a role for Mus81-Eme1 in meiosis. Mol Cell 12:761–774
Romanienko PJ, Camerini-Otero RD (2000) The mouse Spo11 gene is required for meiotic chromosome synapsis. Mol Cell 6:975–987
Sanchez-Moran E, Santos JL, Jones GH, Franklin FC (2007) ASY1 mediates AtDMC1-dependent interhomolog recombination during meiosis in Arabidopsis. Genes Dev 21:2220–2233
Scherthan H (2007) Telomere attachment and clustering during meiosis. Cell Mol Life Sci 64:117–124
Smith AV, Roeder GS (1997) The yeast Red1 protein localizes to the cores of meiotic chromosomes. J Cell Biol 136:957–967
Stacey NJ, Kuromori T, Azumi Y, Roberts G, Breuer C, Wada T, Maxwell A, Roberts K, Sugimoto-Shirasu K (2006) Arabidopsis SPO11-2 functions with SPO11-1 in meiotic recombination. Plant J 48:206–216
Sugawara H, Iwabata K, Koshiyama A, Yanai T, Daikuhara Y, Namekawa SH, Hamada FN, Sakaguchi K (2009) Coprinus cinereus Mer3 is required for synaptonemal complex formation during meiosis. Chromosoma 118:127–139
Sugimoto-Shirasu K, Stacey NJ, Corsar J, Roberts K, McCann MC (2002) DNA topoisomerase VI is essential for endoreduplication in Arabidopsis. Curr Biol 12:1782–1786
Sym M, Roeder GS (1995) Zip1-induced changes in synaptonemal complex structure and polycomplex assembly. J Cell Biol 128:455–466
Tanaka K, Miyamoto N, Shouguchi-Miyata J, Ikeda JE (2006) HFM1, the human homologue of yeast Mer3, encodes a putative DNA helicase expressed specifically in germ-line cells. DNA Seq 17:242–246
Wang K, Tang D, Wang M, Lu J, Yu H, Liu J, Qian B, Gong G, Wang X, Chen J et al (2009) MER3 is required for normal meiotic crossover formation, but not for presynaptic alignment in rice. J Cell Sci 122:2055–2063
Wang M, Wang K, Tang D, Wei C, Li M, Shen Y, Chi Z, Gu M, Cheng Z (2010) The central element protein ZEP1 of synaptonemal complex regulates the number of crossovers during meiosis in rice. Plant Cell 22:417–430
Watanabe Y, Nurse P (1999) Cohesin Rec8 is required for reductional chromosome segregation at meiosis. Nature 400:461–464
Wojtasz L, Daniel K, Roig I, Bolcun-Filas E, Xu H, Boonsanay V, Eckmann CR, Cooke HJ, Jasin M, Keeney S et al (2009) Mouse HORMAD1 and HORMAD2, two conserved meiotic chromosomal proteins, are depleted from synapsed chromosome axes with the help of TRIP13 AAA-ATPase. PLoS Genet 5:e1000702
Zhang D, Yang Q, Bao W, Zhang Y, Han B, Xue Y, Cheng Z (2005a) Molecular cytogenetic characterization of the Antirrhinum majus genome. Genetics 169:325–335
Zhang W, Yi C, Bao W, Liu B, Cui J, Yu H, Cao X, Gu M, Liu M, Cheng Z (2005b) The transcribed 165-bp CentO satellite is the major functional centromeric element in the wild rice species Oryza punctata. Plant Physiol 139:306–315
Acknowledgement
We thank Jason G. Walling and Jiming Jiang for critical reading of the manuscript. This work was supported by grants from the Ministry of Sciences and Technology of China (2008ZX08009), and the National Natural Science Foundation of China (30770131 and 30921061).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by S. Keeney
Hengxiu Yu and Mo Wang contributed equally to this work
Electronic supplementary materials
ESM Fig. S1
The T-DNA structure of the binary vector pBSPO11. OCS, terminator of octopinesynthase gene; Act1, promoter of the rice actin 1 gene; CaMV 35S pro and 35S polyA, promoter and terminator of CaMV 35S gene; nptII, neomycin phosphotransferase gene; RB and LB, right and left borders of T-DNA region, respectively. (GIF 10 kb; GIF 2 kb)
ESM Fig. S2
PCR analyses of the transgenic rice plants. Lane M, DNA molecular marker. Lane P, plasmid pBSPO11. Lane W, nontransformed wild-type Yandao 8. Lanes 1 and 2, two transgenic plants. (GIF 9 kb)
ESM Fig. S3
Down regulation of OsSPO11-1 gene in the transgenic plants. UBQ gene amplification was used as the positive control for each sample. Line WT, the wild-type plant of Yandao 8. Lanes 1 and 2, two transgenic plants from two independent transformants. (GIF 10 kb)
Rights and permissions
About this article
Cite this article
Yu, H., Wang, M., Tang, D. et al. OsSPO11-1 is essential for both homologous chromosome pairing and crossover formation in rice. Chromosoma 119, 625–636 (2010). https://doi.org/10.1007/s00412-010-0284-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-010-0284-7