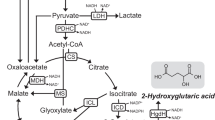
Abstract
Formamide is rarely used as nitrogen source by microorganisms. Therefore, formamide and formamidase have been used as protection system to allow for growth under non-sterile conditions and for non-sterile production of acetoin, a product lacking nitrogen. Here, we equipped Corynebacterium glutamicum, a renowned workhorse for industrial amino acid production for 60 years, with formamidase from Helicobacter pylori 26695, enabling growth with formamide as sole nitrogen source. Thereupon, the formamide/formamidase system was exploited for efficient formamide-based production of the nitrogenous compounds L-glutamate, L-lysine, N-methylphenylalanine, and dipicolinic acid by transfer of the formamide/formamidase system to established producer strains. Stable isotope labeling verified the incorporation of nitrogen from formamide into biomass and the representative product L-lysine. Moreover, we showed ammonium leakage during formamidase-based access of formamide to be exploitable to support growth of formamidase-deficient C. glutamicum in co-cultivation and demonstrated that efficient utilization of formamide as sole nitrogen source benefitted from overexpression of formate dehydrogenase.
Key points
• C. glutamicum was engineered to access formamide.
• Formamide-based production of nitrogenous compounds was established.
• Nitrogen cross-feeding supported growth of a formamidase-negative strain.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nitrogen is an essential main building block of life for its incorporation into numerous cellular components such as amino acids, nucleic acids, and purines (Zerkle and Mikhail 2017). Consequently, biotechnological production processes require the supplementation with an accessible nitrogen source for growth of the producer organism and biosynthesis of nitrogenous target compounds (Kampen 2014). Many bacteria, fungi, and algae naturally assimilate ammonium (NH4+) or nitrate as nitrogen source (Kampen 2014) with NH4+ as preferred nitrogen source for fast bacterial growth (Reitzer 1996). In addition, assimilation pathways for use of less common nitrogen sources have evolved, e.g., for non-peptide carbon–nitrogen nitriles (Chen et al. 2007; Duca et al. 2014; Seffernick et al. 2001). Their breakdown into NH4+ and the respective carboxylic acid is catalyzed by either nitrilase or nitrile hydratase and amidase (Willison 1993). Nitrilase activity was reported for not only filamentous fungi, yeasts, and plants, but also bacteria belonging to various genera, including Corynebacterium (Gong et al. 2012; Egelkamp et al. 2017). Access to non-native nitrogen sources has been attained, but these often feature unfavorable properties, which complicate efficient economic production and purification processes. Examples are Escherichia coli and cyanobacterial strains engineered to access melamine (Shaw et al. 2016; Selão et al. 2019), categorized as group 2B carcinogen. By contrast, formamide, the simplest naturally occurring (monocarboxylic acid) amide, shows low toxicity (Kennedy 2014). It can be considered as promising alternative nitrogen source for use in biotechnological processes, and for some bacteria even as combined nitrogen, carbon and energy source. Formamide is found in nature and in industry and is best known for its applications as solvent, catalyst, or as intermediate in the production of chemicals, mainly heterocyclic compounds, copolymers, or pharmaceuticals (Bipp and Kieczka 2011). Its market reaches an annual global size in the million ton range (Meng et al. 2022). Currently, formamide is predominantly produced from carbon monoxide and ammonia in a high temperature and pressure process (Meng et al. 2022; Bipp and Kieczka 2011).
Formamide is also ubiquitously found in the entire universe (Saladino et al. 2012), and some microbes have evolved the ability to use it as nitrogen source by hydrolysis into formate and ammonia (NH3), catalyzed by formamidase, an amidase with restricted substrate specificity for formamide (Hung et al. 2007). Amidases are generally widespread among bacteria and often involved in detoxification processes for the degradation of toxic amides (Newton et al. 2000; Fournand and Arnaud 2001; Liu et al. 2020), but reports on characterization of formamidases, formamide-specific amidases, are scarce (Brown et al. 1986; Skouloubris et al. 2001; Rath et al. 2010; Soriano-Maldonado et al. 2011). Today, only 14 protein structures are available and even less formamidases have been functionally characterized in detail (Mahenthiralingam et al. 1993; Parish et al. 1997; Skouloubris et al. 2001; Rath et al. 2010; Soriano-Maldonado et al. 2011), although first records of research on formamide utilization date back to 1976, when Pseudomonas SL-4 was described for its ability to utilize formamide as sole nitrogen, carbon, and energy source and for its involvement in the ecological carbon–nitrogen cycle (Thatcher and Weaver 1976).
Heterologous expression of the amiF gene from Helicobacter pylori in the bacteria E. coli, Bacillus subtilis, and Klebsiella pneumoniae enabled formamide degradation (Guo et al. 2020a, b; Ou et al. 2019). The amidase AmiF from H. pylori is one of the best characterized formamidases with formamide as its only identified substrate (Hung et al. 2007). H. pylori is known as human gastric pathogen, causing chronic gastritis and is involved in peptic ulcer disease as well as gastric carcinoma and lymphoma (Dunn et al. 1997). In that environment, NH4+ is generated by urease, AmiF, and its paralog aliphatic amidase AmiE. This primarily serves to neutralize the prevalent gastric acidity to ensure the bacterium’s survival and thriving in the human stomach (Skouloubris et al. 2001). The optimal pH for growth of H. pylori is at 4–6 (Dunn et al. 1997).
The model organism and well-established industrial workhorse Corynebacterium glutamicum is primarily used for production of nitrogenous compounds such as L-lysine since decades (Klaffl and Eikmanns 2010). Its product portfolio has been expanded to include halogenated amino acids, diamines, alkylamines, etc. (Wendisch and Lee 2020; Tsuge and Matsuzawa 2021). The Gram-positive endotoxin-free C. glutamicum (Srivastava and Deb 2005) is preferred over Gram-negative bacteria such as E. coli that often suffer from endotoxin contamination (Lee et al. 1999; Valappil et al. 2007) Its products may gain GRAS status (Wolf et al. 2021), as manifested by its deployment for the production of food-grade sugars and sugar alcohols (Hu et al. 2022), amino acids, feed, and pharmaceutical products for several decades (Leuchtenberger et al. 2005). Moreover, its advantageous high stress tolerance renders C. glutamicum robust for application in fermentation processes (Liu et al. 2021). In addition to its naturally broad carbon source spectrum (Zhang et al. 2020), the synthetically attained access to numerous non-native carbon sources along with its capacity to co-utilize different carbon sources (Zahoor et al. 2012; Schneider et al. 2011; Rittmann et al. 2008; Wendisch et al. 2000) make C. glutamicum an efficient and versatile platform organism for biotech processes.
By contrast, C. glutamicum shows a more restricted spectrum of nitrogen sources since it lacks secreted proteases or peptidases for extracellular degradation of nitrogenous compounds (Wendisch et al. 2022). Cultivation processes commonly rely on the provision of nitrogen in form of ammonium salts or urea (Wendisch et al. 2022). Alternatively, nitrogen can be provided in form of amino acids such as L-glutamine (Rehm et al. 2010), the key metabolite for nitrogen assimilation and transferase reactions, which is taken up by a Na+-dependent glutamine uptake system (Siewe et al. 1995). Another notable exception are hexosamines. While glucosamine is accessible to C. glutamicum wild type (WT) (Uhde et al. 2013), metabolic engineering has broadened the substrate spectrum to N-acetylmuramic acid (Sgobba et al. 2018a, b). Beyond, metabolic engineering has achieved the harnessing of nitrogen from monomethylamine (MMA) for the formation of the N-methylated amino acids sarcosine (N-methylglycine) (Mindt et al. 2019), N-methyl-L-alanine (Mindt et al. 2018), and N-methyl-L-phenylalanine (Kerbs et al. 2021) by overexpression of an imine reductase gene from Pseudomonas putida in strains overproducing the precursor oxoacid.
In this work, we extended the spectrum of nitrogen sources available to C. glutamicum by recruitment of a formamidase for growth and production of several target compounds with formamide as sole or combined nitrogen source. Overexpression of formate dehydrogenase gene emerged as valid tool to improve growth with formamide by counteracting toxicity of formate, the second product of formamidase.
Material and methods
Bacterial strains and growth conditions
All plasmids and strains used in this work are listed in Table 1.
E. coli was grown at 37 °C and 180 rpm in 50-mL baffled flasks on a rotary shaker for plasmid amplification. C. glutamicum was grown at 30 °C. If not indicated otherwise, precultures and production experiments of C. glutamicum strains were performed in 50-mL baffled flasks on a rotary shaker at 120 rpm. Growth was monitored by the measurement of optical density at 600 nm (OD600), using a V-1200 Spectrophotometer (VWR, Radnor, PA, USA). OD600 was converted into cell dry weight concentrations (CDW) according to a previously determined factor of an OD600 of 1 to correspond to 0.25 g L−1 CDW.
Routinely, growth experiments of C. glutamicum strains were performed in a BioLector microcultivation system (m2p-labs, Aachen, Germany) in volumes of 1 mL in a 48-well flower plate at 85% humidity and 1100 rpm. Growth was followed using the backscatter light signal at 620 nm, given in arbitrary units (a.u.).
Precultures were inoculated from fresh lysogeny broth (LB) agar plates and grown in LB or brain–heart infusion broth (BHI) (ROTH, Karlsruhe, Germany) overnight. Main cultures for growth or production experiments were inoculated to an OD600 of 1, using overnight precultures, which were washed and resuspended in CgXII minimal medium without accessible nitrogen (N-CgXII, lacking the nitrogen sources urea and (NH4)2SO4 as compared to CgXII minimal medium) (Eggeling and Bott 2005). Routinely, cultivations were performed in CgXII minimal medium or N-CgXII, supplemented the indicated concentrations of nitrogen in form of urea and (NH4)2SO4 or formamide with 40 g L−1 glucose as carbon source. Indicated nitrogen concentrations in form of urea and (NH4)2SO4 correspond to their ratio in standard CgXII minimal medium. Deviating, MePhe5*-FORM was grown with 20 g L−1 glucose and supplemented with 0.35 M monomethylamine (MMA), 0.2 mM L-tryptophan, and 0.8 mM L-phenylalanine, L-isoleucine, and L-leucine, respectively (Kerbs et al. 2021).
When appropriate, precultures of E. coli strains were supplemented with 10 µg mL−1 tetracycline, 50 µg mL−1 kanamycin, or 100 µg mL−1 ampicillin and pre- and main cultures of C. glutamicum with 5 µg mL−1 tetracycline, 25 µg mL−1 kanamycin, and 100 µg mL−1 spectinomycin. For the induction of the expression plasmids pVWEx1 and pEKEx3 in main cultures, 1 mM Isopropyl-β-D-1-thiogalactopyranoside (IPTG) was added. Glutamate production was elicited by exposure to 8 µg mL −1 ciprofloxacin during mid-exponential growth, when an OD600 of 9 was reached (Lubitz and Wendisch 2016).
Genetic engineering for plasmid and strain construction
All listed oligonucleotides for DNA amplification and sequencing (Supplemental Tab. S1) were purchased from Metabion (Planegg/Steinkirchen, Germany) or Sigma-Aldrich (Ulm, Germany). All commercially available kits and enzymes were used according to the suppliers` instructions. Plasmids and DNA were isolated and purified using the GeneJET plasmid miniprep kit (Thermo Fisher Scientific, Schwerte, Germany) and the NucleoSpin® Gel and PCR Clean-up kit (MACHEREY–NAGEL GmbH & Co. KG, Düren, Germany). Genomic DNA of C. glutamicum was isolated by salt precipitation (Eikmanns et al. 1994). DNA concentrations were measured at 260 nm by a V-1200 Spectrophotometer (VWR, Radnor, PA, USA).
Genes were amplified from plasmids or genomic DNA of C. glutamicum ATCC13032 with the respective oligonucleotides (Supplemental Tab. S1) by ALLin™ HiFi DNA Polymerase (highQu GmbH, Kraichtal, Germany). Primer overhangs were used to insert a consensus ribosome binding site (RBS) sequence (GAAAGGAGGCCCTTCAG) in front of the genes amiF, gfpUV, crimson, and fdhPs, and an optimized RBS (CTGAAGGGCCTCCTTTC, designed using the Salislab software (Cetnar and Salis 2021)) was inserted in front of the gene fdhCg.
The plasmids pECXT_Psyn and pECXT_Psyn-amiF were linearized by restriction with BamHI or XbaI, respectively (New England Biolabs, Frankfurt, Germany), followed by dephosphorylation (Antarctic phosphatase, New England Biolabs, Frankfurt, Germany). Plasmids were assembled via the method of Gibson (Gibson et al. 2009). Newly constructed expression plasmids were sequenced with respective oligonucleotides (Supplemental Tab. S1) for verification of correct assembly and sequence validity of inserted genes.
Standard molecular genetic techniques followed previously described procedures (Sambrook et al. 1989). The CaCl2 method was applied to prepare chemocompetent E. coli cells for transformation by heat shock at 42 °C (Sambrook et al. 1989). Electrocompetent C. glutamicum cells were transformed by electroporation, followed by a heat shock at 46 °C (van der Rest et al. 1999). Recombinant clones were confirmed by colony PCR.
Stable isotope labeling
For isotope-tracing of nitrogen, 60 mM nitrogen in the form of 15N labeled (NH4)2SO4 and/or formamide (Sigma-Aldrich, Ulm, Germany) and/or respective unlabeled substrates were provided as nitrogen sources for cultivation in N-CgXII. Cells were grown in a BioLector microcultivation system for 48 h and harvested by centrifugation. For analysis of lysine production, supernatants were immediately stored at − 20 °C, while cell pellets were washed and resuspended in H2O prior to storage at − 20 °C until analysis.
Proteins were hydrolyzed at 95 °C for 24 h using 6 M HCl (You et al. 2012), which was removed by evaporation under stream of N2 at 95°. Samples were resuspended in H2O and centrifuged for the removal of insoluble compounds for analysis of the supernatants by UPLC-ESI–MS (Giavalisco et al. 2011). Amino acid mass analysis was performed with a Waters Acquity UPLC system (Waters, Eschborn, Germany) equipped with a HSS T3 C18 reversed-phase column (100 mm × 2.1 mm, 1.8 µm; Waters, Eschborn, Germany), using 0.1% formic acid in H2O (A) and 0.1% formic acid in acetonitrile (B) at a flow rate of 0.4 mL min−1 and the following gradient: 0–1 min 99% A; 1–5 min linear gradient 99 to 82% A; 5–6 min linear gradient 82 to 1% A; 6–8 min 1% A; 8–8.5 min linear gradient 1 to 99% A; and 8.5–11 min re-equilibration.
An exactive mass spectrometer (Thermo Fisher Scientific, Schwerte, Germany) was used in positive ionization mode at a scan range of 50.0–300.0 m/z for acquisition of mass spectra, and the Xcalibur™ software (Thermo Fisher Scientific, Schwerte, Germany) was used for data analysis. Retention times were determined by analyzing amino-acid standards (Sigma-Aldrich) under the same conditions.
Assay for ammonium detection
For the measurement of NH4+, cells were grown in N-CgXII, supplemented with 60 mM nitrogen in form of urea and (NH4)2SO4 or formamide for 24 h. Samples were harvested by centrifugation and the NH4+content of the supernatants was quantified with an ammonia assay kit (Megazyme, Bray, Ireland).
Enzyme activity assay for formamidase AmiF
Crude extracts were gained from cells, grown in 50 mL LB in shake flasks overnight and harvested by centrifugation (3100 g, 7 min) at 4 °C. All following steps were performed at 4 °C. Cells were washed 3 times in assay buffer (50 mM phosphate, pH 7), resuspended in 5 mL and lysed by sonication (UP 200S, Dr. Hielscher GmbH, Teltow, Germany), applying ultrasound at an amplitude of 60% and a duty cycle of 0.5 s for 9 min. The protein containing supernatant was attained by centrifugation (25,000 g, 1 h), and the total protein concentrations of the crude extracts were analyzed by the method of Bradford, using bovine serum standard as reference (Bradford 1976).
Formamidase activity was measured by means of formamide hydrolysis, which was detected by the formation of free NH4+ as described previously (Ou et al. 2019; Weatherburn 1967). The reaction was carried out in 1 mL assay buffer, containing 2.5 mg L−1 total crude extract protein, started by the addition of 200 mM formamide, and run for 1 h at 30 °C. The dying was implemented as described elsewhere (Weatherburn 1967), but incubated for 35 min. One unit (U) of formamidase activity was defined as the quantity of total protein required to deplete 1 μM formamide min−1.
Fluorescence analysis by flow cytometry
The cellular composition of co-cultures was analyzed by means of fluorescence using flow cytometry (flow cytometer Gallios™, Beckman Coulter, Krefeld, Germany). Samples of co-cultures were taken after 24 h and diluted to an OD600 of 0.1 in N-CgXII for immediate analysis of 20 000 cells per sample. Fluorescence was excited at 405 nm by a blue solid-state laser for monitoring of the forward- (FSC) and side-scatter (SSC) signals. The GfpUV and Crimson signals were detected via 525/50 nm and 660/20 nm band-pass filters, respectively. WT-EV cells served as reference for preliminary adjustment for autofluorescence.
Product quantification by HPLC analysis
Extracellular DPA and amino acids glutamate, lysine, and NMePhe were quantified by use of a high-pressure liquid chromatography system (HPLC) (1200 series, Agilent Technologies Deutschland GmbH, Böblingen, Germany). Cells were grown in N-CgXII, supplemented with 60 mM nitrogen in form of urea and (NH4)2SO4 or formamide as indicated. Samples were taken after 48 or 72 h and centrifuged for the storage of supernatants at − 20 °C until analysis.
Amino acids were separated by reversed phase HPLC with a pre- (LiChrospher 100 RP18 EC-5µ (40 × 4 mm), CS Chromatographie Service, Langerwehe, Germany), and a main-column (LiChrospher 100 RP18 EC-5µ, 125 × 4.6 mm, CS Chromatographie Service, Langerwehe, Germany) and detected by a fluorescence detector (FLD G1321A, 1200 series, Agilent Technologies, Böblingen, Germany).
For detection of lysine and glutamate, samples were derivatized with ortho-phthaldialdehyde (OPA). Analysis followed previously described procedures with asparagine as internal standard (Schneider and Wendisch 2010). Compounds were detected at 230 nm excitation and 450 nm emission wavelengths. For detection of NMePhe, samples were derivatized with fluorenylmethyl chloroformate (FMOC) (Karl Roth, Karlsruhe, Germany)(Schneider et al. 2012). Compounds were separated at a flow rate of 1.2 mL min−1 with sodium acetate (50 mM, pH 4.2) (A) and acetonitrile (B) as eluents, applying the following gradient: 0 min 38% B, 5 min 38% B, 10 min 48% B, 12 min 52% B, 13 min 57% B, 14 min 63% B, 16 min 68, 17 min 76% B and 20 min 38%. L-proline served as internal standard, and fluorescence was detected at 250 nm excitation and 410 nm emission wavelengths (Kerbs et al. 2021). DPA was separated with an amino exchange column (Aminex, 300 × 8 mm, 10 µm particle size, 25 Å pore diameter, CS Chromatographie Service, Langerwehe, Germany) under isocratic conditions, using 5 mM H2SO4 as mobile phase at a flow rate of 0.8 mL min−1 for 30 min and detected by the refractive index signal (RID G1362A, 1200 series, Agilent Technologies, Böblingen, Germany) as described previously (Schwardmann et al. 2022).
Results
Assessment of the suitability of C. glutamicum for formamide utilization
Formamide is an organic nitrogen source that is degraded by formamidase to yield formate and NH4+. To determine if formamide or formate inhibit growth of C. glutamicum in minimal medium, the empty vector control carrying strain C. glutamicum ATCC13032 (named WT-EV) was grown in CgXII minimal medium in the presence of 0–160 mM formamide or sodium formate.
The exposure to increasing formamide concentrations from 0 to 160 mM had only a minor effect on biomass formation of strain WT-EV, but slightly decreased the maximal specific growth rate µmax by up to 14% from 0.42 ± 0.00 to 0.36 ± 0.00 h−1 (Fig. 1a). A reduction of the growth rate to half maximal was calculated to be reached at a concentration of 690 mM, which exceeds the routinely used nitrogen content of 468 mM in form of urea and (NH4)2SO4 in CgXII minimal medium and demonstrates high tolerance. The presence of sodium formate is known to inhibit growth of C. glutamicum (Witthoff et al. 2012; Ryabchenko et al. 2020). When cultivated in CgXII minimal medium in the presence of 0–160 mM sodium formate, the growth rate was reduced to half-maximal at 227 ± 3 mM formate (Fig. 1d) and lag phase duration increased proportionally to the formate concentration (Fig. 1d). By contrast, this was not observed for addition of 160 mM sodium ions as Na2SO4 salt (data not shown). The presence of formate increased the maximal biomass concentration, probably due to NADH formation as consequence of its oxidation to carbon dioxide by formate dehydrogenase. In conclusion, the high tolerance of C. glutamicum towards formamide provides a promising basis for its use as amine source by engineered strains.
Influence of formamide and formate on the growth of C. glutamicum strains. Biomass (dark blue circles), µmax (light blue triangles), and lag phase (purple squares) of strains WT-EV (open symbols) and FORM (closed symbols), cultivated in CgXII minimal medium (a, b, d, e) or N-CgXII (c) supplemented with 40 g L−1 glucose and either 0–160 mM formamide (a, b, c) or 0–160 mM sodium formate (d, e) in a BioLector microcultivation system for 96 h. Values represent means with standard deviations of triplicate cultivations
Establishment of formamide assimilation by engineered C. glutamicum
Inspection of the C. glutamicum ATCC13032 genome did not identify homologs of AmiF. Additionally, no growth was observed when incubating C. glutamicum with formamide as sole nitrogen (N) source. Hence, with the goal to make formamide an accessible nitrogen source for C. glutamicum WT, it was equipped with a plasmid for constitutive expression of codon optimized version of the gene amiF, coding for formamidase from H. pylori 26695, resulting in strain C.glutamicum WT(pECXT_Psyn-amiF), from here on referred to as FORM.
First, formamidase activity was analyzed in crude extracts of strains FORM and the empty vector control strain WT-EV, obtained after growth in LB by conducting an enzymatic activity assay. The detection of 6.0 ± 0.8 U mgtotal protein−1 in crude extract of strain FORM, but no detectable activity (< 0.4 U mg−1) for the empty vector carrying control strain WT-EV (Fig. 1a) demonstrated functional expression of amiF in C. glutamicum.
Next, we grew both strains in regular CgXII minimal medium containing urea and (NH4)2SO4 as nitrogen sources with added 15N-labeled formamide. When 60 mM labeled (15N) formamide was provided in CgXII minimal medium in addition to 468 mM of N in form of unlabeled (14N) urea and (NH4)2SO4, 17% of lysine molecules of strain FORM were labeled once, but lysine of WT-EV lacked detectable 15N-labeling (Fig. 2b). This revealed that the ammonium from formamide was only accessible for strain FORM, even at abundant availability of natively accessible nitrogen (Fig. 2b).
AmiF activity in crude extracts (a), 15N labeling of L-lysine from 15N-labeled formamide (b), and growth (c) of formamidase-expressing strain FORM (blue) and WT-EV (grey) with formamide as sole nitrogen source. Cells were grown in LB over night for crude extract preparation (a). Cells were grown in CgXII minimal medium containing 468 mM unlabeled (14N) N in form of urea and (NH4)2SO4, supplemented with 60 mM 15N labeled formamide (15N) and 40 g L−1 glucose, in a BioLector microcultivation system for 24 h. The patterns depict the fractions of 0 (grey), 1 (red), or 2 (purple) labeled nitrogen atoms per molecule of lysine in the biomass. Standard deviations refer to triplicate cultivations. For growth assessment, cells were grown in N-CgXII minimal medium, supplemented with 40 mM formamide for 28 h (c). Values represent means with standard deviations of triplicate measurements or cultivations. *n.d., not detectable (< 0.4 U mg.−1)
Additionally, strains WT-EV and FORM were cultivated in CgXII minimal medium without nitrogen in form of urea or (NH4)2SO4 (N-CgXII), supplemented with 40 mM formamide as sole potential nitrogen source to evaluate the capacity of AmiF activity to support growth with formamide as sole nitrogen source. A formamide concentration of 40 mM allowed to monitor growth with hardly any inhibitory effects due to formamide or formate. While WT-EV was unable to grow in N-CgXII with formamide, formamide supported growth of strain FORM to a maximal biomass concentration of 6.3 ± 0.5 g L−1 with a maximal growth rate of 0.17 ± 0.01 h−1 (Fig. 2c). When strain FORM was grown with 40 mM nitrogen in form of urea and (NH4)2SO4, growth to a biomass concentration of 5.2 ± 0.1 g L−1 was faster (0.28 ± 0.02 h−1) than with formamide (data not shown).
Finally, both strains were cultivated in N-CgXII minimal medium supplemented with 60 mM unlabeled or 15N-labeled ammonium sulfate and/or formamide as nitrogen sources and the labeling pattern in the biomass was measured for the five representative proteinogenic amino acids L-lysine, L-alanine, L-serine, L-proline, and L-tyrosine. When strain FORM was cultivated with 15N-labeled ammonium sulfate or 15N-labeled formamide, almost complete labeling of the single nitrogen atoms was observed for alanine, serine, proline, and tyrosine, while both nitrogen atoms of lysine were labeled (Fig. 3).
15N labeling of L-lysine, L-alanine, L-serine, L-proline, and L-tyrosine in strain FORM. Cells were grown in N-CgXII, supplemented with 60 mM 15N-labeled (NH4)2SO4 or formamide and 40 g L−1 glucose, in a BioLector microcultivation system for 24 h. The patterns depict the fractions of 0 (grey), 1 (red), or 2 (purple) labeled nitrogen atoms per molecule of lysine in the biomass. Standard deviations refer to triplicate cultivations
Both strains showed double 15N-labeled lysine with 15N-labeled (NH4)2SO4, but lysine was unlabeled when unlabeled (NH4)2SO4 was provided (Fig. 4a). Strain WT-EV was unable to utilize formamide, but provision of 15N-labeled formamide as sole nitrogen source to strain FORM allowed growth and resulted in double 15N-labeled lysine (Fig. 4a). This demonstrated that strain FORM assimilated formamide as sole nitrogen source as well as ammonium sulfate. This notion was further supported when growth of C. glutamicum FORM with a mixture of 30 mM 15N-labeled formamide and unlabeled ammonium sulfate, respectively, was compared to growth with a mixture of unlabeled formamide and 15N-labeled ammonium sulfate since the labeling patterns of lysine were comparable under both conditions (Fig. 4b).
15N labeling of lysine in WT-EV and FORM from 15N-labeled ammonium sulfate or formamide. Cells were grown in N-CgXII, supplemented with 60 mM unlabeled (114N) or labeled (15N) (NH4)2SO4 or formamide (a), or with 30 mM labeled or unlabeled (NH4)2SO4 and formamide, respectively (b), and 40 g L−1 glucose, in a BioLector microcultivation system for 24 h. The patterns depict the fractions of 0 (grey), 1 (red), or 2 (purple) labeled nitrogen atoms per molecule of lysine in the biomass. Standard deviations refer to triplicate cultivations. *n.g., no growth with formamide
Taken together, we have shown that the engineered C. glutamicum strain FORM efficiently utilizes formamide as sole or combined nitrogen source to support growth.
Co-culturing formamidase-positive with formamidase-negative strains
Since formamidase hydrolyzes formamide to yield formate and ammonium, we tested if ammonium leakage out of the cell was exploitable in a co-culture approach using a formamidase-positive and a formamidase-deficient strain. To allow distinction of the used strains by fluorescence microscopy and FACS analysis, the fluorescence reporter Crimson was constitutively expressed in formamidase-negative C. glutamicum WT-EV, while GFP was expressed as part of a synthetic operon with amiF in strain FORM, resulting in strains WT-crimson and FORM-gfp, respectively. To test if the formamidase-positive strain FORM-gfp can provide ammonium from formamide as sole nitrogen source in the growth medium not only for itself, but also for formamidase-negative strain WT-crimson by ammonium export or leakage, supernatants of FORM-gfp grown with 60 mM N either provided as urea and (NH4)2SO4 or as formamide were analyzed for NH4+. With urea and (NH4)2SO4, less than 2 mM NH4+ were detected (Fig. 5b), while 10.0 ± 0.8 mM NH4+ was detected in the supernatants of strain FORM-gfp grown with formamide (Fig. 5b).
Free ammonium in supernatants of cells of WT-crimson (a) and FORM-gfp (b). Cells were either grown separately (a, b) or in co-culture inoculated with 70% WT-crimson and 30% FORM-gfp (c), supplemented with 60 mM N in form of urea and (NH4)2SO4 (grey) or with 60 mM formamide (blue) and 40 g L−1 glucose as carbon source, for 24 h. Values represent means of triplicate measurements with standard deviations. *n.g., no growth with formamide
Next, a co-cultivation of formamidase-negative WT-crimson and formamidase-positive FORM-gfp was performed, and the inoculum ratio was varied to contain 0, 10, 30, 50, or 70% of FORM-gfp (with 100, 90, 70, 50, or 30% WT-crimson, respectively). In CgXII minimal medium containing 60 mM N as urea and (NH4)2SO4, both strains grew from 0.25 to 15 g L−1 biomass in 24 h and roughly maintained a ratio of 50%/50% throughout cultivation (Fig. 6). In the formamide containing medium N-CgXII, WT-crimson alone (0% FORM-gfp) could not grow as expected (Fig. 6). Remarkably, all co-cultivations of WT-crimson with FORM-gfp resulted in growth of both strains to combined biomass concentrations of about 10 g L−1 (Fig. 6). This indicated that nitrogen released by cleavage of formamide to formate and NH4+ supported growth of formamidase-negative WT-crimson. For example, in a co-cultivation inoculated with 30% of FORM-gfp cells and 70% WT-crimson cells to a combined biomass concentration of about 0.25 g L−1, the biomass after 24 h consisted of about 1.5 g L−1 FORM-gfp cells and 8.5 g L−1 WT-crimson cells (Fig. 6). Similar to the single cultivation of FORM-gfp, supernatants of this co-cultivation showed about 11 mM of extracellular free ammonium (Fig. 5c). Thus, FORM-gfp grown with formamide produced sufficient ammonium from formamide to support growth of a strain unable to use formamide.
Co-cultures of strains WT-crimson (red) and FORM-gfp (green) in varied inoculum ratios. To test if FORM-gfp can provide nitrogen from formamide for growth of amiF-deficient WT-crimson, cells were grown with 60 mM formamide as sole nitrogen source in N-CgXII, whereas a control cultivation in CgXII minimal medium supplemented with 60 mM N in form of urea and (NH4)2SO4 was also made (grey background). 40 g L-1 glucose was added as carbon source. The inoculum contained the indicated percentage of FORM-gfp with the rest to 100% being WT-crimson cells. Biomass formation (black diamond) and culture compositions (stacked red and green bars) were determined after 0, 8, and 24 h by OD600 and FACS analysis. Values represent means with standard deviations of triplicate cultivations
Establishing L-lysine overproduction with formamide as sole nitrogen source
The overproduction of nitrogenous compounds such as amino acids, in particular lysine that contains two nitrogen atoms, requires higher nitrogen concentrations in the growth medium. Since we observed slight growth inhibition of the formamidase-negative wild type-derived strain (Fig. 1a), it was tested first, if formamide affects the formamidase-positive strain FORM differently than a formamidase-negative strain. Growth experiments with strain FORM in CgXII containing 0, 20, 40, 60, 80, 120, or 160 mM formamide in addition to the regular ammonium and urea concentrations showed roughly similar effects on the growth rate, lag phase, and biomass concentration as observed for the formamidase-negative strain WT-EV (Fig. 1b). When formamide was used as sole nitrogen source in the growth medium N-CgXII, FORM grew with the highest growth rate at 60 mM formamide, which complies with the reported affinity of AmiF for formamide (Km = 32 ± 8.7 mM; Skouloubris et al. 2001). The growth rate was reduced to half-maximal at about 200 mM (Fig. 1b, c). However, the use of formamide as sole nitrogen source increased the lag phases with increasing formamide concentrations to a larger extent than as combined nitrogen source (Fig. 1b, c). By contrast, although growth was slowed, the attained maximal biomass concentrations increased with increasing formamide concentrations from 20 to 160 mM (Fig. 1c). In sum, our observations emphasize a clear tradeoff between immediate fast growth and high biomass formation with formamide as sole nitrogen source. To compromise for sufficient nitrogen provision to allow high biomass formation at an adequate growth rate, all following cultivations were performed using 60 mM formamide.
As it is known that formate, the second product of the formamidase reaction besides NH4+, has an inhibitory effect on growth of C. glutamicum (Witthoff et al. 2012; Ryabchenko et al. 2020), we compared its growth inhibitory effect for formamidase-negative strain WT-EV and formamidase-positive strain FORM. The effect of formate on the growth rate was comparable between the two strains (Fig. 1d, e); however, lag phases were longer for strain FORM. Moreover, a positive effect of formate on the biomass concentration of strain FORM was observed only until 40 mM formate, when a plateau was reached, while the formate concentration correlated with increased biomass formation by strain WT-EV (Fig. 1d, e). Thus, higher formate concentrations are tolerated less when formamidase gene amiF is expressed.
Formate is oxidized to carbon dioxide by formate dehydrogenase with concomitant formation of a reduced redox cofactor. Endogenous formate dehydrogenase FdhCg is NAD+-dependent, while an NADP+-dependent variant of Pseudomonas sp. 101 (FdhPs) has been described (Calzadiaz-Ramirez et al. 2020). Both genes were overexpressed in strain FORM as part of a synthetic operon with amiF, yielding strains FORM-FdhCg and FORM-FdhPs.
When these strains were cultivated in N-CgXII supplemented with 60, 120, or 160 mM formamide, comparable effects on the growth rate and maximal biomass concentration were observed (Fig. 7a, b). However, strain FORM-FdhCg grew faster than the other strains (at 160 mM formamide µmax was 13% higher) and at 160 mM formamide showed a lag phase shorter by 40% (Fig. 7b, c). These improved growth characteristics revealed overexpression of native formate dehydrogenase as an effective strategy to alleviate the formate-mediated growth perturbation of C. glutamicum with formamide and formamidase.
Biomass formation (a), maximal growth rates µmax (b), and lag phases (c) of FORM (blue) and formate dehydrogenase overexpressing strains FORM-FdhCg (green) and FORM-FdhPs (orange). Strains were cultivated in N-CgXII, supplemented with 40 g L−1 glucose and 60, 120, or 160 mM formamide in a BioLector microcultivation system for 96 h. Values represent means with standard deviations of triplicate cultivations
Therefore, based on the lysine producer C. glutamicum GRLys1ΔsugRΔldhA (Pérez-García et al. 2016), referred to as Lys, we constructed the formamidase-positive derivatives Lys-FORM, Lys-FORM-FdhCg, and Lys-FORM-FdhPs. Lysine production by strains Lys and Lys-FORM was compared in a 15N-labeling experiment. Strain Lys could not grow with formamide, but when 15N-labeled ammonium sulfate was provided, > 99% of nitrogen atoms of lysine in biomass and supernatants were labeled, whereas no labeling was detectable upon cultivation with unlabeled nitrogen sources (Fig. 8). Lysine in the biomass of strain Lys-FORM as well as in the culture supernatants was uniformly labeled from 15N-formamide (Fig. 8), demonstrating that formamide was utilized for growth and lysine production. A comparison of lysine production by strains Lys-FORM, Lys-FORM-FdhCg, and Lys-FORM-FdhPs revealed that Lys-FORM-FdhCg produced less lysine from 120 mM formamide (6.96 ± 0.57 g L−1) than strain Lys-FORM produced in medium with ammonium and urea (10.62 ± 0.93 g L−1), while strain Lys-FORM-FdhPs produced more lysine from 120 mM formamide (11.20 ± 0.90 g L−1; Fig. 9d). The lysine titer of Lys-FORM-FdhPs was not significantly higher than that of strain Lys-FORM and occurred only after a very long lag phase (Fig. 9c, d).
15N labelling of lysine in biomass and supernatants of strains Lys and Lys-FORM. Cells were grown in N-CgXII, supplemented with 60 mM unlabeled (14N) or labeled (15N) (NH4)2SO4 or formamide, using 40 g L−1 glucose, in a BioLector microcultivation system for 24 h. The patterns depict the fractions of 0 (grey), 1 (red), or 2 (purple) of labeled nitrogen atoms per molecule of lysine in the biomass (filled bars) or the supernatant (dashed bars). Values represent means with standard deviations of triplicate cultivations. *n.g., no growth with formamide
Biomass formation (a), maximal growth rates µmax (b), lag phases (c), and L-lysine titers (d) for strains Lys-FORM (blue), Fdh-overexpressing Lys-FORM-FdhPs (orange), and Lys-FORM-FdhCg (green). Strains were cultivated in N-CgXII, supplemented with 60 or 120 mM nitrogen in form of urea and (NH4)2SO4 (filled bars) or formamide (dashed bars) and 40 g L−1 glucose, in a BioLector microcultivation system for 96 h. Values represent means with standard deviations of triplicate cultivations
Formamide-based production of L-glutamate, N-methyl-L-phenylalanine, and dipicolinate
To test if the developed strategy for formamide utilization can be transferred to production of other compounds, production of L-glutamate, N-methyl-L-phenylalanine (NMePhe), and dipicolinate (DPA) was tested with formamide as sole carbon source. Addition of ciprofloxacin at mid-exponential growth was used to elicit glutamate production (Lubitz and Wendisch 2016). This led to the accumulation of 6.51 ± 0.49 g L−1 L-glutamate from 60 mM formamide (Table 2). The use of 60 mM nitrogen as urea and (NH4)2SO4 resulted in a lower L-glutamate titer, but a higher biomass concentration (Table 2). Next, NMePhe and DPA producing strains MePhe5* (Kerbs et al. 2021) and Dpa1 (Schwardmann et al. 2022) were equipped with a plasmid for formamidase expression. The resulting strains MePhe5*-FORM and Dpa1-FORM grew in N-CgXII, supplemented with 60 mM formamide as sole nitrogen source (Table 2), though slower and to lower total biomass concentrations (Fig. 1b). N provision in form of formamide instead of urea and (NH4)2SO4 reduced biomass formation by up to 60% (Table 2), while the NMePhe and DPA product titers from formamide surpassed those from urea and (NH4)2SO4 by up to 80% and amounted to 1.68 ± 0.20 g L−1 NMePhe, and 0.56 ± 0.01 g L−1 DPA, respectively (Table 2).
Taken together, formamide supported production of L-glutamate, L-lysine, NMePhe, and DPA and the observed biomass specific product yields (YP/X) were approximately 1.4- to threefold than with the same nitrogen provision from urea and (NH4)2SO4 (Table 2). Thus, the developed strategy was successfully transferred to the production of various nitrogenous compounds, and formamide was revealed as superior amine source regarding the attainable biomass specific product yields.
Discussion
This study provides the first report on formamide-based growth of C. glutamicum. The activity of the synthetic pathway was substantiated by detection of formamidase activity and stable isotope labeling. In co-culture, ammonium released from a formamidase-positive strain supported growth of a formamidase-deficient strain when formamide was the only nitrogen source. Notably, transfer of this metabolic pathway and substitution of regular nitrogen sources for formamide enabled production of various nitrogenous compounds.
Besides, wild-type C. glutamicum growing well in the presence of at least 160 mM formamide, i.e., higher than the concentration tolerated by the naturally amiF harboring H. pylori 26695 (Vliet et al. 2003) and amiF expression in C. glutamicum strain FORM, led to formamidase activity of 6.0 ± 0.8 U mg−1 which was comparable to the donor H. pylori (approximately 2.7 to 5 U mg−1 total crude extract protein) (Vliet et al. 2003; Bury-Moné et al. 2004). This activity exceeded that reported for heterologous overexpression of the amiF gene in B. subtilis (1.2 U mg−1) (Guo et al. 2020a, b).
The release of NH4+ by H. pylori in its natural acidic environment may be disadvantageous for other organisms by decoupling the transmembrane pH gradient (Kleiner 1981). Nevertheless, formamide cleavage by strain FORM provided sufficient ammonium to support growth of a co-cultured formamidase-deficient C. glutamicum strain. While the uptake system for formamide in C. glutamicum is not known, ammonium uptake is mediated by the membrane potential-dependent ammonium transporters AmtA and AmtB (Siewe et al. 1998; Jakoby et al. 2000). A similar observation was made in co-cultures of E. coli with one of the cognate amino acids arginine or glutamate serving as sole nitrogen source and leakage of ammonium enabling growth of the co-cultured competitor strain (Wang et al. 2016). However, unwanted background growth by the parental E. coli strain with the provided amino acids as sole nitrogen source was possible (Wang et al. 2016), but was not observed here as C. glutamicum WT-EV showed no detectable growth with formamide. Thus, formamide utilization as alternative nitrogen source is suited to maintain synthetic consortia with strict mandatory dependency.
The controlled feeding of formamide to a co-culture consisting of formamidase-positive and -negative strains grown with either ammonium and/or urea will favor the formamidase-positive strains (compare Fig. 1a, b), while the provision of formamide as sole nitrogen source will favor formamidase-negative strains (compare Fig. 6) to regulate co-culture dynamics. This approach may complement the strategic inoculation to regulate co-cultivation dynamics as shown for C. glutamicum strains utilizing different carbon sources for production of riboflavin in co-culture (Pérez-García et al. 2021). Alternatively, the trait of formamide utilization may be used to design mutualistic synthetic consortia such as shown for co-culturing E. coli with C. glutamicum for production of cadaverine and L-pipecolic acid from starch (Sgobba et al. 2018a, b). In this context, the choice of a non-nitrogenous product might be preferable to prevent nitrogen limitation perturbing production or the process design would require accurate fine-tuning for balanced and sufficient nitrogen availability.
Previously, anticontamination systems for naturally formamidase deficient E. coli and B. subtilis strains have been engineered by heterologous expression of amiF and use of formamide as sole nitrogen source (Guo et al. 2020a, b; Ou et al. 2019). This type of contamination-free cultivation under non-sterile conditions can also be envisioned for C. glutamicum to avoid of time-, and cost-, and resource-intensive sterilization processes and antibiotic addition. However, the ammonium leakage observed for C. glutamicum strain FORM may pose a problem as described for the use of phosphite as sole phosphorus source when the gene ptxD for phosphite dehydrogenase from Pseudomonas stutzeri was expressed (Guo et al. 2020a, b). Hence, the amiF/formamide system alone cannot ensure contamination-free non-sterile cultivation. However, the combined use of the amiF/formamide and ptxD/phosphite systems, as employed for non-sterile cultivation of E. coli and B. subtilis strains (Guo et al. 2020a, b; Ou et al. 2019), may be required to avoid contaminations.
Formate generated by formamidase from formamide is inhibiting growth to a much larger extent than the other product of formamidase, NH4+, that is growth inhibitory only at very high extracellular concentrations, supposedly due to elevated ionic strength or osmolarity (Müller et al. 2006). Formate tolerance was suggested to be connected to the activity level of formate oxidizing formate dehydrogenase (Cotton et al. 2020). C. glutamicum naturally possesses formate dehydrogenase FdhF (encoded by cg0618) that reduces growth retardation due to 100 mM extracellular formate (Witthoff et al. 2012). Here, we showed that overexpression of genes for native NAD-dependent FdhF as well as NADP-dependent Pseudomonas Fdh (Calzadiaz-Ramirez et al. 2020) alleviated growth inhibition by formate generated in the AmiF reaction. However, under the chosen conditions, provision of NADPH by FdhPs instead of NADH by native FdhF did not improve lysine production, a biosynthesis pathway requiring 4 NADPH molecules per lysine molecule. Such an advantage was only seen when the gene for the native Fdh of a C. glutamicum lysine producer strain was disrupted (Ryabchenko et al. 2020), indicating sufficient formate dehydrogenase activity in wild-type C. glutamicum.
To the best of our knowledge, formamide was hitherto only used as xenobiotic nitrogen source for the production of nitrogen-free compounds, namely acetoin and 2,3-butanediol by engineered strains of B. subtilis and K. pneumoniae, respectively (Guo et al. 2020a, b). For the first time, we demonstrated formamide-based overproduction of four nitrogenous compounds. Notably, while DPA yields were comparable with 60 mM formamide to 60 mM nitrogen in form of urea and (NH4)2SO4, formamide-based production of glutamate, lysine, and NMePhe was superior to production with 60 mM nitrogen in form of urea and (NH4)2SO4. The superior yields from formamide were observed at comparatively low nitrogen concentrations. A balance between the concentrations for carbon and nitrogen sources is not only important to increase titers, but may also change product to by-product ratios as observed with 47 mM nitrogen for NMePhe and DPA production, which led to lower formation of byproducts N-methylalanine and lysine, respectively (Kerbs et al. 2021; Schwardmann et al. 2022).
Data availability
All data are present in the manuscript and its Supplement.
Code availability
Not applicable.
References
Bipp H, Kieczka H (2011) Formamides. In: Wiley-VCH (eds) Ullmann’s encyclopedia of industrial chemistry, 7th edn. Wiley-VCH, Weinheim, pp 720–741. https://doi.org/10.1002/14356007.a12_001.pub2
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1006/abio.1976.9999
Brown D, Hitchcock MJM, Katz E (1986) Purification and characterization of kynurenine formamidase activities from Streptomyces parvulus. Can J Microbiol 32:465–472. https://doi.org/10.1139/m86-086
Bury-Moné S, Thiberge J-M, Contreras M, Maitournam A, Labigne A, De Reuse H (2004) Responsiveness to acidity via metal ion regulators mediates virulence in the gastric pathogen Helicobacter pylori. Mol Microbiol 53:623–638. https://doi.org/10.1111/j.1365-2958.2004.04137.x
Calzadiaz-Ramirez L, Calvó-Tusell C, Stoffel GMM, Lindner SN, Osuna S, Erb TJ, Garcia-Borràs M, Bar-Even A, Acevedo-Rocha CG (2020) In vivo selection for formate dehydrogenases with high efficiency and specificity toward NADP+. ACS Catal 10:7512–7525. https://doi.org/10.1021/acscatal.0c01487
Cetnar DP, Salis HM (2021) Systematic quantification of sequence and structural determinants controlling mRNA stability in bacterial operons. ACS Synth Biol 10:318–332. https://doi.org/10.1021/acssynbio.0c00471
Chen XH, Koumoutsi A, Scholz R, Eisenreich A, Schneider K, Heinemeyer I, Morgenstern B, Voss B, Hess WR, Reva O, Junge H, Voigt B, Jungblut PR, Vater J, Süssmuth R, Liesegang H, Strittmatter A, Gottschalk G, Borriss R (2007) Comparative analysis of the complete genome sequence of the plant growth-promoting bacterium Bacillus amyloliquefaciens FZB42. Nat Biotechnol 25:1007–1014. https://doi.org/10.1038/nbt1325
Cotton CA, Claassens NJ, Benito-Vaquerizo S, Bar-Even A (2020) Renewable methanol and formate as microbial feedstocks. Curr Opin Biotech 62:168–180. https://doi.org/10.1016/j.copbio.2019.10.002
Duca D, Rose DR, Glick BR (2014) Characterization of a nitrilase and a nitrile hydratase from Pseudomonas sp. strain UW4 that converts indole-3-acetonitrile to indole-3-acetic acid. Appl Environ Microb 80:4640–4649. https://doi.org/10.1128/AEM.00649-14
Dunn BE, Cohen H, Blaser MJ (1997) Helicobacter pylori. Clin Microbiol Rev 10:720–741. https://doi.org/10.1128/CMR.10.4.720
Egelkamp R, Schneider D, Hertel R, Daniel R (2017) Nitrile-degrading bacteria isolated from compost. Front Environ Sci 5:56. https://doi.org/10.3389/fenvs.2017.00056
Eggeling L, Bott M (2005) Handbook of Corynebacterium glutamicum, 1st edn. CRC Press, Boca Raton
Eikmanns BJ, Thum-Schmitz N, Eggeling L, Lüdtke K-U, Sahm H (1994) Nucleotide sequence, expression and transcriptional analysis of the Corynebacterium glutamicum gltA gene encoding citrate synthase. Microbiology 140:1817–1828. https://doi.org/10.1099/13500872-140-8-1817
Fournand D, Arnaud A (2001) Aliphatic and enantioselective amidases: from hydrolysis to acyl transfer activity. J Appl Microbiol 91:381–393. https://doi.org/10.1046/j.1365-2672.2001.01378.x
Giavalisco P, Li Y, Matthes A, Eckhardt A, Hubberten H-M, Hesse H, Segu S, Hummel J, Köhl K, Willmitzer L (2011) Elemental formula annotation of polar and lipophilic metabolites using 13C, 15N and 34S isotope labelling, in combination with high-resolution mass spectrometry. Plant J 68:364–376. https://doi.org/10.1111/j.1365-313X.2011.04682.x
Gibson DG, Young L, Chuang R-Y, Venter JC, Hutchison CA, Smith HO (2009) Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6:343–345. https://doi.org/10.1038/nmeth.1318
Gong J-S, Lu Z-M, Li H, Shi J-S, Zhou Z-M, Xu Z-H (2012) Nitrilases in nitrile biocatalysis: recent progress and forthcoming research. Microb Cell Fact 11:142. https://doi.org/10.1186/1475-2859-11-142
Guo Z-W, Ou X-Y, Liang S, Gao H-F, Zhang L-Y, Zong M-H, Lou W-Y (2020a) Recruiting a phosphite dehydrogenase/formamidase-driven antimicrobial contamination system in Bacillus subtilis for nonsterilized fermentation of acetoin. ACS Synth Biol 9:2537–2545. https://doi.org/10.1021/acssynbio.0c00312
Guo Z-W, Ou X-Y, Xu P, Gao H-F, Zhang L-Y, Zong M-H, Lou W-Y (2020b) Energy- and cost-effective non-sterilized fermentation of 2,3-butanediol by an engineered Klebsiella pneumoniae OU7 with an anti-microbial contamination system. Green Chem 22:8584–8593. https://doi.org/10.1039/D0GC03044A
Hanahan D (1983) Studies on transformation of Escherichia coli with plasmids. J Mol Biol 166:557–580. https://doi.org/10.1016/s0022-2836(83)80284-8
Henke NA, Krahn I, Wendisch VF (2021) Improved plasmid-based inducible and constitutive gene expression in Corynebacterium glutamicum. Microorganisms 9:204. https://doi.org/10.3390/microorganisms9010204
Hu M, Liu F, Wang Z, Shao M, Xu M, Yang T, Zhang R, Zhang X, Rao Z (2022) Sustainable isomaltulose production in Corynebacterium glutamicum by engineering the thermostability of sucrose isomerase coupled with one-step simplified cell immobilization. Front Microbiol 13:979079. https://doi.org/10.3389/fmicb.2022.979079
Hung C-L, Liu J-H, Chiu W-C, Huang S-W, Hwang J-K, Wang W-C (2007) Crystal structure of Helicobacter pylori formamidase AmiF reveals a cysteine-glutamate-lysine catalytic triad. J Biol Chem 282:12220–12229. https://doi.org/10.1074/jbc.M609134200
Jakoby M, Nolden L, Meier-Wagner J, Krämer R, Burkovski A (2000) AmtR, a global repressor in the nitrogen regulation system of Corynebacterium glutamicum. Mol Microbiol 37:964–977. https://doi.org/10.1046/j.1365-2958.2000.02073.x
Kalinowski J, Bathe B, Bartels D, Bischoff N, Bott M, Burkovski A, Dusch N, Eggeling L, Eikmanns BJ, Gaigalat L, Goesmann A, Hartmann M, Huthmacher K, Krämer R, Linke B, McHardy AC, Meyer F, Möckel B, Pfefferle W, Pühler A, Rey DA, Rückert C, Rupp O, Sahm H, Wendisch VF, Wiegräbe I, Tauch A (2003) The complete Corynebacterium glutamicum ATCC 13032 genome sequence and its impact on the production of L-aspartate-derived amino acids and vitamins. J Biotechnol 104:5–25. https://doi.org/10.1016/S0168-1656(03)00154-8
Kampen WH (2014) Nutritional requirements in fermentation processes. In: Vogel HC, Todaro CM (eds) Fermentation and biochemical engineering handbook, 3rd edn. William Andrew Publishing, Boston, pp 37–57. https://doi.org/10.1016/B978-1-4557-2553-3.00004-0
Kennedy GL (2014) Formamide. In: Wexler P (ed) Encyclopedia of toxicology 3rd edn. Academic Press, Oxford, pp 657–658. https://doi.org/10.1016/B978-0-12-386454-3.00023-3
Kerbs A, Mindt M, Schwardmann L, Wendisch VF (2021) Sustainable production of N-methylphenylalanine by reductive methylamination of phenylpyruvate using engineered Corynebacterium glutamicum. Microorganisms 9:824. https://doi.org/10.3390/microorganisms9040824
Klaffl S, Eikmanns BJ (2010) Genetic and functional analysis of the soluble oxaloacetate decarboxylase from Corynebacterium glutamicum. J Bacteriol 192:2604–2612. https://doi.org/10.1128/JB.01678-09
Kleiner D (1981) The transport of NH3 and NH4+ across biological membranes. Biochim Biophys Acta 639:41–52. https://doi.org/10.1016/0304-4173(81)90004-5
Lee SY, Choi J, Han K, Song JY (1999) Removal of endotoxin during purification of poly(3-hydroxybutyrate) from Gram-negative bacteria. Appl Environ Microb 65:2762–2764. https://doi.org/10.1128/AEM.65.6.2762-2764.1999
Leuchtenberger W, Huthmacher K, Drauz K (2005) Biotechnological production of amino acids and derivatives: current status and prospects. Appl Microbiol Biot 69:1–8. https://doi.org/10.1007/s00253-005-0155-y
Liu J, Zhang Y, Feng K, Liu X, Li J, Li C, Zhang P, Yu Q, Liu J, Shen G, He L (2020) Amidase, a novel detoxifying enzyme, is involved in cyflumetofen resistance in Tetranychus cinnabarinus (Boisduval). Pestic Biochem Phys 163:31–38. https://doi.org/10.1016/j.pestbp.2019.10.001
Liu X-X, Li Y, Bai Z-H (2021) Corynebacterium glutamicum as a robust microbial factory for production of value-added proteins and small molecules: fundamentals and applications. In Microbial cell factories engineering for production of biomolecules; Singh, V (Ed), Academic Press; 235–263
Lubitz D, Wendisch VF (2016) Ciprofloxacin triggered glutamate production by Corynebacterium glutamicum. BMC Microbiol 16:235. https://doi.org/10.1186/s12866-016-0857-6
Mahenthiralingam E, Draper P, Davis EO, Colston MJ (1993) Cloning and sequencing of the gene which encodes the highly inducible acetamidase of Mycobacterium smegmatis. J Gen Microbiol 139:575–583. https://doi.org/10.1099/00221287-139-3-575
Meng N, Shao J, Li H, Wang Y, Fu X, Liu C, Yu Y, Zhang B (2022) Electrosynthesis of formamide from methanol and ammonia under ambient conditions. Nat Commun 13:5452. https://doi.org/10.1038/s41467-022-33232-w
Mindt M, Risse JM, Gruß H, Sewald N, Eikmanns BJ, Wendisch VF (2018) One-step process for production of N-methylated amino acids from sugars and methylamine using recombinant Corynebacterium glutamicum as biocatalyst. Sci Rep 8:12895. https://doi.org/10.1038/s41598-018-31309-5
Mindt M, Heuser M, Wendisch VF (2019) Xylose as preferred substrate for sarcosine production by recombinant Corynebacterium glutamicum. Bioresource Technol 281:135–142. https://doi.org/10.1016/j.biortech.2019.02.084
Müller T, Walter B, Wirtz A, Burkovski A (2006) Ammonium toxicity in bacteria. Curr Microbiol 52:400–406. https://doi.org/10.1007/s00284-005-0370-x
Newton GL, Av-Gay Y, Fahey RC (2000) A novel mycothiol-dependent detoxification pathway in Mycobacteria involving mycothiol S-conjugate amidase. Biochemistry 39:10739–10746. https://doi.org/10.1021/bi000356n
Ou X-Y, Wu X-L, Peng F, Zeng Y-J, Li H-X, Xu P, Chen G, Guo Z-W, Yang J-G, Zong M-H, Lou W-Y (2019) Metabolic engineering of a robust Escherichia coli strain with a dual protection system. Biotechnol Bioeng 116:3333–3348. https://doi.org/10.1002/bit.27165
Parish T, Mahenthiralingam E, Draper P, Davis EO, Colston EO (1997) Regulation of the inducible acetamidase gene of Mycobacterium smegmatis. Microbiology 143:2267–2276. https://doi.org/10.1099/00221287-143-7-2267
Pérez-García F, Peters-Wendisch P, Wendisch VF (2016) Engineering Corynebacterium glutamicum for fast production of L-lysine and L-pipecolic acid. Appl Microbiol Biot 100:8075–8090. https://doi.org/10.1007/s00253-016-7682-6
Pérez-García F, Burgardt A, Kallman DR, Wendisch VF, Bar N (2021) Dynamic co-cultivation process of Corynebacterium glutamicum strains for the fermentative production of riboflavin. Fermentation 7:11. https://doi.org/10.3390/fermentation7010011
Rath M, Salas J, Parhy B, Norton R, Menakuru H, Sommerhalter M, Hatlstad G, Kwon J, Allan D, Vance C, Uhde-Stone C (2010) Identification of genes induced in proteoid roots of white lupin under nitrogen and phosphorus deprivation, with functional characterization of a formamidase. Plant Soil 334:137–150. https://doi.org/10.1007/s11104-010-0373-7
Rehm N, Georgi T, Hiery E, Degner U, Schmiedl A, Burkovski A, Bott M (2010) L-Glutamine as a nitrogen source for Corynebacterium glutamicum: derepression of the AmtR regulon and implications for nitrogen sensing. Microbiology 156:3180–3193. https://doi.org/10.1099/mic.0.040667-0
Reitzer LJ (1996) Sources of nitrogen and their utilization. In: Neidhardt FC (ed) Escherichia coli and Salmonella: cellular and molecular biology, 2nd edn. ASM Press, Washington, DC, pp 380–390
Rittmann D, Lindner SN, Wendisch VF (2008) Engineering of a glycerol utilization pathway for amino acid production by Corynebacterium glutamicum. Appl Environ Microb 74:6216–6222. https://doi.org/10.1128/AEM.00963-08
Ryabchenko LE, Leonova TE, Shustikova TE, Gerasimova TV, Ivankova TA, Sidorenko KV, Yanenko AS (2020) Expression of the NADPH+-dependent formate-dehydrogenase gene from Pseudomonas increases lysine production in Corynebacterium glutamicum. Appl Biochem Microbiol 56:828–836. https://doi.org/10.1134/S0003683820080086
Saladino R, Crestini C, Pino S, Costanzo G, Di Mauro E (2012) Formamide and the origin of life. Phys Life Rev 9:84–104. https://doi.org/10.1016/j.plrev.2011.12.002
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press
Schneider J, Wendisch VF (2010) Putrescine production by engineered Corynebacterium glutamicum. Appl Microbiol Biot 88:859–868. https://doi.org/10.1007/s00253-010-2778-x
Schneider J, Niermann K, Wendisch VF (2011) Production of the amino acids L-glutamate, L-lysine, L-ornithine and L-arginine from arabinose by recombinant Corynebacterium glutamicum. J Biotechnol 154:191–198. https://doi.org/10.1016/j.jbiotec.2010.07.009
Schneider J, Eberhardt D, Wendisch VF (2012) Improving putrescine production by Corynebacterium glutamicum by fine-tuning ornithine transcarbamoylase activity using a plasmid addiction system. Appl Microbiol Biot 95:169–178. https://doi.org/10.1007/s00253-012-3956-9
Schwardmann LS, Dransfeld AK, Schäffer T, Wendisch VF (2022) Metabolic engineering of Corynebacterium glutamicum for sustainable production of the aromatic dicarboxylic acid dipicolinic acid. Microorganisms 10:730. https://doi.org/10.3390/microorganisms10040730
Seffernick JL, de Souza ML, Sadowsky MJ, Wackett LP (2001) Melamine deaminase and atrazine chlorohydrolase: 98 percent identical but functionally different. J Bacteriol 183:2405–2410. https://doi.org/10.1128/JB.183.8.2405-2410.2001
Selão TT, Włodarczyk A, Nixon PJ, Norling B (2019) Growth and selection of the cyanobacterium Synechococcus sp. PCC 7002 using alternative nitrogen and phosphorus sources. Metab Eng 54:255–263. https://doi.org/10.1016/j.ymben.2019.04.013
Sgobba E, Blöbaum L, Wendisch VF (2018a) Production of food and feed additives from non-food-competing feedstocks: valorizing n-acetylmuramic acid for amino acid and carotenoid fermentation with Corynebacterium glutamicum. Front Microbiol 9:2046. https://doi.org/10.3389/fmicb.2018.02046
Sgobba E, Stumpf AK, Vortmann M, Jagmann N, Krehenbrink M, Dirks-Hofmeister ME, Moerschbacher B, Philipp B, Wendisch VF (2018b) Synthetic Escherichia coli-Corynebacterium glutamicum consortia for L-lysine production from starch and sucrose. Bioresource Technol 260:302–310. https://doi.org/10.1016/j.biortech.2018.03.113
Shaw AJ, Lam FH, Hamilton M, Consiglio A, MacEwen K, Brevnova EE, Greenhagen E, LaTouf WG, South CR, van Dijken H, Stephanopoulos G (2016) Metabolic engineering of microbial competitive advantage for industrial fermentation processes. Science 353:583–586. https://doi.org/10.1126/science.aaf6159
Siewe RM, Weil B, Krämer R (1995) Glutamine uptake by a sodium-dependent secondary transport system in Corynebacterium glutamicum. Arch Microbiol 164:98–103. https://doi.org/10.1007/BF02525314
Siewe RM, Weil B, Burkovski A, Eggeling L, Krämer R, Jahns T (1998) Urea uptake and urease activity in Corynebacterium glutamicum. Arch Microbiol 169:411–416. https://doi.org/10.1007/s002030050591
Skouloubris S, Labigne A, De Reuse H (2001) The AmiE aliphatic amidase and AmiF formamidase of Helicobacter pylori: natural evolution of two enzyme paralogues. Mol Microbiol 40:596–609. https://doi.org/10.1046/j.1365-2958.2001.02400.x
Soriano-Maldonado P, Martínez-Gómez AI, Andújar-Sánchez M, Neira JL, Clemente-Jiménez JM, Las Heras-Vázquez FJ, Rodríguez-Vico F, Martínez-Rodríguez S (2011) Biochemical and mutational studies of the Bacillus cereus CECT 5050T formamidase support the existence of a C-E-E-K tetrad in several members of the nitrilase superfamily. Appl Environ Microb 77:5761–5769. https://doi.org/10.1128/AEM.00312-11
Srivastava P, Deb JK (2005) Gene expression systems in corynebacteria. Protein Express Purif 40:221–229. https://doi.org/10.1016/j.pep.2004.06.017
Thatcher RC, Weaver TL (1976) Carbon-nitrogen cycling through microbial formamide metabolism. Science 192:1234–1235. https://doi.org/10.1126/science.192.4245.1234
Tsuge Y, Matsuzawa H (2021) Recent progress in production of amino acid-derived chemicals using Corynebacterium glutamicum. World J Microb Biotechnol 37:49. https://doi.org/10.1007/s11274-021-03007-4
Uhde A, Youn J-W, Maeda T, Clermont L, Matano C, Krämer R, Wendisch VF, Seibold GM, Marin K (2013) Glucosamine as carbon source for amino acid-producing Corynebacterium glutamicum. Appl Microbiol Biot 97:1679–1687. https://doi.org/10.1007/s00253-012-4313-8
Valappil SP, Boccaccini AR, Bucke C, Roy I (2007) Polyhydroxyalkanoates in Gram-positive bacteria: insights from the genera Bacillus and Streptomyces. Antonie Van Leeuwenhoek 91:1–17. https://doi.org/10.1007/s10482-006-9095-5
van der Rest ME, Lange C, Molenaar D (1999) A heat shock following electroporation induces highly efficient transformation of Corynebacterium glutamicum with xenogeneic plasmid DNA. Appl Microbiol Biot 52:541–545. https://doi.org/10.1007/s002530051557
van Vliet AHM, Stoof J, Poppelaars SW, Bereswill S, Homuth G, Kist M, Kuipers EJ, Kusters JG (2003) Differential regulation of amidase- and formamidase-mediated ammonia production by the Helicobacter pylori Fur Repressor. J Biol Chem 278:9052–9057. https://doi.org/10.1074/jbc.M207542200
Wang J, Yan D, Dixon R, Wang Y-P (2016) Deciphering the principles of bacterial nitrogen dietary preferences: a strategy for nutrient containment. mBio 7:e00792-16. https://doi.org/10.1128/mBio.00792-16
Weatherburn MW (1967) Phenol-hypochlorite reaction for determination of ammonia. Anal Chem 39:971–974. https://doi.org/10.1021/ac60252a045
Wendisch VF, De Graaf AA, Sahm H, Eikmanns BJ (2000) Quantitative determination of metabolic fluxes during coutilization of two carbon sources: comparative analyses with Corynebacterium glutamicum during growth on acetate and/or glucose. J Bacteriol 182:3088–3096. https://doi.org/10.1128/JB.182.11.3088-3096.2000
Wendisch VF, Nampoothiri KM, Lee J-H (2022) Metabolic engineering for valorization of agri- and aqua-culture sidestreams for production of nitrogenous compounds by Corynebacterium glutamicum. Front Microbiol 13:835131. https://doi.org/10.3389/fmicb.2022.835131
Wendisch VF, Lee J-H (2020) Metabolic engineering in Corynebacterium glutamicum. In: Inui, M., Toyoda, K. (Eds.), Corynebacterium glutamicum - biology and biotechnology, microbiology monographs. Springer International Publishing, 287–322. https://doi.org/10.1007/978-3-030-39267-3_10
Willison JC (1993) Biochemical genetics revisited: the use of mutants to study carbon and nitrogen metabolism in the photosynthetic bacteria. FEMS Microbiol Rev 10:1–38. https://doi.org/10.1111/j.1574-6968.1993.tb05862.x
Witthoff S, Eggeling L, Bott M, Polen T (2012) Corynebacterium glutamicum harbours a molybdenum cofactor-dependent formate dehydrogenase which alleviates growth inhibition in the presence of formate. Microbiology 158:2428–2439. https://doi.org/10.1099/mic.0.059196-0
Wolf S, Becker J, Tsuge Y, Kawaguchi H, Kondo A, Marienhagen J, Bott M, Wendisch VF, Wittmann C (2021) Advances in metabolic engineering of Corynebacterium glutamicum to produce high-value active ingredients for food, feed, human health, and well-being. Essays Biochem 65:197–212. https://doi.org/10.1042/EBC20200134
You L, Page L, Feng X, Berla B, Pakrasi HB, Tang YJ (2012) Metabolic pathway confirmation and discovery through (13)C-labeling of proteinogenic amino acids. JoVE 26(59):e3583 https://doi.org/10.3791/3583
Zahoor A, Lindner SN, Wendisch VF (2012) Metabolic engineering of Corynebacterium glutamicum aimed at alternative carbon sources and new products. Comput Struct Biotechnol J 3:e201210004. https://doi.org/10.5936/csbj.201210004
Zerkle AL, Mikhail S (2017) The geobiological nitrogen cycle: from microbes to the mantle. Geobiology 15:343–352. https://doi.org/10.1111/gbi.12228
Zhang B, Jiang Y, Li Z, Wang F, Wu X-Y (2020) Recent progress on chemical production from non-food renewable feedstocks using Corynebacterium glutamicum. Front Bioeng Biotechnol 8:606047. https://doi.org/10.3389/fbioe.2020.606047
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was supported in part by the BMBF project ForceYield (031B0825C).
Author information
Authors and Affiliations
Contributions
L.S.S. and A.K.D. cloned strains and performed experiments. T.W. analyzed stable isotope labeling samples. V.F.W., L.S.S. and S.N.L. conceived and designed the study. V.F.W. acquired funding. L.S.S., V.F.W. and S.N.L. drafted the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schwardmann, L.S., Wu, T., Dransfeld, A.K. et al. Formamide-based production of amines by metabolically engineering Corynebacterium glutamicum. Appl Microbiol Biotechnol 107, 4245–4260 (2023). https://doi.org/10.1007/s00253-023-12592-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-023-12592-3