Abstract
This review presents an analysis of formamide, focussing on its occurrence in nature, its functional roles, and its promising applications in the context of the bioeconomy. We discuss the utilization of formamide as an innovative nitrogen source achieved through metabolic engineering. These approaches underscore formamide’s potential in supporting growth and production in biotechnological processes. Furthermore, our review illuminates formamide’s role as a nitrogen source capable of safeguarding cultivation systems against contamination in non-sterile conditions. This attribute adds an extra layer of practicality to its application, rendering it an attractive candidate for sustainable and resilient industrial practices. Additionally, the article unveils the versatility of formamide as a potential carbon source that could be combined with formate or CO2 assimilation pathways. However, its attributes, i.e., enriched nitrogen content and comparatively limited energy content, led to conclude that formamide is more suitable as a co-substrate and that its use as a sole source of carbon for biomass and bio-production is limited. Through our exploration of formamide’s properties and its applications, this review underscores the significance of formamide as valuable resource for a large spectrum of industrial applications.
Key points
• Formidases enable access to formamide as source of nitrogen, carbon, and energy
• The formamide/formamidase system supports non-sterile fermentation
• The nitrogen source formamide supports production of nitrogenous compounds
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Formamide occurrence in nature
Formamide, also known as methanamide, is the simplest naturally occurring (monocarboxylic acid) amide and the smallest molecule with a peptide bond. Its composition includes hydrogen, oxygen, carbon, and nitrogen atoms, which belong to the seven most prevalent elements of the universe (Heiserman 1991), where it ubiquitously occurs (Saladino et al. 2012b). It is present in interstellar clouds (Solomon 1973) and on comets (Despois et al. 2002), estimated to constitute 0.015% of cometary ice in relation to H2O. Furthermore, formamide is a common molecule in star-forming regions in the galactic habitable zone in dense molecular clouds (Adande et al. 2013) and was detected at the galactic center of the Milky Way (Rubin et al. 1971; Gottlieb et al. 1973). Databased hypotheses suggested the presence of liquid formamide in a stratosphere under the frozen mantle surface of celestial bodies of our solar system, including some of the largest icy moons such as Saturn’s satellite Titan (Parnell et al. 2006) and Jupiter’s satellite Europa (Levy et al. 2000; Borucki et al. 2002).
The role of formamide in prebiotic chemistry and the origin of life is controversially discussed (Saladino et al. 2012a). Formamide may have been a precursor for the synthesis of a broad variety of biogenic molecules such as nucleoside bases (Ferus et al. 2015), sugars (Saladino et al. 2015), carboxylic acids, or amino acids (Saladino et al. 2013), with energy provided in form of heat or UV radiation (Saladino et al. 2012b) in the presence of mineral catalysts (Bizzarri et al. 2021).
In some microorganisms, formamide occurs as a degradation product of histidine and cyanide (Wachsman and Barker 1955; Ferber et al. 1988; Kunz et al. 1994). Formamide is a rare metabolite in microbes, e.g., while its role in nitrogen metabolism of Heliobacter pylori as a nitrogen source as well as for protection at acidic conditions is clear, the source of formamide in its habitat, the human gastro-intestinal system, is currently unknown (Skouloubris et al. 2001).
Formamide has an annual market estimated to reach US $270 million by 2027 with the four major uses as feedstock for producing agrochemicals (pesticides, herbicides), plastics, paper, and card board or as solvent in plasticizers for making concrete (IndustyArc, 2022). Chemical synthesis of formamide proceeds by direct carbonylation of ammonia or by aminolysis of methyl formate, and both processes occur under high-temperature and high-pressure conditions. Very recently, electrochemical synthesis of formamide under ambient conditions has been described from either ammonia and methanol (Meng et al. 2022), ammonia and carbon dioxide (Li and Kornienko, 2022), and nitrate or ammonia, or carbon monoxide and nitrite (Lan et al. 2023) in a similar manner as electrosynthesis of methylamine from carbon dioxide and nitrate (Wu et al. 2021). Its uses comprise industrial production of hydrogen cyanide, its application as cryoprotectant or as ionizing solvent in aqueous buffers, but also molecular biology uses such as destabilizing double helices in RNA gel electrophoresis are established (Böckler et al. 2018).
Formamide as nitrogen and/or carbon source for bacteria
Formamide belongs to the reduced C1 nitrogen compounds. While ammonium carbamate and carbamoyl phosphate readily decompose to yield ammonia (also catalyzed by carbamate kinase; EC 2.7.2.2; Pols et al. 2021), the liberation of nitrogen from formamide and monomethylamine, a related C1 nitrogen source, occurs by enzyme catalyzed reactions (Fig. 1). In the case of formamide, the straightforward amide hydrolysis reaction catalyzed by formamidase (EC 3.5.1.49; AmiF/FmdA) yields ammonia and formate. In the case of monomethylamine, nitrogen can be liberated either as ammonia or as L-glutamate. Oxidative deamination of monomethylamine by monomethylamine oxidase (EC 1.4.9.1; MAO) in Gram-positive methylotrophs (Iersel et al. 1986; Dooley et al. 1990; Cai and Klinman 1994) or by a periplasmic methylamine dehydrogenase (Eady and Large 1968; Chistoserdov 1991) yields ammonia and formaldehyde. Alternatively, monomethylamine can first be converted to formaldehyde via N-methyl-L-glutamate in methylotrophs and non-methylotrophs (Chen et al. 2010). Two options for the synthesis of N-methyl-L-glutamate from monomethylamine and 2-oxoglutarate exist. On the one hand, N-methylglutamate synthase (MGS) reductively methylaminates 2-oxoglutarate to N-methyl-L-glutamate in a reaction comparable to glutamate dehydrogenase (GDH, EC 1.4.1.2/3/4). On the other hand, N-methyl-L-glutamate is formed in a two-step sequence involving γ-glutamylmethylamide synthase (GmaS, EC 6.3.4.12) for ATP-dependent methylamidation comparable to glutamine synthetases (GS) followed by MGS operating comparable to glutamine oxoglutarate aminotransferase GOGAT (EC 1.4.1.13/14) (Bamforth and O’Connor 1979). Thus, synthesis of N-methyl-L-glutamate from monomethylamine and 2-oxoglutarate is similar to the well-known GDH and GS/GOGAT reactions for L-glutamate synthesis from ammonia and 2-oxoglutarate (Mindt et al. 2018). Subsequently, N-methyl-L-glutamate is oxidized by N-methyl-L-glutamate dehydrogenase (EC 1.5.99.5, MGD) to yield formaldehyde and L-glutamate. L-glutamate is either oxidatively deaminated to 2-oxoglutarate and ammonia or can be used in transamination reactions directly.
Catabolism of the reduced C1 nitrogen compounds monomethylamine and formamide. Enzymes are boxed in dark grey, liberation of nitrogen as either ammonia or L-glutamate is indicated by boxing in green, linear dissimilation of methanol to carbon dioxide is shaded in light grey, and assimilation of formaldehyde, formate, and carbon dioxide is indicated by blue arrows. Abbreviations: 2 e-, transfer of 2 electrons from various redox cofactors; 2OG, 2-oxoglutarate; AmiF, formamidase (EC 3.5.1.49); FADH, formaldehyde dehydrogenase; FDH, formate dehydrogenase; Glut, L-glutamate; GmaS, γ-glutamylmethylamide synthase (EC 6.3.4.12); MAO, monomethylamine oxidase (EC 1.4.9.1); MDH, methanol dehydrogenase, MGS, N-methylglutamate synthase (operating comparable to either glutamate dehydrogenase (EC 1.4.1.2/3/4) or glutamine oxoglutarate aminotransferase GOGAT (EC 1.4.1.13/14)); MGD, N-methyl-L-glutamate dehydrogenase (EC 1.5.99.5)
While utilization of monomethylamine as C source may occur by assimilation of the generated formaldehyde (e.g., in the ribulosemonophosphate cycle or the serine cycle), formamide yields formate that can be assimilated or oxidized to carbon dioxide, which in turn may be assimilated (e.g., via the Calvin-Benson-Bassham cycle; see the “Formamide as potential C source in biotechnology” section). Efficient utilization of monomethylamine and formamide as nitrogen sources requires prompt dissimilation and/or assimilation of formaldehyde and formate to avoid growth inhibition, which is particularly relevant in the case of formaldehyde.
Since the first observation of formamide utilization in 1976 for Pseudomonas SL-4, which can utilize formamide as sole nitrogen, carbon, and energy source as part of an ecological carbon-nitrogen cycle (Thatcher and Weaver 1976), this trait has been found in other bacteria such as Paracoccus aminophilus and Pseudomonas putida. However, formamide utilization by bacteria is rare as compared to the abundance of bacteria utilizing ammonia, nitrite, and nitrate as nitrogen sources. Thus, transfer of this rare metabolic trait to industrially relevant bacteria offers unique application opportunities.
Formamidase: phylogeny, enzyme activity, biochemical and genetic regulation
The enzyme sub-class EC 3.5.1 comprises many enzymes hydrolyzing linear C–N bonds other than peptide bonds. Some enzymes are active on N-formylated amino acids such as N-formyl-L-aspartate, N-formyl-L-methionine, N-methyl-anthranilate, N-formyl-L-kynurenine, or on 10-formyltetrahydrofolate. Formamidases (EC 3.5.1.49), also known as formamide aminohydrolases, catalyze the hydrolysis of the amide bond in formamide to release formate and ammonia.
Although amidases are widespread among bacteria for degradation of toxic amides (Newton et al. 2000; Fournand and Arnaud 2001; Liu et al. 2020), only few formamide-specific amidases have been identified. Formamidases were detected in Helicobacter pylori (Wyborn et al. 1996; Skouloubris et al. 2001), Bacillus cereus (Soriano-Maldonado et al. 2011), Streptomyces parvulus (Brown et al. 1986), Methylophilus methylotrophus (Wyborn et al. 1994), Cupriavidus necator (formerly known as Ralstonia eutropha) (Friedrich and Mitrenga 1981), and Sinorhizobium meliloti (Yurgel et al. 2022), but only some of these have been structurally and biochemically characterized. Beyond that, formamidases are found in fungi like Aspergillus nidulans (Hynes 1975; Fraser et al. 2001) or Paracoccidioides brasiliensis (Borges et al. 2005) and plants like Arabidopsis thaliana (Fraser et al. 2001) or in the roots of white lupin (Lupinus albus L.) (Rath et al. 2010). Their unambiguous classification is complicated by the ability of other enzymes such as acetamidase from Mycobacterium smegmatis (Draper 1967) and some aliphatic amidases (Egorova et al. 2004; Makhongela et al. 2007; Engelhardt et al. 2009) to hydrolyze formamide and led to the incorrect annotation of > 20 proteins (Soriano-Maldonado et al. 2011).
Phylogenetic analysis revealed that formamidases belong to at least two different groups of enzymes, namely the acetamidase/formamidase super family (FmdA-AmdA, Pfam PF03069), which also includes amidohydrolases of acetamide (Draper 1967), and the nitrilase family, which is a subfamily of the carbon-hydrogen nitrilase superfamily and hydrolyses various nitriles, producing ammonia and the respective carboxylic acid (Bessonnet et al. 2021; Teepakorn et al. 2021). Although both have certain sequence similarities with members of the major amidase families like aliphatic amidases, acylamide aminohydrolases, and nitrilase/cyanide hydratases, these two groups are demarcated (Fig. 2). Between each other, they possess less than 10% sequence similarity (Soriano-Maldonado et al. 2011) and differ in their molecular masses of around 45 kDa (FmdA-AmdA) (Wyborn et al. 1994; Wyborn et al. 1996; Borges et al. 2010) and 34 kDa (nitrilases) (Skouloubris et al. 2001; Thuku et al. 2009; Soriano-Maldonado et al. 2011), respectively. The low similarity between the amino acid sequences of the formamidases of the Fmda-AmdA superfamily member Mycobacterium smegmatis and the nitrilase superfamily member Methylophilus methylotrophus led to the hypothesis of their evolutionary emergence from a common ancestral protein or by early horizontal gene transfer (Wyborn et al. 1996). H. pylori formamidase AmiF (EC 3.5.1.49) was discovered as putative paralogue to aliphatic amidase AmiE of H. pylori and was the first described formamidase of the nitrilase family. AmiF and AmiE share 34% of the amino acid sequence but differ in substrate specificity. AmiF activity is restricted to formamide, whereas AmiE possesses a broader substrate spectrum and was demonstrated to act on propionamide, acetamide, and acrylamide (Skouloubris et al. 2001). Thus, AmiE and AmiF of H. Pylori have supposedly evolved after ancestral gene duplication and represent specialized paralogues (Skouloubris et al. 2001).
Phylogenetic tree of formamidase AmiF (H. pylori 26695) homologues depicted with the protein structure of AmiF of H. pylori. The tree includes the 100 most homologous proteins to formamidase AmiF from H. pylori 26695 (HELPY), identified by PSI-BLAST, and 7 further formamidase proteins of biotechnological interest. The identified enzymes are assigned to at least 9 different enzyme classes (outlined) and some lack classification. Labels refer to the host organism, as defined in Tab. S1. The crystal structure 2E2L of AmiF from H. pylori (Hung et al. 2007) is depicted
Homology database searches using the FmdA-AmdA superfamily formamidase sequence of A. nidulans found that highly conserved formamidase-like sequences are not restricted to microorganisms but also distributed among eukaryotes like A. thaliana and archaea like Aeropyrum pernix (Fig. 2). Functional formamidase expression was exemplarily verified for A. thaliana and the fission yeast Schizosaccharomyces pombe (Fraser et al. 2001).
While the enzyme activities of some formamidases have been characterized in fair detail, a systematic analysis of the substrate specificity beyond formamide is absent. Among the characterized formamidases of the nitrilase family, none acted on urea. However, while H. pylori formamidase only showed activity with formamide but not with acetamide, acrylamide, or propionamide (Skouloubris et al. 2001), M. methylotrophus formamidase hydrolyzed acetamide, butyramide, propionamide, and acrylamide (Wyborn et al. 1996), and B. cereus formamidase also accepted acetamide besides formamide but not alaninamide, butyramide, isobutyramide, leucinamide, glycinamide, or propionamide (Soriano-Maldonado et al. 2011). These divergences are reflected in the kinetic parameters of these enzymes as summarized in Table 1. From a thermodynamic perspective, the formamide cleavage direction is highly favored over the reverse reaction (Beber et al. 2022). Additionally, formamidase’s specific enzyme activities (as shown in Table 1) are in the range of the activities of the average central metabolism enzyme (e.g., kcat of 64 s−1 for M. methylotrophus Wyborn et al. 1994) (Soares et al. 2011) and, most importantly, exceed the activities reported for glutamate dehydrogenase (GDH), the primary NH3 assimilation reaction (e.g., 0.18 mmol min−1 mg−1 of GDH of E. coli Sakamoto et al. 1975).
AmiF of H. pylori and B. cereus belong to the nitrilase superfamily. A characteristic of this family is the presence of a C-E-K (Cys-Glu-Lys) triad in which the active cysteine acts as the nucleophile, glutamate mediates the proton transfer, and lysine stabilizes the tetrahedral transition state (Hung et al. 2007). The hydroxylation of formamide by AmiF from H. pylori comprises two phases. (1) During the acylation reaction, formamide diffuses into the pocket and binds onto the C-E-K triad. Glu60 mediates the proton transfers from the –SH group of C166 to initiate the first attack to the carbon atom of formamide. The transition state negatively charged intermediate is formed, and the instability of the substrate carbonyl oxygen results in the collapse of this intermediate. This leads to the production of an acyl-enzyme intermediate, breaking the C–N bond, and the release of an NH3 molecule. (2) The acyl-enzyme intermediate is deacylated. Glu60 deprotonates a water molecule to start the second nucleophilic attack. Again, a tetrahedral intermediate is formed, the collapse of the unstable intermediate yields a formic acid molecule, and the enzyme is regenerated. In AmiF of B. cereus, W136 is essential for the conformational stability of the enzyme (Soriano-Maldonado et al. 2011), while D168 is the essential residue in AmiF of H. pylori (Skouloubris et al. 2001). The exchange of E140D prohibited enzymatic activity while the binding of formamide was not affected, indicating a key role of E140 for hydrolysis. For H. pylori, it was suggested that this amino acid residue maintains the side chain geometry of the catalytic C-E-K triad and facilitates the docking of the substrate. The amino acid residues Trp137 and Tyr192 generate an exterior wall and therefore form a small room at the active site, which only allows formamide as substrate (Hung et al. 2007).
Biochemical regulation of formamidase activity is known. Compounds such as Hg2+ ions, iodoacetamide, or iodoacetate inhibit the formamidases of B. cereus and H. pylori as they react with a catalytic cysteine residue (Skouloubris et al. 2001; Soriano-Maldonado et al. 2011). Urea and thiourea, which resemble formamide, inhibit the formamidases of B. cereus and M. methylotrophus (Wyborn et al. 1996; Martínez-Rodríguez et al. 2019), and the presence of Ni2+ ions activates the H. pylori enzyme (Bury-Moné et al. 2004).
Genetic regulation of formamidase genes has been described in response to the substrate formamide, the product nitrogen, iron, or carbon source availability. The formamidase gene fmdS from A. nidulans is regulated by AreA-dependent nitrogen metabolite repression and contains multiple GATA sequences in the promoter region for AreA binding (Hynes 1972; Fraser et al. 2001). No induction by formamide was observed, while the response to nitrogen limitation was reduced when carbon was also limited as the activation by AreA is lost (Hynes 1972; Fraser et al. 2001). Bacterial amidases are often induced by the presence of their amide substrates. For example, in M. methylotrophus, the gene cluster fmdCABDEF, which encodes a formamidase, a putative positive regulator, an outer-membrane porin for short-chain amides and urea, and the three subunits for binding protein-dependent high-affinity uptake of short-chain amides and urea (Wyborn et al. 1996), is induced by formamide and urea and repressed by high concentrations of ammonia (Mills et al. 1998). In the gastric pathogen H. pylori, amiF is not substrate-inducible, but the genes amiE, amiF, and ureA are transcriptionally upregulated by acid exposure (Merrell et al. 2003; Bury-Moné et al. 2004). Two metal-dependent transcriptional regulators, nickel homeostasis activator NikR and ferric uptake repressor FurR, are directly or indirectly involved in the acid induction of urease, amidase, and formamidase genes, although both amidases do not contain metal ions. At acidic pH, amiF is derepressed by Fur. Fur is epistatic on NikR, which represses fur. NikR directly responds to changes in cytosolic pH during acid acclimation as it shows pH-dependent DNA binding to its target promoter sequences (Jones et al. 2018). In addition, amiF is controlled by a yet unknown third regulator (Bury-Moné et al. 2004).
Formamide for contamination-free, non-sterile cultivation
Microbial contamination constitutes a major obstacle to the stable performance of bioprocesses, may hamper their economically competitive implementation, and is a threat to product quality and safety (Neu 1992). Typically, this risk is encountered by the addition of antimicrobial agents and sterilization of fermentation vessels, laboratory equipment, and cultivation media (Guo et al. 2020b). However, these measures are expensive in cost, resources, energy, and time and favor the emergence of drug-resistant strains (Neu 1992; Guo et al. 2020b). Therefore, their replacement for innovative, more sustainable strategies is highly desirable.
Recently, some biofuels and chemicals have been successfully produced under non-sterile conditions by exploiting the capacity of extraordinary microbes to withstand inhospitable conditions or to assimilate uncommon macronutrient sources as a selective trait (Thorwall et al. 2020). Most successful approaches relied on the former, e.g., cultivation at elevated temperature of ≥ 50 °C enabled the non-sterile production of poly-γ-glutamate, acetoin, and lactic acid by thermophilic Bacilli strains (Zeng et al. 2013; Zhang et al. 2014; Xiao et al. 2017) and an evolved thermotolerant strain of Thermoanaerobacterium aotearoense (Yang et al. 2015). The salt and pH tolerance of Halomonas bluephagenesis and Halomonas campaniensis LS21 was exploited for the non-sterile production of poly-3-hydroxybutyrate-co-4-hydroxybutyrate (Ye et al. 2018) and poly-3-hydroxybutyrate (Jiang et al. 2017). The non-sterile production of an anticancer polysaccharide by an evolved methanol-tolerant mutant strain of Chaetomium globosum serves as another example (Wang et al. 2019). However, this strategy is limited to some extraordinary microbes as it is prone to slight environmental changes (Barig et al. 2011), and unwanted mutations may arise (Ling et al. 2014). Another limitation is that the desired product must tolerate the extraordinary culture conditions (Thorwall et al. 2020).
Alternatively, a competitive advantage can be conferred by engineering the target strain to harness a rare xenobiotic compound as an essential growth nutrient, which is not accessible to (most) competing microbes and thus enables cultivation and production in the respective auxotrophic medium. To construct an antimicrobial contamination system, naturally formamidase-deficient B. subtilis and E. coli were equipped with codon-optimized versions of formamidase genes from H. pylori 26695 and Paenibacillus pasadenensis CS0611, respectively. However, slight growth of the formamidase-deficient E. coli control strain with formamide was observed (Ou et al. 2019), which was in accordance with little growth of wild-type strains, when melamine or cyanamide assimilation was introduced as selective advantage (Shaw et al. 2016). Therefore, a second key nutritional constraint in the form of phosphite dehydrogenase-mediated phosphite utilization was added (Ou et al. 2019), yielding a dual protection system, which was similarly introduced in B. subtilis (Guo et al. 2020a). The power of the dual protection system was demonstrated by outcompeting representative eukaryotic and bacterial competitors since the engineered target strains constituted > 90% of the final culture composition after 30–35 h (Ou et al. 2019; Guo et al. 2020a). The metabolic selection pressure was sufficient to ensure the maintenance of the phosphite dehydrogenase-encoding plasmid for 17 serial dilutions, whereas the same plasmid was lost when commonly accessible phosphate was provided (Schwardmann et al. 2022), and enabled a non-sterile fed-batch fermentation for acetoin production (Guo et al. 2020a).
The presence of undesired competitor organisms was hypothesized to be due to leaked phosphite and formamide degradation products. This was demonstrated regarding ammonium leakage of ammonium from a formamidase-positive C. glutamicum strain that allowed growth of a second formamidase-deficient strain in co-cultivation (Schwardmann et al. 2023b). High formamidase activity (4-fold higher in crude extracts of C. glutamicum as compared to B. subtilis; 6 compared to 1.2 U mg−1) may have led to surplus ammonium formation (Guo et al. 2020a; Schwardmann et al. 2023b), indicating that the catalytic activity must be fine-tuned to the utilization capacity of the host to avoid nutrient leakage (Shaw et al. 2016).
Besides formamide, melamine and cyanamide present the only uncommon nitrogen sources exploited for contamination control in the cyanobacterium Synechococcus sp. PCC 7002 (Selão et al. 2019) and E. coli, Saccharomyces cerevisiae, and Yarrowia lipolyptica, respectively (Shaw et al. 2016), and shown to be applicable to prevent contamination (Shaw et al. 2016; Selão et al. 2019).
Taken together, the utilization of formamide as a rare xenobiotic nutrient provides a promising tool to ensure ample dominance of a formamidase-positive strain, although it cannot guarantee completely contamination-free non-sterile cultivation and necessitates appropriate expression levels.
Formamide as potential C source in biotechnology
Formamide has the potential to serve as a source of carbon, nitrogen, and energy, supporting the growth of microorganisms for biotechnological applications. The enzymatic breakdown of formamide by formamidase yields formate, which is a natural carbon source and source of reducing power for certain formate-utilizing microorganisms. These organisms either utilize formate directly through pathways (Fig. 3) like the reductive acetyl-CoA pathway or the serine cycle or first oxidize formate to CO2 and use less efficient CO2 assimilation pathways such as the reductive pentose phosphate pathway (Calvin-Benson-Bassham cycle) (Bar-Even 2016). In recent years, the use of formate as a CO2-based renewable and scalable feedstock was proposed with the aim of promoting a sustainable bioeconomy (Yishai et al. 2016).
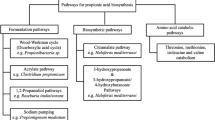
Formamide as a formate source to support growth and production via formate and CO2 assimilation pathway. Schemes illustrate formate (blue) and CO2 (orange) utilization via the reductive Acetyl-CoA pathway, the reductive Glycine pathway, the Serine Cycle, and the Calvin-Benson-Bassham (CBB) cycle (from left to right)
Consequently, there has been a concerted effort to introduce both natural and engineered routes for the incorporation of formate through direct formate or CO2 assimilation into biotechnologically significant bacteria and yeast (Fig. 3) (Yu and Liao 2018; Gleizer et al. 2019; Claassens et al. 2020; Kim et al. 2020; Turlin et al. 2022; Wenk et al. 2022; Bruinsma et al. 2023). These projects have led to the creation of synthetic autotrophs and formatotrophs. Here, particularly noteworthy is the successful implementation of the highly ATP-efficient synthetic reductive glycine pathway in E. coli (Claassens et al. 2020; Kim et al. 2020; Turlin et al. 2022; Bruinsma et al. 2023), which enables fast growth and high biomass yields of the engineered E. coli strains, outperforming the growth performance of some natural formatotrophic strains (Cotton et al. 2020; Kim et al. 2023). Nonetheless, a significant limitation of relying on formate as a sole feedstock arises from the necessity to oxidize a substantial portion of it to CO2 in order to generate the required reduction power to attain the reduction state of cellular biomass or the targeted bioproduct. This loss of feedstock for assimilation poses a challenge by influencing the carbon footprint and the resource efficiency of the process. To mitigate these shortcomings, CO2 recycling can contribute to enhance the sustainability of the process. In this context, feedstocks with higher levels of reduction, such as methanol, could potentially offer a more advantageous alternative.
When originating from formamide, a 1:1 proportion of formate to ammonium is produced. However, the elemental composition of a bacterial cell exhibits a carbon and nitrogen content of 4:1 (Milo and Phillips 2015). Thus, the equimolar provision of carbon and nitrogen by formamide is not ideal to support growth. By comparison, glucosamine represents a much more suitable sole source of carbon, nitrogen, and energy for C. glutamicum (Uhde et al. 2013). To address these limitations, an optimal strategy involves co-feeding formamide alongside supplementary carbon sources. This approach would result in reduced formate-derived CO2 loss and a more efficient utilization of the excess nitrogen inherent in formamide as a feedstock. Formate already has applications as a co-substrate, by either providing reducing power or by enhancing the carbon yield. For example, formate has been used as co-substrate together with glucose in order to balance reducing equivalents for anaerobic succinate production by C. glutamicum (Litsanov et al. 2012), as well as its co-substrate role with glucose in Ustilago cynodontis for itaconate production (Ullmann et al. 2022). Additionally, formate-derived CO2 was used to enhance ethanol production from glucose in E. coli (Tseng et al. 2018) and from glucose and galactose in S. cerevisiae (Guadalupe-Medina et al. 2013).
However, some organisms do not easily support carbon source co-utilization due to metabolite repression mechanisms that lead to sequential carbon source utilization. This behavior is observed in bacteria such as E. coli (Alva et al. 2020) and B. subtilis (Fujita 2009). Overcoming this limitation necessitates metabolic engineering strategies (Wendisch et al. 2016). Consequently, organisms that effectively co-utilize diverse carbon sources, e.g., C. glutamicum (Blombach and Seibold 2010; Teramoto et al. 2011), provide a clear process development advantage.
Taken together, while the use of formamide as nitrogen source is relevant for biotechnology applications, using formamide as sole carbon source is of limited value if the surplus of energy-intensive ammonium is not utilized. Here, supplementation of additional formate or other C1-compounds could be of value, similar co-feeding strategies with, e.g., sugar-based feedstocks.
Formamide as N source in biotechnology
Biotechnological processes commonly rely on inorganic ammonia, ammonium salts, urea, or organic substrates like peptone or yeast extract as nitrogen sources because they support fastest growth (Reitzer 1996). In contrast to the extensive engineering efforts to broaden the carbon source spectrum for major biotechnological workhorses (Wendisch et al. 2016), only few alternative, non-conventional nitrogen sources have been made accessible. However, their utilization often is unfavorable due, e.g., to the carcinogenicity of melamine (Shaw et al. 2016). Formamide by contrast shows low toxicity (Kennedy 2014), and its presence in concentration of up to 160 and 700 mM only had a minor inhibitory effect on growth of E. coli and C. glutamicum, respectively (Ou et al. 2019; Schwardmann et al. 2023b). Moreover, the use of formamide does not suffer from nitrogen repression if supplemented as the sole source of nitrogen, but not as the primary carbon source (Guo et al. 2020a; Schwardmann et al. 2023b). Both studies recruited formamidase AmiF from H. pylori (Van Vliet et al. 2003; Guo et al. 2020a; Schwardmann et al. 2023b), whereas the anticontamination system for E. coli relied on AmiF from P. pasadenensis (Guo et al. 2017; Ou et al. 2019), that only share 48% identity of their amino acid sequences.
In the context of exploiting the formamidase/formamide strategy as anticontamination system, non-sterile fermentation of a B. subtilis acetoin producer strain, supplemented with 60 mM formamide as sole source of nitrogen, yielded a titer of about 25 g L−1 (Table 2), which is comparable to those from conventional media (Guo et al. 2020a). As acetoin does not contain nitrogen atoms, nitrogen from formamide was exclusively allocated to the synthesis of biomass and natural metabolites but did not end up in the product. Thus, restricted nitrogen availability did not limit production. The growth limitation upon nitrogen deprivation provides a potential tool to perform a two-stage cultivation with growth-accompanied production during the first phase until nitrogen is depleted, followed by sole product synthesis in the second phase. This concept of nutrient content–dependent two-stage cultivation was successfully used to decouple amino acid production from growth in C. glutamicum strains, engineered to prevent glucose utilization for growth, allocating it exclusively to production after acetate depletion (Blombach et al. 2008). Similarly, it could be elaborated for formamide to completely decouple biomass and biosynthesis of nitrogen-free products. For the bioconversion of xylose to produce xylitol and xylonate in C. glutamicum (Schwardmann et al. 2023a), nitrogen starvation–inducible promoters combined with balanced provision of ammonia allowed for growth-decoupled production. What these approaches have in common is that they only work for non-nitrogenous target compounds such as acetoin or xylitol. Notably, nitrogen limitation has often been used to improve polyhydroxybutyrate production by strains of C. necator due to the increased NADPH availability (Zhang et al. 2022). Therefore, the implementation of the formamide/formamidase system as a selective trait in this organism may be interesting for polyhydroxybutyrate production under non-sterile conditions in a two-stage process.
To target formamide-based production of nitrogenous compounds such as amines and amino acids, high concentrations of formamide have to be supplemented as nitrogen source for growth and production. This has been achieved for C. glutamicum (Schwardmann et al. 2023b). Stable isotope labeling using 15N-labeled formamide or ammonium sulfate confirmed the incorporation of nitrogen from formamide into both biomass and the exemplary nitrogenous product L-lysine (Schwardmann et al. 2023b). Notably, the simultaneous provision with natively accessible ammonium sulfate and xenobiotic formamide revealed similar acceptance and incorporation of nitrogen from both substrates into biomass (Schwardmann et al. 2023b). Beyond formamide-based production of the feed amino acid L-lysine, the system was transferred to established producer strains for formamide-based production of the food amino acid L-glutamate, the N-alkylated amino acid N-methylphenylalanine, and the aromatic dicarboxylate dipicolinic acid (Schwardmann et al. 2023b) (Fig. 4A, Table 2). Formamide was even superior to the standard nitrogen source mixture of urea and ammonium sulfate for yielding up to 80% increased titers of all four products (Schwardmann et al. 2023b).
Engineering of C. glutamicum for formamide-based production of nitrogenous compounds (A) and for co-cultivation with a formamidase-deficient C. glutamicum strain (B). C. glutamicum was engineered to overexpress formamidase (AmiF) gene from H. pylori 26695 (FORM, blue) to produce L-glutamate, L-lysine, dipicolinic acid, and N-methylphenylalanine in nitrogen-substituted minimal salt medium using glucose and formamide as sole sources of carbon and nitrogen. Ammonium release from formamide hydrolysis by AmiF by a formamidase-positive C. glutamicum strain (FORM) supports growth of a formamidase-deficient C. glutamicum strain (WT) in co-cultivation. For differentiation, strains were overexpressing genes either for the fluorescence protein GfpUV (green) or Crimson (red) (B)
Formamide-based production using recombinant E. coli and C. glutamicum strains revealed a clear tradeoff between immediate fast growth at low formamide concentrations and the support of higher biomass formation with increasing formamide concentrations (Guo et al. 2020a; Schwardmann et al. 2023b). Accumulation of formate as a degradation product from formamide hydrolysis was detected, and its growth inhibitory effect (Witthoff et al. 2012; Schwardmann et al. 2023b) limited cultivation at higher formamide concentrations. The formate problem was solved by oxidation to carbon dioxide. To this end, either native NAD-dependent formate dehydrogenase (Witthoff et al. 2012) or heterologous NADP-dependent formate dehydrogenase variant from Pseudomonas sp. 101 (Calzadiaz-Ramirez et al. 2020) was used (Schwardmann et al. 2023b). Probably reflecting the catabolic nature of formamide utilization as nitrogen source, NAD-dependent formate dehydrogenase was superior to improve growth characteristics when higher formamide concentrations were used as nitrogen source (Schwardmann et al. 2023b).
A completely different application of formamide offers its use in synthetic microbial consortia, since these typically depend on the (mutual inter-)dependency of both microbial partner strains, e.g., based on cross-feeding of essential nutrients (Sgobba and Wendisch 2020). In nature, ammonium cross-feeding occurs between algae and N2-fixing or methylamine-degrading bacteria (Suleiman et al. 2016; Zecher et al. 2020; Ambrosio and Curatti 2021). In contrast to the exploitation of formamidase-driven formamide degradation as selective trait in E. coli and B. subtilis (Ou et al. 2019; Guo et al. 2020a), formamidase overexpression in C. glutamicum led to ammonium leakage into the medium. The leaked ammonium was sufficient to support growth of a second formamidase-deficient strain in co-cultivation with formamide as the sole nitrogen source (Schwardmann et al. 2023b). Differentiation between formamidase-positive and -deficient strains by the expression of either of the genes for the fluorescence proteins GfpUV and Crimson revealed that inoculation with about 10% formamidase-positive cells and 90% formamidase-negative cells was sufficient to support growth of both strains (Schwardmann et al. 2023b).
The demonstrated strict obligatory intra-species dependency (Fig. 4B) provides a promising basis for using nitrogen cross-feeding from formamide in combination with a second conversely cross-fed metabolite. This could be achieved in inter- or intra-species consortia. However, while these kinds of synthetic consortia are intensely discussed for potential application in biotechnology (McCarty and Ledesma-Amaro 2019; Sgobba and Wendisch 2020; Cao et al. 2022), a number of limitations have to be considered. Often, the metabolic costs of transport processes are neglected although primary or secondary active transport entails a metabolic burden (or cost in the form of ATP, ion gradients, etc.) and uptake/export of charged/uncharged species of a molecule imparts cost in form of the decoupled transmembrane pH/ion gradient perturbing ATP synthesis, e.g., by FoF1 ATP synthase (Krämer 1994). However, the metabolic cost associated with transport processes may be used in the design of synthetic microbial consortia by transport engineering (Pérez-García and Wendisch 2018). Thus, the demonstrated strict obligatory intra-species dependency via formamide/formamidase (Fig. 4B) is an interesting design option for synthetic consortia, but it has to be kept in mind that division of labor in a synthetic consortium must outcompete monocultures that perform all labor by one cell to achieve applicability in biotechnology.
Outlook
The utilization of formamide/formamidase shows application potential for non-sterile fermentations, as rare nitrogen source for fermentative production under growth and non-growth conditions, as well as in synthetic microbial consortia. However, some inherent limitations characterize the formamide/formamidase system. Although formamide shows low toxicity towards cells as it affects DNA and RNA helicity (Blake and Delcourt 1996), it inhibits bacterial oxidases (Gupta and Mazumdar 2008) and dissociates and inactivates some enzymes such as E. coli alkaline phosphatase (Falk et al. 1982). For applications in thermophiles, additional considerations have to be taken since hot formamide introduces formyl groups into proteins to generate formyl-glycyl and diformyl-lysyl residues (Perkins 1965). Addition of amino acids and/or purines to the growth medium may alleviate some toxic effects as shown for E. coli (Wheeler and Grammer 1960).
Inhibition due to formate generated from formamide by formamidase was overcome by the overexpression of formate dehydrogenase genes in engineered bacteria. This mimics the fast assimilation or dissimilation of formate in natural formamidase-positive bacteria. However, since some formamidase enzymes show slower hydrolysis of other short-chain amides such as acetamide, propanamide, and butanamide, these side reactions may perturb growth and production with formamide (Clarke 1969; Wyborn et al. 1996; Soriano-Maldonado et al. 2011).
The enzymatic activity of formamidase may be relevant in cell-free biocatalaysis or whole-cell biotransformation. Purified B. cereus formamidase was used in cross-linked enzyme crystals (CLECs) in a microfluidic setup (Conejero-Muriel et al. 2015). Here, its side activity as acyl-transferase was used to produce acetohydroxamic acid, an inhibitor of urease used to treat chronic urea-splitting urinary infection (Lithostat®). It may be possible to use B. cereus formamidase in whole-cell biotransformation under non-growth conditions provided that acetohydroxamic acid may be exported by the producing cells. Another enzyme application may be similar to the use of the phosphite dehydrogenase/phosphite system. Fusions between the NADPH-dependent monooxygenase P450BM3 with phosphite dehydrogenase accepted phosphite as cheap electron donor for monooxygenase reactions (Beyer et al. 2017). In principle, it is conceivable to use formamidase to provide ammonia in situ for reactions of ammonia-dependent enzymes. In this respect, urease has been used to provide ammonia via hydrolysis of urea in situ for the synthesis of polyhydroquinoline and polyhydroacridine derivatives. The reactions of the aryl aldehydes with dimedone or ethyl acetoacetate and the in situ generated ammonia occurred in water under mild green conditions (Zhu and Li 2021). Besides ammonia, urease generates carbon dioxide gas but formamidase formic acid; thus, the application of these enzymes in bio-chemo-catalysis has to consider the respective by-products with regard to possible perturbation or beneficial aspects.
While first steps to exploit the formamide/formamidase system in fermentative production and enzyme catalysis, for non-sterile fermentations, and to tune synthetic microbial consortia have been taken, the full potential has yet to be tapped by future research.
Data availability
All data are present in the manuscript and its Supplement.
Code availability
Not applicable.
References
Adande GR, Woolf NJ, Ziurys LM (2013) Observations of interstellar formamide: availability of a prebiotic precursor in the galactic habitable zone. Astrobiology 13:439–453. https://doi.org/10.1089/ast.2012.0912
Alva A, Sabido-Ramos A, Escalante A, Bolívar F (2020) New insights into transport capability of sugars and its impact on growth from novel mutants of Escherichia coli. Appl Microbiol Biotechnol 104:1463–1479. https://doi.org/10.1007/s00253-019-10335-x
Ambrosio R, Curatti L (2021) Deferred control of ammonium cross-feeding in a N2-fixing bacterium-microalga artificial consortium. Appl Microbiol Biotechnol 105:2937–2950. https://doi.org/10.1007/s00253-021-11210-4
Bamforth CW, O’Connor ML (1979) The isolation of pleiotropic mutants of Pseudomonas aminovorans deficient in the ability to grow on methylamine and an examination of their enzymic constitution. J Gen Microbiol 110:143–149. https://doi.org/10.1099/00221287-110-1-143
Bar-Even A (2016) Formate Assimilation: The metabolic architecture of natural and synthetic pathways. Biochemistry 55:3851–3863. https://doi.org/10.1021/acs.biochem.6b00495
Barig S, Alisch R, Nieland S, Wuttke A, Gräser Y, Huddar M, Schnitzlein K, Stahmann K-P (2011) Monoseptic growth of fungal lipase producers under minimized sterile conditions: cultivation of Phialemonium curvatum in 350 L scale. Eng Life Sci 11:387–394. https://doi.org/10.1002/elsc.201000219
Beber ME, Gollub MG, Mozaffari D, Shebek KM, Flamholz AI, Milo R, Noor E (2022) eQuilibrator 3.0: a database solution for thermodynamic constant estimation. Nucleic Acids Res 50:D603–D609. https://doi.org/10.1093/nar/gkab1106
Bessonnet T, Mariage A, Petit J-L, Pellouin V, Debard A, Zaparucha A, Vergne-Vaxelaire C, de Berardinis V (2021) Purification and characterization of Nitphym, a robust thermostable nitrilase from Paraburkholderia phymatum. Front Bioeng Biotechnol 9:2296–4185. https://doi.org/10.3389/fbioe.2021.686362
Beyer N, Kulig JK, Bartsch A, Hayes MA, Janssen DB, Fraaije MW (2017) P450BM3 fused to phosphite dehydrogenase allows phosphite-driven selective oxidations. Appl Microbiol Biotechnol 101:2319–2331. https://doi.org/10.1007/s00253-016-7993-7
Bizzarri BM, Saladino R, Delfino I, García-Ruiz JM, Di Mauro E (2021) Prebiotic organic chemistry of formamide and the origin of life in planetary conditions: what we know and what is the future. Int J Mol Sci 22:917. https://doi.org/10.3390/ijms22020917
Blake RD, Delcourt SG (1996) Thermodynamic effects of formamide on DNA stability. Nucleic Acids Res 24:2095–2103. https://doi.org/10.1093/nar/24.11.2095
Blombach B, Schreiner ME, Bartek T, Oldiges M, Eikmanns BJ (2008) Corynebacterium glutamicum tailored for high-yield l-valine production. Appl Microbiol Biotechnol 79:471–479. https://doi.org/10.1007/s00253-008-1444-z
Blombach B, Seibold GM (2010) Carbohydrate metabolism in Corynebacterium glutamicum and applications for the metabolic engineering of l-lysine production strains. Appl Microbiol Biotechnol 86:1313–1322. https://doi.org/10.1007/s00253-010-2537-z
Böckler F, Dill B, Eisenbrand G, Faupel F, Fugmann B, Gamse T, Matissek R, Pohnert G, Rühling A, Schmidt S, Sprenger G (2018) Formamid. https://roempp.thieme.de/lexicon/RD-06-01643 (accessed 13.08.2023)
Borges CL, Parente JA, Barbosa MS, Santana JM, Báo SN, De Sousa MV, De Almeida Soares CM (2010) Detection of a homotetrameric structure and protein-protein interactions of Paracoccidioides brasiliensis formamidase lead to new functional insights. FEMS Yeast Res 10:104–113. https://doi.org/10.1111/j.1567-1364.2009.00594.x
Borges CL, Pereira M, Felipe MSS, de Faria FP, Gomez FJ, Deepe GS, Soares CMA (2005) The antigenic and catalytically active formamidase of Paracoccidioides brasiliensis: protein characterization, cDNA and gene cloning, heterologous expression and functional analysis of the recombinant protein. Microbes Infect 7:66–77. https://doi.org/10.1016/j.micinf.2004.09.011
Borucki JG, Khare B, Cruikshank DP (2002) A new energy source for organic synthesis in Europa’s surface ice. J Geophys Res Planets 107:24-1–24–5. https://doi.org/10.1029/2002JE001841
Brown D, Hitchcock MJM, Katz E (1986) Purification and characterization of kynurenine formamidase activities from Streptomyces parvulus. Can J Microbiol 32:465–472. https://doi.org/10.1139/m86-086
Bruinsma L, Wenk S, Claassens NJ, Martins dos Santos VAP (2023) Paving the way for synthetic C1 - metabolism in Pseudomonas putida through the reductive glycine pathway. Metab Eng 76:215–224. https://doi.org/10.1016/j.ymben.2023.02.004
Bury-Moné S, Thiberge J-M, Contreras M, Maitournam A, Labigne A, De Reuse H (2004) Responsiveness to acidity via metal ion regulators mediates virulence in the gastric pathogen Helicobacter pylori: regulation of the adaptive response of H. pylori to moderate acidity. Mol Microbiol 53:623–638. https://doi.org/10.1111/j.1365-2958.2004.04137.x
Cai D, Klinman JP (1994) Copper amine oxidase: heterologous expression, purification, and characterization of an active enzyme in Saccharomyces cerevisiae. Biochemistry 33:7647–7653. https://doi.org/10.1021/bi00190a019
Calzadiaz-Ramirez L, Calvó-Tusell C, Stoffel GMM, Lindner SN, Osuna S, Erb TJ, Garcia-Borràs M, Bar-Even A, Acevedo-Rocha CG (2020) In vivo selection for formate dehydrogenases with high efficiency and specificity toward NADP+. ACS Catal 10:7512–7525. https://doi.org/10.1021/acscatal.0c01487
Cao Z, Yan W, Ding M, Yuan Y (2022) Construction of microbial consortia for microbial degradation of complex compounds. Front Bioeng Biotechnol 10:1051233. https://doi.org/10.3389/fbioe.2022.1051233
Chen Y, McAleer KL, Murrell JC (2010) Monomethylamine as a nitrogen source for a nonmethylotrophic bacterium, Agrobacterium tumefaciens. Appl Environ Microbiol 76:4102–4104. https://doi.org/10.1128/AEM.00469-10
Chistoserdov AY (1991) Genetic organization of methylamine utilization genes from Methylobacterium extorquens AMI. J Bacteriol 173:5901–5908. https://doi.org/10.1128/jb.173.18.5901-5908.1991
Claassens NJ, Bordanaba-Florit G, Cotton CAR, De Maria A, Finger-Bou M, Friedeheim L, Giner-Laguarda N, Munar-Palmer M, Newell W, Scarinci G, Verbunt J, de Vries ST, Yilmaz S, Bar-Even A (2020) Replacing the Calvin cycle with the reductive glycine pathway in Cupriavidus necator. Metab Eng 62:30–41. https://doi.org/10.1016/j.ymben.2020.08.004
Clarke PH (1969) The aliphatic amidases of Pseudomonas aeruginosa. Adv Microb Physiol 4:179–222. https://doi.org/10.1016/S0065-2911(08)60442-7
Conejero-Muriel M, Rodríguez-Ruiz I, Martínez-Rodríguez S, Llobera A, Gavira JA (2015) McCLEC, a robust and stable enzymatic based microreactor platform. Lab Chip 15:4083–4089. https://doi.org/10.1039/C5LC00776C
Cotton CA, Claassens NJ, Benito-Vaquerizo S, Bar-Even A (2020) Renewable methanol and formate as microbial feedstocks. Curr Opin Biotechnol 62:168–180. https://doi.org/10.1016/j.copbio.2019.10.002
Despois D, Crovisier J, Bockelée-Morvan D, Biver N (2002) Comets and prebiotic chemistry: the volatile component. In: Pole R (ed) First European workshop on exo-astrobiology, Graz, Austria, pp 123–127
Dooley DM, McIntire WS, McGuirl MA, Cote CE, Bates JL (1990) Characterization of the active site of Arthrobacter P1 methylamine oxidase: evidence for copper-quinone interactions. J Am Chem Soc 112:2782–2789. https://doi.org/10.1021/ja00163a047
Draper P (1967) The aliphatic acylamide amidohydrolase of Mycobacterium smegmatis: its inducible nature and relation to acyl-transfer to hydroxylamine. Microbiology 46:111–123. https://doi.org/10.1099/00221287-46-1-111
Eady RR, Large PJ (1968) Purification and properties of an amine dehydrogenase from Pseudomonas AM1 and its role in growth on methylamine. Biochem J 106:245–255. https://doi.org/10.1042/bj1060245
Egorova K, Trauthwein H, Verseck S, Antranikian G (2004) Purification and properties of an enantioselective and thermoactive amidase from the thermophilic actinomycete Pseudonocardia thermophila. Appl Microbiol Biotechnol 65:38–45. https://doi.org/10.1007/s00253-004-1607-5
Engelhardt BE, Jordan MI, Repo ST, Brenner SE (2009) Phylogenetic molecular function annotation. J Phys Conf Ser 180:012024. https://doi.org/10.1088/1742-6596/180/1/012024
Falk MC, Bethune JL, Vallee BL (1982) Formamide-induced dissociation and inactivation of Escherichia coli alkaline phosphatase. Metal-dependent reassociation and restoration of activity from isolated subunits. Biochemistry 21:1471–1478. https://doi.org/10.1021/bi00536a001
Ferber DM, Khambaty F, Ely B (1988) Utilization of histidine by Caulobacter crescentus. Microbiology 134:2149–2154. https://doi.org/10.1099/00221287-134-8-2149
Ferus M, Nesvorný D, Šponer J, Kubelík P, Michalčíková R, Shestivská V, Šponer JE, Civiš S (2015) High-energy chemistry of formamide: a unified mechanism of nucleobase formation. Proc Natl Acad Sci 112:657–662. https://doi.org/10.1073/pnas.1412072111
Fournand D, Arnaud A (2001) Aliphatic and enantioselective amidases: from hydrolysis to acyl transfer activity. J Appl Microbiol 91(3):381–393. https://doi.org/10.1046/j.1365-2672.2001.01378.x
Fraser JA, Davis MA, Hynes MJ (2001) The formamidase gene of Aspergillus nidulans : regulation by nitrogen metabolite repression and transcriptional interference by an overlapping upstream gene. Genetics 157:119–131. https://doi.org/10.1093/genetics/157.1.119
Friedrich CG, Mitrenga G (1981) Utilization of aliphatic amides and formation of two different amidases by Alcaligenes eutrophus. Microbiology 125:367–374. https://doi.org/10.1099/00221287-125-2-367
Fujita Y (2009) Carbon catabolite control of the metabolic network in Bacillus subtilis. Biosci Biotechnol Biochem 73:245–259. https://doi.org/10.1271/bbb.80479
Gleizer S, Ben-Nissan R, Bar-On YM, Antonovsky N, Noor E, Zohar Y, Jona G, Krieger E, Shamshoum M, Bar-Even A, Milo R (2019) Conversion of Escherichia coli to generate all biomass carbon from CO2. Cell 179:1255–1263.e12. https://doi.org/10.1016/j.cell.2019.11.009
Gottlieb CA, Palmer P, Rickard LJ, Zuckerman B (1973) Studies of interstellar formamide. Astrophys J 182:699–710. https://doi.org/10.1086/152178
Guadalupe-Medina V, Wisselink HW, Luttik MA, de Hulster E, Daran J-M, Pronk JT, van Maris AJ (2013) Carbon dioxide fixation by Calvin-Cycle enzymes improves ethanol yield in yeast. Biotechnol Biofuels 6:125. https://doi.org/10.1186/1754-6834-6-125
Guo X, Xu P, Zong M, Lou W (2017) Purification and characterization of alkaline chitinase from Paenibacillus pasadenensis CS0611. Chin J Catal 38:665–672. https://doi.org/10.1016/S1872-2067(17)62787-6
Guo Z-W, Ou X-Y, Liang S, Gao H-F, Zhang L-Y, Zong M-H, Lou W-Y (2020a) Recruiting a phosphite dehydrogenase/formamidase-driven antimicrobial contamination system in Bacillus subtilis for nonsterilized fermentation of acetoin. ACS Synth Biol 9:2537–2545. https://doi.org/10.1021/acssynbio.0c00312
Guo Z-W, Ou X-Y, Xu P, Gao H-F, Zhang L-Y, Zong M-H, Lou W-Y (2020b) Energy- and cost-effective non-sterilized fermentation of 2,3-butanediol by an engineered Klebsiella pneumoniae OU7 with an anti-microbial contamination system. Green Chem 22:8584–8593. https://doi.org/10.1039/D0GC03044A
Gupta S, Mazumdar S (2008) Inhibition of bacterial oxidases by formamide and analogs. Biol Chem 389:599–607. https://doi.org/10.1515/BC.2008.059
Heiserman D (1991) Exploring chemical elements and their compounds. McGraw-Hill Companies, Incorporated
Hung C-L, Liu J-H, Chiu W-C, Huang S-W, Hwang J-K, Wang W-C (2007) Crystal structure of Helicobacter pylori formamidase AmiF reveals a cysteine-glutamate-lysine catalytic triad. J Biol Chem 282:12220–12229. https://doi.org/10.1074/jbc.M609134200
Hynes MJ (1972) Mutants with altered glucose repression of amidase enzymes in Aspergillus nidulans. J Bacteriol 111:717–722. https://doi.org/10.1128/jb.111.3.717-722.1972
Hynes MJ (1975) Amide utilization in Aspergillus nidulans: evidence for a third amidase enzyme. J Gen Microbiol 91:99–109. https://doi.org/10.1099/00221287-91-1-99
Iersel J, Meer RA, Duine JA (1986) Methylamine oxidase from Arthrobacter P1. A bacterial copper-quinoprotein amine oxidase. Eur J Biochem 161:415–419. https://doi.org/10.1111/j.1432-1033.1986.tb10461.x
IndustryArc (2022) Formamide market report, 2022-2027. https://www.industryarc.com/Research/Global-Formamide-Market-Research-511870 (accessed 17.08.2023).
Jiang X-R, Yao Z-H, Chen G-Q (2017) Controlling cell volume for efficient PHB production by Halomonas. Metab Eng 44:30–37. https://doi.org/10.1016/j.ymben.2017.09.004
Jones MD, Li Y, Zamble DB (2018) Acid-responsive activity of the Helicobacter pylori metalloregulator NikR. Proc Natl Acad Sci 115:8966–8971. https://doi.org/10.1073/pnas.1808393115
Kennedy GL (2014) Formamide. In: Wexler P (ed) Encyclopedia of toxicology. Academic Press, pp 657–658
Kim S, Giraldo N, Rainaldi V, Machens F, Collas F, Kubis A, Kensy F, Bar-Even A, Lindner SN (2023) Optimizing E. coli as a formatotrophic platform for bioproduction via the reductive glycine pathway. Front Bioeng Biotechnol 11:2296–4185. https://doi.org/10.3389/fbioe.2023.1091899
Kim S, Lindner SN, Aslan S, Yishai O, Wenk S, Schann K, Bar-Even A (2020) Growth of E. coli on formate and methanol via the reductive glycine pathway. Nat Chem Biol 16:538–545. https://doi.org/10.1038/s41589-020-0473-5
Krämer R (1994) Functional principles of solute transport systems: concepts and perspectives. Biochim Biophys Acta BBA - Bioenerg 1185:1–34. https://doi.org/10.1016/0005-2728(94)90189-9
Kunz DA, Wang C-S, Chen J-L (1994) Alternative routes of enzymic cyanide metabolism in Pseudomonas fluorescens NCIMB 11764. Microbiology 140:1705–1712. https://doi.org/10.1099/13500872-140-7-1705
Lan J, Wei Z, Lu YR, Chen D, Zhao S, Chan TS, Tan Y (2023) Efficient electrosynthesis of formamide from carbon monoxide and nitrite on a Ru-dispersed Cu nanocluster catalyst. Nat Commun 14:2870. https://doi.org/10.1038/s41467-023-38603-5
Levy M, Miller SL, Brinton K, Bada JL (2000) Prebiotic synthesis of adenine and amino acids under Europa-like conditions. Icarus 145:609–613. https://doi.org/10.1006/icar.2000.6365
Li J, Kornienko N (2022) Electrochemically driven C–N bond formation from CO2 and ammonia at the triple-phase boundary. Chem Sci 13:3957–3964. https://doi.org/10.1039/D1SC06590D
Ling H, Teo W, Chen B, Leong SSJ, Chang MW (2014) Microbial tolerance engineering toward biochemical production: from lignocellulose to products. Curr Opin Biotechnol 29:99–106. https://doi.org/10.1016/j.copbio.2014.03.005
Liu J, Zhang Y, Feng K, Liu X, Li J, Li C, Zhang P, Yu Q, Liu J, Shen G, He L (2020) Amidase a novel detoxifying enzyme is involved in cyflumetofen resistance in Tetranychus cinnabarinus (Boisduval). Pestic Biochem Physiol 163:31–38. https://doi.org/10.1016/j.pestbp.2019.10.001
Litsanov B, Brocker M, Bott M (2012) Toward homosuccinate fermentation: metabolic engineering of Corynebacterium glutamicum for anaerobic production of succinate from glucose and formate. Appl Environ Microbiol 78:3325–3337. https://doi.org/10.1128/AEM.07790-11
Makhongela HS, Glowacka AE, Agarkar VB, Sewell BT, Weber B, Cameron RA, Cowan DA, Burton SG (2007) A novel thermostable nitrilase superfamily amidase from Geobacillus pallidus showing acyl transfer activity. Appl Microbiol Biotechnol 75:801–811. https://doi.org/10.1007/s00253-007-0883-2
Martínez-Rodríguez S, Conejero-Muriel M, Gavira JA (2019) A novel cysteine carbamoyl-switch is responsible for the inhibition of formamidase, a nitrilase superfamily member. Arch Biochem Biophys 662:151–159. https://doi.org/10.1016/j.abb.2018.12.008
McCarty NS, Ledesma-Amaro R (2019) Synthetic biology tools to engineer microbial communities for biotechnology. Trends Biotechnol 37:181–197. https://doi.org/10.1016/j.tibtech.2018.11.002
Meng N, Shao J, Li H, Wang Y, Fu X, Liu C, Yu Y, Zhang B (2022) Electrosynthesis of formamide from methanol and ammonia under ambient conditions. Nat Commun 13:5452. https://doi.org/10.1038/s41467-022-33232-w
Merrell DS, Goodrich ML, Otto G, Tompkins LS, Falkow S (2003) pH-regulated gene expression of the gastric pathogen Helicobacter pylori. Infect Immun 71:3529–3539. https://doi.org/10.1128/IAI.71.6.3529-3539.2003
Mills J, Wyborn NR, Greenwood JA, Williams SG, Jones CW (1998) Characterisation of a binding-protein-dependent, active transport system for short-chain amides and urea in the methylotrophic bacterium Methylophilus methylotrophus. Eur J Biochem 251:45–53. https://doi.org/10.1046/j.1432-1327.1998.2510045.x
Milo R, Phillips R (2015) Cell biology by the numbers, 1st edn. Garland Science
Mindt M, Walter T, Risse JM, Wendisch VF (2018) Fermentative production of N-methylglutamate from glycerol by recombinant Pseudomonas putida. Front Bioeng Biotechnol 6:1–11. https://doi.org/10.3389/fbioe.2018.00159
Neu HC (1992) The crisis in antibiotic resistance. Science 257:1064–1073. https://doi.org/10.1126/science.257.5073.1064
Newton GL, Av-Gay Y, Fahey RC (2000) A novel mycothiol-dependent detoxification pathway in mycobacteria involving mycothiol s-conjugate amidase. Biochemistry 39(35):10739–10746. https://doi.org/10.1021/bi000356n
Ou X, Wu X, Peng F, Zeng Y, Li H, Xu P, Chen G, Guo Z, Yang J, Zong M, Lou W (2019) Metabolic engineering of a robust Escherichia coli strain with a dual protection system. Biotechnol Bioeng 116:3333–3348. https://doi.org/10.1002/bit.27165
Parnell J, Baron M, Lindgren P (2006) Potential for irradiation of methane to form complex organic molecules in impact craters: implications for Mars, Titan and Europa. J Geochem Explor 89:322–325. https://doi.org/10.1016/j.gexplo.2005.11.024
Pérez-García F, Wendisch VF (2018) Transport and metabolic engineering of the cell factory Corynebacterium glutamicum. FEMS Microbiol Lett 365:1–11. https://doi.org/10.1093/femsle/fny166
Perkins H (1965) The action of hot formamide on bacterial cell walls. Biochem J 95:876–882. https://doi.org/10.1042/bj0950876
Pols T, Singh S, Deelman-Driessen C, Gaastra BF, Poolman B (2021) Enzymology of the pathway for ATP production by arginine breakdown. FEBS J 288:293–309. https://doi.org/10.1111/febs.15337
Rath M, Salas J, Parhy B, Norton R, Menakuru H, Sommerhalter M, Hatlstad G, Kwon J, Allan DL, Vance CP, Uhde-Stone C (2010) Identification of genes induced in proteoid roots of white lupin under nitrogen and phosphorus deprivation, with functional characterization of a formamidase. Plant Soil 334:137–150. https://doi.org/10.1007/s11104-010-0373-7
Reitzer LJ (1996) Sources of nitrogen and their utilization. In: Escherichia coli and Salmonella: cellular and molecular biology. ASM Press, Washington, D.C., pp 380–390
Rubin RH, Swenson GW Jr, Benson RC, Tigelaar HL, Flygare WH (1971) Microwave detection of interstellar formamide. Astrophys J 169:L39. https://doi.org/10.1086/180810
Sakamoto N, Kotre AM, Savageau MA (1975) Glutamate dehydrogenase from Escherichia coli: purification and properties. J Bacteriol 124:775–783. https://doi.org/10.1128/jb.124.2.775-783.1975
Saladino R, Botta G, Delfino M, Di Mauro E (2013) Meteorites as catalysts for prebiotic chemistry. Chem – Eur J 19:16916–16922. https://doi.org/10.1002/chem.201303690
Saladino R, Botta G, Pino S, Costanzo G, Mauro ED (2012a) Genetics first or metabolism first? The formamide clue. Chem Soc Rev 41:5526–5565. https://doi.org/10.1039/C2CS35066A
Saladino R, Carota E, Botta G, Kapralov M, Timoshenko GN, Rozanov AY, Krasavin E, Di Mauro E (2015) Meteorite-catalyzed syntheses of nucleosides and of other prebiotic compounds from formamide under proton irradiation. Proc Natl Acad Sci 112. https://doi.org/10.1073/pnas.1422225112
Saladino R, Crestini C, Pino S, Costanzo G, Di Mauro E (2012b) Formamide and the origin of life. Phys Life Rev 9:84–104. https://doi.org/10.1016/j.plrev.2011.12.002
Schwardmann LS, Dransfeld AK, Schäffer T, Wendisch VF (2022) Metabolic engineering of Corynebacterium glutamicum for sustainable production of the aromatic dicarboxylic acid dipicolinic acid. Microorganisms 10:730. https://doi.org/10.3390/microorganisms10040730
Schwardmann LS, Rieks M, Wendisch VF (2023a) Nitrogen-controlled valorization of xylose-derived compounds by metabolically engineered Corynebacterium glutamicum. Synth Biol Eng 1:1–14. https://doi.org/10.35534/sbe.2023.10009
Schwardmann LS, Wu T, Dransfeld AK, Lindner SN, Wendisch VF (2023b) Formamide-based production of amines by metabolically engineering Corynebacterium glutamicum. Appl Microbiol Biotechnol. 107:4245–4260. https://doi.org/10.1007/s00253-023-12592-3
Selão TT, Włodarczyk A, Nixon PJ, Norling B (2019) Growth and selection of the cyanobacterium Synechococcus sp. PCC 7002 using alternative nitrogen and phosphorus sources. Metab Eng 54:255–263. https://doi.org/10.1016/j.ymben.2019.04.013
Sgobba E, Wendisch VF (2020) Synthetic microbial consortia for small molecule production. Curr Opin Biotechnol 62:72–79. https://doi.org/10.1016/j.copbio.2019.09.011
Shaw AJ, Lam FH, Hamilton M, Consiglio A, MacEwen K, Brevnova EE, Greenhagen E, LaTouf WG, South CR, Van Dijken H, Stephanopoulos G (2016) Metabolic engineering of microbial competitive advantage for industrial fermentation processes. Science 353:583–586. https://doi.org/10.1126/science.aaf6159
Skouloubris S, Labigne A, De Reuse H (2001) The AmiE aliphatic amidase and AmiF formamidase of Helicobacter pylori: natural evolution of two enzyme paralogues: study of the two paralogous amidases of H. pylori. Mol Microbiol 40:596–609. https://doi.org/10.1046/j.1365-2958.2001.02400.x
Soares AS, Engel MA, Stearns R, Datwani S, Olechno J, Ellson R, Skinner JM, Allaire M, Orville AM (2011) Acoustically mounted microcrystals yield high-resolution X-ray structures. Biochemistry 50(21):4399–4401. https://doi.org/10.1021/bi200549x
Solomon PM (1973) Interstellar molecules. Phys Today 26:32–40. https://doi.org/10.1063/1.3127983
Soriano-Maldonado P, Martínez-Gómez AI, Andújar-Sánchez M, Neira JL, Clemente-Jiménez JM, Las Heras-Vázquez FJ, Rodríguez-Vico F, Martínez-Rodríguez S (2011) Biochemical and mutational studies of the Bacillus cereus CECT 5050T formamidase support the existence of a C-E-E-K tetrad in several members of the nitrilase superfamily. Appl Environ Microbiol 77:5761–5769. https://doi.org/10.1128/AEM.00312-11
Suleiman M, Zecher K, Yücel O, Jagmann N, Philipp B (2016) Interkingdom cross-feeding of ammonium from marine methylamine-degrading bacteria to the diatom Phaeodactylum tricornutum. Appl Environ Microbiol 82:7113–7122. https://doi.org/10.1128/AEM.01642-16
Teepakorn C, Zajkoska P, Cwicklinski G, De Berardinis V, Zaparucha A, Nonglaton G, Anxionnaz-Minvielle Z (2021) Nitrilase immobilization and transposition from a micro-scale batch to a continuous process increase the nicotinic acid productivity. Biotechnol J 16:2100010. https://doi.org/10.1002/biot.202100010
Teramoto H, Inui M, Yukawa H (2011) Transcriptional regulators of multiple genes involved in carbon metabolism in Corynebacterium glutamicum. J Biotechnol 154:114–125. https://doi.org/10.1016/j.jbiotec.2011.01.016
Thatcher RC, Weaver TL (1976) Carbon-nitrogen cycling through microbial formamide metabolism. Science 192:1234–1235. https://doi.org/10.1126/science.192.4245.1234
Thorwall S, Schwartz C, Chartron JW, Wheeldon I (2020) Stress-tolerant non-conventional microbes enable next-generation chemical biosynthesis. Nat Chem Biol 16:113–121. https://doi.org/10.1038/s41589-019-0452-x
Thuku RN, Brady D, Benedik MJ, Sewell BT (2009) Microbial nitrilases: versatile, spiral forming, industrial enzymes. J Appl Microbiol 106:703–727. https://doi.org/10.1111/j.1365-2672.2008.03941.x
Tseng I-T, Chen Y-L, Chen C-H, Shen Z-X, Yang C-H, Li S-Y (2018) Exceeding the theoretical fermentation yield in mixotrophic Rubisco-based engineered Escherichia coli. Metab Eng 47:445–452. https://doi.org/10.1016/j.ymben.2018.04.018
Turlin J, Dronsella B, De Maria A, Lindner SN, Nikel PI (2022) Integrated rational and evolutionary engineering of genome-reduced Pseudomonas putida strains promotes synthetic formate assimilation. Metab Eng 74:191–205. https://doi.org/10.1016/j.ymben.2022.10.008
Uhde A, Youn J-W, Maeda T, Clermont L, Matano C, Krämer R, Wendisch VF, Seibold GM, Marin K (2013) Glucosamine as carbon source for amino acid-producing Corynebacterium glutamicum. Appl Microbiol Biotechnol 97:1679–1687. https://doi.org/10.1007/s00253-012-4313-8
Ullmann L, Guntermann N, Kohl P, Schröders G, Müsgens A, Franciò G, Leitner W, Blank LM (2022) Improved itaconate production with Ustilago cynodontis via co-metabolism of CO2-derived formate. J Fungi 8:1277. https://doi.org/10.3390/jof8121277
Van Vliet AHM, Stoof J, Poppelaars SW, Bereswill S, Homuth G, Kist M, Kuipers EJ, Kusters JG (2003) Differential regulation of amidase- and formamidase-mediated ammonia production by the Helicobacter pylori Fur Repressor. J Biol Chem 278:9052–9057. https://doi.org/10.1074/jbc.M207542200
Wachsman JT, Barker HA (1955) The accumulation of formamide during the fermentation of histidine by Clostridium tetanomorphum. J Bacteriol 69:83–88. https://doi.org/10.1128/jb.69.1.83-88.1955
Wang Z, Chen X, Liu S, Zhang Y, Wu Z, Xu W, Sun Q, Yang L, Zhang H (2019) Efficient biosynthesis of anticancer polysaccharide by a mutant Chaetomium globosum ALE20 via non-sterilized fermentation. Int J Biol Macromol 136:1106–1111. https://doi.org/10.1016/j.ijbiomac.2019.06.186
Wendisch VF, Brito LF, Gil Lopez M, Hennig G, Pfeifenschneider J, Sgobba E, Veldmann KH (2016) The flexible feedstock concept in industrial biotechnology: metabolic engineering of Escherichia coli, Corynebacterium glutamicum, Pseudomonas, Bacillus and yeast strains for access to alternative carbon sources. J Biotechnol 234:139–157. https://doi.org/10.1016/j.jbiotec.2016.07.022
Wenk S, Rainaldi V, He H, Schann K, Bouzon M, Döring V, Lindner SN, Bar-Even A (2022) Synthetic carbon fixation via the autocatalytic serine threonine cycle. https://doi.org/10.1101/2022.09.28.509898
Wheeler GP, Grammer MG (1960) Prevention of the inhibitory effects of urethan, formamide, and N-methylformamide on the growth of Escherichia coli. Biochem Pharmacol 3:316–327. https://doi.org/10.1016/0006-2952(60)90097-6
Witthoff S, Eggeling L, Bott M, Polen T (2012) Corynebacterium glutamicum harbours a molybdenum cofactor-dependent formate dehydrogenase which alleviates growth inhibition in the presence of formate. Microbiology 158:2428–2439. https://doi.org/10.1099/mic.0.059196-0
Wu Y, Jiang Z, Lin Z, Liang Y, Wang H (2021) Direct electrosynthesis of methylamine from carbon dioxide and nitrate. Nat Sustain 4:725–730. https://doi.org/10.1038/s41893-021-00705-7
Wyborn NR, Mills J, Williams SG, Jones CW (1996) Molecular characterisation of formamidase from Methylophilus methylotrophus. Eur J Biochem 240:314–322. https://doi.org/10.1111/j.1432-1033.1996.0314h.x
Wyborn NR, Scherr DJ, Jones CW (1994) Purification, properties and heterologous expression of formamidase from Methylophilus methylotrophus. Microbiology 140:191–195. https://doi.org/10.1099/13500872-140-1-191
Xiao Z, Gu R, Hou X, Zhao J, Zhu H, Lu JR (2017) Non-sterilized fermentative production of acetoin with 2,3-butanediol as a main byproduct from maize hydrolysate by a newly isolated thermophilic Bacillus strain. J Chem Technol Biotechnol 92:2845–2852. https://doi.org/10.1002/jctb.5301
Yang X, Zhu M, Huang X, Lin CSK, Wang J, Li S (2015) Valorisation of mixed bakery waste in non-sterilized fermentation for l-lactic acid production by an evolved Thermoanaerobacterium sp. strain. Bioresour Technol 198:47–54. https://doi.org/10.1016/j.biortech.2015.08.108
Ye J, Hu D, Che X, Jiang X, Li T, Chen J, Zhang HM, Chen G-Q (2018) Engineering of Halomonas bluephagenesis for low cost production of poly(3-hydroxybutyrate-co-4-hydroxybutyrate) from glucose. Metab Eng 47:143–152. https://doi.org/10.1016/j.ymben.2018.03.013
Yishai O, Lindner SN, Gonzalez de la Cruz J, Tenenboim H, Bar-Even A (2016) The formate bio-economy. Curr Opin Chem Biol 35:1–9. https://doi.org/10.1016/j.cbpa.2016.07.005
Yu H, Liao JC (2018) A modified serine cycle in Escherichia coli coverts methanol and CO2 to two-carbon compounds. Nat Commun 9:3992. https://doi.org/10.1038/s41467-018-06496-4
Yurgel SN, Johnson SA, Rice J, Sa N, Bailes C, Baumgartner J, Pitzer JE, Roop RM, Roje S (2022) A novel formamidase is required for riboflavin biosynthesis in invasive bacteria. J Biol Chem 298:102377. https://doi.org/10.1016/j.jbc.2022.102377
Zecher K, Hayes KR, Philipp B (2020) Evidence of interdomain ammonium cross-feeding from methylamine- and glycine betaine-degrading Rhodobacteraceae to diatoms as a widespread interaction in the marine phycosphere. Front Microbiol 11:1–15. https://doi.org/10.3389/fmicb.2020.533894
Zeng W, Li W, Shu L, Yi J, Chen G, Liang Z (2013) Non-sterilized fermentative co-production of poly(γ-glutamic acid) and fibrinolytic enzyme by a thermophilic Bacillus subtilis GXA-28. Bioresour Technol 142:697–700. https://doi.org/10.1016/j.biortech.2013.05.020
Zhang L, Jiang Z, Tsui TH, Loh KC, Dai Y, Tong YW (2022) A review on enhancing Cupriavidus necator fermentation for poly(3-hydroxybutyrate) (PHB) production from low-cost carbon sources. Front Bioeng Biotechnol 10:946085. https://doi.org/10.3389/fbioe.2022.946085
Zhang Y, Chen X, Qi B, Luo J, Shen F, Su Y, Khan R, Wan Y (2014) Improving lactic acid productivity from wheat straw hydrolysates by membrane integrated repeated batch fermentation under non-sterilized conditions. Bioresour Technol 163:160–166. https://doi.org/10.1016/j.biortech.2014.04.038
Zhu G, Li Y (2021) Urease: a highly efficient biocatalyst for synthesis of polyhydroquinolines and polyhydroacridines from the ammonia formed in situ. Mol Divers 25:2149–2159. https://doi.org/10.1007/s11030-020-10109-y
Funding
Open Access funding enabled and organized by Projekt DEAL. Open access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
L.S.S., L.B. S.N.L., and V.F.W. drafted the manuscript. L.S.S., L.B. S.N.L., and V.F.W. finalized the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
The online version contains supplementary material available at ## to be entered after revision ##. (PDF 672 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schwardmann, L.S., Benninghaus, L., Lindner, S.N. et al. Prospects of formamide as nitrogen source in biotechnological production processes. Appl Microbiol Biotechnol 108, 105 (2024). https://doi.org/10.1007/s00253-023-12962-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-023-12962-x