Abstract
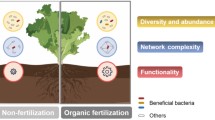
Organic farming can enhance biodiversity and soil health and is a sustainable alternative to conventional farming. Yet, soil protists especially protistan predators, have received inadequate attention, and their contributions to the sustainability of organic farming remained underexplored. In this study, we examined soil microbial communities from 379 samples, including both organic and chemically fertilized soils from China. Our findings revealed higher bacterial diversity and increases in plant-beneficial bacteria in organically farmed soils. Notably, organic farming systems facilitated dynamic predator-prey interactions, which may be disrupted by the application of chemical fertilizers. Additionally, organic farming enriched protistan predators, enhancing the relative abundance of functional PGPR, thus improving soil health. We further conducted a case study highlighting the critical role of organic matter in sustaining protistan predator populations and their interactions with bacteria. We propose the crucial contributions of organic inputs for supporting protistan predators and the interplay of predator-prey, ultimately enhancing soil functions and promoting agricultural sustainability.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Conventional agricultural practices, relying heavily on intensive cropping, pesticides, and chemical fertilizers, play a crucial role in enhancing crop production to meet the growing food demands of the increasing human population (Azarbad 2022; Gabriel et al. 2013). However, agricultural intensification derived from for example overuse of chemical fertilizers in conventional agricultural practices, has led to potential soil issues, including accumulation of plant pathogens, severe soil degradation, and losses of biodiversity and as well as ecosystem services (Kumar Bhatt et al. 2019; Lichtenberg et al. 2017; Sánchez-Bayo and Wyckhuys 2019). Organic farming practices supported by organic fertilizers (such as composts and straws) are sustainable alternatives to conventional farming in food production and strive for environmental benefits (Gomiero et al. 2011). The use of organic fertilizers can provide external nutrients to soils similar to their chemical counterparts and also enhance the soil ecosystem's capacity to retain nutrients in agriculture (Bhogal et al. 1997; Diacono and Montemurro 2011; Siedt et al. 2021). Additionally, organic farming can increase soil organic matter content, positively affecting soil physical properties and structure, which benefits plant growth (Lehmann et al. 2020). For example, organic farming can ameliorate the negative impacts of soil compaction caused by intensive tillage and promote plant root growth (Abdollahi et al. 2014; Bu et al. 2023). Furthermore, organic matter amendments can increase water holding capacity and water use efficiency, which are crucial for plant growth (Wang et al. 2014).
Organic farming practices can promote the development of a diverse and active microbial community in soils (Maeder et al. 2002), which drives various soil functions (Luo et al. 2018; Naylor et al. 2020) and sustain soil health (Dubey et al. 2019). It has been reported that the use of organic fertilizers can increase the microbial biomass C and N (Rong et al. 2018) as well as bacterial and fungal diversities (Bebber and Richards 2022; Luo et al. 2023; Ma et al. 2023) compared to chemical fertilizer in soils. Organic farming can also improve plant performance by inducing compositional shifts in microbial communities residing in the rhizosphere of plants and fostering the interactions between soil microbiomes and plants (Li et al. 2024b; Liu et al. 2018; Röös et al. 2018; Schmidt et al. 2019). For example, the application of organic fertilizers, particularly when combined with beneficial microorganisms, could enhance soil suppression against plant pathogens by enriching pathogen-inhibiting microorganisms (Chen et al. 2020; Deng et al. 2022). Nevertheless, existing research has primarily focused on bacteria and fungi when investigating and comparing soil microbial communities between organic farming soils and their conventional counterparts.
Soil protists, an often overlooked but important component of the soil microbial communities, present wide functional diversities including predators, phototrophs, animal parasites, and plant pathogens (Gao et al. 2019; Geisen et al. 2018). Soil protists, in particular predatory protists, can function as dynamic hubs within microbial communities and drive the microbial composition in soil ecosystems (Chowdhury et al. 2022; Fujino et al. 2023) and thereby microbiome functions via top-down regulations (Gao et al. 2019; Geisen et al. 2018). Predatory protists through selective predatory activities on their microbial preys contribute to various ecological functions in soils, such as soil nutrient cycling and plant performance (Geisen et al. 2018; Nguyen et al. 2023; Thakur and Geisen 2019). These microbiome predators are also directly affected by the changes in the physicochemical properties of soils (Hu et al. 2022; Oliverio et al. 2020; Xiong et al. 2021). Protistan predators are more sensitive to organic fertilizer application in agricultural ecosystems compared with other microbial groups (e.g., bacteria and fungi) (Guo et al. 2022). The applications of organic fertilizers may trigger changes in the diversity and taxonomic as well as functional compositions of soil protist communities (Guo et al. 2018; Ren et al. 2023). This is expected to enhance the suppression against soil-borne plant pathogens by directly consuming pathogens and indirectly increasing the presence of pathogen-suppressive bacteria in the soil by predatory activities of soil protists (Guo et al. 2022, 2024). Despite the important role of protists in agricultural ecosystems and their high sensitivity to agricultural management, only very few studies have investigated the influence of different fertilization regimes on the diversity and composition of soil protists, including protistan predators, as well as on the difference in top-down regulations by protistan predators over bacteria.
In this work, we selected 379 soil samples where both bacterial and protistan amplicon data are available from diverse agricultural systems (i.e., 200 organic fertilizer-applied soils and 179 conventional chemical fertilizer-applied soils) in China (Xiong et al. 2021). We hypothesized that 1) organic fertilizer farming can enrich soil protistan predators and orchestrate bacterial community compositions and functionalities; 2) organic farming soils can lead to a shift in trophic structure between protistan predators and bacteria within the soil microbiome; and 3) variations in soil properties mainly organic matters in soils resulting from different fertilizer managements contribute to changes in protistan predators and their interrelationships with bacteria. Here, we explored the often overlooked protistan predators and their interrelationships with bacteria in soils under different fertilizer regimes and aimed to (at least partly) explain important ecological benefits in sustainable agriculture from a protist standpoint.
Materials and methods
Datasets collection
Datasets collection for bacterial communities (by 16S rRNA genes) and protistan communities (by 18S rRNA genes) were carried out through a meta-analysis in previous research (Xiong et al. 2021). Our focus in this study was comparing the diversity and functional profiles of microbial communities (i.e., bacterial and protistan communities) between organic (200 samples) and conventional (179 samples) farming systems (Xiong et al. 2021). We thus further selected five datasets comprising in total of 379 soil samples obtained from two agricultural ecosystems (i.e., conventional agricultural ecosystem and organic agricultural ecosystem) across China, including three already published works (Guo et al. 2021; Qiao et al. 2019; Wei et al. 2019) and two still on-going not yet published projects (please see details including their geographic locations, cultivation histories and experiment designs in Table S1). Amplicon raw sequencing raw data have been deposited under the Sequence Read Archive accession number SRP090147 in the DDBJ, under the BioProject PRJNA525676 and PRJNA599073 in the NCBI.
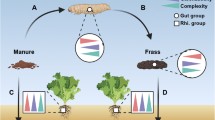
In addition, we selected a long-term continuous cropping field located at the Nanjing Institute of Vegetable Science, Jiangsu province, China (31° 43′ N, 118° 46′ E) as a case study to examine how soil properties resulting from different fertilizer management practices contribute to changes in bacteria and protistan predators (Guo et al. 2021). The field underwent three years of continuous cucumber cultivation from March 2014 to October 2016 under organic and chemical fertilizer application. In this case study, soil pH, organic matter (OM), total nitrogen (TN), total phosphorus (TP), total potassium (TK), available nitrogen (AN), available phosphorus (AP), and available potassium (AK) were measured (Table S5).
Bioinformatic analyses for amplicon data
All the 379 samples were reanalyzed using a unified pipeline in the study. In brief, pair-end reads were merged with USEARCH v11 (Edgar 2010), and merged sequences for each sample with expected errors > 1.0 or a length shorter than 300 bp (protist V4 region) or 200 bp (bacteria V4 region) were discarded. The two primer sets: 616*f (TTAAARV GYTCGTAGTYG)/TAReukREV3 (ACTTTCGTTCTTGATYRA), and 520F (AYTGGGYDTAAAGNG) / (TACNVGGGTATC TAATCC) were utilized to extract the targeted protist V4 and bacterial V4 regions by the “search_pcr2” command in USEARCH, respectively. To obtain an equivalent sequencing depth for later analyses, we rarefied each sample to 10,000 reads (but kept the samples if they did not contain 10,000 reads). Both bacterial and protistan ASV tables were further rarefied to 1,000 sequences for each sample to calculate the diversity and community composition of bacteria and protists.
We then identified amplicon sequence variants (ASV) or so-called “zOTU” with UNOISE3 (Edgar 2016) methods in USEARCH, with chimeras being removed. Obtained reprehensive sequences of ASVs were further taxonomically annotated utilizing PR2 databases (Guillou et al. 2012) for 18S rRNA sequences to obtain protist sequence annotation and RDP databases for 16S rRNA sequences to obtain bacterial sequence annotation. We selected richness (number of observed ASV) as a commonly used biodiversity metric to evaluate the diversity of bacteria and protists in different agricultural ecosystems.
Functional prediction for bacterial and protist communities
Protistan ASVs were then assigned into different functional groups according to their predicted feeding mode (i.e., phagotrophs, parasites, phototrophs, plant pathogens, and saprotrophs) (Adl et al. 2019; Xiong et al. 2021). Potential bacterial pathogens were predicted by aligning bacteria ASV sequences against the MBPD database which was designed to detect bacterial pathogens for One Health practices (Yang et al. 2023) utilizing the usearch_global command in VSEARCH (Li et al. 2023a) with identity parameter 0.99. Putative Plant Growth-Promoting Rhizobacteria (PGPR) in bacterial communities were predicted utilizing a database derived from Milke et al. (2024) via also a similarity-based method with an identity parameter setting to 0.97.
The Functional Annotation of Prokaryotic Taxa (FAPROTAX, http://mem.rcees.ac.cn:8080/) pipeline was used to extrapolate bacterial community functions relating to nutrient cycling (i.e., C, N, and S in the study) (Djemiel et al. 2022; Ling et al. 2022; Louca et al. 2016). For P cycling pathways of bacterial communities, those genes that were reported to be engaged in P cycling pathways (Zeng et al. 2022) were retrieved using KO identifiers from Tax4Fun2 analysis (Wemheuer et al. 2020). Characteristics of bacterial community related to dormancy potential and growth rate have important implications for the restoration and maintenance of functionality of ecosystems (Ling et al. 2022). The dormancy potential of bacterial communities thus was measured by summing the relative abundances of genes [K07699 (spo0A), K10715 (rpfC), K07154 (hipA), K03830 (yafP), K07172 (mazE), K06218 (relE), K01451 (hipO), K07473 (dinJ), K07171 (mazF), and K00951 (relA)] that confer dormancy strategies in different agricultural management (Kearns and Shade 2018) in Tax4Fun2 analysis. We also calculated the weighted mean ribosomal operon copy number (rrn) of each bacterial community to indicate community-level growth rate by summing the relative abundance of each ASV multiplied by its potential ribosomal operon copy number that was obtained by the PICRUSt2 pipeline (Douglas et al. 2020).
Statistical analyses
We employed the Wilcoxon test in R (version 4.2.2) (if not otherwise specified) to contrast the differences in different parameters between organic and conventional farming soil samples. Bray–Curtis distance metrics calculated based on the relative abundance of protist or bacterial ASVs were utilized to explore microbial community dissimilarity between organic and conventional farming soil samples. For a comprehensive comparison of the dissimilarity, one-way analyses of variance (ANOVA) were executed, followed by Tukey’s HSD tests for multiple comparisons of community dissimilarity. The STAMP package (Parks et al. 2014) was used to compare the differences in the relative abundance of main bacterial genera as well as normalized soil properties between organic and conventional farming soil samples in our case study.
We quantified the relative importance of the abundant bacterial genera in predicting the relative abundance of protistan predators based on the percentage increase in mean square error (%IncMSE) in random forest regression implemented by the “randomForest” package in R (Breiman 2001). Spearman’s rank correlations were computed correlations between the abundant bacterial genera and protistan predators, as well as soil properties and microbial parameters. We performed the redundancy analysis (RDA) in the case study to find associations between soil properties and protistan functional groups. Linear regression analysis was performed using the “lm” function in R. Spearman’s rank correlation analysis was performed to decipher the interrelationships between genera of protistan predators and bacteria by controlling for the influence of organic matter concentration, utilizing the “pcor.test” function from the “ppcor” package (Kim 2015).
Results and Discussion
Differences in bacterial communities under the two agricultural ecosystems
Bacterial richness in organic farming soils was significantly higher than that in conventional farming soils (Fig. 1a; p < 0.001, Wilcoxon test). This finding aligns with a previous meta-analysis (Bebber and Richards 2022) and a long-term field experiment (Hartmann et al. 2015), both demonstrating that organic farming led to a higher richness of soil microbiome in comparison to conventional farming. The increased prokaryote diversity under the usage of organic fertilizers can be partly attributed to ample nutrients provided by carbon-rich contents from organic fertilizers for thriving bacterial communities (Li et al. 2017) and also results from changes in soil physicochemical properties. For example, the usage of organic fertilizers in soils increased pH to a level that supports a higher diversity of soil bacteria compared to chemical fertilizers that acidified soils (Fig. S7) (Heinze et al. 2010; Laurent et al. 2020; Shen et al. 2020). Organic farming soils were exhibited to have more heterogeneous communities of both bacteria and protistan predators than conventional farming soils (Fig. S1). This could be explained by the reduction of ecological niches due to intensive agriculture with long-term chemical fertilizer application, which led to a homogenization of soil microbial communities (Lupatini et al. 2017).
Bacterial diversity and the main bacterial taxa in the two fertilization regimes. The richness (number of observed ASVs) of bacteria in organic fertilizer and chemical fertilizer soils (a) and the differences of the main bacterial genera in organic fertilizer and chemical fertilizer soils by STAMP analysis (b); the relative abundance of putative PGPR in organic fertilizer and chemical fertilizer soils (c). In panel b, only the significantly (Wilcoxon test, p < 0.050) differential bacterial genera between organic fertilizer and chemical fertilizer soils were shown. In panel c, “*” means p < 0.05, “**” means p < 0.01 and “***” means p < 0.001 by Wilcoxon test
We then focused on the impact of the two agricultural systems on specific bacterial genera (Fig. 1 and Table S1). Our results indicated significant enrichment of Gp6 from Acidobacteria, Lysobacter, Neobacillus, and Tumebacillus in organic farming soils compared to conventional farming systems (Fig. 1b, p < 0.050, Wilcoxon test). These results are in line with previous work showing a high relative abundance of the subgroup Gp6 in soils with high levels of organic matter (Navarrete et al. 2010). Additionally, these results also corroborated our prior observations that Lysobacter was enriched in soils treated with organic fertilizer and bio-organic fertilizers (Xiong et al. 2017), which is in accordance with the capacity of Lysobacter species to decompose organic matter (Reichenbach 2006). The two bacterial genera, Gp6 (Rosenzweig et al. 2012) and Lysobacter (Folman et al. 2004), enriched by organic farming, may be also indicative of soil health in agricultural ecosystems. In addition, two bacterial genera from the Bacillales order, namely Neobacillus and Tumebacillus, enriched by organic fertilizer inputs in this study, have been reported to also contribute to multiple important soil ecological functions, including enhancing nutrient cycling and conferring (abiotic and biotic) stress tolerance to plants (Saxena et al. 2020).
Potential bacterial pathogens (Table S2) and putative PGPR (Table S3 and Fig. S2) were identified in the two agricultural systems. The relative abundances of overall and each type of putative bacterial PGRP were significantly higher in organic farming ecosystems (Fig. 1c, p < 0.001, Wilcoxon test) with an exception from PGPR with the biostimulation effects (p =0.998, Wilcoxon test). The relative abundance of the overall PGPR showed opposite significant linear relationships with that of potential plant pathogens under the two farming systems (Fig. 3a). All three main PGPR bacteria types (putative PGPR with bioprotection effects, biofertilization effects, and biostimulation effects) showed potential phytopathogenic bacteria suppression capacity in the organic farming system as they established significantly negative relationships with bacterial plant pathogens while only bioprotection PGPR did so in conventional farming systems (Fig. S3). This is because different crop management leads to physiological differences in potential PGPR (Melo et al. 2016), and partly explains the opposite response of PGPR to potential plant pathogens in the two agricultural systems.
Differences in protistan communities under the two agricultural ecosystems
The richness of entire protists exhibited a decline in organic farming soils compared to conventional farming soils (Fig. 2a, p < 0.001, Wilcoxon test), while the richness of protistan predators remained stable (Fig. 2b, p = 0.077, Wilcoxon test). The decreased richness of entire protists in organic farming soils compared to conventional farming soils might be explained by the notable reduction in protist plant pathogens (Fig. 2c, p < 0.050, Wilcoxon test) and phototrophic protists (Fig. 2c, p < 0.001, Wilcoxon test). These findings underscore the capacity of organic amendments to bolster soil health by reducing protistan plant pathogens such as Pythium (Xiong et al. 2018).
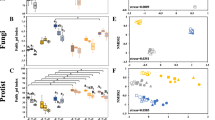
The diversity and main functional compositions of protists and the potential interrelationships between protistan predators and bacterial preys in the two fertilization regimes. The richness of entire protists in organic fertilizer and chemical fertilizer soils (a); the richness of protistan predators in organic fertilizer and chemical fertilizer soils (b); the main protistan functional groups in organic fertilizer and chemical fertilizer soils (c); the relative importance of the main bacterial genera in predicting the relative abundance of protistan predators and Spearman’s rank correlations between each bacterial genus and protistan predators across all the samples (d); the linear relationship between the richness of bacteria and the relative abundance of protistan predators in organic fertilizer and chemical fertilizer soils (e). In panel c, “*” means p < 0.05, “**” means p < 0.01 and “***” means p < 0.001 by Wilcoxon test; the “↑” indicates the enrichment and “↓” indicates the deletion of protistan functional groups in organic fertilizer compared with chemical fertilizer soils. In panel d, %IncMSE (percentage increase in the Mean Squared Error) indicates the relative importance of the variables
It is noteworthy that protistan predators occupied a dominant position in total protistan communities in both organic farming and conventional agricultural soils, aligning with globally reported biogeographic patterns of soil protistan communities (Oliverio et al. 2020). Our study further exhibited that organic farming primarily influenced the composition rather than the diversity of predatory protists. Specifically, we observed a significant enrichment in the relative abundance of protistan predators in organic farming soils (Fig. 2c, p < 0.001, Wilcoxon test), with an average value of 66.85%, in contrast to conventional farming soils with an average value of 60.28%. This observation is in line with prior research indicating that long-term application of organic fertilizers substantially increases the proportions of protistan predators (Zhang et al. 2022) and also aligns with the beneficial effects of organic farming on soil fauna (Betancur-Corredor et al. 2023). This can be explained by the fact that the primary food sources of predatory protists, namely soil bacteria (Gao et al. 2019), were significantly enriched by organic fertilizer inputs, as previously discussed (Fig. 1a).
Associations between protistan predators and bacteria
We examined the potential associations between protistan predators and bacterial communities within the two agricultural ecosystems. By random forest analysis, we identified the primary predictors (with %IncMSE > 20%) of bacterial genera influencing the relative abundance of protistan predators. We found that Streptomyces, Neobacillus, and Tumebacillus emerged as strong predictors positively influencing the relative abundance of protistan predators (Fig. 2d). Conversely, Pseudomonas, Sphingomonas, Gammaproteobacteria_unclassified (unclassified genus from Gammaproteobacteria) and Chitinophagaceae_unclassified (unclassified genus from Gammaproteobacteria) were identified as bacterial predictors exerting negative effects on protistan predators. These findings indicated intricate trophic interactions within soil microbial communities, in particular between protistan predators and metabolic active bacteria (Gao et al. 2019; Jiang et al. 2024; Nguyen et al. 2020; Thakur and Geisen 2019). Additionally, we found a negative correlation between the relative abundance of protistan predators and bacterial richness in organic farming soils (Fig. 2e; R2 = 0.193, p < 0.001), confirming that protistan predators predominantly feed on bacteria (Geisen 2016; Geisen et al. 2018). Our results also revealed that conventional farming might decouple the trophic structure between protistan predators and bacterial preys within the soil microbiome, as this significant correlation between the relative abundance of protistan predators and bacterial richness in organic farming soils, was not observed in conventional soils (Fig. 2e). In other words, organic farming soils might support trophic structure between protistan predators and bacteria within soil microbiome.
The weighted mean ribosomal operon copy number (rrn) for the bacterial community in each sample was also measured in the study. The trait has been reported to be related to bacterial life-history strategy, growth rate, and nutrient demand (Lin et al. 2023; Tianjiao et al. 2022). For example, the fast-growing bacteria [most are copiotrophs (r-strategists)] produce many ribosomes to keep up their high growth rate and thus have more rrn copy number compared with low-growing bacteria (Klappenbach et al. 2000; Roller et al. 2016). Our analysis found that community-level rrn copy number was significantly higher in bacterial communities residing in organic farming soils compared to those in conventional farming soils, indicating a higher growth rate of bacterial communities under organic farming (Fig. S5a). This agrees with a study revealing that organic fertilization accelerated the resilience of copiotrophic bacteria to biodiversity loss (Luo et al. 2023). Because the growth rate of bacteria is constrained by carbon: nitrogen: phosphorus stoichiometry in soils, the growth rate of bacteria in particularly fast-growing copiotrophic bacteria relies heavily on external nutrients supplies (e.g., phosphorus and nitrogen inputs) to support phosphorus-rich ribosomal RNA production (Elser et al. 2000; Wang et al. 2024). We thus attributed the higher rrn copy number in the organic farming system to the ample nutrients (e.g., available phosphorus, total nitrogen) in organic fertilizer-supplied soils (Fig. S7). We also found that the community-level rrn copy number was significantly correlated with the relative abundance of predatory protists in both agricultural ecosystems (Fig. S5b). This indicates the potential influence of predation pressure by protistan predators on the life-history strategy of bacterial communities. Given the enrichment of protistan predators in soils supplied with organic fertilizers, the higher rrn copy number may also result from the higher relative abundance of protistan predators in the organic farming system.
In the study, the bacterial functions associated with community-wide dormancy capacity in the two agricultural ecosystems were also studied. Although no statistical differences in the relative abundance of two pathways related to bacterial dormancy capacity were observed (Table S4), we found distinct linear relationships between the relative abundance of genes in the bacterial toxin-antitoxin system and the relative abundance of predatory protists (Fig. S7). As the relative abundance of predatory protists increased, the relative abundance of toxin-antitoxin pathways decreased in organic farming systems but increased in conventional farming systems. Bacterial toxin–antitoxin systems were considered beneficial for bacterial adaptation and growth under stress conditions by regulating reversible cell growth inhibition (González Barrios et al. 2006; Jurėnas et al. 2022; Page and Peti 2016). The decrease of bacterial toxin–antitoxin systems with the increase of predatory protists under organic farming thus may contribute to the higher bacteria growth rate under the usage of organic fertilizers revealed by rrn copy number analysis. In addition, the different relationships between bacterial toxin-antitoxin systems and predatory protists again supported that protistan predators established different trophic interactions with soil bacteria under the two agricultural ecosystems. Furthermore, the toxin-antitoxin systems have been reported to be associated with bacterial pathogenesis (Gu et al. 2021; Ramage et al. 2009; Yao et al. 2015). The differing patterns between toxin-antitoxin systems and predatory protists in the two agricultural ecosystems may underscore the potential role of protistan predators in suppressing phytopathogens by regulating bacterial pathogenesis, although this has not been well studied in agricultural ecosystems.
Associations between protistan predators with PGPR and bacterial plant pathogens
We next examined the relationships between protistan predators and specific bacterial groups (i.e., PGPR and bacteria pathogens). Linear regression analysis confirmed the positive association between protistan predators and PGPR in organic farming systems (Fig. 3b and Fig. S4), highlighting the positive interaction between protistan predators and PGPR (Asiloglu et al. 2020; Jousset 2017) and the potential mechanisms by which organic farming enhance the relative abundance of PGPR in soils. Given the significant negative correlation between the relative abundance of PGPR, in particular, those with bioprotection effects and bacterial plant pathogens, our results collectively suggested that protistan predators could suppress bacterial plant pathogens via indirectly enhancing PGPR that inhibit bacterial plant pathogens (Guo et al. 2022, 2024).
The linear relationships between relative abundance of different bacteria and predatory protists. The linear relationships between the relative abundance of putative PGPR (a) and predatory protists (c) with that of putative bacterial plant pathogens; the linear relationships between the relative abundance of putative PGPR (b) and protist plant pathogens (d) with that of predatory protists. In panel b and c, dashed lines indicate non-significant linear relationships
However, we did not observe significant correlations between protistan predators with the overall PGPR (Fig. 3b) or each PGPR group (Fig. S4, except biostimulation PGPR) in conventional farming systems. The discrepancy in association between protistan predators and PGPR under different farming systems may resulted from the influence of different fertilizers (i.e., organic fertilizers and chemical fertilizers) on the taxonomic and functional compositions of soil protists (Xiong et al. 2018; Zhao et al. 2020). Because predation pressure by protists regulates plant growth-promoting traits of PGPR (Levrat et al. 1992; Mazzola and Bruijin 2009; Jousset and Bonkowski 2010; Chandarana and Amaresan 2023) as well as their survival rate (Jousset et al. 2008), the dysfunction of the interaction between predatory protists and PGRP may partly account for the non-significant or weak association between bacteria PGPR with putative bacteria pathogens in conventional farming systems (Fig. 3a and Fig. S3). In addition, we also found significantly negative correlations between protistan predators with putative phytopathogenic bacteria in organic farming systems (Fig. 3c) and protist plant pathogens (Fig. 3d) in both two agricultural ecosystems. The observations suggest that protistan predators may directly prey on plant pathogens, thereby enhancing plant health (Guo et al. 2022, 2024; Ren et al. 2024).
Differences in nutrient cycling of bacterial communities under the two agricultural ecosystems and their association with predatory protists
Previous local-scale studies highlight the influence of organic farming on the bacterial taxonomical composition and thereby nutrient cycling in soils (Ding et al. 2019; Fernandez et al. 2016). In agreement with these results, we found differences in farming management could lead to significant changes in nutrient cycling (Fig. 4 and Fig. 5). For example, we found chitinolysis that is included in C cycling pathways was significantly improved by the usages of organic fertilizers (Fig. 4, p<0.05, Wilcoxon test), in line with previous studies (Li et al. 2023b; Liu et al. 2021). This pathway was reported to have a positive correlation with nitrogen use efficiency and beneficial bacterial lineages like Bacillus (Li et al. 2024a). Due to the contribution of chitin to morphogenesis and maintenance of the cellular shape for protists (Steinfeld et al. 2019), chitinolysis and protists may thus have negative correlations (Fig. 4, p<0.05, Pearson correlation).
C, N, S cycling from soils supplied with organic fertilizer or conventional chemical fertilizer. “*” means p < 0.05, “**” means p < 0.01 and “***” means p < 0.001 by Wilcoxon test; colours in pathway labels indicates enrichment in organic farming systems (blue) or conventional farming systems (yellow); the red “↑” indicates positive and red “↓” indicates negative correlation between the relative abundance of pathways and that of predatory protists by Pearson correlation
P cycling from soils supplied with organic fertilizer or conventional chemical fertilizer and their linear relationships with predatory protists. Pathways included in P cycling with significant differences in their relative abundance between two agricultural systems (a); the linear relationships between the relative abundance of these pathways and that of predatory protists (b). In panel a, the colour of shadows indicates enrichment in organic farming systems (blue) or conventional farming systems (yellow). In panel b, dashed lines indicate non-significant linear relationships
We noticed that bacterial communities residing in soils from organic farming systems exhibited higher denitrification and nitrate-reduction potential than those from conventional system soils (Fig. 4, p<0.05, Wilcoxon test), which has been reported previously (Butterbach-Bahl and Dannenmann 2011; He et al. 2023; Hu et al. 2024; Kramer et al. 2006). Denitrification pathways enriched in organic farming systems showed no significant correlation with the relative abundance of predatory protists, while the activity or presence of predatory protists may decrease the rate of nitrate reduction (Fig. 4, p<0.05, Pearson correlation). This implies the activity of predatory protists may be involved in the N cycling in the soils.
As expected, organically fertilized soils exhibited different P cycling patterns compared with conventionally fertilized soils (Fig. 5a, p<0.05, Wilcoxon test). This is in line with previous work where long-term sustainable agricultural practices lead to great changes in P cycling (Dai et al. 2020). We noticed that 8 out of 10 pathways included in P cycling showed significant linear relationships with predatory protists (Fig. 5a, p<0.05, linear regression analysis). This may be because under different P limitation conditions, the trophic interactions between predatory protists and bacteria change (Guillonneau et al. 2022), which may ultimately result in shifts in the taxonomic composition of bacteria related to P cycling.
The impacts of soil properties on protists and the association between soil protists and bacteria
To understand the impacts of soil properties on protists and their interrelationships with bacteria in different fertilization regimes, we used one field experiment as a case study with available information of soil properties (Table S5) (Guo et al. 2021). As expected, the application of different fertilizers led to discernible variations in soil properties, particularly enriched organic matter in organic farming soils (Fig. S7), consistent with previous studies (Crystal-Ornelas et al. 2021; Marinari et al. 2000). The changes in organic matter, are significantly and positively linked with the richness and the relative abundance of protistan predators (Fig. 6a and Fig. 6b), as well as protistan predators of Trinematidae, Sandonidae and Euglyphida (Fig. S8). Our results also highlighted a more pronounced positive effect of organic matter on protistan predators (with a higher proportion of explained variance) in conventional farming soils compared to organic farming soils (Fig. 6c and Fig. 6d).
The correlations between soil properties and microbial parameters. Spearman’s rank correlation between soil microbial parameters and soil properties including pH, OM (organic matter), TP (total P), TK (total K), TN (total N), AP (available P) , AK (available K) and AN (available N) (a); redundancy analysis (RDA) of the protistan functional groups and soil properties for soil samples (b); and the linear relationships between the content of organic matters and the richness (c) also relative abundance (d) of predatory protists. The data for Figure 3 were selected from one field experiment site with three years continuously planted cucumber as explained in Materials and Methods section
Additionally, our findings indicated that organic farming soils tend to facilitate trophic structure between protistan predators and bacteria, as evidenced by more positive interconnections (45 significant with 29 positive correlations in organic farming soils, 17 significant with 5 positive correlations in conventional soils) from partial correlation where we controlled the effects of organic matter between major protistan predators and bacterial groups (Fig. S9). Furthermore, we found Cercomonas and Sandona positively correlated with plant-beneficial bacteria of Sphingomonas and Pseudomonas in organic farming soils, but not in conventional farming soils (Fig. S9), these findings indicate that organic inputs promote beneficial interrelationships between protistan predators and bacterial communities. This observation aligns with prior research indicating that organic amendments promote microbial cooperation (Banerjee et al. 2016), also reflected in predator-prey interactions.
Conclusion
Our results highlighted the enhanced bacterial diversity and the higher relative abundance of potentially plant-beneficial bacteria in organic farming soils compared to their conventional counterparts. Furthermore, organic farming soils enriched a higher relative abundance of protistan predators with the decreased plant pathogens of protists. Interestingly, our results highlighted intensive agricultural management by chemical fertilizer application might decouple the stable trophic structure between protistan predators and bacterial prey. This disruption might be attributed, at least in part, to the deficiency of organic matter in conventional chemical fertilizer-applied systems. Thus, we propose that organic farming with organic fertilizer applications improve soil ecologic functions and thereby the sustainability of agricultural ecosystems, through increasing beneficial protistan predators and sustaining predator-prey interrelationships within soil microbiome.
Data availability
The data that support the findings of this study are openly available in the figshare repository: https://doi.org/10.6084/m9.figshare.26028745.
References
Abdollahi L, Schjønning P, Elmholt S, Munkholm LJ (2014) The effects of organic matter application and intensive tillage and traffic on soil structure formation and stability. Soil Till Res 136:28–37. https://doi.org/10.1016/j.still.2013.09.011
Adl SM, Bass D, Lane CE, Lukeš J, Schoch CL, Smirnov A, Agatha S, Berney C, Brown MW, Burki F, Cárdenas P, Čepička I, Chistyakova L, Del Campo J, Dunthorn M, Edvardsen B, Eglit Y, Guillou L, Hampl V, Heiss AA, Hoppenrath M, James TY, Karnkowska A, Karpov S, Kim E, Kolisko M, Kudryavtsev A, Lahr DJG, Lara E, Le Gall L, Lynn DH, Mann DG, Massana R, Mitchell EAD, Morrow C, Park JS, Pawlowski JW, Powell MJ, Richter DJ, Rueckert S, Shadwick L, Shimano S, Spiegel FW, Torruella G, Youssef N, Zlatogursky V, Zhang Q (2019) Revisions to the Classification, Nomenclature, and Diversity of Eukaryotes. J Eukaryot Microbiol 66:4–119. https://doi.org/10.1111/jeu.12691
Asiloglu R, Shiroishi K, Suzuki K (2020) Protist-enhanced survival of a plant growth promoting rhizobacteria, Azospirillum sp. B510, and the growth of rice (Oryza sativa L.) plants. Appl Soil Ecol 154. https://doi.org/10.1016/j.apsoil.2020.103599
Azarbad H (2022) Conventional vs. Organic Agriculture–Which One Promotes Better Yields and Microbial Resilience in Rapidly Changing Climates? Front Microbiol 13: 903500. https://doi.org/10.3389/fmicb.2022.903500
Banerjee S, Kirkby CA, Schmutter D, Bissett A, Kirkegaard JA, Richardson AE (2016) Network analysis reveals functional redundancy and keystone taxa amongst bacterial and fungal communities during organic matter decomposition in an arable soil. Soil Biol Biochem 97:188–198. https://doi.org/10.1016/j.soilbio.2016.03.017
Bebber DP, Richards VR (2022) A meta-analysis of the effect of organic and mineral fertilizers on soil microbial diversity. Appl Soil Ecol 175:104450. https://doi.org/10.1016/j.apsoil.2022.104450
Betancur-Corredor B, Lang B, Russell DJ (2023) Organic nitrogen fertilization benefits selected soil fauna in global agroecosystems. Biol Fertil Soils 59:1–16. https://doi.org/10.1007/s00374-022-01677-2
Bhogal A, Young SD, Sylvester-Bradley R (1997) Straw incorporation and immobilization of spring-applied nitrogen. Soil Use Manag 13:111–116. https://doi.org/10.1111/j.1475-2743.1997.tb00568.x
Breiman L (2001) Random Forests. Mach Learn 45:5–32. https://doi.org/10.1023/A:1010933404324
Bu R, Li M, Cheng W, Han S, Wang H, Tang S, Lu C, Wu J (2023) Subsoil Tillage and Organic Fertilization Benefit Rice Root Growth and Yield by Ameliorating Soil Compaction and Fertility. J Soil Sci Plant Nutr 23:6114–6124. https://doi.org/10.1007/s42729-023-01468-0
Butterbach-Bahl K, Dannenmann M (2011) Denitrification and associated soil N2O emissions due to agricultural activities in a changing climate. Curr Opin Environ Sustain 3:389–395. https://doi.org/10.1016/j.cosust.2011.08.004
Chandarana KA, Amaresan N (2023) Predation pressure regulates plant growth promoting (PGP) attributes of bacterial species J Appl Microbiol 134: lxad083. https://doi.org/10.1093/jambio/lxad083
Chen D, Wang X, Zhang W, Zhou Z, Ding C, Liao Y, Li X (2020) Persistent organic fertilization reinforces soil-borne disease suppressiveness of rhizosphere bacterial community. Plant Soil 452:313–328. https://doi.org/10.1007/s11104-020-04576-3
Chowdhury SA, Kaneko A, Baki MZI, Takasugi C, Wada N, Asiloglu R, Harada N, Suzuki K (2022) Impact of the chemical composition of applied organic materials on bacterial and archaeal community compositions in paddy soil. Biol Fertil Soils 58:135–148. https://doi.org/10.1007/s00374-022-01619-y
Crystal-Ornelas R, Thapa R, Tully KL (2021) Soil organic carbon is affected by organic amendments, conservation tillage, and cover cropping in organic farming systems: A meta-analysis. Agric Ecosyst Environ 312:107356. https://doi.org/10.1016/j.agee.2021.107356
Dai Z, Liu G, Chen H, Chen C, Wang J, Ai S, Wei D, Li D, Ma B, Tang C, Brookes PC, Xu J (2020) Long-term nutrient inputs shift soil microbial functional profiles of phosphorus cycling in diverse agroecosystems. ISME J 14:757–770. https://doi.org/10.1038/s41396-019-0567-9
Deng X, Zhang N, Li Y, Zhu C, Qu B, Liu H, Li R, Bai Y, Shen Q, Falcao Salles J (2022) Bio-organic soil amendment promotes the suppression of Ralstonia solanacearum by inducing changes in the functionality and composition of rhizosphere bacterial communities. New Phytol 235:1558–1574. https://doi.org/10.1111/nph.18221
Diacono M, Montemurro F (2011) Long-Term Effects of Organic Amendments on Soil Fertility. In: Lichtfouse E, Hamelin M, Navarrete M, Debaeke P (eds) Sustainable Agriculture, vol 2. Springer. Netherlands, Dordrecht, pp 761–786
Ding G-C, Bai M, Han H, Li H, Ding X, Yang H, Xu T, Li J (2019) Microbial taxonomic, nitrogen cycling and phosphorus recycling community composition during long-term organic greenhouse farming. FEMS Microbiol Ecol 95:fiz042. https://doi.org/10.1093/femsec/fiz042
Djemiel C, Maron P-A, Terrat S, Dequiedt S, Cottin A, Ranjard L (2022) Inferring microbiota functions from taxonomic genes: a review. Gigascience 11:giab090. https://doi.org/10.1093/gigascience/giab090
Douglas GM, Maffei VJ, Zaneveld JR, Yurgel SN, Brown JR, Taylor CM, Huttenhower C, Langille MG (2020) PICRUSt2 for prediction of metagenome functions. Nat Biotechnol 38:685–688. https://doi.org/10.1038/s41587-020-0548-6
Dubey A, Malla MA, Khan F, Chowdhary K, Yadav S, Kumar A, Sharma S, Khare PK, Khan ML (2019) Soil microbiome: a key player for conservation of soil health under changing climate. Biodivers Conserv 28:2405–2429. https://doi.org/10.1007/s10531-019-01760-5
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Edgar RC (2016) UNOISE2: improved error-correction for Illumina 16S and ITS amplicon sequencing. BioRxiv, 081257.
Elser JJ, Sterner RW, Gorokhova E, Fagan WF, Markow TA, Cotner JB, Harrison JF, Hobbie SE, Odell GM, Weider LW (2000) Biological stoichiometry from genes to ecosystems. Ecol Lett 3:540–550. https://doi.org/10.1111/j.1461-0248.2000.00185.x
Fernandez AL, Sheaffer CC, Wyse DL, Staley C, Gould TJ, Sadowsky MJ (2016) Associations between soil bacterial community structure and nutrient cycling functions in long-term organic farm soils following cover crop and organic fertilizer amendment. Sci Total Environ 566–567:949–959. https://doi.org/10.1016/j.scitotenv.2016.05.073
Folman LB, De Klein MJEM, Postma J, van Veen JA (2004) Production of antifungal compounds by Lysobacter enzymogenes isolate 3.1T8 under different conditions in relation to its efficacy as a biocontrol agent of Pythium aphanidermatum in cucumber. Biol Control 31:145–154. https://doi.org/10.1016/j.biocontrol.2004.03.008
Fujino M, Suzuki K, Harada N, Asiloglu R (2023) Protists modulate active bacterial community composition in paddy field soils. Biol Fertil Soils 59:709–721. https://doi.org/10.1007/s00374-023-01733-5
Gabriel D, Sait SM, Kunin WE, Benton TG (2013) Food production vs. biodiversity: comparing organic and conventional agriculture. J Appl Ecol 50:355–364. https://doi.org/10.1111/1365-2664.12035
Gao Z, Karlsson I, Geisen S, Kowalchuk G, Jousset A (2019) Protists: Puppet Masters of the Rhizosphere Microbiome. Trends Plant Sci 24:165–176. https://doi.org/10.1016/j.tplants.2018.10.011
Geisen S (2016) The bacterial-fungal energy channel concept challenged by enormous functional versatility of soil protists. Soil Biol Biochem 102:22–25. https://doi.org/10.1016/j.soilbio.2016.06.013
Geisen S, Mitchell EAD, Adl S, Bonkowski M, Dunthorn M, Ekelund F, Fernández LD, Jousset A, Krashevska V, Singer D, Spiegel FW, Walochnik J, Lara E (2018) Soil protists: a fertile frontier in soil biology research. FEMS Microbiol Rev 42:293–323. https://doi.org/10.1093/femsre/fuy006
Gomiero T, Pimentel D, Paoletti MG (2011) Is There a Need for a More Sustainable Agriculture? Crit Rev Plant Sci 30:6–23. https://doi.org/10.1080/07352689.2011.553515
González Barrios AF, Zuo R, Hashimoto Y, Yang L, Bentley WE, Wood TK (2006) Autoinducer 2 Controls Biofilm Formation in Escherichia coli through a Novel Motility Quorum-Sensing Regulator (MqsR, B3022). J Bacteriol 188:305–316. https://doi.org/10.1128/jb.188.1.305-316.2006
Gu Q, He P, Wang D, Ma J, Zhong X, Zhu Y, Zhang Y, Bai Q, Pan Z, Yao H (2021) An Auto-Regulating Type II Toxin-Antitoxin System Modulates Drug Resistance and Virulence in Streptococcus suis. Front Microbiol 12:671706. https://doi.org/10.3389/fmicb.2021.671706
Guillonneau R, Murphy ARJ, Teng Z-J, Wang P, Zhang Y-Z, Scanlan DJ, Chen Y (2022) Trade-offs of lipid remodeling in a marine predator–prey interaction in response to phosphorus limitation. Proc Natl Acad Sci 119:e2203057119. https://doi.org/10.1073/pnas.2203057119
Guillou L, Bachar D, Audic S (2012) The Protist Ribosomal Reference database (PR2): a catalog of unicellular eukaryote small sub-unit rRNA sequences with curated taxonomy. Nucleic Acids Res 41:597–604
Guo S, Xiong W, Xu H, Hang X, Liu H, Xun W, Li R, Shen Q (2018) Continuous application of different fertilizers induces distinct bulk and rhizosphere soil protist communities. Eur J Soil Biol 88:8–14. https://doi.org/10.1016/j.ejsobi.2018.05.007
Guo S, Xiong W, Hang X, Gao Z, Jiao Z, Liu H, Mo Y, Zhang N, Kowalchuk GA, Li R, Shen Q, Geisen S (2021) Protists as main indicators and determinants of plant performance. Microbiome 9:64. https://doi.org/10.1186/s40168-021-01025-w
Guo S, Tao C, Jousset A, Xiong W, Wang Z, Shen Z, Wang B, Xu Z, Gao Z, Liu S, Li R, Ruan Y, Shen Q, Kowalchuk GA, Geisen S (2022) Trophic interactions between predatory protists and pathogen-suppressive bacteria impact plant health. ISME J 16:1932–1943. https://doi.org/10.1038/s41396-022-01244-5
Guo S, Jiao Z, Yan Z, Yan X, Deng X, Xiong W, Tao C, Liu H, Li R, Shen Q, Kowalchuk GA, Geisen S (2024) Predatory protists reduce bacteria wilt disease incidence in tomato plants. Nat Commun 15:1–12. https://doi.org/10.1038/s41467-024-45150-0
Hartmann M, Frey B, Mayer J, Mäder P, Widmer F (2015) Distinct soil microbial diversity under long-term organic and conventional farming. ISME J 9:1177–1194. https://doi.org/10.1038/ismej.2014.210
He Z, Ding B, Pei S, Cao H, Liang J, Li Z (2023) The impact of organic fertilizer replacement on greenhouse gas emissions and its influencing factors. Sci Total Environ 905:166917. https://doi.org/10.1016/j.scitotenv.2023.166917
Heinze S, Raupp J, Joergensen RG (2010) Effects of fertilizer and spatial heterogeneity in soil pH on microbial biomass indices in a long-term field trial of organic agriculture. Plant Soil 328:203–215. https://doi.org/10.1007/s11104-009-0102-2
Hu Z, Yao J, Chen X, Gong X, Zhang Y, Zhou X, Guo H, Liu M (2022) Precipitation changes, warming, and N input differentially affect microbial predators in an alpine meadow: Evidence from soil phagotrophic protists. Soil Biol Biochem 165:108521. https://doi.org/10.1016/j.soilbio.2021.108521
Hu W, Zhang Y, Rong X, Zhou X, Fei J, Peng J, Luo G (2024) Biochar and organic fertilizer applications enhance soil functional microbial abundance and agroecosystem multifunctionality. Biochar 6:3. https://doi.org/10.1007/s42773-023-00296-w
Jiang G, Liu C, Xiong W, Shen Q, Wei Z (2024) Protist predation selects for the soil resistome. ISME J 18(1): wrad007. https://doi.org/10.1093/ismejo/wrad007
Jousset A, Bonkowski M (2010) The model predator Acanthamoeba castellanii induces the production of 2,4, DAPG by the biocontrol strain Pseudomonas fluorescens Q2–87. Soil Biol Biochem 42:1647–1649. https://doi.org/10.1016/j.soilbio.2010.05.018
Jousset A, Scheu S, Bonkowski M (2008) Secondary metabolite production facilitates establishment of rhizobacteria by reducing both protozoan predation and the competitive effects of indigenous bacteria. Funct Ecol 22:714–719. https://doi.org/10.1111/j.1365-2435.2008.01411.x
Jousset A (2017) Application of Protists to Improve Plant Growth in Sustainable Agriculture. Rhizotrophs: Plant growth promotion to bioremediation, 263–273
Jurėnas D, Fraikin N, Goormaghtigh F, Van Melderen L (2022) Biology and evolution of bacterial toxin–antitoxin systems. Nat Rev Microbiol 20:335–350. https://doi.org/10.1038/s41579-021-00661-1
Kearns PJ, Shade A (2018) Trait-based patterns of microbial dynamics in dormancy potential and heterotrophic strategy: case studies of resource-based and post-press succession. ISME J 12:2575–2581. https://doi.org/10.1038/s41396-018-0194-x
Kim S (2015) ppcor: An R Package for a Fast Calculation to Semi-partial Correlation Coefficients. Commun Stat Appl Methods 22:665–674. https://doi.org/10.5351/CSAM.2015.22.6.665
Klappenbach JA, Dunbar JM, Schmidt TM (2000) rRNA Operon Copy Number Reflects Ecological Strategies of Bacteria. Appl Environ Microbiol 66:1328–1333. https://doi.org/10.1128/AEM.66.4.1328-1333.2000
Kramer SB, Reganold JP, Glover JD, Bohannan BJM, Mooney HA (2006) Reduced nitrate leaching and enhanced denitrifier activity and efficiency in organically fertilized soils. Proc Natl Acad Sci 103:4522–4527. https://doi.org/10.1073/pnas.0600359103
Kumar Bhatt M, Labanya R, Joshi HC (2019) Influence of Long-term Chemical fertilizers and Organic Manures on Soil Fertility - A Review. Univers J Agric Res 7:177–188. https://doi.org/10.13189/ujar.2019.070502
Laurent C, Bravin MN, Crouzet O, Pelosi C, Tillard E, Lecomte P, Lamy I (2020) Increased soil pH and dissolved organic matter after a decade of organic fertilizer application mitigates copper and zinc availability despite contamination. Sci Total Environ 709:135927. https://doi.org/10.1016/j.scitotenv.2019.135927
Lehmann J, Bossio DA, Kögel-Knabner I, Rillig MC (2020) The concept and future prospects of soil health. Nat Rev Earth Environ 1:544–553. https://doi.org/10.1038/s43017-020-0080-8
Levrat P, Pussard M, Alabouvette C (1992) Enhanced bacterial metabolism of a Pseudomonas strain in response to the addition of culture filtrate of a bacteriophagous amoeba. Eur J Protistol 28:79–84. https://doi.org/10.1016/S0932-4739(11)80322-6
Li F, Chen L, Zhang J, Yin J, Huang S (2017) Bacterial Community Structure after Long-term Organic and Inorganic Fertilization Reveals Important Associations between Soil Nutrients and Specific Taxa Involved in Nutrient Transformations. Front Microbiol 8:187. https://doi.org/10.3389/fmicb.2017.00187
Li C, Gillings MR, Zhang C, Chen Q, Zhu D, Wang J, Zhao K, Xu Q, Leung PH, Li X, Liu J, Jin L (2023a) Ecology and risks of the global plastisphere as a newly expanding microbial habitat. The Innovation 5:100543. https://doi.org/10.1016/j.xinn.2023.100543
Li G, Niu W, Ma L, Du Y, Zhang Q, Sun J, Siddique KHM (2023b) Legacy effects of wheat season organic fertilizer addition on microbial co-occurrence networks, soil function, and yield of the subsequent maize season in a wheat-maize rotation system. J Environ Manage 347:119160. https://doi.org/10.1016/j.jenvman.2023.119160
Li D, Duan W, Guo H, Zong J, Chen J, Wang J (2024a) High-nitrogen organic fertilizer promotes the growth of bermudagrass (Cynodon dactylon), zoysiagrass (Zoysia japonica) and paspalum grass (Paspalum vaginatum) by enhancing nitrogen use efficiency associated with Bacillus-stimulated bacterial community. Sci Hortic 329:113027. https://doi.org/10.1016/j.scienta.2024.113027
Li W-J, Zhou X-Y, An X-L, Li L-J, Lin C-S, Li H, Li H-Z (2024b) Enhancement of beneficial microbiomes in plant–soil continuums through organic fertilization: Insights into the composition and multifunctionality. Soil Ecol Lett 6:230223. https://doi.org/10.1007/s42832-023-0223-1
Lichtenberg EM, Kennedy CM, Kremen C, Batáry P, Berendse F, Bommarco R, Bosque-Pérez NA, Carvalheiro LG, Snyder WE, Williams NM, Winfree R, Klatt BK, Åström S, Benjamin F, Brittain C, Chaplin-Kramer R, Clough Y, Danforth B, Diekötter T, Eigenbrode SD, Ekroos J, Elle E, Freitas BM, Fukuda Y, Gaines-Day HR, Grab H, Gratton C, Holzschuh A, Isaacs R, Isaia M, Jha S, Jonason D, Jones VP, Klein A-M, Krauss J, Letourneau DK, Macfadyen S, Mallinger RE, Martin EA, Martinez E, Memmott J, Morandin L, Neame L, Otieno M, Park MG, Pfiffner L, Pocock MJO, Ponce C, Potts SG, Poveda K, Ramos M, Rosenheim JA, Rundlöf M, Sardiñas H, Saunders ME, Schon NL, Sciligo AR, Sidhu CS, Steffan-Dewenter I, Tscharntke T, Veselý M, Weisser WW, Wilson JK, Crowder DW (2017) A global synthesis of the effects of diversified farming systems on arthropod diversity within fields and across agricultural landscapes. Glob Change Biol 23:4946–4957. https://doi.org/10.1111/gcb.13714
Lin R, Wu H, Kong X, Ren H, Lu Z (2023) Ribosomal RNA gene operon copy number, a functional trait indicating the hydrocarbon degradation level of bacterial communities. J Hazard Mater 459:132100. https://doi.org/10.1016/j.jhazmat.2023.132100
Ling N, Wang T, Kuzyakov Y (2022) Rhizosphere bacteriome structure and functions. Nat Commun 13:836. https://doi.org/10.1038/s41467-022-28448-9
Liu H, Xiong W, Zhang R, Hang X, Wang D, Li R, Shen Q (2018) Continuous application of different organic additives can suppress tomato disease by inducing the healthy rhizospheric microbiota through alterations to the bulk soil microflora. Plant Soil 423:229–240. https://doi.org/10.1007/s11104-017-3504-6
Liu J, Shu A, Song W, Shi W, Li M, Zhang W, Li Z, Liu G, Yuan F, Zhang S, Liu Z, Gao Z (2021) Long-term organic fertilizer substitution increases rice yield by improving soil properties and regulating soil bacteria. Geoderma 404:115287. https://doi.org/10.1016/j.geoderma.2021.115287
Louca S, Parfrey LW, Doebeli M, (2016) Decoupling function and taxonomy in the global ocean microbiome. Science 353:1272–1277. https://doi.org/10.1126/science.aaf4507
Luo G, Li L, Friman V-P, Guo J, Guo S, Shen Q, Ling N (2018) Organic amendments increase crop yields by improving microbe-mediated soil functioning of agroecosystems: A meta-analysis. Soil Biol Biochem 124:105–115. https://doi.org/10.1016/j.soilbio.2018.06.002
Luo J, Banerjee S, Ma Q, Liao G, Hu B, Zhao H, Li T (2023) Organic fertilization drives shifts in microbiome complexity and keystone taxa increase the resistance of microbial mediated functions to biodiversity loss. Biol Fertil Soils 59:441–458. https://doi.org/10.1007/s00374-023-01719-3
Lupatini M, Korthals GW, de Hollander M, Janssens TKS, Kuramae EE (2017) Soil Microbiome Is More Heterogeneous in Organic Than in Conventional Farming System. Front Microbiol 7:2064
Ma Y, Shen S, Wan C, Wang S, Yang F, Zhang K, Gao W (2023) Organic fertilizer substitution over six years improves the productivity of garlic, bacterial diversity, and microbial communities network complexity. Appl Soil Ecol 182:104718. https://doi.org/10.1016/j.apsoil.2022.104718
Maeder P, Fliessbach A, Dubois D, Gunst L, Fried P, Niggli U (2002) Soil Fertility and Biodiversity in Organic Farming. Science 296:1694–1697. https://doi.org/10.1126/science.1071148
Marinari S, Masciandaro G, Ceccanti B, Grego S (2000) Influence of organic and mineral fertilisers on soil biological and physical properties. Bioresour Technol 72:9–17. https://doi.org/10.1016/S0960-8524(99)00094-2
Mazzola M, Bruijn I, Cohen MF (2009) Protozoan-induced regulation of cyclic lipopeptide biosynthesis is an effective predation defense mechanism for Pseudomonas fluorescens. Appl Environ Microbiol J 75:6804–6811. https://doi.org/10.1128/AEM.01272-09
Melo J, Carolino M, Carvalho L, Correia P, Tenreiro R, Chaves S, Meleiro AI, de Souza SB, Dias T, Cruz C, Ramos AC (2016) Crop management as a driving force of plant growth promoting rhizobacteria physiology. SpringerPlus 5:1574. https://doi.org/10.1186/s40064-016-3232-z
Milke F, Rodas-Gaitan H, Meissner G, Masson V, Oltmanns M, Möller M, Wohlfahrt Y, Kulig B, Acedo A, Athmann M, Fritz J (2024) Enrichment of putative plant growth promoting microorganisms in biodynamic compared with organic agriculture soils. ISME Commun 4:ycae021. https://doi.org/10.1093/ismeco/ycae021
Navarrete AA, Cannavan FS, Taketani RG, Tsai SM (2010) A Molecular Survey of the Diversity of Microbial Communities in Different Amazonian Agricultural Model Systems. Diversity 2:787–809. https://doi.org/10.3390/d2050787
Naylor D, Sadler N, Bhattacharjee A, Graham EB, Anderton CR, McClure R, Lipton M, Hofmockel KS, Jansson JK (2020) Soil Microbiomes Under Climate Change and Implications for Carbon Cycling. Annu Rev Environ Resour 45:29–59. https://doi.org/10.1146/annurev-environ-012320-082720
Nguyen B-AT, Chen Q-L, He J-Z, Hu H-W (2020) Microbial regulation of natural antibiotic resistance: Understanding the protist-bacteria interactions for evolution of soil resistome. Sci Total Environ 705:135882. https://doi.org/10.1016/j.scitotenv.2019.135882
Nguyen TB-A, Bonkowski M, Dumack K, Chen Q-L, He J-Z, Hu H-W (2023) Protistan predation selects for antibiotic resistance in soil bacterial communities. ISME J 1–8. https://doi.org/10.1038/s41396-023-01524-8
Oliverio AM, Geisen S, Delgado-Baquerizo M, Maestre FT, Turner BL, Fierer N (2020) The global-scale distributions of soil protists and their contributions to belowground systems. Sci Adv 6:eaax8787. https://doi.org/10.1126/sciadv.aax8787
Page R, Peti W (2016) Toxin-antitoxin systems in bacterial growth arrest and persistence. Nat Chem Biol 12:208–214. https://doi.org/10.1038/nchembio.2044
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30:3123–3124. https://doi.org/10.1093/bioinformatics/btu494
Qiao C, Penton CR, Xiong W, Liu C, Wang R, Liu Z, Xu X, Li R, Shen Q (2019) Reshaping the rhizosphere microbiome by bio-organic amendment to enhance crop yield in a maize-cabbage rotation system. Appl Soil Ecol 142:136–146. https://doi.org/10.1016/j.apsoil.2019.04.014
Ramage HR, Connolly LE, Cox JS (2009) Comprehensive Functional Analysis of Mycobacterium tuberculosis Toxin-Antitoxin Systems: Implications for Pathogenesis, Stress Responses, and Evolution. PLOS Genet 5:e1000767. https://doi.org/10.1371/journal.pgen.1000767
Reichenbach H (2006) The Genus Lysobacter. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The Prokaryotes: A Handbook on the Biology of Bacteria, vol 6. Proteobacteria: Gamma Subclass. Springer, New York, NY, pp 939–957
Ren P, Sun A, Jiao X, Shen J-P, Yu D-T, Li F, Wu B, He J-Z, Hu H-W (2023) Predatory protists play predominant roles in suppressing soil-borne fungal pathogens under organic fertilization regimes. Sci Total Environ 863:160986. https://doi.org/10.1016/j.scitotenv.2022.160986
Ren X, Zhou Z, Liu M, Shen Z, Wang B, Jousset A, Geisen S, Ravanbakhsh M, Kowalchuk GA, Li R, Shen Q, Xiong W (2024) Intercropping with Trifolium repens contributes disease suppression of banana Fusarium wilt by reshaping soil protistan communities. Agric Ecosyst Environ 361:108797. https://doi.org/10.1016/j.agee.2023.108797
Roller BRK, Stoddard SF, Schmidt TM (2016) Exploiting rRNA operon copy number to investigate bacterial reproductive strategies. Nat Microbiol 1:1–7. https://doi.org/10.1038/nmicrobiol.2016.160
Rong Q, Li R, Huang S, Tang J, Zhang Y, Wang L (2018) Soil microbial characteristics and yield response to partial substitution of chemical fertilizer with organic amendments in greenhouse vegetable production. J Integr Agric 17:1432–1444. https://doi.org/10.1016/S2095-3119(18)61946-X
Röös E, Mie A, Wivstad M, Salomon E, Johansson B, Gunnarsson S, Wallenbeck A, Hoffmann R, Nilsson U, Sundberg C, Watson CA (2018) Risks and opportunities of increasing yields in organic farming. A review. Agron Sustain Dev 38:14. https://doi.org/10.1007/s13593-018-0489-3
Rosenzweig N, Tiedje JM, Quensen JF, Meng Q, Hao JJ (2012) Microbial Communities Associated with Potato Common Scab-Suppressive Soil Determined by Pyrosequencing Analyses. Plant Dis 96:718–725. https://doi.org/10.1094/PDIS-07-11-0571
Sánchez-Bayo F, Wyckhuys KAG (2019) Worldwide decline of the entomofauna: A review of its drivers. Biol Conserv 232:8–27. https://doi.org/10.1016/j.biocon.2019.01.020
Saxena AK, Kumar M, Chakdar H, Anuroopa N, Bagyaraj DJ (2020) Bacillus species in soil as a natural resource for plant health and nutrition. J Appl Microbiol 128:1583–1594. https://doi.org/10.1111/jam.14506
Schmidt JE, Kent AD, Brisson VL, Gaudin ACM (2019) Agricultural management and plant selection interactively affect rhizosphere microbial community structure and nitrogen cycling. Microbiome 7:146. https://doi.org/10.1186/s40168-019-0756-9
Shen B, Wang X, Zhang Y, Zhang M, Wang K, Xie P, Ji H (2020) The optimum pH and Eh for simultaneously minimizing bioavailable cadmium and arsenic contents in soils under the organic fertilizer application. Sci Total Environ 711:135229. https://doi.org/10.1016/j.scitotenv.2019.135229
Siedt M, Schäffer A, Smith KEC, Nabel M, Roß-Nickoll M, van Dongen JT (2021) Comparing straw, compost, and biochar regarding their suitability as agricultural soil amendments to affect soil structure, nutrient leaching, microbial communities, and the fate of pesticides. Sci Total Environ 751:141607. https://doi.org/10.1016/j.scitotenv.2020.141607
Steinfeld L, Vafaei A, Rösner J, Merzendorfer H (2019) Chitin Prevalence and Function in Bacteria, Fungi and Protists. In: Yang Q, Fukamizo T (eds) Targeting Chitin-containing Organisms. Springer, Singapore, pp 19–59
Thakur MP, Geisen S (2019) Trophic Regulations of the Soil Microbiome. Trends Microbiol 27:771–780. https://doi.org/10.1016/j.tim.2019.04.008
Tianjiao D, Wen D, Bates C, Wu L, Guo X, Liu S, Su Y, Lei J, Zhou J, Yang Y (2022) Nutrient supply controls the linkage between species abundance and ecological interactions in marine bacterial communities. Nat Commun 13:175. https://doi.org/10.1038/s41467-021-27857-6
Wang X, Jia Z, Liang L (2014) Effect of straw incorporation on soil moisture, evapotranspiration, and rainfall-use efficiency of maize under dryland farming. J Soil Water Conserv 69:449–455. https://doi.org/10.2489/jswc.69.5.449
Wang C, Shi Z, Li A, Geng T, Liu L, Liu W (2024) Long-term nitrogen input reduces soil bacterial network complexity by shifts in life history strategy in temperate grassland. iMeta 3:e194. https://doi.org/10.1002/imt2.194
Wei Z, Gu Y, Friman V-P, Kowalchuk GA, Xu Y, Shen Q, Jousset A (2019) Initial soil microbiome composition and functioning predetermine future plant health. Sci Adv 5: eaaw0759. https://doi.org/10.1126/sciadv.aaw0759
Wemheuer F, Taylor JA, Daniel R, Johnston E, Meinicke P, Thomas T, Wemheuer B (2020) Tax4Fun2: prediction of habitat-specific functional profiles and functional redundancy based on 16S rRNA gene sequences. Environ Microbiome 15:11. https://doi.org/10.1186/s40793-020-00358-7
Xiong W, Guo S, Jousset A, Zhao Q, Wu H, Li R, Kowalchuk GA, Shen Q (2017) Bio-fertilizer application induces soil suppressiveness against Fusarium wilt disease by reshaping the soil microbiome. Soil Biol Biochem 114:238–247. https://doi.org/10.1016/j.soilbio.2017.07.016
Xiong W, Jousset A, Guo S (2018) Soil protist communities form a dynamic hub in the soil microbiome. ISME J 12:634–638. https://doi.org/10.1038/ismej.2017.171
Xiong W, Jousset A, Li R, Delgado-Baquerizo M, Bahram M, Logares R, Wilden B, de Groot GA, Amacker N, Kowalchuk GA, Shen Q, Geisen S (2021) A global overview of the trophic structure within microbiomes across ecosystems. Environ Int 151:106438. https://doi.org/10.1016/j.envint.2021.106438
Yang X, Jiang G, Zhang Y, Wang N, Zhang Y, Wang X, Zhao F-J, Xu Y, Shen Q, Wei Z (2023) MBPD: A multiple bacterial pathogen detection pipeline for One Health practices. iMeta 2:e82. https://doi.org/10.1002/imt2.82
Yao X, Chen T, Shen X, Zhao Y, Wang M, Rao X, Yin S, Wang J, Gong Y, Lu S, Le S, Tan Y, Tang J, Fuquan H, Li M (2015) The chromosomal SezAT toxin–antitoxin system promotes the maintenance of the SsPI-1 pathogenicity island in epidemic Streptococcus suis. Mol Microbiol 98:243–257. https://doi.org/10.1111/mmi.13116
Zeng J, Tu Q, Yu X, Qian L, Wang C, Shu L, Liu F, Liu S, Huang Z, He J, Yan Q, He Z (2022) PCycDB: a comprehensive and accurate database for fast analysis of phosphorus cycling genes. Microbiome 10:101. https://doi.org/10.1186/s40168-022-01292-1
Zhang S, Zhang H, Liu H, Wang H, Xiu W, Li G, Zhang G, Zhou Z, Jiang N, Zhang H, Zhao J, Yang D (2022) Fertilization drives distinct biotic and abiotic factors in regulating functional groups of protists in a 5-year fertilization system. Front Microbiol 13:036362
Zhao Z-B, He J-Z, Quan Z, Wu C-F, Sheng R, Zhang L-M, Geisen S (2020) Fertilization changes soil microbiome functioning, especially phagotrophic protists. Soil Biol Biochem 148:107863. https://doi.org/10.1016/j.soilbio.2020.107863
Acknowledgements
This project received fundings from National Natural Science Foundation of China (42107141 and 42377296), National Key Research and Development Program of China (2023YFD1901402 and 2023YFD1901105), and Fundamental Research Funds for the Central Universities (YDZX2023023 and XUEKEN2023039).
Author information
Authors and Affiliations
Contributions
Chen Liu: Conceptualization, Methodology, Software, Visualization, Writing – review & editing. Zeyuan Zhou: Methodology, Writing – review & editing. Shiqi Sun: Writing – review & editing. Qi Zhang: Writing – review & editing. Shuo Sun: Resources, Writing – review & editing. Xinnan Hang: Resources. Mohammadhossein Ravanbakhsh: Writing – review & editing. Zhong Wei: Resources, Supervision. Shimei Wang: Resources, Conceptualization, Methodology, Supervision. George A. Kowalchuk: Resources, Supervision. Wu Xiong: Investigation, Conceptualization, Methodology, Visualization, Writing – review & editing, Supervision, Funding acquisition. Qirong Shen: Resources, Supervision.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
374_2024_1845_MOESM1_ESM.pdf
Supplementary file1 Figure S1. Community dissimilarity of bacteria and protistan predators between organic and chemical fertilizer soil samples. Figure S2. Predicted putative PGPR and their potential PGP effects. Figure S3. The linear relationships between putative PGPR of different effects and putative bacterial plant pathogens. Figure S4. The linear relationships between putative PGPR of different effects and predatory protists. Figure S5. The differences in ribosomal RNA operon (rrn) copy number of bacterial communities in different agricultural ecosystems (a) and their correlations with the relative abundance of predatory protists in soils. Figure S6. The correlations between the relative abundance of Toxin–antitoxin systems among all bacterial pathways predicted by tax2fun2 with the relative abundance of predatory protists in soil protist community. Figure S7. The differences in soil properties including pH, OM (organic matter), TP (total P), TK (total K), TN (total N), AP (available P), AK (available K) and AN (available N) between organic fertilizer and chemical fertilizer soils by STAMP analysis. Figure S8. The correlations between the abundant microbial taxa and soil properties. Figure S9. Spearman’s rank correlations between protistan predators and bacteria genus in the two fertilization regimes with controlling for the influence of organic matter. (PDF 1162 KB)
374_2024_1845_MOESM2_ESM.xlsx
Supplementary file2 Table S1. Details about 379 samples included in the study. Table S2. Predicted putative bacterial pathogens in the two agricultural systems. Table S3. Predicted putative PGPR in the two agricultural systems. Table S4. Differences in the relative abundance of pathways associated with bacterial dormancy. Table S5. Soil properties for samples in the case study. (XLSX 5032 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, C., Zhou, Z., Sun, S. et al. Investigating protistan predators and bacteria within soil microbiomes in agricultural ecosystems under organic and chemical fertilizer applications. Biol Fertil Soils (2024). https://doi.org/10.1007/s00374-024-01845-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00374-024-01845-6