Abstract
Crown rust, caused by Puccinia coronata f. sp. avenae (Pca), is a significant impediment to global oat production. Some 98 alleles at 92 loci conferring resistance to Pca in Avena have been designated; however, allelic relationships and chromosomal locations of many of these are unknown. Long-term monitoring of Pca in Australia, North America and elsewhere has shown that it is highly variable even in the absence of sexual recombination, likely due to large pathogen populations that cycle between wild oat communities and oat crops. Efforts to develop cultivars with genetic resistance to Pca began in the 1950s. Based almost solely on all all-stage resistance, this has had temporary benefits but very limited success. The inability to eradicate wild oats, and their common occurrence in many oat growing regions, means that future strategies to control Pca must be based on the assumption of a large and variable prevailing pathogen population with high evolutionary potential, even if cultivars with durable resistance are deployed and grown widely. The presence of minor gene, additive APR to Pca in hexaploid oat germplasm opens the possibility of pyramiding several such genes to give high levels of resistance. The recent availability of reference genomes for diploid and hexaploid oat will undoubtedly accelerate efforts to discover, characterise and develop high throughput diagnostic markers to introgress and pyramid resistance to Pca in high yielding adapted oat germplasm.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
“The extremely variable heteroecious fungus, Puccinia coronata, attacks many Gramineae and causes economically important crown rust of oat. Such a high level of specialization is exhibited on the 190 species of susceptible Gramineae and 59 species of Rhamnus and 6 other genera of Rhamnaceae and Eleagnaceae that a satisfactory sub-specific classification of the pathogen has still to be devised. Despite this difficulty, breeding programs have developed temporarily useful commercial resistant varieties for the major oat-growing areas of the world. The use of multiline resistant cultivars seems particularly promising for control. This disease thus illustrates the continuing race between the plant breeder and the pathogen”. This introductory paragraph from a monograph on crown rust of oats and grasses by Simons (1970) provides deep insight into the status of knowledge and progress in breeding oat for resistance to the destructive crown rust disease in 1970.
Among cereals, oat ranks seventh in the world in terms of production, after wheat, maize, rice, barley, sorghum and millet. This ranking is achieved largely through the use of improved cultivars of Avena sativa for livestock feed and human consumption, and to some extent cultivars of A. strigosa grown mostly in South America (Suttie and Renolds 2004). Awareness of the nutritional and health benefits of oat continues to increase, especially since the discovery of phytoalexins such as avenanthramides (Collins 1989) in oat groats. These phenolic compounds are reported to possess antioxidant (Emmons and Peterson 1999) and cholesterol (LDL) lowering properties (Chen et al. 2004). Minimising losses to diseases will be critical in meeting projected increases in future global demand for cereals such as oat. Crown rust (caused by Puccinia coronata f. sp. avenae; Pca) and stem rust (caused by Puccinia graminis f. sp. avenae; Pga) occur in all major oat growing regions of the world and have caused significant yield losses in oat crops. Pca is widespread globally (Savile 1984) and is considered one of the most important oat diseases and an important factor limiting oat production world-wide (Harder and Haber 1992; Sebesta et al. 2003; Nazareno et al. 2018). It was considered by Leonard et al. (2004) as the most serious threat to oat production in North America, with records of epidemics dating back more than 100 years. From 1918 to 1930, annual estimated losses in the USA averaged greater than 13.7 million bushels (Murphy 1935). Simons and Murphy (1961) reported annual losses of up to 30% in years when crown rust was severe. Severe damage due to Pca has also occurred in coastal areas of Australia in years favourable to the disease (Waterhouse 1952; Luig 1985). Grain yields are negatively correlated with crown rust severity, and may be reduced by as much as 50% in susceptible cultivars (Leonard and Martinelli 2005). In addition to causing reductions in grain yield, severe crown rust can also reduce the yield and quality of forage (Simons 1985).
A great deal of effort has been made to investigate the genetics of resistance to Pca in Avena, quite possibly more than any other pathogen of oat, with a view to developing crown rust resistant oat cultivars. However, the lack of a physical map for the oat genome for many years limited advances in the genetics of resistance to Pca. Some 98 alleles at 92 loci that confer resistance to Pca in Avena have been catalogued and accepted, however the chromosomal location of many of these genes remains unknown. In addition, studies on the pathogenicity of Pca on these alleles have shown that virulence for most exists, making breeding for resistance to this rust pathogen very challenging. Tellingly, as quoted above, Simons (1970) stated that “breeding programs have developed temporarily useful commercial resistant varieties”, a testament to the propensity of Pca to change and evade detection by resistance genes. This review summarises the current state of knowledge of the genetic basis of resistance to Pca in oat, how it has been applied to resistance breeding, and looks to future advances that we believe will lead to more sustained and environmentally sound approaches to control this globally damaging pathogen.
Advances in oat genomics
Avena sativa has a large genome (ca. 12.5 Gb) comprising 21 chromosome pairs from three separate ancestral genomes, with the genome constitution of AACCDD. Physical rearrangements among the ancestral genomes (e.g. Chen and Armstrong 1994; Leggett and Markhand 1995) have impeded the development of linkage maps for A. sativa, and at times have made the assignment of loci to chromosomes very difficult. The first oat consensus map was developed by Oliver et al. (2013), based on 985 SNPs assayed on 390 recombinant inbred lines (RILs) derived from six hexaploid oat biparental populations and SNP deletion analysis in a set of monosomic stocks. This map provided substantial improvements over previous maps because of low error rates in the scoring of SNP markers, and the joint mapping of markers assayed in parallel across multiple populations. Further improvements were made by integrating high-density SNPs discovered via genotype-by-sequencing (GBS) (Tinker et al. 2014).
In a further attempt to develop a single high density consensus linkage map representative of the majority of commonly grown oat cultivars, Chaffin et al. (2016) not only saturated the map with additional populations and genetic markers but also corrected errors in the previous map and gained a better understanding of biological deviations among existing maps. Homoeologous regions among oat chromosomes and matches to orthologous regions of rice (Oryza sativa) revealed that the hexaploid oat genome is highly rearranged compared to the diploid genome as a consequence of frequent translocation among chromosomes and such subgenome rearrangements probably accounted for the failure of certain linkage groups to correspond to the consensus map. In other advances in oats genomics, Yan et al. (2016) performed high-density marker profiling and genome analysis of 27 oat species and further identified ancestral origins of 21 mapped chromosomes in hexaploid oat, and deduced the likely pathway from which hexaploid oat originated by sequential polyploidization events. In a further de novo GBS analysis of 4,657 accessions of cultivated oat, Bekele et al. (2018) discovered 164,741 tag-level genetic variants comprising 241,224 SNPs that were used to expand marker density of the existing oat consensus map by the addition of more than 70,000 GBS loci (Tinker et al. 2016). This high-density marker system was used to construct the first oat haplotype map.
GBS marker analysis by Latta et al. (2019) allowed additional improvement in the maps of diploid A genome (A. strigosa x A. wiestii, 2n = 14), and tetraploid AB genome (A. barbata 2n = 28). Seven linkage groups in the tetraploid showed considerably greater homology and synteny with the A genome diploids than did the other seven, suggesting an allopolyploid hybrid origin of A. barbata rather than autopolyploid genome doubling within a single species.
Bekele et al. (2020) enhanced GBS in oat by developing the targeted assay ‘Rapture’, based on an additional bait-based capture of specific DNA fragments representing close to 10 K loci. The assay accomplished deeper sequence coverage of target markers by including only those fragments that provided effective polymorphic markers. The authors argued that Rapture was cost-effective (40% less than GBS) and consistently outperformed the GBS or SNP assays when filtering markers at 80% completeness or greater, even though the total number of reads per sample of Rapture was only a quarter of that of GBS.
Although accomplishment of a full genome assembly of hexaploid oats is extremely challenging because of its highly repetitive large genome size with numerous major intra – and intergenomic arrangements, Maughan et al. (2019) developed fully annotated, chromosome scale assemblies for the extant progenitor species of the As- and Cp-subgenomes, A. atlantica and A. eriantha, respectively. These genome assemblies span 3.69 Gb with an N50 of 513 Mb (A. atlantica) and 3.78 Gb with an N50 of 535 Mb (A. eriantha) annotating approximately 50 K gene models in each species that include ~ 3 K resistance gene analogs. By systematically comparing consensus maps and analysing syntenic relationships with other cereals and homeology within oat, the authors found orthologous relationships within Pooideae and subgenome origins for each of the 21 hexaploid linkage groups.
In 2020, Pepsico in partnership with several public and private research organizations unveiled the first ever A. sativa genome assembly v1 for use in open-source applications (https://wheat.pw.usda. gov/GG3/graingenes_downloads/oat-ot3098-pepsico). Based on the oat genotype OT3098, this assembly represents a major breakthrough in oat genomics research and will open new horizons in spurring genetic improvement of traits of interest in oat including high throughput mapping and cloning of crown rust resistance genes. Jiang et al. (2021) predicted the chromosomal distribution of tandem repeats of all the 21 individual chromosomes by analysing data from the OT3098 reference assembly v1. Eight new oligonucleotide probes were then designed for non-denaturing fluorescence in situ hybridization, which were used along with 11 existing probes for chromosome karyotyping on mitotic metaphase spreads of six different species (A. brevis, A. nuda, A. wiestii, A. ventricosa, A. fatua and A. sativa). Breakage points in some chromosome translocations in A. sativa were also identified.
An advanced version (v2) of the OT3098 assembly has also been released, incorporating gap filling as well as reorientation/flipping of certain chromosomes. As the entire chromosome will be inverted/flipped, the positions for a subset of the chromosomes will change but this chromosome orientation will ensure consistent nomenclature of current and future A. sativa assemblies.
A new convention for numbering chromosomes in oat was accepted by the International Oat Nomenclature Committee (https://protect-au.mimecast.com/s/XLulC2xMQzip25rzMcnoWMY?domain=wheat.pw.usda.gov). The convention is based on an in-depth phylogeny-based analysis that established the presence of conserved core regions with collinearity across Avena, barley and wheat, using a phylogeny-based numbering and orientation system in which hexaploid, tetraploid, and diploid Avena chromosomes are numbered and oriented based on an ancestral core region that is consistent with the Triticeae. In the remainder of this manuscript, the IONC accepted chromosome nomenclature is provided for any locus with a published genetic location using previous systems, using the format [chr. xx].
The genetics and durability of resistance to rust in cereals
Genetic analysis of disease resistance in plants began when Biffen (1905) demonstrated Mendelian inheritance of stripe rust resistance in wheat. Since then, many genes conferring resistance in plants have been characterised, and the inheritance of pathogenicity (virulence/ avirulence) has been investigated in many plant pathogens including several that cause rust in cereals. Based on studies of the genetic basis of the interaction between flax and the flax rust pathogen, Flor (1955) developed the gene-for-gene hypothesis, which has provided an important framework for much of the genetic work on plant: pathogen interactions since.
Resistance to many plant pathogens has been defined on the basis of genetics or function. In genetic terms, it is generally defined on the basis of inheritance: oligogenic, being controlled by one or few genes of major effect; polygenic, being controlled by many genes of individually small phenotypic effect. In functional terms, resistance is defined based on epidemiological (e.g. slow rusting, rate reducing) or developmental (e.g. all-stage resistance (ASR), also referred to as seeding or major gene resistance versus adult plant resistance (APR), also often referred to as minor gene resistance) criteria.
Most of the rust resistance genes in cereals that have been catalogued confer ASR. The process of identifying and designating rust resistance loci in cereals involves initially showing that a given resistance is distinct from previously catalogued resistances based on rust isolate specificity and/or genomic location, and then showing that the new resistance is either a locus independent of or allelic to previously catalogued alleles prior to a gene designation being assigned (see McIntosh et al. 1995). A critical part of this process is the nomination of a reference genetic stock carrying the gene, which can be accessed by the international scientific community for further study and use in resistance breeding. In studies where more than one resistance gene is found in a specific genotype, single gene progeny should be generated for each and identified as single gene reference stocks.
The concept of durable resistance was introduced in 1978 by Dr Roy Johnson, following the failure of resistance to stripe rust in some English winter wheats and the continued effectiveness of resistance in others (Johnson 1978). Genetic studies since then have dissected durable resistance to rust in several wheat genotypes, and genes conferring either ASR or APR to rust are being isolated from wheat and barley genomes. These studies have shown a greater tendency for lack of durability in ASR genes encoding nucleotide-binding and leucine-rich repeat (NLR) receptor proteins compared to other non-NLR-encoding resistance genes. Of the 17 ASR NLR-encoding rust resistance genes isolated from wheat (Zhang et al. 2020) and two isolated from barley (Dracatos et al. 2019; Chen et al. 2021) to date, only the ASR stem rust resistance gene Sr26 (Zhang et al. 2021) has been deployed widely and remained effective, being used in Australian agriculture in more than 30 wheat cultivars since it was first deployed in the cultivar Eagle in 1969 (Park 2007).
Two loci, Lr34/Yr18/Sr57 (Krattinger et al. 2009), and Lr67/Yr46/Sr55 (Moore et al. 2015), confer APR and encode an ATP-binding cassette transporter, and a protein that has lost hexose transport function, respectively. The loci Lr34/Yr18/Sr57 and Lr67/Yr46/Sr55, along with Lr46/Yr29/Sr58, confer resistance against all three rust pathogens, and are associated with a linked morphological trait known as leaf tip necrosis. Experience to date suggests that these pleiotropic APR genes have some intrinsic durability, and that combinations of multiple effective resistance genes contribute to durability by lowering the chance of virulence matching gene combinations developing. Pleiotropic APR genes have been used to great effect in conjunction with ASR genes, providing backbone resistance that may also contribute to increasing the durability of ASR genes (Park 2015).
Resistance to Puccinia coronata f. sp. avenae in Avena
When catalogued, genes conferring resistance to Pca in oat are given the designation “Pc”. Although Simons et al. (1978) used the designation “Pc-”, most publications since have used simply “Pc”. At this time, 92 loci that confer resistance to Pca in Avena have been designated and accepted (Pc1-Pc85, Pc91-Pc96, Pc98). Six loci are reported as conferring APR (Pc27, Pc28, Pc69, Pc72, Pc73, Pc74) and the remaining 86 conferring ASR. Multiple alleles at five loci have been reported: Pc2 (Pc2, Pc2b), Pc3 (Pc3, Pc3c), Pc4 (Pc4, Pc4c), Pc6 (Pc6, Pc6c, Pc6d) and Pc9 (Pc9 and Pc9c). Four pairs of complementary genes have been designated (Pc3 + Pc4, Pc3c + Pc4c, Pc7 + Pc8, Pc24 + Pc25; Tables 1 and 2). The Pc genes designated to date have been found in cultivated and wild hexaploid oat, A. sativa and A. sterilis, respectively, and other related AACCDD sub-species, A. strigosa (AA, diploid), A. abyssinica (AABB, tetraploid), and A. magna (CCDD, tetraploid). A. sterilis has proven a particularly rich source of resistance, contributing some 44 of the resistance loci catalogued to date. This may be in part because it is a sexually compatible hexaploid, making it far easier to transfer this resistance into cultivated oat.
In addition to designated genes, J Chong (unpublished) gave temporary designations to eight resistance genes based on rust isolate specificity and preliminary genetic analyses (viz. Pc97-temp, Pc99-temp through Pc105-temp; Chong J unpublished). Toporowska et al. (2021) mapped a gene present in a selection of the Pc50 differential, given the temporary designation Pc50-5, to chr. 6A. Further work is needed to determine the relationships of all of these genes to those already designated.
The ability to identify and work with catalogued Pc resistance genes depends on the existence of single gene reference stocks for each, and isolates of Pca that carry matching avirulence. Although reference stocks have been nominated for all Pc genes designated so far, 35 are not available in single gene or single resistance specificity (in the case of complementary genes) stocks (Table 1). In these cases it is very difficult or impossible to know which of the genes present in a given stock correlates to which designated Pc gene. Of note in this regard is the diploid A. strigosa accession CI 3815, which was originally reported to carry Pc19 (Simons et al. 1959) and Pc30 (Marshall and Myers 1961). Subsequent studies provided evidence of five tightly linked genes, designated Pc81–85 (Yu and Wise 2000). The genetic relationships between Pc19, Pc30 and the Pc81–85 complex are not known, and the designations Pc81-85 were allocated without considering the potential precedence of Pc19 and Pc30 for at least two of these. Furthermore, unpublished studies in Australia have provided convincing evidence that CI 3815 carries Pc94, meaning also that the designation of Pc94 is possibly incorrect (Park RF unpublished). Gene Pc94 was introgressed into hexaploid oats by Aung et al. (1996) from the diploid A. strigosa (accession RL1697), and in later studies (J Chong unpublished), no segregation in crown rust response was observed in over 800 F2 plants derived from an intercross between RL1697 and CI 3815. The gene Pc92, present in the hexaploid lines Obee (Rothman 1984) and Obee/Midsouth (Rooney et al. 1994), is thought to have come from one of three autotetraploid A. strigosa accessions that included colchicine doubled Saia (Pc15, Pc16, Pc17) and CI 3815. Based on multipathotype tests (Park RF unpublished), gene Pc92 is distinct from Pc15, Pc16 and Pc17 but may be present in CI 3815. The absence of single gene reference stocks for the seven genes designated from CI 3815 makes it very difficult to resolve the genetic basis of resistance to crown rust in this accession and to verify which Pc gene is which. A sequencing and mutational genetics approach may resolve the situation in future.
Single gene stocks have been nominated for the remaining 59 genes (this includes the complementary gene pair Pc3+Pc4 in Bond), of which 12, two and three are available in NILs in the backgrounds of Pendek, Fraser and Makuru, respectively (Table 2). The genes Pc14, Pc36, Pc50, Pc51, Pc52, Pc53, Pc57, Pc70, Pc71 and several other unknown crown rust resistance genes were also backcrossed into one or more of four backgrounds (CI 7555, CI 7970, CI 8044, Lang) by Browning and colleagues at Iowa Agriculture and Home Economics Experimental Station and USDA-ARS as part of a program to develop oat multiline cultivars (Supplementary Table S1). All of these NIL stocks have between two to six backcrosses to the recurrent parent. Because even reputedly single gene stocks may carry additional resistance genes, any assessment of virulence using these NIL stocks must always include the recurrent parent to exclude the presence of additional genes in the background.
A further complicating factor with the designation of Pc genes is that the allelic relationships between many are unknown. Evidence has been obtained of tight linkage or allelism between: Pc35 and Pc54 (McKenzie and Martens pers. comm. to Simons et al. (1978)); Pc39 and Pc55; Pc38, Pc62 and Pc63; Pc44, Pc46, Pc50, Pc68, Pc95 and PcX; Pc45 and Pc53. The latter four linkage groups are discussed in greater detail in the following section. About 60 or so of the designated Pc ASR genes have been shown to confer different specificities using Australian isolates of Pca (Park RF unpublished) and hence represent distinct resistance genes/ alleles.
Most crown rust resistance genes are dominant (e.g. Pc68; Wong et al. 1983), some are incompletely dominant (e.g. Pc56, Pc64, Pc65 and Pc66), and some recessive (e.g. Pc54 and Pc55). The action of some genes may be influenced by genetic background. For example, Pc55 was recessive in crosses with Pendek (Kiehn et al. 1976), but dominant in crosses with Algerian, TAM-O-312 and a number of Pendek isolines carrying other genes (Brouwer 1983). Resistances due to complementary gene pairs were identified in cultivars Bond (Pc3 and Pc4, Baker and Upadhyaya 1966), Santa Fe (Pc7 and Pc8, Osler and Hayes 1953), Garry (Pc24 and Pc25, Upadhyaya and Baker 1960) and Ukraine (Pc3c and Pc4c, Upadhyaya and Baker 1962). Cruz et al. (2001) identified one dominant resistance gene in UFRGS 881,920 and two complementary resistance genes to crown rust in UFRGS 15.
Mapped catalogued ASR genes
Few of the 92 catalogued loci conferring ASR to Pca have been mapped to a chromosome. Some of these studies were only able to assign the genes to linkage groups that did not provide an indication of physical location in the oat genome. Mapped catalogued ASR genes were identified using a number of different molecular methods including: SCAR (Sequence Characterised Amplified region) markers adapted from probes developed for other cereals; RFLP (Restriction Fragment Length Polymorphism) makers based on short fragments of digested DNA; RAPD (Random Amplified Polymorphic DNA) fragments from PCR amplification of random segments of genomic DNA with single primer of arbitrary nucleotide sequences; AFLP (amplified fragment length polymorphism) based on the detection of genomic restriction fragments by PCR amplification; DArT (Diversity Arrays Technology) marker microarray hybridizations based on restriction enzymes; and cDNA-derived single-nucleotide polymorphism (SNP) markers. The genes for which mapping and/or marker development have been undertaken are detailed as follows:
Pc38, Pc62 and Pc63 Harder et al. (1980) conducted genetic analyses of these three genes and found that they were either linked or allelic. Wight et al. (2004) mapped Pc38 to a region homologous to KO (Kanota/ Ogle) group 17 (Mrg02, [chr. 7D]) in two F2:3 populations.
Pc39, Pc55, and Pc71 Kiehn et al. (1976) found that Pc39 and Pc55 are either very closely linked or allelic. Based on very similar responses of differentials carrying Pc39, Pc55, or Pc71 to 130 isolates of Pca collected off A. sterilis and A. barbata in Israel and 683 isolates from A. sativa and 73 aecial collections from buckthorn in the USA, Leonard et al. (2005) concluded that these genes are either identical or allelic. Chong and Seaman (1989) similarly reported complete association between virulence for Pc39 and Pc55 in Canada. This is consistent with results from surveys of virulence in Pca in Australia, where identical responses were documented for differentials carrying each of the three resistance genes in tests of over 1,400 isolates of Pca collected between 2000 and 2020, and 152 historical isolates dating back to the 1950s (Park RF, unpublished).
Wight et al. (2004) mapped Pc39 to a region homologous to KO group 37. Two more recent studies mapped Pc39 to Mrg11 [chr. 4C]. Sowa and Paczos-Grzeda (2020) mapped Pc39 to Mrg11. Six tightly linked PCR-based SCAR markers were validated on a set of 22 oat cultivars, confirming the presence of Pc39 in 16 of these (e.g. cvv. Celer (Pc39), AC Assiniboia (Pc38, Pc39, Pc68), AC Medallion (Pc38, Pc39, Pc68), Riel (Pc38, Pc39), Stainless (Pc38, Pc39, Pc68, Pc91)), and its absence in three (Bingo, Borys and Dragon). Zhao et al. (2020a) again mapped Pc39 to Mrg11 [chr. 4C]. Kompetitive Allele-Specific PCR assays for two Pc39-linked SNP loci accurately predicted Pc39 in two genotypes, and its absence in 72 diverse oat genotypes.
Bush and Wise (1998) used a high density RFLP map to localise Pc71 to a group of 12 linked RFLP markers within 6 cM either proximal or distal to the gene. All markers were positioned in the centre of linkage group 11 of the Kanota/Ogle map, which Sowa and Paczos-Grzeda (2020) attributed to Mrg21 [chr. 4D]. Esvelt Klos et al. (2017) mapped Pc71 to Mrg05 [chr. 6A]. Should Pc71 be located on chr. 4D or chr. 6A, it cannot be allelic with Pc55. If these genes prove to be genetically independent, they do nonetheless confer the same resistance specificity and are of equal value in resistance breeding. Clearly, more work is needed to resolve the positions and genetic relationships between genes Pc39, Pc55 and Pc71.
Pc45 and Pc53 Kebede et al. (2019) examined the genetic relationship between a tentatively designated crown rust resistance gene PcKM, previously found and mapped in the oat cultivars Kame and AC Morton (Gnanesh et al. 2015) and Pc45. They mapped both resistances across five populations and localised both on Mrg08 [chr. 2D], concluding that they were one and the same. They identified two codominat KASP markers as being the most accurate, predicting the presence of Pc45 and PcKM in two oat genotypes, and their absence in 74 diverse oat genotypes. Esvelt Klos et al. (2017) also mapped PcKM to Mrg08 [chr. 2D].
Admassu-Yimer et al. (2018a) demonstrated close linkage between Pc45 and Pc53 of within 1 cM on Mrg08 [chr. 2D], suggesting they are either tightly linked or allelic.
Pc44, Pc46, Pc50, Pc68, Pc95, and PcX A series of studies that led to the designation of these genes provided evidence that they form a large cluster along with the stem rust resistance genes Pg3 and Pg9. Initial studies by McKenzie and Green (1965) demonstrated repulsion linkage or allelism between Pg3 (gene “E”) and Pg9 (gene “H”), and subsequent studies showed linkage between Pg9 and Pc44 (Martens et al. 1968), Pg9 and PcX (Chong et al. 1994), and Pg3 and Pc95 (Harder et al. 1995). Chong et al. (1994) also established linkage between Pg9, PcX and Pc68, Wong et al. (1983) between Pc68 and Pc46, and Fleischmann et al. (1971) between Pc46 and Pc50, indicating that Pc46, Pc50 and Pc68 are also part of this linkage group. Chong et al. (1994) considered PcX to be either Pc6c or Pc9. Given that both the Pc6 and Pc9 loci comprise multiple alleles (Tables 1 and 2), either would add to the complexity of this resistance gene cluster.
Based on RFLP probes that were previously shown to be closely linked to Pg9, Chen et al. (2006) developed two SNP markers and an STS marker that were linked with Pc68 at recombinational distances of 4.2 cM and 6.7 cM in two populations. Based on the original RFLP markers, Pg9 (and hence Pc68) were located to the Kanota/Ogle linkage group 4_12 (spread across Mrg19 and 23; chr. 3D and chr. 3A, respectively). As found by Chen et al. (2006), Kulchelski et al. (2010) mapped Pc68 to linkage group 4_12 of the Kanota/Ogle linkage map. The GWAS study of Esvelt Klos et al. (2017) confimed the location of Pc68 on Mrg19 [chr. 3D], and also identified a SNP marker that could be suitable for MAS of this gene.
Pc48 Wight et al. (2004) mapped Pc48 to a region corresponding to the KO group 22_44 + 18 based on linakge with five RFLP markers (Mrg20 [chr. 4A]). Interestingly, they found differences in marker linkage in two populations based on the resistant parent Pendek-48. Further genotyping of different plants within Pendek-48 showed that it comprised at least two genotypes carrying different sized introgressions with Pc48 from A. sterilis. GWAS studies by Esvelt Klos et al. (2017) confirmed the location of Pc48 on Mrg20 [chr. 4A].
Pc54 Admassu-Yimer et al. (2022) mapped Pc54 to Mrg02 [chr. 7D] in two biparental RIL populations. The location of 60–84 cM on Mrg02 was considered by Adamssu-Yimer et al. (2022) as possibly overlapping with that reported for Pc38 by Wight et al. (2004).
Pc58 Hoffman et al. (2006) demonstrated the presence of three ASR genes in TAM-O-301 in crown rust tests of an Ogle/TAM-O-301 F6:7 RIL population. Because the resistance in TAM-O-301 had been previously designated Pc58, the authors designated these genes Pc58a, Pc58b, Pc58c. In wheat, such designations would indicate allelism. The three loci were however mapped to a region spanning 41.0 cM as defined by RFLP markers, with gene Pc58a and Pc58c being closely linked and Pc58b being independent of these two loci. Overall, the three genes were mapped to two previously described linkage groups, OT32 and OT33, both of which correspond to KO linkage group 17 (hence Mrg02 [chr. 7D]). The chr. 7D location was confirmed in the GWAS analysis by Esvelt Klos et al. (2017).
Jackson et al. (2008) used the Ogle/TAM-O-301 F6:7 RIL population to map ASR to two pathotypes of Pca that were virulent on Ogle but avirulent on TAM-O-301. They mapped a single gene that conferred a “bleached fleck” phenotype, different to that conferred by Pc58, to both pathotypes to the linkage group OT-11 (homologous to KO_4). Detailed measurements of uredinial length (UL) identified major QTL on linkage groups OT-11 and OT-32. The latter was mapped very closely (0.3 cM) to the previously determined location for Pc58a. The same pathotypes were used to phenotype the population in the field at adult plant growth stages for infection type, UL and relative fungal DNA (FDNA). Measurements of FDNA in both greenhouse and field experiments similarly mapped OT-32 very close to Pc58a. It was concluded that the reductions in UL and FDNA were due to either Pc58a or genes tightly linked to this locus. Three minor QTL were also detected, on linkage groups OT-8 and OT-15 (from TAM-O-301) and OT-27 (from Ogle).
Jackson et al. (2007) also investigated resistance to Pca in Ogle in the Ogle/TAM-O-301 F6:7 RIL population using an isolate of Pca that was avirulent on Ogle and virulent on TAM-O-301. Based on greenhouse assessments and artificially inoculated field plots, a single gene for resistance derived from Ogle was mapped to linkage group OT6 (homologous to linkage group KO). A previous study by Bush and Wise (1996) mapped two unlinked resistance genes derived from Ogle to the linkage groups KO_4 and KO_13. Based on pedigree analysis, Jackson et al. (2007) concluded that the gene detected in their study was likely derived from either Landhafer (Pc4, Pc5) or Victoria (Pc2, Pc11, Pc12).
Pc91 Using aneuploid stocks, Rooney et al. (1994) localised Pc91 to chromosome 18. The gene was further mapped by McCartney et al. (2011), who developed five robust PCR-based SCAR markers that were validated on 23 North American oat cultivars/ breeding lines to confirm its presence in cultivars HiFi, Souris and Stainless carry Pc91, and absence in Ogle, Leggett, Ronald CDC Sol-Fi, CDC Minstrel, CDC Dancer and CDC Weaver. Aligning the linkage map generated with the Kanota/Ogle linkage maps indicated that Pc91 was likely located on either chromosome 7C [chr. 1C] or 17A [chr. 1A].
In a subsequent study, Gnanesh et al. (2013) developed three KASP assays that accurately predicted the presence/ absence of Pc91. Further mapping located Pc91 to the intergenomic translocation involving chromosomes 7C and 17A that is present in most spring oat genotypes.
Esvelt Klos et al. (2017) mapped Pc91 to Mrg18 [chr 1A] and this was further validated by Maughan et al. (2019)
Pc92 This gene was identified by Rooney et al. (1994) in the hexaploid oat breeding line Obee/Midsouth. Using backcross-derived lines, RFLP markers linked to Pc92 were identified. The locus was then mapped using segregating F2 populations, and an attempt to determine chromosomal location using aneuploid stocks was unsuccessful presumably because of the lack of the corresponding aneuploid stock.
Pc94 Chong et al. (2004) and Chen et al. (2007) undertook genetic studies of Pc94 and although they did not map the gene they did develop linked markers for use in resistance breeding.
Pc98 Zhao et al. (2020b) mapped Pc98 in two F2:3 populations based on genotypic data generated with the 6 K Infinium iSelect SNP array, and using the consensus map developed by Chaffin et al. (2016) and Bekele et al. (2018), mapped Pc98 to Mrg20 [chr. 4A].
To further identify probable differences from the current consensus map that may be attributable to segregating translocations and to map genes or quantitative trait loci (QTL) affecting oat crown rust resistance, Sunstrum et al. (2019) developed a de novo genetic linkage map in a population based on a breeding line from southern USA (TX07CS-1948) and a Canadian spring oat line (SA04213) using high‐density GBS markers. This linkage map served as a useful tool in detecting four QTL associated with resistance to Pca, of which two (on linkage groups Mrg02 and Mrg19) corresponded with QTL detected in a previous GWAS by Esvelt Klos et al. (2017). These QTL can be reconciled to known Pc genes loci or resistance gene clusters including Pc38 (Wight et al. 2004) and Pc58a (Hoffman et al. 2006; Jackson et al. 2007) on Mrg02, and Pc68 (Kulcheski et al. 2010) on Mrg19. The collation of marker details can be used as a tool to assist in reconciling the most probable location for a given gene. Marker sequences of probes associated with the mapped catalogued ASR genes detailed above were retrieved from the GrainGenes database (wheat.pw.usda.gov; Blake et al. (2019)) and the inbuilt Basic Local Alignment Search Tool (BLAST) used to assign physical positions in chromosomes of the PepsiCo OT3098 Hexaploid Oat v2 pseudomolecules (2021) (Table 3 as Electronic supplementary material). Although the genome assembly is advanced, it is still being resolved and functional homologs have been detected. Due to the short read length of probes reported in previous studies and the highly repetitive nature of the oat genomes, the presence of multiple complementary sequences with reported markers is expected. Therefore, the top three the BLAST hits with E values below 10–5 are presented for each marker. Descriptions of the markers linked to Pc genes, mapping populations and sequence accession are also presented in Table 3 (as Electronic supplementary material).
Mapping studies of uncharacterised “field” resistance to Pca at adult plant growth stages
QTL mapping and GWAS approaches have been used to map resistance to rust in wheat, barley and oat based on rust response data from field plots. The resistance observed in many of these studies is referred to as “field” resistance as it is effective at adult plant growth stages under field conditions. In most cases, the rust pathotype(s) present in the field plots were however unknown, meaning that it is impossible to attribute the field resistance documented to either ASR, APR, or both.
The crown rust resistance of the hexaploid line MAM17-5, believed to carry two dominant resistance genes to Pca derived from one or both of two A. strigosa accessions (viz. CI 3436 and CI 3820), was studied in an F5:6 RIL population based on the cross MAM17-5/ Ogle (Zhu and Kaeppler 2003). The population was assessed under field conditions in Wisconsin across two years, using natural and artificial inoculation of a composite mixture of Pca pathotypes common to the Midwest of the USA at the time. Two QTL, referred to by the authors as “field” resistance, were detected and mapped to linkage groups but with no reference to other consensus maps. In subsequent studies of the same population, Zhu et al. (2003) mapped ASR resistance to three North American pathotypes of Pca. Combined, these two studies confirmed the presence of two QTL, one of which (Pcq1) mainly controlled field resistance and one of which (Pcq2) mainly controlled greenhouse ASR. Given that the resistance examined in these two studies was derived from A. strigosa accession CI 3436 or CI 3820, and that the ASR gene Pc23 was described from A. strigosa accession CD3820 (Table 2), it could be that at least one of the loci mapped is in fact Pc23.
Barbosa et al. (2006) characterised resistance to Pca in the oat genotype UFRGS910906, which the authors stated carried partial resistance to Pca. UFRGS910906 was crossed to the Pca susceptible UFRGS7 and assessed under field conditions with natural infection at the F2 in 1999 and at the F6 in 2000. Five QTL were detected at the F2, none of which were detected at F6, in which three different QTL were found. The simplest explanation for these results is that different Pca pathotypes prevailed in each year. Given that the identity of the pathotypes present were unknown, and that the ASR status of both parents is unknown, it is impossible to attribute the QTL detected to either ASR or APR.
Jackson et al. (2007) evaluated the ability of three different assessment methods (quantitative real-time q-PCR to estimate fungal growth in the host, digital image analysis, and visual ratings) to map crown rust resistance in cvv. Ogle and TAM O-301. A major gene in Ogle was identified and mapped by all three assessment methods to linkage group OT6 although the resolution produced by q-PCR enabled more precise mapping. Data generated by the latter method permitted identification of QTL on linkage groups OT32 and OT2.
The field resistances of CDC Boyer and the oat breeding line 9417A1-9–2-2–2-5 were studied by Babiker et al. (2015) in three F6:8 RIL populations across two field sites (Louisiana State University Baton Rouge (2 years) and the Cereal Disease Laboratory (CDL) St Paul, MN (3 years)). The authors refer to the resistance assessed as “partial” resistance, but it is impossible to know if it was ASR or APR in the absence of information of the pathotypes present. Four QTLs were detected: three from CDC Boyer (two on chromosome 19A [chr. 4A] and one on 12D [chr. 2D]), and one from 9417A1-9–2-2–2-5 (on 13A [chr. 7A]). Only one of these, Qcr.cdl9lsu9-19A, was detected at more than one field site. To date, two ASR genes have been mapped to chr. 4A (viz. Pc48 and Pc98), and two to chr. 2D (Pc45 and Pc53). In the absence of information regarding the virulence of the pathotypes present in the nurseries, it is not possible to discount one or more of these gene as contributing to the resistances identified.
Montilla-Bascón et al. (2015) used a GWAS approach to map ASR resistance in 177 oat accessions to a single pathotype of Pca with broad virulence collected from Spain. Positive associations with five markers were found, two of which were located on KO linkage groups 32 [chr. 5C] and 17 [chr. 7D]. The identities of these resistances are unknown; Pc38 and Pc58 have been mapped to chr. 7D, however the genes detected cannot be these because the pathotype used was virulent for both.
A set of 1,000 A. sativa genotypes collected prior to 1930 and landraces collected in the 19th and 20th Centuries was assessed for crown rust response in the CDL St Paul Buckthorn nursery in 1994 and 1997 (Winkler et al. 2016). The response data were not consistent across the years, but singly provided evidence of associations with five linkage groups, two of which could be associated with catalogued Pc genes of known map location (viz. Mrg08 (possibly PcKM/Pc45), Mrg28 (possibly Pc91), Mrg20 (possibly Pc48)), and two on Mrg23 and Mrg01 that were of unknown identity. All loci had closely linked SNPs.
Esvelt Klos et al. (2017) used a GWAS approach to map ASR and field resistance to Pca in a set of 631 oat lines representing modern elite germplasm and 31 Pca differential genotypes. All were genotyped using the Illumina oat iSelect 6 K beadchip array, tested as seedlings for response to 10 pathotypes of Pca, and field tested as adult plants in 10 location/ years in the USA and Canada across six field sites (2010 and 2011: Castroville Texas, Fargo North Dakota, St Paul Minnesota (in the CDL buckthorn nursery), Winnipeg Manitoba; 2011 only: Baton Rouge Louisiana, Ottawa Ontario). A total of 29 SNPs in 12 linkage groups were predictive of crown rust response across seedling and adult plant tests. The authors conceded that interpreting these associations with respect to the type of resistance (ASR vs APR) and identity with respect to previously designated Pc genes was not simple. Analyses of the 31 Pca differential genotypes did not provide useful information in relating the QTL detected in the 631 oat lines to catalogued Pc genes. Ten SNPs were primarily associated with seedling response and were regarded as ASR genes that are no longer effective in the field in North America due to matching virulence. Six SNPs were exclusively associated with adult plant field response to crown rust; however, none were detected at all year/ locations: one field site (1 QTL); two year/locations (3 QTL); five year/ locations (1 QTL on Mrg18 at 68 cM [chr. 1A]); 8 year/ locations (1 QTL on Mrg06 at 74 cM [chr. 5D]). The remaining 13 SNPs were associated with both seeding and adult plant response, and it was suggested that these could represent ASR genes that were effective at some field sites in some years.
Genetic studies of adult plant resistance to Pca in oat
Much less is known about the genetic basis of APR to rust pathogens in cereals compared to ASR. For example, while it is known that many Australian wheat cultivars carry APR to stripe rust, only two of the genes responsible for this can be reliably identified (viz. Yr18 and Yr29; Park et al. 2020). The principal reason behind this is that APR is technically more difficult to work with than ASR: phenotyping must be undertaken on adult plants and is only reliable if it is conducted in field nurseries that have been artificially inoculated with well-defined rust pathotypes that are virulent for any ASR genes present in the material being assessed. Added complications are that the resistance conferred by APR is often intermediate in effect, the onset of expression of different APR varies with growth stage, and the level of expression can be affected by inoculum load and environment.
Catalogued genes conferring APR
Of the 92 loci conferring resistance to Pca in Avena that have been catalogued, six confer APR. The chromosomal locations of these genes are unknown.
Studies conducted at the University of Sydney by Upadhyaya and Baker (1960) found that the oat cultivars Victoria and Garry (a Victoria derivative) carried the same resistance genes. Two complementary genes conferred seedling resistance to Pca (Pc24 + Pc25), but these genes were not effective at adult plant growth stages. APR to Pca in Garry was attributed to two genes that were designated Pc27 and Pc28. A third gene conferring ASR in Garry, Pc26, was later reported by Upadhyaya and Baker (1962). Field tests of the adult plant crown rust response of Garry at the Plant Breeding Institute Cobbitty have shown that it is quite resistant (10MR), despite being susceptible in greenhouse seedling tests (Roake J and Park RF, unpublished). Further genetic studies are required to determine the inheritance of this resistance.
Harder et al. (1984) reported APR to crown rust in A. sterilis accession CAV 1387. The inheritance of the resistance was studied by crossing CAV 1387 with the A. sativa cultivar Fraser, and shown to be governed by a partially dominant gene designated Pc69. Further rust testing of a putative single gene line derived from CAV 1387 showed heterogeneity among plants at both seedling and adult plant growth stages, and did not detect APR (Chong J, unpublished).
Three genes from A. sterilis were reported by Simons and Michel (personal communication to https://www.ars.usda.gov/midwest-area/stpaul/cereal-disease-lab/docs/cereal-rusts/oat-crown-rust/unpublished) as conferring APR to crown rust: Pc72 from A. sterilis accession PI 298,129. Pc73 and Pc74 from A. sterilis accession PI 309,560, tester line being IA H592. The level of protection afforded by these resistances has not been assessed critically; accessions believed to carry Pc72 or Pc73 were completely susceptible in the field in Poland (Paczos-Grzeda E and Sowa S, unpublished), and line IA H611-447 (Pc72) was rated as 5S under field conditions in Australia (Roake J and Park RF, unpublished).
Uncatalogued genes conferring APR
Several oat cultivars or accessions have been reported to carry uncharacterised APR to crown rust; however, the genetic basis of the resistance in these is largely unknown. APR to crown rust was reported in the cultivars Santa Fe and Ukraine by Upadhyaya and Baker (1965); however, the genetic basis of these resistances was not studied and it is unknown if it is distinct from the APR in Garry reported previously by these authors. Both cutivars also carry designated ASR genes (Pc6, Pc7, Pc8 and Pc21 in Santa Fe; Pc3c, Pc4c, Pc6c and Pc9 in Ukraine). Red Rustproof oats were introduced into southern USA in the mid 1800s, from which improved “strains” were selected (Coffman et al. 1961). Luke et al. (1972) found that the slow rusting of Red Rustproof 14 (CI 4876), a selection made from Red Rustproof around 1865, was stable over an 11 year period, and later showed high broad-sense heritability and was governed by an estimated 2.16 genes (Luke et al. 1975). The cultivars Ajax, Beaver and Erban (Welsh et al. 1953), and Lodi, Portage and Rodney (Heagle and Moore 1970), were also reported to carry uncharacterised APR to Pca. By using specific pathotypes of Pca in greenhouse seedling and adult plant field tests, Simons (1961) identified three “strains” with high levels of APR (viz. PI 174,545, PI 185,783, and PI 197,278). While all three have been assessed under Australian conditions, only one has shown good levels of APR (PI 174,545) (Roake J and Park RF, unpublished). In a subsequent study, Simons (1975) undertook genetic studies of the APR in two of these lines (viz. PI 174,545 and PI 185,783) and two others that were found in the 1961 study to have low levels of APR (viz. PI 174,544 and PI 19,727), and found continuous variation in response that was suggestive of polygenic inheritance.
Briére et al. (1994) assessed the adult plant field responses of 33 oat genotypes over two years. The plots were inoculated with a pathotype of Pca that developed sporulating pustules on all genotypes under controlled environmental conditions. Two breeding lines (OA 712-17 and OA 712-33) and four cultivars (Glen, Woodstock, Sylvya, Fidler) had high levels of what the authors termed partial resistance. Genetic studies have shown that both Fidler and Woodstock carry Pc39 (Chong J, unpublished), and it is unclear whether this ASR gene was effective or not in the studies by Briére et al. (1994).
As mentioned previously, Jackson et al. (2008) mapped resistance to Pca in TAM-O-301 based on an F6:7 RIL population (Ogle/TAM-O-301). Seedling tests with two pathotypes of Pca identified two ASR genes, one of which was considered to be Pc58 (“Pc58a”). Field tests at adult plant growth stages using the same two pathotypes allowed the identification of two minor QTL from TAM-O-301 and one from Ogle.
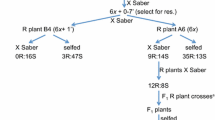
The USDA Cereal Disease Laboratory (CDL) at St Paul Minnesota established a buckthorn (Rhamnus carthartica) nursery in 1953. Named after the pathologist who established it, the Matt Moore Buckthorn Nursery comprises hedges of R. carthartica between which plots of oat can be sown and assessed for response to a multitude of pathotypes of Pca arising from sexual recombination on its aecial host. The nursery was used by Stuthman and Moore (1970) and coworkers to select crown rust resistant lines from breeding populations based on crosses designed to combine different types of resistance from diverse sources. Leonard (2002) undertook detailed greenhouse (seedling) and field (adult plant) studies of the crown rust resistance in 14 of the lines produced by Stuthman and Moore that were resistant in the buckthorn nursery in annual tests over an 8 year period. The cultivar Portage, known to carry moderate levels of slow rusting to Pca, was included as a control. Portage also demonstrates slow rusting under Australian conditions (Roake J and Park RF, unpublished). All lines were resistant to crown rust across field tests over seven years, many with resistance superior to that of Portage. Although greenhouse studies indicated that at least some of the resistance observed was likely due to ASR, four lines were identifed with APR, two of which had levels of APR adequate enough to protect against Pca in the absence of ASR (viz. MN841801 (pedigree: Florad/Coker 58–7/3/CI7558//Black Mesdag/Aberdeen 101 [65B663/65B1362]; MN841804 (pedigree: CI7683//Black Mesdag/Aberdeen 101/3/Rodney/4/CI7558//Black Mesdag/Aberdeen 101 [65B1286/65B1362]). Given the similarity in the pedigrees of these lines, there may be some commonality in the genes underlying the APR they carry. Both lines have displayed good levels of APR to crown rust under Australian field conditions in artificially inoculated nurseries over at least 5 years (Roake J and Park RF unpublished). Several studies have been conducted to investigate the genetic basis of APR in MN841801, these are described below.
Chong (2000) conducted crown rust tests of adult plants of an F7:9 RIL population based on MN841801 in a growth chamber to show that this line carried two complementary (or additive) genes for APR. Lin et al. (2014) phenotyped the same RIL population and two others based on MN841801 for adult plant response to crown rust in the field over nine site/years. All plots were inoculated with the same pathotype used previously in the growth chamber study (Chong 2000). A major effect QTL (RMR to MR/MS), QPc.crc-14D, was detected in all three populations in all environments. QPc.crc-14D was mapped to chromosome 14 D [chr. 5D]. Two KASP markers were identified that provided strong evidence for the presence of QPc.crc-14D in three oat genotypes (viz. 02P07-BC1A, W020203 and Clav 8361). Comparative mapping suggested that Qpc.crc-14D is homologous to the wheat stripe rust resistance gene Yr16, which was mapped in two independent studies to wheat chromosome 2D (Worland and Law 1986; Agenbeg et al. 2012). Additional seedling tests of the RILs of the three populations with pathotypes virulent on AC Assiniboia, AC Medallion and Makuru showed the MN841801 parent was heterogeneous for ASR, carrrying either two genes or one gene depending on the RIL population used (J. Chong and J. Lin, unpublished).
Portyanko et al. (2005) tested an F6:8 RIL population based on the crown rust susceptible genotype Noble-2 and MN841801-1 in the field twice at St Paul Minnesota (1997 and 1998; plots inoculated with a bulk population of Pca) and once at Rosemount Minnesota (1998, natural inoculum), and under greenhouse conditions as adult plants using a Pca pathotype that was virulent on both parents at seedling growth stages. A total of seven QTL were detected, accounting for up to half the variation associated with resistance of MN84801-1, four of which were considered major QTL of which one (Prq2 on linkage group 26) had the most consistent effect. A lack of common markers across different studies made comparisons difficult, but the QTL Prq1b was considered to be associated with Pc38/ Pc62/ Pc63 ([chr. 7D]).
The population developed by Portyanko et al. (2005) was advanced to F6:9 by Acevedo et al. (2010) and phenotyped for adult plant response to Pca over six sites/ years and in the greenhouse. Three field sites and the greenhouse tests used two pathotypes of Pca that were virulent on seedlings of MN841801-1 and Noble-2, one of which was the same pathotype used by Chong (2000) and Lin et al. (2014). Overall, the study detected the same seven QTL as reported by Portyanko et al. (2005), plus an additional one (Prq8) that was detected at two field sites only. It was concluded that four QTL (viz. Prq1a, Prq2, Prq7, Prq8) were responsible for the partial resistance of MN841801-1 to crown rust. As was found by Portyanko et al. (2005), Prq2 had the most consistent effect; this along with Prq1a accounted for 30.1% of the variation associated with the observed resistance to Pca. The region containing QTL Prq1a (linkage group MN3) is homologous to the linkage group KO-17 (Mrg02 [chr. 7D]).
Differences in findings from the above studies of the resistance of MN841801 clearly demonstrate the difficulties of genetic studies of rust resistance, especially APR. The number of genomic regions associated with APR to Pca in these studies varied from one (Lin et al. 2014), to two (Chong 2000), to four (Portyanko et al. 2005; Acevedo et al. 2010). The lack of congruity between the above APR studies could relate to genetic heterogeneity in the original accession of MN841801, pathogen virulence, the different methods (e.g. growth chamber vs field) used to assess crown rust response, and/or environment. Of particular significance among these is pathogen virulence, as using the correct rust isolates is the only way APR can be distinguished from ASR. The MN841801 line used as parent by Chong (2000) and Lin et al. (2014) was shown to be heterogeneous for ASR. The discrepancy in findings between the studies by Chong (2000) and Lin et al. (2014) could be due to the different methods used. The detection of two additive APR genes by Chong (2000) was based on a single inoculation in growth chamber experiments. Rust reactions from a single inoculation would be useful for identification of infection types but would not be useful for detecting resistance that expresses quantitatively over multiple disease cycles in field experiments. Alternatively, it is possible that a second APR QTL was not detected by Lin et al. (2014) because their linkage map was incomplete. Differences in findings between the studies by Lin et al. (2014) and Acevedo et al. (2010) are more difficult to explain because the same pathotype of Pca was used in both studies.
The Matt Moore Buckthorn Nursery was also used by Admassu-Yimer et al. (2018b), who undertook adult plant tests on 607 A. sativa accessions from 44 countries in the nursery and at Baton Rouge Louisiana. A total of 36 genotypes were identified that were resistant at both field sites, and these were tested a second time at both field sites. Seedling tests of the 36 genotypes identified three that were seedling susceptible to all eight pathotypes used. These three genotypes were tested for adult plant response to each of the eight pathotypes in growth cabinets in two experiments, and ranged in response from 5 to 30MR (Clav 3390), from 5MR to 40MS (Clav 2272), and from TrMR to 30MS (PI 285,583), confirming the presence of good levels of APR in all. The genetic basis of the resistance in the lines is unknown as no genetic analyses were undertaken.
All seven designated Pc genes conferring APR to Pca are from the hexaploid species A. sativa (Pc27 and Pc28) and A. sterilis (Pc69, Pc72, Pc73, Pc74). Using controlled inoculations of seedling and adult plants with specific pathotypes, Cabral et al. (2011) identified six diploid A. strigosa accessions (CIav6956, CIav7280, CIav8089, CIav9020, PI292226, PI436082) and one tetraploid A. barbata accession (PI337865) that carried APR to Pca. The APR of three diploid accessions of diverse geographic origins (Canada, Chile, USA) was shown to be governed by single dominant genes, the genetic relationship between which was not investigated. Although being expressed at adult plant growth stages only, the three genes conferred a hypersensitive phenotype typical of many non-durable ASR genes. This could suggest a lack of durability of these APR genes, as was found in wheat with the hypersensitive APR gene Lr12 (Park and McIntosh 1994).
Breeding for resistance to P. coronata f. sp. avenae
Early efforts
Simons (1970, 1985) and Coffman et al. (1961) provided comprehensive summaries of the early use of resistance to control Pca in oat in North America. Two important cultivars in this were Victoria and Bond. The latter was developed in Australia from a cross between the Swedish cultivar Golden Rain and a selection from the genotype Red Algerian (Baker 1966). Baker and Upadhyaya (1966) and Baker (1966) identified two complementary resistance genes in Bond, later designated Pc3 and Pc4. Although virulence to Bond was first detected in Australia in 1948 by Waterhouse (1952), the frequency of virulence remained low for some time after that and the resistance remained useful (Baker and Upadhyaya 1966). Virulence to the Bond resistance became common in the USA along with the adoption of Bond-derived cultivars (e.g. Bonda, Andrew, Camelia; Simons 1970), to the point where by 1953 this resistance was of almost no value at all (Simons 1970). The cultivar Victoria was introduced to North America from South America in 1927. North American studies of the genetic basis of resistance to Pca in Victoria reported the presence of one (Murphy et al. 1937; designated Pc2) or two (Chang and Sadanga 1964; Pc2 and Pc11) resistance genes, of which Pc2 was used in developing many oat cultivars released during the 1940s (e.g. Tama, Vicland, Vikota, Cedar; Coffman et al. 1961), which later succumbed to Victoria blight caused by Helminothosporium victoriae because of the association between Pc2 and susceptiblility to this pathogen caused by the pleiotropic or tightly linked locus Hv-1. The spread of Victoria blight was rapid and losses were estimated in the order of hundreds of millions of dollars (Coffman et al. 1961).
The cultivars Bond and Victoria were the sole sources of resistance to Pca used in Australia prior to 1960 (Upadhyaya and Baker 1960). The development of virulence for both cultivars there and the emergence of Pca race 45 in the USA, which rendered Bond and its derivatives susceptible and caused high losses in southern USA (Coffman et al. 1961), led to searches for new resistance to Pca. Studies of the genetic basis of resistance to Pca in Bond, Victoria, Garry (a Victoria derivative), Landhafer (originating from Germany), Santa Fe (Argentina), Trispernia (Romania) and Ukraine (Russia) (Upadhyaya and Baker 1960, 1962, 1965; Baker and Upadhyaya 1966, Baker 1966) demonstrated considerable genetic diversity in both ASR and APR to Pca. Landhafer, Santa Fe and Trispernia were used as sources of crown rust resistance in North America in the 1950s (e.g. Clintafe, Seminole, Fayette, AB 110 from Santa Fe; Floriland, Sunland, Clintland, Minhafer, Radar 1 and Radar 2, from Landhafer; Moregrain, Suregrain from Trispernia; Coffman et al. 1961). Two varieties based on Landhafer were released in Australia (Minhafer, Stout; Fitzsimmons et al. 1983). Pathogenicity surveys in the USA by Simons and Murphy (1955) detected virulence on Landhafer, Santa Fe and Trispernia. Based on pathogenicity surveys in Australia between 1952 and 1954 in which virulence for the ASR in all of these oat genotypes and Klein 69B (Pc10) was detected, Baker and Upadhyaya (1955) stated that the “situation as regards the future of resistance breeding is not particularly straightforward”, and recommended assessing the value of combining different resistances to reduce the probability of the pathogen acquiring virulence, mutation to induce new resistance specificities, and wide crossing to introgress new resistance.
Alien species as sources of new resistance to crown rust
Of the 92 designated loci conferring resistance to Pca, only 25 come from the hexaploid species A. sativa and A. byzantina, highlighting the importance of other Avena species as sources of resistance (Tables 1 and 2). During the 1960s, attention turned to the wild hexaploid species A. sterilis as a source of resistance to Pca, with most of the genes designated after Pc34 originating from this species (Table 2). Although 19 loci have been designated from the diploid A. strigosa, only three of these, Pc23 (Dyck and Zillinsky 1963), Pc92 (Rooney et al. 1994) and Pc94 (Aung et al. 1996) are available in stable hexaploid lines. The breeding line WIX4361-9 (“WIX”) was shown by Bonnett (1996) to carry two resistance genes. Based on rust isolate specificity and pedigree (WIX has the pedigree N569-42–51/Froker/2/Froker/WIX1274-2, with N569-42–51 being an accession of A. strigosa), one of these two genes is likely Pc15, Pc16 or Pc17 derived from Saia, and the second Pc39 (Park RF unpublished). Gene Pc91 originated from the tetraploid species A. magna (Rooney et al. 1994). A. magna accession CI 8330 was crossed by the diploid A. longiglumis (accession CW 57), and after treating with colchicine a single F1 seed was obtained from which the synthetic hexaploid line Amagalon was generated. The A. magna accession CI 8330 is also listed as the source of a second gene, Pc93, (https://www.ars.usda.gov/midwest-area/stpaul/cereal-disease-lab/docs/resistance-genes/resistance-genes/), however no further details are available concerning this resistance gene.
Deployment of Pc genes and the emergence of matching virulence
The crown rust resistances of Victoria, Bond, Landhafer, Santa Fe, and Trispernia provided transient, short termed resistance only in both North America and Australia, with the development of matching virulence in Pca following soon after cultivar release and often significant losses resulting. The deployment of many different ASR genes since then has similarly met with the rapid appearance of matching virulence in Pca populations; Carson (2009) for example reported that oat cultivars with major gene resistance to crown rust in the USA were generally rendered susceptible by the emergence of matching virulence in Pca five years or less after release, and similar observations have been made in Canada and in Australia. At least 10 resistance genes derived from A. sterilis have been deployed in pure line oat cultivars in Australia, Canada, USA, and elsewhere (viz. Pc38, Pc39, Pc48, Pc50, Pc56, Pc58, Pc59, Pc61, Pc62, Pc68; Supplementary Table S1), and in all cases matching virulence was detected very soon following deployment.
The emergence of virulences for genes Pc50, Pc68, and Pc91 in Australia provide well documented examples of oat cultivars that were resistant to crown rust at the time of deployment but succumbed soon thereafter. All three genes were deployed only in northern regions of the eastern Australian cereal belt. The dramatic selective force exerted on populations of Pca there by cultivars carrying these genes is apparent from the rapid appearance of virulence for each gene, the rapid increase in virulence frequency following gene deployment, and the lack of detectable virulence for each gene in Western Australia where they have not yet been deployed. Gene Pc68 was first deployed in eastern Australia in 1998 in the forage oat cultivars Graza 68 (syn. AC Assiniboia) and Moola (syn. Dumont 68), both of which likely carry Pc38 and Pc39 as well as Pc68 (Supplementary Table S1). Monitoring for virulence on Pc68 began in 1993 (Bonnett 1996), and virulence was first detected in 1999 only a year after the gene was first deployed, in a pathotype that combined virulence for Pc38, Pc39 and Pc68 (Park RF, unpublished). Since then, more than 50 pathotypes virulent for Pc68 have been detected, all of which are also virulent on Pc38 and Pc39. Monitoring for virulence on Pc50 began in 1977, and between then and 2007 only a single isolate of one pathotype of Pca was detected that was virulent on this gene (Brouwer 1983; Bonnett 1996; Park RF unpublished). Gene Pc50 was first deployed in Australia in the cultivar Volta, a forage oat released in the east in 2003 (Supplementary Table S1). In 2008, two pathotypes with virulence for Pc50 were detected, with six additional Pc50-virulent pathotypes being detected up to 2020 (Park RF unpublished). Monitoring for virulence on Pc91 began in 1993 (Bonnett 1996), the gene was first deployed in the forage oat cultivar Drover in 2006, and viruence was first detected in 2010 (Park, 2013). Two years later, two further virulent pathotypes had been detected and approximately 35% of isolates analysed from eastern Australia were virulent for Pc91, and by 2020 a further seven pathotypes virulent for Pc91 had been detected (Park RF unpublished). All pathotypes detected virulent on Pc91 were also found to be virulent on Pc38, Pc39 and Pc68, and one combined virulence on all of these genes with Pc50, showing the tremendous evolutionary potential of this asexually reproducing pathogen population. At least 10 other oat cultivars with known or unknown seedling resistance genes to Pca were developed and released in Australia from 1991 to 2020 (Amby, Barcoo, Bettong, Cleanleaf, Comet, Culgoa, Gwydir, Nobby, Taipan, Warrego) that were regarded as resistant to crown rust at the time of release but succumbed to the emergence of pathotypes with matching virulence soon after. Multipathotype testing with an array of pathotypes of Pca including pathotypes virulent on these cultivars has supported the presence of resistances based on genes Pc38, Pc39, Pc48, Pc52, Pc56, Pc59 or Pc61 singly or in various combinations (Supplementary Table S1). Of interest, genes Pc38, Pc39, Pc50, Pc52, Pc56, Pc68 and Pc91 all appear to have been deployed in various combinations (e.g. cultivars Cleanleaf, Drover, Graza 68, Riel, Volta) and genes Pc48, Pc59, Pc60 and Pc61 separately in other combinations (e.g. cultivars Amby, Barcoo, Bettong, Nobby). A similar situation was reported for the genes Pc58, Pc59, Pc60 and Pc61 in South America by Leonard and Martinelli (2005), who stated that these genes are most likely to have been introduced into South American oat cultivars from germplasm derived from Texas A&M University and the Coker Seed Company that was distributed in the Quaker Oat International Nursery. In Australia, deploying these two groups of genes separately was paralleled by the emergence of two dominant groups of many pathotypes that are characterised by combinations of virulence for each (Park RF unpublished).
Multiline cultivars
Multiline cultivars are “re-constitutable composites of phenotypically similar, genetically dissimilar lines” (Jensen 1965). By increasing crop genetic diversity and ensuring that mixture components are relevant to prevailing pathogen populations, such mixtures are believed to buffer against abiotic and biotic stresses (Mundt 2002). Browning and colleagues at Iowa Agriculture and Home Economics Experimental Station and USDA-ARS applied the concept of multiline cultivars to crown rust control in oats by backcrossing different Pc genes into a common genetic background to produce component isolines. Resulting BC6 (backcross) fixed lines were mixed to produce a cultivar comprised of between 7 and 12 agronomically similar isolines with different crown rust resistance derived from either A. sativa (Pc5, Pc14) or A. sterilis (Pc36, Pc46, Pc51, Pc52, Pc53, Pc57, Pc58, Pc71; Leonard 2003) (Supplementary Table S1). Fourteen multiline cultivars were developed, registered and released in the 1960s and 1970s under the “E” series (early maturing; Frey et al. 1971a, 1985; Frey and Browning 1976b) and “M” series (midseason maturity; Frey et al. 1971b; Frey and Browning 1976a) plus the multiline cultivar Webster (Frey et al. 1988) (Supplementary Table S1). The multilines were grown on up to 0.4 million hectares in Iowa in the 1970s with no reported losses due to rust (Mundt and Browning 1985). According to Leonard (2003), Multiline E cultivars were still being grown in Iowa on about 15–20% of the acreage sown to oat, Webster continued to be grown in Minnesota into the 1990s, but series “E” cultivars were discontinued due to grower dissatisfaction. Although increases in virulence for genes Pc36, Pc57, Pc70 and Pc71 were seen in some parts of the USA during the 1980s and 1990s, Leonard (2003) speculated that this was not due to the cultivation of multiline cultivars but due to the possible deployment of these genes individually as a result of the use of the isolines used in Webster in breeding programs. This could also account for the use of gene Pc52 in cultivar Cleanleaf in Australia (Supplementary Table S1).
The role of wild Avena species in the epidemiology of Pca and implications for resistance breeding
Genetic diversity in pathogen populations and the size of pathogen populations have both been important determinants of the success in the genetic control of rust pathogens. The eradication of barberry in the USA from 1918 to 1980 was shown to have a large impact on both the oat stem rust pathogen Pga and the wheat stem rust pathogen P. graminis f. sp. tritici (Pgt), by delaying epidemic onset, reducing initial inoculum levels, vastly reducing the number of pathotypes, and stabilizing the pathogen population (Roelfs 1982). Long-term studies of Pgt in Australia, where sexual recombination is absent, have shown a strong association between the amount of stem rust present and the number of pathotypes detected (Zwer et al. 1992; Park 2007). Pca is a heteroecious macrocyclic rust pathogen, which alternates between Avena species in the telial stage and Rhamnus species (buckthorn) in the aecial stage. While sexual recombination has been documented in the Middle East, Europe and North America (Dinoor 1977; Simons 1985; Wahl 1970), it is absent in Australia where the pathogen reproduces solely via clonal asexual means. Pathogenicity surveys of Pca in Australia have been conducted at the University of Sydney since 1936 (Waterhouse, 1952; Baker and Upadhyaya 1955; Luig, 1985; Brouwer and Oates 1980; Park et al. 2000; Park RF, unpublished), and although not continuous, data from the past 70 years have clearly established that Australian populations of this pathogen are pathogenically very diverse despite the absence of sexual recombination (Luig 1985). Over the 40 year period 1979 to 2019, 107 pathotypes of Pca were detected compared to 23 of the oat stem rust pathogen P. graminis f. sp. avenae (Pga), 25 of the wheat stem rust pathogen P. graminis f. sp. tritici, 25 of the wheat leaf rust pathogen P. triticina, 24 of the wheat stripe rust pathogen Pst, and 31 of the barley leaf rust pathogen P. hordei (Park, RF unpublished).
The high degree of pathogenic diversity in Pca in Australia in the absence of sexual recombination is most likely a function of the large pathogen populations maintained on extensive wild oat communities. In addition to A. sativa, Pca also infects ubiquitous wild oats (A. barbata, found mainly along roadsides and waste areas; A. fatua, which predominates in southern Australia; A. sterilis subsp. ludoviciana, which is more prevalent in northern New South Wales (NSW) and Queensland (Qld)). Studies have shown that the composition of isolates of Pca originating from cultivated and wild oats in Qld and NSW do not differ significantly (Oates et al. 1983; Park et al. 2000), showing the importance of both host systems in the epidemiology and population dynamics of Pca there. For example, the cultivar Cleanleaf (Pc38, Pc39, Pc52; Supplementary Table S1) was released in this region in 1992. Virulence on Cleanleaf was first detected in two pathotypes in 1995, and by 1999 a further four had been detected (Park et al. 2000). Following initial detection, virulence for Cleanleaf increased rapidly in Qld and northern NSW, with at least 40% of the isolates identified from 1996 to 1998 being virulent on this cultivar (Park et al. 2000). Furthermore, from 1996 to 1999 the frequency of isolates virulent on Cleanleaf among isolates of Pca obtained from cultivated oats and wild oats were very similar, clearly illustrating the close relationship between populations of Pca on these two host groups (Park et al. 2000).
Detailed tests of Australian pathotypes of Pca have shown the presence of virulence for most designated ASR genes (Tables 1 and 2). Apart from stocks carrying 17 ASR genes that have not been tested, virulence has been detected for all others except Glabrota (Pc18, Pc29) and CI 3815 (Pc19, Pc30, Pc81-85). Many of the designated Pc genes for which virulence has been detected have not been deployed in agriculture, not only in existing pathotypes but also in newly detected presumed mutational derivatives of existing pathotypes. In some cases, virulence for these genes has been detected rarely (e.g. Pc63, Pc94), whereas in other cases it has become very common after being first detected (e.g. Pc36, Pc46, Pc92). The presence of these virulences could be due to random mutation followed by hitch-hiking, the increased frequency of certain virulence combinations due to association with other advantageous triats under direct selection, during asexual reproduction (Brown et al. 1997), or due to the presence of the genes in wild oat populations. Burdon et al. (1983) demonstrated the common occurrence of uncharacterised resistance to four pathotypes of Pca in 21 populations of wild tetraploid and hexaploid oats collected from 13 sites across NSW, with populations from the more rust prone northern NSW being significantly more resitant compared to those from the south. Such resistance in wild oat communities would be expected to exert selection pressure on populations of Pca in much the same way as is exerted by cultivated hexaploid oat.
Challenges, future directions, and prospects
The problem of pathogenic diversity in Pca
In many regions where race analysis is undertaken for cereal rust pathogens, Pca is the most pathogenically variable. Even in places such as Australia where sexual recombination is absent, far more pathotypes of Pca are detected than for any other cereal rust pathogen. The detection of virulence for Pc68 in Australia in 1999, only a year after it was first deployed, and of more than 50 Pc68-virulent pathotypes over the ensuing 20 years, is a clear example of the variation this pathogen can generate in the absence of sexual recombination. In comparison, virulence in Australia in the similarly asexually reproducing wheat stem rust pathogen Pgt for gene Sr38 was first detected in 2001; although this was just three years after Sr38 was first deployed in Australia, only five further pathotypes with virulence on this gene had been detected up to 2020 (Park RF, unpublished) despite similar numbers of differentials being used to identify pathotypes in both pathogens (30 for Pca, 29 for Pgt). The extent of variation in Pca in the absence of sexual recombination is staggering, and must be principally due to the large pathogen populations harboured by naturalised weedy wild oats.
The removal of more than 99 million barberry bushes in the USA between 1917 and 1967 and its effect on stem rust on both wheat and oat demonstrate that it is possible to undertake large-scale eradication of noxious weeds to improve disease control. In regions where buckthorn plays a role in the life cycle of Pca, large-scale eradication could be undertaken, but as in the case of the barberry eradication campaign, would require significant resourcing and ongoing efforts to ensure awareness of the danger posed to cereal production to prevent reestablishment (Leonard 2001). Even if alternate hosts such as buckthorn are eradicated in oat growing regions, it is hard to see how extensive natural and naturalised wild oat populations that occur in many oat growing regions could be removed or reduced to an extent that would have a significant impact on Pca. In the Australian wheat: Pgt system, there is evidence that the widespread cultivation of genetically resistant wheat cultivars reduced pathogen population size and diversity, stabilising resistance and the pathogen population (Zwer et al. 1992). Given that few non-cereal grass species are susceptible to Pgt in Australia, resistant wheat cultivars not only control rust in crops, but also reduce opportunities for oversummering (the “green bridge”) and early season buildup (the “green ramp”). In contrast, the widespread occurrence of wild oats as ancillary hosts means that any strategy to control Pca must be based on the assumption of a continuing, large and variable prevailing pathogen population with high evolutionary potential.
Monitoring pathogen virulence and the promise of sequence-base pathogenomics
Race (pathotype) surveys of cereal rust pathogens have been conducted in many parts of the world since the early 1900s. The only way to identify rust pathotypes remains greenhouse virulence testing using genotypes (“differentials”) with different resistance genes. In view of the very high level of pathogenic variability in Pca, and the lack of durability of ASR genes, the question could be asked as to whether pathotype (race) analysis should be undertaken for this pathogen at all. Identifying what pathotypes are present where and in what frequencies by either greenhouse testing, or in future possibly genotyping, while important, is only part of the role pathotype surveys play in rust control. When conducted in a robust and relevant way, surveys underpin all gene-based rust control strategies, from devising breeding strategies, to gene discovery, to post-release stewardship of resistance resources (Park 2015).
Rust response phenotypes are relatively easy to record and high throughput phenotyping systems have been in place in rust laboratories around the world for many years. To be relevant to agriculture, such phenotyping systems are entirely reliant on pathotype surveys to provide the most important, fully characterised isolates (defined pathotypes) for use in identifying new sources of resistance and screening breeding material. The use of minor gene APR in breeding wheat for resistance to all three rust pathogens has received increased attention over the past 50 years (e.g. Singh et al. 2011). The most efficient way to separate the effects of ASR from APR has been to use pathotypes that render any ASR genes present ineffective by matching virulence. This approach has been used extensively by staff and students at the Plant Breeding Institute University of Sydney in screening tens of thousands of lines of oats to identify those carrying useful levels of APR to crown rust. Being able to monitor and know what ASR genes are present in breeding programs is also important to prevent the inadvertent selection of ASR genes that are known to be non-durable. These include “fossil” resistance genes, genes present in germplasm that are occasionally revealed by the recurrence of previously rare avirulence or occurrence of unusual avirulences. For example, a pathotype of P. triticina first identified in Australia in 2004 was found to be avirulent on an ASR gene, Lr73, present in the rust susceptible genotype Morocco and nine Australian wheat cultivars (Park et al. 2014). Being able to recognise Lr73 has been important in preventing its unintentional selection in resistance breeding. Knowing what pathotypes are present and in what frequency will also be critical if multilines are used in future (see below).
It is impractical to include all currently designated/ temorarily designated alleles conferring resistance to Pca in a differential set that is used routinely. It is also of little value as many of the genes are defeated or have not been used in agriculture. As discussed in detail by McIntosh et al. (1995), the reasons for including or excluding differential genotypes in a defined differential set are varied, and factors such as geographical differences in virulence frequencies and the resistance genes deployed in agriculture limit international agreement on what genotypes should be used. Chong et al. (2000) proposed a set of 16 differential genotypes as the basis of a North American system for naming pathotypes of Pca. Most if not all of these are also used as differentials in Australia (Park RF unpublished), the Czech Republic (Klenová-Jiráková et al. 2010), Poland (Paczos-Grzeda and Sowa 2019), South Africa (Boshoff et al. 2020) and Spain (Sánchez-Martín et al. 2012), and hence can be considered as core international pathogenicity testers for Pca. Excluding Pc54 and Pc64, the differential gene in 14 of these genotypes has been deployed in agriculture in at least one cultivar (Supplementary Table S1). Genes that are not represented, but which have been deployed, include Pc1, Pc3+Pc4, Pc10, Pc13, Pc14, Pc61, Pc91, Pc94, plus genes in older sources of resistance (Anthony/Bond//Boone, Bond, Garry, Landhafer, Santa Fe, Trispernia, Ukraine, Victoria). Many of these are either present in currently grown oat cultivars or still provide useful discrimination of local populations of Pca, and are thus used by some groups as regional supplemental differentials. Should APR genes be deployed in commercial oat cultivars, it may be necessary to monitor virulence on these as not all APR genes are durable (Park and McIntosh 1994).
Designated Pc loci
Based on the resistance loci catalogued to date, it would appear that the diversity of resistance to Pca in A. sativa is low, and to Pga it appears to be even lower with only 12 loci being designated from genotypes of A. sativa (Gnanesh et al. 2014). The 92 loci that confer resistance to Pca in Avena that have been designated so far may seem to be a large number, however to date 78 distinct loci conferring resistance to P. triticina have so far been mapped robustly and designated in wheat.
Allelic relationships between many of the designated Pc genes are unknown, and it is likely that at least some are duplicates, or even triplicates. Although 19 loci have been designated from the diploid A. strigosa, only three or possibly four are available in stable hexaploid lines and there is convincing evidence that some of these represent the same gene, raising the question as to how diverse resistance to Pca in A. strigosa actually is. The recent release of chromosome scale assemblies for the two diploid species A. atlantica and A. eriantha (Maughan et al. 2019) and the hexaploid A. sativa genotype OT3098 (https://wheat.pw.usda.gov/GG3/graingenes_downloads/oat-ot3098-pepsico) have provided invaluable genetic resources that will unquestionably accelerate efforts to map all designated Pc genes for which unambigiuous genetic stocks exist, and to assess potential allelism between them. It is to be expected that this will lead to changes in the numbering of at least some designated loci, and possible deletion of others from the catalogue, as has occurred in wheat (e.g. the deletion of Sr1 due to synonymy with Sr9d, and of Sr3 and Sr4 due to a lack of single gene stocks; McIntosh et al. 1995).
Searches for new sources of resistance
Simons (1985) stated that “It is now apparent that there is little point in studying the genetics of resistance to P. coronata unless the resistance has some practical usefulness.” Most efforts to find new sources of resistance to Pca until now have focused on ASR genes, and most of these genes have come from species of Avena other than A. sativa. Searches for new sources of resistance in alien species will no doubt uncover new ASR genes. Cabral and Park (2014) examined resistance to Pca in 97 accessions of A. strigosa from North America, South America and Europe, finding that 20 lacked detectable resistance to the pathotypes used, 18 carried the Saia resistance, one Pc94, and only three that were resistant to all pathotypes used. A second study using SSRs showed that the A. strigosa accessions were 86% genetically similar, so this study may not be a true reflection of the genetic variability available in this species (Cabral et al. 2013). The accessions proved to be very difficult to work with and very few introgression stocks could be produced (Aung et al. 2010; Cabral et al. 2011; Cabral and Park 2014, 2016). Any attempt to introgress rust resistance from diploid or tetraploid Avena species to hexaploid oat, in the absence of transgenic approaches, will take at least 10 years to be of benefit to oat growers. It should also be noted that any resistance genes sourced from Avena species that are weeds could well be already present in weedy wild oat communities in regions where the resistance is deployed in agriculture.
Sánchez-Martín et al. (2012) screened 159 Avena accessions including A. sativa (subspp. sativa and strigosea) landraces and cultivars. Eight landraces and four cultivars showed an elevated degree of resistance according to macroscopic components at seedling stage under controlled conditions and further histological analysis showed a set of different resistance responses acting at susccesive stages of the infection process. Later, field experiments of the commercial varieties at six contrasting locations along Mediterranean Basin, including Spain, Tunisia, Egypt and Palestinian Territories, showed that some of these varieties were consistently resistant across these different environments (Sánchez-Martín et al. 2014). Interestingly, these accessions were characterised by a high level of pre-penetration resistance and/ or early aborted colonies not associated with host cell necrosis. This study highlights how knowledge of resistance mechanisms/responses may enable the use of plant genotypes able to stop the pathogen at different developmental stages reducing the pressure on ASR genes (usually related to hypersensitive response) and may prove more durable and difficult to overcome by evolving pathogenic races (Rubiales and Niks 2000).
Six designated loci (Pc27, Pc28, Pc69, Pc72, Pc73, Pc74) are reported as conferring APR to crown rust. Several other oat cultivars (Ajax, Beaver, Erban, Lodi, Portage, Red Rustproof, Rodney, Santa Fe, Ukraine) and accessions (PI 174545, PI 18783, PI 197278, PI 174544, PI 179279, MN841801, MN841804) have been reported to carry uncharacterised APR to crown rust. The latter are especially promising as potential sources of effective and durable resistance given that they displayed stable resistance in a buckthorn nursery over seven years (Leonard 2002), and one or both lines have shown good levels of APR under field conditions in Canada, the USA and Australia in subsequent studies. Both lines have also been shown to induce premature telial formation, truncating urediniospore production, reducing inoculum load, and hence functioning as a form of resistance (viz. MN841801 and MN841804). These lines were distributed with the 2000 Quaker International Oat Nursery and were shown to induce premature telial formation under Australian field conditions (Roake J and Park RF unpublished). This type of resistance may hold some promise in future resistance breeding efforts, especially as a component of polygenic minor APR.
In addition to these sources of APR to Pca, integrated greenhouse seedling tests and field adult plant tests of more than 100,000 oat genotypes at the University of Sydney’s Plant Breeding Institute over the past 25 years have identified 385 lines of A. sativa with reasonable agronomic performance that carry levels of APR to Pca varying from low to very high (Haque 2004; Issa 2005; Park RF, Singh D, and Roake J, unpublished). Combined, these studies demonstrate that APR to crown rust in hexaploid oat is not uncommon, and although the genetic diversity of the resistance found to date is not yet well understood, it is known that multiple genes of additive effects are involved in some cases and that some of these have been stable over time and in different environments, making them potentially valuable in breeding for resistance to Pca.
Studies under controlled conditions have shown that APR genes are switched on at different growth stages, some soon after seedling growth stages and others not until flag leaf emergence (Park and McIntosh 1994). Just how and at what growth stages resistance genes are expressed are important considerations in ensuring adequate protection from diseases, especially so if APR is used in grazing oat, in which the early onset of resistance will be essential because of the implications of withholding periods for fungicides in grazing.
Resistance gene deployment
The concept of combining resistance genes to bestow greater durability was proposed by researchers many years ago (e.g. Watson and Singh 1952). The development of markers linked to resistance genes continues to improve the efficiency by which multiple gene resistances can be assembled (e.g. Pc68, Chen et al. (2006), Satheeskumar et al. (2011); Pc71 (Bush and Wise 1998); Pc92 (Rooney et al. 1994); Pc94 (Chong et al. 2004) and Pc91 (McCartney et al. 2011)). The problem of pathogen variability, however remains; if this approach is applied to pyramiding genes conferring ASR to Pca, it will be of critical importance to ensure that none are deployed singly or occur in a natural state in wild oat populations. The use of single genes provides the “stepping stones” needed for the pathogen to acquire the virulences needed to render multigenic resistances ineffective. This was documented in Australia when cultivars carrying Pc39 (Quamby, released 1990), Pc52 (Cleanleaf, 1992), Pc38 and Pc39 (Riel, 1993), Pc68 (Graza 68, 1998), Pc50 (Volta, 2003), and Pc91 (Drover, 2006) were released and Pca acquired sequential matching virulence resulting in a single pathotype combining virulence for all six ASR genes in about 25 years (Park RF unpublished). Although no rust resistance gene has been isolated from oat to date, the availability of reference genomes and the availability of non-map-based approaches to gene cloning (e.g. Steuernagel et al. 2017) will no doubt fast-track efforts to do so. The recent availability of reference genomes for hexaploid and diploid oat will undoubtedly also accelerate efforts to discover, characterise and develop high throughput diagnostic markers to introgress resistance to Pca into high yielding adapted oat germplasm. The common occurrence of minor gene, additive APR to Pca in hexaploid oat germplasm raises the possibility of pyramiding several such genes to give high levels of resistance, an effort that will be greatly expedited by the development of high throughput diagnostic markers based on gene sequences.
Luo et al. (2021) recently published a protocol to stack cloned ASR and APR genes to Pgt in a single cassette and transform them into wheat as a single locus. The authors stressed the importance of strategic deployment of such stacks to retain their effectiveness, highlighted by the detection of a pathotype of Pgt combining virulence for three of the four stacked ASR genes in Europe despite none having been deployed in agriculture.
An important consideration in combining resistance genes is resistance gene suppression. Hurni et al. (2014) demonstrated that the susceptibility allele Pm3C, and three resistance alleles (Pm3a, Pm3b and Pm3f) all suppressed the rye derived resistance gene Pm8, which had been introgressed into wheat. The strength of the suppression was shown to vary with the Pm3 allele used, raising the possibility that gene editing could be used to modify alleles so that suppression did not occur. Wilson and McMullen (1997) showed that Pc38 suppressed Pc62, and Chong and Aung (1996) found that Pc38 suppressed Pc94. While these examples may be the exception, it remains an important consideration in control strategies based on combining resistance genes.
An alternative to multi-gene cassettes in a single cultivar is to use transformation to introduce different genes or gene combinations into a single background, and mix to produce multiline cultivars. The oat multilines with resistance to crown rust developed by the Iowa group were based on common backgrounds that were developed well before the multiline was release. For example, multilines E76 and E77 were based on the recurrent parental line CI 8044, developed from the cross Clintland/Garry 5, with the original cross being made in 1954 and eventual release of the multiline cutivars in 1976 and 1977 (Frey et al. 1985). Shortening the 20 year period between making the first cross and cultivar release by using transformation technologies would ensure future multiline cultivars compete with genetic gains made in pure line cultivar breeding programs, and ensure quality traits matched to market demands. An efficient transformation system and supply of cloned resistance genes in oat may make this possible. In the absence of acceptance of transgenic oat, accelerated generation breeding approaches (Gonzalez-Barrios et al. 2020) combined with marker assisted selection and genomic selection/ prediction (Asoro et al. 2011; Rio et al. 2021) could be used.
Mundt (2002) stressed the importance of functional diversity in multiline cultivars; randomly chosen mixtures will not necessarily provide adequate disease control, and in order to be effective mixture components need to be relevant, or functional, against a pathogen population. Of the nine genes used to develop the oat multilines at Iowa, combined virulence exists for six in a single pathotype in Australia despite none of these genes being deployed there (Park RF unpublished). Understanding the virulence compositon of populations within an epidemiological region is therefore critical in ensuring that the components and their frequencies within mixtures bestow the best protection possible against prevailing pathotypes. Integrating pathogen surveillance with gene deployment in this way would be analogous to the approach used to mass vaccinate chickens against prevailing variants of the infectious bronchitis virus (Jordan 2017).
There is evidence that genetic “background” can influence durability (eg. The pvr23 gene in pepper, Palloix et al. 2009; Lr24 in wheat in Australia, Park et al. 2000), supporting the strategy of using reputedly durable sources of rust resistance as “background” resistance, to which other resistance genes are added (McIntosh 1988). The discovery of three durable pleiotropic minor gene (APR) loci in wheat that provide resistance to three rust pathogens and powdery mildew, and the development of diagnostic gene-based markers (Lr67/Yr46/Sr55; Lr34/Yr18/Sr57) or closely linked markers (Lr46/Yr29/Sr58) has been a huge advance in efforts to breed for durable rust resistance in this crop. These markers have allowed for the selection of these minor APR genes in breeding populations and their use as background or backbone resistance on top of which ASR genes have been pyramided. These markers have also made discovering other sources of minor APR faster by allowing genotypes carrying APR to be genotyped and attention focused on lines lacking these genes but carrying useful levels of APR. Whether or not such pleiotropic genes exist in oat is unknown, but given their importance in wheat efforts to discover new sources of resistance in oat should take this possibility into consideration.
Enduring rust control
As stated by Browing and Frey (1969), it is populations of plants that feed increasing populations of people, not individual plants. While the development and use of oat cultivars with reputedly durable sources of resistance to crown rust will be foundational in attempts to achieve enduring crown rust control, it must be noted that not all durable resistance protects against yield loss (e.g. Ma and Singh 1996), and that durable resistance may not remain so forever (Johnson 1984). Ensuring that the durable resistance sources used to develop crown rust resistant oat cultivars are genetically diverse and bestow levels of resistance that prevent yield loss and slow epidemic development will be vital in reducing the impact of this disease in oat production, and in particular buffering against potential future changes in Pca.
The use of diverse sources of resistance has been a critical aspect of enduring stem rust control in wheat in Australia for the past 50 years (Park 2007). Stem rust resistance breeding has been underpinned by close monitoring of the pathogen, and integrated with practices including avoiding the cultivation of highly susceptible cultivars, not overlapping crops and optimising the break between cropping cycles, green bridge destruction, and at times the use of fungicides. Achieving enduring control of oat crown rust will require a similar integrated whole of system approach based on a foundation of durable resistance. This could include manipulating sowing date and crop phenology, which have been shown to reduce crown rust severity. Simons and Michel (1968) found that early maturing oat varieties tend to be less damaged than do later maturing varieties, probably because they ripen before the disease becomes fully established. Fleischmann and McKenzie (1965) reported less crown rust damage in the oat cultivar Garry, which carries the APR genes Pc27 and Pc28, when it was planted earlier than usual. The durability of any resistance to crown rust deployed in agriculture will continue to be challenged by the ongoing presence of large and variable populations of Pca in wild oat communities in many of the world’s oat growing regions and of a functional sexual stage in some regions. The oat: Pca system may therefore also provide important opportunities to understand how best to integrate durable resistance with non-gene-based control strategies to protect important genes and achieve enduring crop disease control.
References
Acevedo M, Jackson EW, Chong J, Rines HW, Harrison S, Bonman JM (2010) Identification and validation of quantitative trait loci for partial resistance to crown rust in oat. Phytopathology 100:511–521
Admassu-Yimer B, Bonman JM, Esvelt Klos K (2018a) Mapping of crown rust resistance gene Pc53 in oat (Avena sativa). PLoS ONE. https://doi.org/10.1371/journal.pone.0209105
Admassu-Yimer B, Gordon T, Harrison S, Kianian S, Bockleman H, Bonman JM, Esvelt Klos K (2018b) New sources of adult plant and seedling resistance to Puccinia coronata f. sp. avenae identified among Avena sativa accessions from the National Small Grains Collection. Plant Dis 102:2180–2186
Admassu-Yimer B, Klos KE, Griffiths I, Cowan A, Howarth C (2022) Mapping of crown rust (Puccinia coronata f. sp. avenae) resistance Gene Pc54 and a novel quantitative trait locus effective against powdery mildew (Blumeria graminis f. sp. avenae) in the Oat (Avena sativa) Line Pc54. Phytopathology. 2022 Apr 25:PHYTO10210445R. doi: https://doi.org/10.1094/PHYTO-10-21-0445-R. Epub ahead of print. PMID: 34982574
Agenbag GM, Pretorius ZA, Boyd LA, Bender CM, Prins R (2012) Identification of adult plant resistance to stripe rust in the wheat cultivar Cappelle-Desprez. Theor Appl Genet 125:109–120
Asoro FG, Newell MA, Beavis WD, Scott WD, Jannink J-L (2011) Accuracy and training population design for genomic selection on quantitative traits in elite North American oats. The Plant Genome 4:132–144
Aung T, Zwer P, Park R, Davies P, Singh P, Dundas I (2010) Hybrids of Avena sativa with two diploid wild oats (Clav6956) and (Clav7233) resistant to crown rust. Euphytica 174:189–198
Aung T, Chong J, Leggett M (1996). The transfer of crown rust resistance gene Pc94 from a wild diploid to cultivated hexaploid oat. In: Proceedings of the 9th European and Mediterranean cereal rusts and powdery mildews conference (pp. 2–6)
Avena sativa – OT3098 v2, PepsiCo, https://wheat.pw.usda.gov/jb?data=/ggds/oat-ot3098v2-pepsico
Babiker EM, Gordon TC, Jackson EW, Chao S, Harrison SA, Carson ML, Obert DE, Bonman JM (2015) Quantitative trait loci from two genotypes of oat (Avena sativa) conditioning resistance to Puccinia coronata. Phytopathology 105:239–245
Baker EP (1966) Isolation of complementary genes conditioning crown rust resistance in the variety Bond. Euphytica 15:313–318
Baker EP, Upadhyaya YM (1966) Studies on the inheritance of rust resistance in oats. IV. Expression of allelomorphism and gene interaction for crown rust resistance in crosses involving the oat variety Bond. Proc Linnean Soc NSW 91:155–175
Baker EP, Upadhyaya YM (1955) Physiologic specialization in crown rust of oats. Proceedings of the Linnnean Society of NSW 80: 240-257
Barbosa MM, Federizzi LC, Milach SCK, Martinelli JA, Thome GC (2006) Molecular mapping and identification of QTLs associated to oat crown rust partial resistance. Euphytica 150:257–269
Bekele WA, Wight CP, Chao S, Howarth CJ, Tinker NA (2018) Haplotype-based genotyping-by-sequencing in oat genome research. Plant Biotechnol J 16:1452–1463
Bekele WA, Itaya A, Boyle B, Yan W, Fetch JM, Tinker NA (2020) A targeted genotyping-by-sequencing tool (rapture) for genomics-assisted breeding in oat. Theor Appl Genet 133:653–664
Biffin RH (1905) Mendel’s laws of inheritance and wheat breeding. J Agric Sci 1:4–48
Blake VC, Woodhouse MR, Lazo GR, Odell SG, Wight CP, Tinker NA, Wang Y, Gu YQ, Birkett CL, Jannink JL, Matthews DE, Hane DL, Michel SL, Yao E, Sen TZ (2019). GrainGenes: centralized small grain resources and digital platform for geneticists and breeders. Database (Oxford). 2019 Jan 1;2019:baz065. doi: https://doi.org/10.1093/database/baz065. PMID: 31210272; PMCID: PMC6580076
Bonnett DG (1996). Host: pathogen studies of oat leaf rust in Australian (Doctoral dissertation, University of Sydney)
Boshoff WHP, Pretorius ZA, Tefere T, Visser B (2020) Occurrence and pathogenicity of Puccinia coronata f. sp. avenae on oat in South Africa. Crop Prot 133:105–144
Briére SC, Kushalappa AC, Mather DE (1994) Screening for partial resistance to an isolate of crown rust (Puccinia coronata f. sp. avenae) race 264 in oat cultivars and breeding lines. Can J Plant Path 16:49–55
Brouwer JB, Oates JD (1980) Regional variation of Puccinia coronata f. sp. avenae in Australia and its implications for resistance breeding. Annal Appl Biol 109:269–277
Brouwer JB (1983) Inheritance and breeding potential of resistances to Puccinia coronata avenae in oats. PhD Thesis. University of Sydney, Australia, 370 pp
Brown CM, Kolb FL (1989a) Registration of ‘Don’ oat. Crop Sci 29:1572–1573
Brown CM, Kolb FL (1989b) Registration of ‘Hazel’ oat. Crop Sci 29:1573
Brown JKM, Foster EM, O’Hara RB (1997) Adaptation of powdery mildew populations to cereal varieties in relation to durable and non- durable resistance. In: Crute IR, Holub EB, Burdon JJ (eds) The Gene- for- Gene Relationship in Plant- Parasite Interactions. CAB International, Wallingford UK, pp 119–138
Brown PD, Duguid SD, Haber S, Chong J, Harder DE, Menzies J, Noll JS, McKenzie RIH (2000) AC Assiniboia oat. Can J Plant Sci 81:77–79
Browning JA, Frey KJ (1969) Multiline cultivars as a means of disease control. Annu Rev Phytopathol 7:355–382
Burdon JJ, Oates JD, Marshall DR (1983) Interactions between Avena and Puccinia species. I. The Wild Hosts: Avena barbata Pott Ex Link, A. fatua LA ludoviciana Durieu. J Appl Ecol 20:571–584
Bush AL, Wise RP (1996) Crown rust resistance loci on linkage groups 4 and 13 in cultivated oat. J Hered 87:427–432
Bush AL, Wise RP (1998) High-resolution mapping adjacent to the Pc71 crown-rust resistance locus in hexaploid oat. Mol Breeding 4:13–21
Cabral AL, Park RF (2014) Seedling resistances to Puccinia coronata f. sp. avenae in Avena strigose, A Barbata and a Sativa. Euphytica 196:385–395
Cabral AL, Park RF (2016) Genetic analysis of seedling resistance to crown rust in five diploid oat (Avena strigosa) accessions. J Appl Genet 57:27–36
Cabral AL, Singh D, Park RF (2011) Identification and genetic characterisation of adult plant resistance to crown rust in diploid and tetraploid accessions of Avena. Annal Appl Biol 159:220–228
Cabral AL, Karaoglu H, Park RF (2013) The use of microsatellite polymorphisms to characterise and compare genetic variability in Avena strigosa and A. barbata. Genet Resour Crop Evol 60:1153–1163
Carson ML (2009) Broad spectrum resistance to crown rust, Puccinia coronata f. sp. avenae, in accessions of the tetraploid slender oat Avena Barbata. Plant Dis 93:363–366
Chaffin S, Huang YF, Smith S, Bekel W, Babiker E, Gnanesh BN, Foresman BJ, Blanchard SG, Jay JJ, Reid RW, Wright CP, Chao S, Oliver R, Islamovic E, Kolb FL, McCartney C, Mitchell-Fetch JW, Beattie AD, Bjornstad A, Bonman JM, Langdon T, Howarth CJ, Brouwer CR, Jellen EN, Klos KE, Poland JA, Hsieh TF, Brown R, Jackson E, Schlueter JA, Tinker NA (2016) A consensus map in cultivated hexaploid oat reveals conserved grass synteny with substantial subgenome rearrangement. Plant Genome. https://doi.org/10.3835/plantgenome2015.10.0102
Chang TD, Sadanga K (1964) Crosses of six monosomics in Avena sativa L. with varieties, species, and chlorophyl mutants. Crop Sci 4:589–593
Chen Q, Armstrong K (1994) Genomic in situ hybridization in Avena sativa. Genome 37:607–612
Chen C-Y, Milbury PE, Kwak H-K, Collins FW, Samuel P, Blumberg JB (2004) Avenanthramides and phenolic acids from oats are bioavailable and act synergistically with Vitamin C to enhance hamster and human LDL resistance to oxidation. J Nutr 134:1459
Chen G, Chong J, Gray M, Prashar S, Procunier D (2006) Identification of single nucleotide polymorphisms linked to resistance gene Pc68 to crown rust in cultivated oat. Can J Plant Path 28:214–222
Chen G, Chong J, Prashar S, Procunier JD (2007) Discovery and genotyping of high-throughput SNP markers for crown rust resistancegene Pc94 in cultivated oat. Plant Breed 126:379–384
Chen CC, Jost M, Clark B, Martin M, Matny O, Steffenson BJ, Franckowiak JD, Mascher M, Singh D, Perovic D, Richardson T, Periyannan S, Lagudah ES, Park RF, Dracatos PM (2021) BED domain containing NLR from wild barley confers resistance to leaf rust. Plant Biotech J. https://doi.org/10.1111/pbi.13542
Chong J (2000) Inheritance of resistance to two Puccinia coronata isolates in a partial resistant oat line MN841801. Acta Phytopathologia Entomologia Hungarica 35:37–40
Chong J, Seaman WL (1989) Virulence and distribution of Puccinia coronata in Canada in 1988. Can J Plant Path 11:439–442
Chong J, Howes NK, Brown PD, Harder DE (1994) Identification of the stem rust resistance gene Pg9 and its association with crown rust resistance and endosperm proteins in ‘Dumont’ oat. Genome 37:440–447
Chong J, Leonard KJ, Salmeron JJ (2000) A North American system of nomenclature for Puccinia coronata f. sp. avenae. Plant Dis 84:580–585
Chong J, Reimer E, Somers D, Aung T, Penner GA (2004) Development of sequence-characterized amplified region (SCAR) markers for resistance gene Pc94 to crown rust in oat. Can J Plant Path 26:89–96
Chong J, Aung T (1996). Interactions of the crown rust resistance gene Pc94 with several Pc genes. Proceedings of the 9th European and Mediterranean Cereal Rusts and Powdery Mildews Conference. 206 September 1996, Lunteren The Netherlands. 172–175
Coffman FA, Murphy HC, Chapman WH (1961). Oat breeding. In: Oat and Oat Improvement (Coffman FA, ed), pp. 263–329. Agronomy 8, The American Society of Agronomy: Madison Wisconsin UAS
Coffman FA (1977) Oat history, identification and classification. USDA ARS Technical Bulletin No. 1516. 356pp
Collins FW (1989) Oat phenolics: avenanthramides, novel substituted N-cinnamoylanthranilate alkaloids from oat groats and hulls. J Agric Food Chem 37:60–66
Derick RA (1953) Oats in Canada. Canadian Department of Agriculture Pub 554: 3–23
Ding Y, Cuddy WS, Wellings CR, Zhang P, Thach T, Hovmøller MS, Qutob D, Brar GS, Kutcher HR, Park RF (2021) Incursions of divergent genoptypes, evolution of virulence and host jumps shape a cpntintntal clonal population of the stripe rust pathogen Puccinia striiformis. Mol Ecol. https://doi.org/10.1111/mec.16182
Dinoor A (1977) Oat crown rust resistance in Israel [Rhamnus palaestina on Avena sterilis and Avena barbata]. Annals of the New York Academy of Sciences (USA)
Dracatos PM, Bartoš J, Elmansour H, Singh D, Karafiátová M, Zhang P, Steuernagel B, Svačina R, Cobbin JCA, Clark B, Hoxha S, Khatkar MS, Doležel J, Wulff BB, Park RF (2019) The coiled-coil NLR Rph1, confers leaf rust resistance in barley cultivar Sudan. Plant Physiol. https://doi.org/10.1104/pp.18.01052
Dyck PL, Zillinsky FJ (1963) Inheritance of crown rust resistance transferred from diploid to hexaploid oats. Can J Genet Cytol 5:398–407
Emmons CL, Peterson DM (1999) Antioxidant activity and phenolic contents of oat groats and hulls. Cereal Chem 76:902–906
Fitzsimmons RW, Roberts GL, Wrigley CW (1983) Australian Oat Varieties. CSIRO Editorial and Publishing Service, Melbourne Australia, p 68
Fleischmann G, McKenzie RIH (1965) Yield losses in Garry oats infected with crown rust. Phytopathology 55:767–770
Fleischmann G, McKenzie RIH, Shipton WA (1971) Inheritance of crown rust resistance in Avena sterilis collections from Israel, Portugal, and Tunisia. Can J Genet Cytol 13:251–255
Flor HH (1955) Host –parasite interaction in flax rust: Its genetics and other implications. Phytopathology 45:680–685
Frey KJ, Browning JA (1976a) Registration of Multiline M72 and Multiline M73 oat cultivars. Crop Sci 16:311
Frey KJ, Browning JA (1976b) Registration of Multiline E72, Multiline E73, and Multiline E74 oat cultivars. Crop Sci 16:311–312
Frey KJ, Browning JA, Grindeland RL (1971a) Registration of Multiline E68, Multiline E69 and Multiline E70 oat cultivars. Crop Sci 11:939–940
Frey KJ, Browning JA, Grindeland RL (1971b) Registration of Multiline M68, Multiline M69 and Multiline M70 oat cultivars. Crop Sci 11:940–941
Frey KJ, Browning JA, Simons MD (1985) Registration of Multiline E76 and Multiline M77 oats. Crop Sci 25:1125
Frey KJ, Simons MD, Michel LJ, Murphy JP, Browning JA (1988) Registration of Webster oat. Crop Sci 28:374–375
Gnanesh BN, Mitchell Fetch J, Menzies JG, Beattie AD, Eckstein PE, McCartney CA (2013) Chromosome location and allele-specific PCR markers for marker-assisted selection of the oat crown rust resistance gene Pc91. Mol Breed 32:679–686
Gnanesh BN, McCartney CA, Eckstein PE, Fetch JWM, Menzies JG, Beattie AD (2015) Genetic analysis and molecular mapping of a seedling crown rust resistance gene in oat. Theor Appl Genet 128:247–258
Gnanesh BN, Fetch JM, Zegeye T, McCartney CA, Fetch T (2014) Oat. Alien Gene Transfer in Crop Plants 2, pp.51–73
Gonzalez-Barrios P, Bhatta M, Halley M, Sandro P, Gutierrez L (2020) Speed breeding and early panicle harvest accelerates oat (Avena sativa L.) breeding cycles. Crop Sci 61:320–330
Haque MS (2004). Rust resistance in oats and variability on the oat stem rust pathogen Puccinia graminis f. sp. avenae. PhD Thesis, Univeristy of Sydney
Harder DE, McKenzie RIH, Martens JW (1980) Inheritance of crown rust resistance in three accessions of Avena sterilis. Can J Genet Cytol 22:27–33
Harder DE, McKenzie RIH, Martens JW (1984) Inheritance of adult plant resistance to crown rust in an accession of Avena sterilis. Phytopathology 74:352–353
Harder DE, Chong J, Brown PD (1995) Stem rust and crown rust resistance in the Wisconsin oat selection X1588–2. Crop Sci 35:1011–1015
Harder DE, Haber S (1992) Oat diseases and pathologic techniques. In: Oat Science and Technology (Marshall, H and Sorells, M, eds), pp. 307–425. Madison, WI: ASA/CSA
Hoffman DL, Chong J, Jackson EW, Obert DE (2006) Characterization and mapping of a crown rust resistance gene complex (Pc58) in TAM O-301. Crop Sci 46:2630–2635
Hurni S, Brunner S, Stirnweis D, Herren G, Peditto D, McIntosh RA, Keller B (2014) The powdery mildew resistance gene Pm8 derived from rye is suppressed by its wheat orthologue Pm3. Plant J 79:904–913
Issa AKA (2005) Crown rust resistance in oats Puccinia coronata f. sp. avenae. MSc Thesis, University of Sydney. 92pp
Jackson EW, Obert DE, Menz M, Hu G, Avant JB, Chong J, Bonman JM (2007) Characterization and mapping of oat crown rust resistance genes using three assessment methods. Phytopathology 97:1063–1070
Jackson EW, Obert DE, Menz M, Hu G, Bonman JM (2008) Qualitative and quantitative trait loci conditioning resistance to Puccinia coronata pathotypes NQMG and LGCG in the oat (Avena sativa L.) cultivars Ogle and TAM-O-301. Theor Appl Genet 116:517–527
Jackson EW, Obert DE, Avant JB, Harrison SA, Chong J, Carson ML, Bonman JM (2010) Quantitative trait loci in the Ogle/TAM O-301 oat mapping population controlling resistance to Puccinia coronata in the field. Phytopathology 100:484–492
Jellen EN, Rooney WL, Phillips RL, Rines HW (1993) Characterization of the hexaploid oat Avena byzantina cv Kanota monosomic series using C-banding and RFLPs. Genome 36:962–970
Jensen NF (1965) Multiline superiority in creals. Crop Sci 5:566–568
Jiang W, Jiang C, Yuan W, Zhang M, Fang Z, Li Y, Li G, Jia J, Yang Z (2021) A universal karyotypic system for hexaploid and diploid Avena species brings oat cytogenetics into the genomics era. BMC Plant Biol 21:1–15
Johnson R (1978) Practical breeding for durable resistance to rust diseases in self- pollinating cereals. Euphytica 27:529–540
Johnson R (1984) A critical analysis of durable resistance. Annu Rev Phytopathol 22:309–330
Jordan B (2017) Vaccination against infectious bronchitis virus: a continuous challenge. Vet Microbiol 206:137–143
Kebede AZ, Friesen-Enns J, Gnanesh BN, Menzies JG, Mitchell Fetch JW, Chong J, Beattie AD, Paczos-Grzeda E, McCartney CA (2019) Mapping oat crown rust resistance gene Pc45 confirms association with PcKM. G3 Genes Genom Genet 9:505–511
Kiehn FA, McKenzie RIH, Harder DE (1976) Inheritance of resistance to Puccinia coronata avenae and its association with seed characteristics in four accessions of Avena sterilis. Can J Genet Cytol 18:717–726
Klenová-Jiráková H, Leišová-Svobodová L, Hanzalová A, Kucera L (2010) Diversity of oat crown rust (Puccinia coronata f. sp. avenae) isolates detected by virulence and AFLP analyses. Plant Prot Sci 46:98–106
Klos KE, Yimer BA, Babiker EM, Beattie AD, Bonman JM, Carson ML, Chong J, Harrison SA, Ibrahim AMH, Kolb FL, McCartney CA, McMullen M, Fetch JM, Mohammadi M, Murphy JP, Tinker NA (2017) Genome-wide association mapping of crown rust resistance in oat elite germplasm. Plant Genome 10:2
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Huerta-Espino J, McFadden H, Bossolini E, Selter LL, Keller B (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363
Kulchelski FR, Graichen FAS, Martinelli JA, Locatelli AB, Federizzi LC, Delatorre CA (2010) Molecular mapping of Pc68, a crown rust resistance genes in Avena sativa. Euphytica 175:423–432
Latta RG, Bekele WA, Wight CP, Tinker NA (2019) Comparative linkage mapping of diploid, tetraploid, and hexaploid Avena species suggests extensive chromosome rearrangement in ancestral diploids. Sci Rep 9:1–12
Leggett JM, Markhand GS (1995) The genomic identification of some monosomics of A. sativa L. cv. Sun II using genomic in situ hybridization. Genome 38:747–751
Leonard KJ (2002) Oat lines with effective adult plant resistance to crown rust. Plant Dis 86:593–598
Leonard KJ (2003) Regional frequencies of virulence in oat crown rust in the United States from 1990 through 2000. Plant Dis 87:1301–1310
Leonard KJ, Martinelli JA (2005) Virulence of oat crown rust in Brazil and Uruguay. Plant Dis 89:802–808
Leonard KJ, Anikster Y, Manisterski J (2004) Patterns of virulence in natural populations of Puccinia coronata on wild oat in Israel and in agricultural populations on cultivated oat in the United States. Phytopathology 94:505–514
Leonard KJ, Anikster Y, Manisterski J (2005) Virulence associations in oat crown rust. Phytopathology 95:53–61
Leonard KJ (2001). Stem rust – Future enemy? In: Stem Rust of Wheat. From Ancient Enemy to Modern Foe (PD Peterson ed), pp. 119–146. APS Press: St Paul Minnesota USA
Lin Y, Gnanesh BN, Chong J, Chen G, Beattie AD, Mitchell Fetch JW, Kutcher HR, Eckstein PE, Menzies JG, Jackson EW, McCartney CA (2014) A major quantitative trait locus conferring adult plant partial resistance to crown rust in oat. BMC Plant Biol 14:250
Loskutov IG, Rines HW (2011) Avena. In: Kole C (ed) Wild Crop relatives: genomic and breeding reources. Springer, Berlin, pp 109–183
Luig NH (1985) Epidemiology in Australia and New Zealand. In: Roelfs AP, Bushnell WR (eds) The cereal rusts, volume ii diseases, distribution, epidemiology, and control. Academic Press Inc., Orlando USA, pp 301–328
Luke HH, Chapman WH, Barnett RD (1972) Horizontal resistance of Red Rustproof oats to crown rust. Phytopathology 62:414–417
Luke HH, Barnett RD, Pfahler PL (1975) Inheritance of horizontal resistance to crown rust in oats. Phytopathology 65:631–632
Luo M, Xie L, Chakraborty S, Wang A, Matny O, Jugovich M, Kolmer JA, Richardson T, Bhatt D, Hoque M, Patpour M (2021) A five-transgene cassette confers broad-spectrum resistance to a fungal rust pathogen in wheat. Nat Biotechnol 39:561–566
Ma H, Singh RP (1996) Contribution of adult plant resistance gene Yr18 in protecting wheat from yellow rust. Plant Dis 80:66–69
Marshall HG, Myers WM (1961) A cytogenetic study of certain interspecific Avena hybrids and the inheritance of resistance in diploid and tetraploid varieties to races of crown rust. Crop Sci 1:29–34
Martens JW, McKenzie RIH, Fleischmann G (1968) The inheritance of resistance to stem rust and crown rust in Kyto oats. Can J Genet Cytol 10:808–812
Maughan PJ, Lee R, Walstead R, Vickerstaff RJ, Fogarty MC, Brouwer CR, Reid RR, Jay JJ, Bekele WA, Jackson EW, Tinker NA (2019) Genomic insights from the first chromosome-scale assemblies of oat (Avena spp.) diploid species. BMC Biol 17:1–19
McCallum BD, Fetch T, Chong J (2007) Cereal rust control in Canada. Aust J Agric Res 58:639–647
McCartney CA, Stonehouse RG, Rossnagel BG, Eckstein PE, Scoles GJ, Zatorski T, Beattie AD, Chong J (2011) Mapping of the oat crown rust resistance gene Pc91. Theor Appl Genet 122:317–325
McIntosh RA, Wellings CR, Park RF (1995) Wheat rusts. An atlas of resistance genes. 200pp. Melbourne: CSIRO
McIntosh RA (1988) The role of specific genes in breeding for durable stem rust resistance in wheat and triticale. Breeding strategies for resistance to the rusts of wheat. CIMMYT, Mexico, pp.1–9
McKenzie RIH, Green GJ (1965) Stem rust resistance in oats. I. The inheritance of resistance to race 6AF in six varieties of oats. Can J Genet Cytol 7:268–274
McMullen MS, Sorrells ME (1991) Registration of ‘Newdak’ oat. Crop Sci 31:1384
McMullen MS, Doehlert DC, Miller JD (1997) Registration of ‘Jerry’ oat. Crop Sci 37:1015–1016
McMullen MS, Doehlert DC, Miller JD (2005) Registration of ‘Hifi’ oat. Crop Sci 45:1664
McNish IG, Zimmer CM, Susko AQ, Heuschele DJ, Tiede T, Case AJ, Smith KP (2020). Mapping crown rust resistance at multiple time points in elite oat germplasm. Plant Genome. 13(1):e20007. doi: https://doi.org/10.1002/tpg2.20007. Epub 2020 Apr 2. PMID: 33016637
Mitchell Fetch JW, Kibite S, Chong J, Clayton GW, Fetch TG, Menzies JG, Turkington TK (2009a) Lee Williams oat. Can J Plant Sci 89:665–669
Mitchell Fetch JW, Kibite S, Clayton GW, Turkington TK, Chong J, Fetch TG, Haber SM, Menzies JG, Ames N (2009b) Lu oat. Can J Plant Sci 89:671–675
Mitchell Fetch JW, Brown PD, Ames N, Chong J, Fetch TG, Haber SM, Menzies JG, Tekauz A, Townley-Smith TF, Stadnyk KD (2011) Summitt oat. Can J Plant Sci 91:787–791
Mitchell Fetch JW, Stanley K, Entz M, Fox SL, Spaner D, Kirk A, Vaisman I, Brown PD, Ames N, Chong J, Fetch TG, Haber SM, McCartney CA, Menzies JG, Tekauz A, Townley-Smith TF, Nilsen KT, Hamilton KD, Green DA (2021) AAC Oravena oat. Can J Plant Sci. https://doi.org/10.1139/CJPS-2021-0036
Montilla-Bascón G, Rispail N, Sánchez-Martín J, Rubiales D, Mur LA, Langdon T, Howarth CJ, Prats E (2015) Genome-wide association study for crown rust (Puccinia coronata f. sp. avenae) and powdery mildew (Blumeria graminis f. sp. avenae) resistance in an oat (Avena sativa) collection of commercial varieties and landraces. Front Plant Sci 6:103
Moore JW, Herrera-Foessel S, Lan C, Schnippenkoetter W, Ayliffe M, Huerto-Espino J, Lillemo M, Viccars L, Periyannan S, Kong X, Spielmeyer W, Talbot M, Bariana H, Patrick JW, Dodds P, Amil S, Lagudah E (2015) A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat Genet. https://doi.org/10.1038/ng.3439
Mundt CC (2002) Use of multiline cultivars and cultivar mixtures for disease management. Annu Rev Phytopathol 40:381–410
Mundt CC, Browning JA (1985) Genetic diversity and cereal rust management. In: Roelfs AP, Bushnell WR (eds) The cereal rusts Volume II diseases, Distribution, Epidemiology and Control. Academic Press, Orlando
Murphy HC (1935) Effects of crown rust infection on yield and water requirement of oats. J Agric Res 50:387–411
Murphy HC, Stanton TR, Stevens H (1937) Breeding winter oats resistant to crown rust, smut and cold. J Am Soci Agron 29:622–637
Nazareno ES, Li F, Smith M, Park RF, Kianian SF, Figueroa M (2018) Puccinia coronata f. sp. avenae: a threat to global oat production. Mol Plant Pathol 19:1047–1060
Oates JD, Burdon JJ, Brouwer JB (1983) Interactions between Avena and Puccinia species. II. The pathogens: Puccinia coronata Cda and P. graminis Pers. f. sp. avenae Eriks. And Henn. J Appl Ecol 20:585–596
O’Donoughue LS, Sorrells ME, Tanksley SD, Autrique E, Deynze AV, Kianian SF, Phillips RL, Wu B, Rines HW, Rayapati PJ, Lee M (1995) A molecular linkage map of cultivated oat. Genome 38:368–380
Oliver RE, Tinker NA, Lazo GR, Chao S, Jellen EN, Carson ML, Rines HW, Obert DE, Lutz JD, Shackelford I, Korol AB (2013) SNP discovery and chromosome anchoring provide the first physically-anchored hexaploid oat map and reveal synteny with model species. PLoS ONE 8:e58068
Osler RD, Hayes HK (1953) Inheritance studies in oats with particular reference to the Santa Fe type of croiwn rust resistance. Agron J 45:49–53
Paczos-Grzeda E, Sowa S (2019) Virulence structure and diversity of Puccinia coronata f. sp. avenae P. Syd. & Syd. In Poland during 2013 to 2015. Plant Dis 103:1559–1564
Palloix A, Ayme V, Moury B (2009) Durability of plant major resistance genes to pathogens depends on the genetic background, experimental evidence and consequences for breeding strategies. New Phytol 183:190–199
Park RF (2007) Stem rust of wheat in Australia. Aust J Agric Res 58:558–566
Park RF (2013) New oat crown rust pathotype with virulence for Pc91. Cereal Rust Rep 11:8–10
Park RF (2015) Long term surveys of pathogen populations underpin sustained control of the rust diseases of wheat in Australia. J Royal Soci New South Wales 148:15–27
Park RF, McIntosh RA (1994) Adult plant resistances to Puccinia recondita f. sp. tritici in wheat. NZ J Crop Hortic Sci 22:151–158
Park RF, Meldrum SM, Oates JD (2000) Recent pathogenic changes in the leaf (brown) rust pathogen of wheat and the crown rust pathogen of oats in Australia in relation to host resistance. Acta Phytopathologica Et Entomol Hungarica 35:387–394
Park RF, Mohler V, Nazari K, Singh D (2014) Characterisation and mapping of gene Lr73 conferring seedling resistance to Puccinia triticina in common wheat. Theor Appl Genet 127:2041–2049
Park RF, Bansal U, Bariana HS, Singh D (2020) Rust resistance genotypes and expected rust responses of Australian common wheat, durum wheat and triticale vvarieties. Cereal Rust Report, 17(3) https://www.sydney.edu.au/science/our-research/research-areas/life-and-environmental-sciences/cereal-rust-research/rust-reports.html
Patterson FL, Schafer JF (1977). Purdue University Agriculture Experimental Station Journal, paper 6683
Portyanko VA, Hoffman DL, Lee M, Holland JB (2001) A linkage map of hexaploid oat based on grass anchor DNA clones and its relationship to other oat maps. Genome 44:249–265
Portyanko VA, Chen G, Rines HW, Phillips RL, Leonard KJ, Ochocki GE, Stuthman DD (2005) Quantitative trait loci for partial resistance to crown rust, Puccinia coronata, in cultivated oat, Avena sativa L. Theor Appl Genet 111:313–324
Rio S, Gallego-Sánchez L, Montilla-Bascón G, Canales FJ, Isidro y Sánchez J, Prats E (2021) Genomic prediction and training set optimization in a structured Mediterranean oat population. Theor Appl Genet 134:3595–3609
Roelfs AP (1982) The effects of barberry eradication on stem rust in the United States. Plant Dis 66:177–181
Rooney WL, Rines HW, Phillips RL (1994) Identification of RFLP markers linked to crown rust resistance genes Pc91 and Pc92 in oat. Crop Sci 34:940–944
Rothman PG (1984) Registration of four stem rust and crown rust resistant oat germplasm lines. Crop Sci 24:1217–1218
Cruz RP, Federizzi LC, Milach SC (2001) Genetics of crown rust resistance in oat. Pesquisa Agropecuária Brasileira 36(9):1127–32
Rubiales D, Niks RE (2000) Combination of mechanisms of resistance to rust fungi as a strategy to increase durability. Options Méditérr 40:333–339
Sánchez-Martín J, Rubiales D, Sillero JC, Prats E (2012) Identification and characterization of sources of resistance in Avena sativa, A. trigose and A. trigose germplasm against a pathotype of Puccinia coronata f. sp. avenae with virulence the Pc94 resistance gene. Plant Pathol 61:315–322
Sánchez-Martín J, Rubiales D, Flores F, Emeran AA, Shtaya MJY, Sillero JC, Allagui MB, Prats E (2014) Adaptation of oat (Avena sativa) cultivars to autumn sowings in Mediterranean environments. Field Crop Res 156:111–122
Satheeskumar S, Sharp PJ, Lagudah ES, McIntosh RA, Molnar SJ (2011) Genetic association of crown rust resistance gene Pc68, storage protein loci, and resistance gene analogues in oats. Genome 54:484–497
Savile DBO (1984) Taxonomy of the cereal rust fungi. In The Cereal Rusts, Vol. I, Origins, Specificity, Structure, and Physiology, pp. 79–112. Eds W.R. Bushnell and A.P. Roelfs. New York: Academic Press Inc
Sebesta J, Zwatz B, Roderik H, Corazza L, Manisterski J, Stojanovic S (2003) Incidence of crown rust and virulence of Puccinia coronata Cda f. sp. avenae Eriks. And the effectiveness of Pc genes for resistance in Europe, Middle East and North Africa. Arch Phytopath Plant Protect 36:179–194
Simons MD (1961) Testing oats in the field with specific races of crown rust. Proceed Iowa Acad Sci 68:119–123
Simons MD (1975) Heritability of field resistance to the oat crown rust fungus. Phytopathology 65:324–328
Simons MD (1985) Crown rust. In: Roelfs AP, Bushnell WR (eds) The Cereal Rusts Volume II Diseases, Distribution, Epidemiology, and Control. Academic Press Inc., Orlando USA, pp 142–172
Simons MD, Michel LJ (1968) Oat maturity and crown rust response 1. Crop Sci 8:254–256
Simons M, Sadanaga K, Murphy HC (1959) Inheritance of resistance to strains of diploid and tetraploid species of oats to races of the crown rust fungus. Phytopathology 49:257–259
Simons MD, Murphy HC (1955) A comparison of certain combinations of oat varieties as crown rust differentials. US Department of Agriculture Technical Bulletin 112: 1–22
Simons MD, Murphy HC (1961). Oat diseases. In: Oat and Oat Improvement (Coffman FA, ed), pp. 330–390. Agronomy 8, The American Society of Agronomy: Madison Wisconsin USA
Simons, M.D., Martens, J.W., McKenzie, R.I.H., Nishiyama, I., Sadanaga, K., Sebesta, J., and Thomas, H. (1978) Oats: a standardized system of nomenclature for genes and chromosomes and catalog of genes governing characters. USDA Agriculture Handbook Number 509, 43pp
Simons MD (1970) Crown rust of oats and grasses. Monograph Number 5, The American Phytopathological Society . The Heffernan Press Inc: Worcester Massechusets USA. 47 pp
Singh RP, Huerta-Espino J, Bhavani S, Hererra-Foessel SA, Singh D, Singh PK, Velu G, Mason RE, Jin Y, Njau P, Crossa J (2011) Race non-specific resistance to rust diseases in CIMMYT spring wheats. Euphytica 179:175–186
Sowa S, Paczos-Grzeda E (2020) Identification of molecular markers for the Pc39 gene conferring resistance to crown rust in oat. Theoret Appl Genet 133:1081–1094
Steuernagel B, Vrána J, Karafiátová M, Wulff BBH, Dolezel J (2017) Rapid gene isolation using MutChromSeq. Methods Mol Biol 1659:231–243
Stuthman DD, Moore MB (1970) A unique approach to crown rust resistance. Oat Newsletter 21:23–24
Sunstrum FG, Bekele WA, Wight CP, Yan W, Chen Y, Tinker NA (2019) A genetic linkage map in southern-by-spring oat identifies multiple quantitative trait loci for adaptation and rust resistance. Plant Breed 138:82–94
Suttie JM, Reynolds SG (2004) Fodder oats: a world overview, vol 33, Plant production and protection series. Food and Agriculture Organization of the United Nations, Rome
Tinker NA, Kilian A, Wight CP, Heller-Uszynska K, Wenzl P, Rines HW, Bjørnstad A, Howarth CJ, Jannink JL, Anderson JM, Rossnagel BG (2009) New DArT markers for oat provide enhanced map coverage and global germplasm characterization. BMC Gen 10:1–22
Tinker NA, Chao S, Lazo GR, Oliver RE, Huang YF, Poland JA, Jellen EN, Maughan PJ, Kilian A, Jackson EW (2014) A SNP genotyping array for hexaploid oat. The Plant Genome. https://doi.org/10.3835/plantgenome2014.03.0010
Tinker NA, Bekele WA, Hattori J (2016) Haplotag: software for haplotype-based genotyping-by-sequencing analysis. G3: Genes, Genomes, Genetics 6:857–863
Toporowska J, Sowa S, Kiklian A, Koroluk A, Paczos-Grzeda E (2021) Discovery and chromosomal location of a highly effective oat crown rust resistance gene Pc50-5. Int J Molec Sci 22:11183
Upadhyaya YM, Baker EP (1960) Studies on the mode of inheritance of Hajira type stem rust resistance and Victoria type crown rust resistance as exhibited in crosses involving the oat variety Garry. Proc Linnean Soc NSW 85:157–179
Upadhyaya YM, Baker EP (1962) Studies on the inheritance of rust resistance in oats. IE The mode of inheritance of crown rust resistance in the varieties Landhafer, Santa Fe, Mutica Ukraine, Trispernia, and Victoria in their crosses with susceptible varieties. Proc Linnean Soc NSW 87:200–219
Upadhyaya YM, Baker EP (1965) Studies of the inheritance of rust resistance in oats. III. Genetic diversity in the varieties Landhafer, Santa Fe, Mutica Ukraine, Trispernia and Victoria for crown rust resistance. Proceed Linnean Soci New Soouth Wales 90:129–151
Van Niekerk BD, Pretorius ZA, Boshoff WHP (2001) Pathogenic variability of Puccinia coronata f. sp. avenae on oat in South Africa. Plant Dis 85:1085–1090
Wahl I (1970) Prevalence and geographic distribution of resistance to crown rust in Avena sterilis. Phytopathology 60:746–749
Waterhouse WL (1952) Australian rust studies. IX. Physiologic race determinations and surveys of cereal rusts. Proceed Linnnean Soci NSW 77:210–258
Watson IA, Singh D (1952) The future for rust resistant wheat in Australia. J Austral Instit Agricult Sci 18:190–197
Welsh JN, Carson RB, Cherewick WJ, Hagborg WAF, Peturson B, Wallace HAH (1953). Oat varieties- past and present. Canada Department of Agriculture, Ottawa, Pub 891
Wight CP, Tinker NA, Kianian SF, Sorrells ME, O’Donoughue LS, Hoffman DL, Groh S, Scoles GJ, Li CD, Webster FH, Phillips RL, Rines HW, Livingston SM, Armstrong KC, Fedak G, Molnar SJ (2003) A molecular marker map in “Kanota” x “Ogle” hexaploid oat (Avena spp.) enhanced by additional markers and a robust framework. Genome 46:28–47
Wight CP, O’Donoughue LS, Chong J, Tinler NA, Molnar SJ (2004) Discovery, localization and sequence characterization of molecaular markers for the crown rust resistance genes Pc38, Pc39 and Pc48 in cultivated oat (Avena sativa L.). Mol Breed 14:349–361
Wilson WA, McMullen MS (1997) Dosage dependent genetic suppression of oat crown rust resistance gene Pc62. Crop Sci 37:1699–1705
Winkler LR, Bonman JM, Chao S, Yimer BA, Bockelman H, Esvelt Klos K (2016) Population structure and genotype–phenotype associations in a collection of oat landraces and historic cultivars. Front Plant Sci. https://doi.org/10.3389/fpls.2016.01077
Wong LSL, McKenzie RIH, Harder DE, Martens JW (1983) The inheritance of resistance to Puccinia coronata and of floret characters in Avena sterilis. Can J Genet Cytol 25:329–335
Worland AJ, Law CN (1986) Genetic analysis of chromosome 2D of wheat. I: The location of genes affecting height, day-I insensitivity, hybrid dwarfism and yellow-rust resistance. Zeitschrift Für Pflanzenzüchtung 96:331–345
Yan H, Bekele WA, Wight CP, Peng Y, Langdon T, Latta RG, Fu YB, Diederichsen A, Howarth CJ, Jellen EN, Boyle B (2016) High-density marker profiling confirms ancestral genomes of Avena species and identifies D-genome chromosomes of hexaploid oat. Theor Appl Genet 129:2133–2149
Yan W, McElroy A, Fregeau-Reid J, Martin R, Pageau D, Xue A, Jakubinek K, deHaan B, Thomas S, Sibbit D, Cumminskey A (2017a) AAC Almonte oat. Can J Plant Sci 97:943–949
Yan W, Fregeau-Reid J, Martin R, Pageau D, Xue A, Jakubinek K, deHaan B, Thomas S, Hayes M, Cumminskey A (2017b) AAC Kolosse oat. Can J Plant Sci 97:352–355
Yan W, Fregeau-Reid J, Martin R, Pageau D, Xue A, Jakubinek K, deHaan B, Thomas S, Hayes M, Cumminskey A (2017c) AAC Noranda oat. Can J Plant Sci 97:928–934
Yu GX, Wise RP (2000) An anchored AFLP- and retrotransposon-based map of diploid Avena. Genome 43:736–749
Zhang J, Zhang P, Dodds P, Lagudah E (2020) How Target-Sequence Enrichment and Sequencing (TEnSeq) pipelines have catalyzed resistance gene cloning in the wheat-rust pathosystem. Front Plant Sci. https://doi.org/10.3389/fpls.2020.00678
Zhang J, Hewitt TC, Boshoff WHP, Dundas I, Upadhyaya N, Li J, Patpour M, Chandramohan S, Pretorius ZA, Hovmoller M, Schnippenkoetter W, Park RF, Mago R, Periyannan S, Bhatt D, Hoxha S, Chakraborty S, Luo M, Dodds P, Steuernagel B, Wulff BBH, Ayliffe M, McIntosh RA, Zhang P, Lagudah E (2021) A recombined Sr26 and Sr61 disease resistance gene stack in wheat encodes unrelated NLR genes. Nat Commun 12:1–8
Zhao J, Kebede AZ, Bekele WA, Menzies JG, Chong J, Mitchell Fetch JW, Tinker NA, Beattie AD, Peng YY, McCartney CA (2020a) Mapping of the oat crown rust resistance gene Pc39 relative to single nucleotide polymorphism markers. Plant Dis 104:1507–1513
Zhao J, Kebede AZ, Menzies JG, Paczos-Grzeda E, Chong J, Mitchell Fetch JW, Beattie AD, Peng YY, McCartney CA (2020b) Chromosomal location of the oat crown rust resistance gene Pc98 in cultivated oat (Avena sativa L.). Theoret Appl Genet 133:1109–1122
Zhu S, Kaeppler H (2003) Identification of quantitative trait loci for crown rust in oat line MAM17-5. Crop Sci 43:358–366
Zhu S, Leonard KJ, Kaeppler HF (2003) Quantitative trait loci associated with seedling resistance to isolates of Puccinia coronata in oat. Phytopathology 93:860–866
Zwer PK, Park RF, McIntosh RA (1992) Wheat stem rust in Australia 1969–1985. Aust J Agric Res 43:399–431
Acknowledgements
We thank Mr Matthew Williams, Ms Bethany Clark and Ms Margerita Pietilainen for technical assistance.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. Financial support from Judith and David Coffey and family, the Grains Research and Development Corporation (GRDC: DAS00133, UOS1707-003RTX, UOS2104-001RTX) and the University of Sydney is gratefully acknowledged. Some of the unpublished research reported on was undertaken as part of a long running program on national cereal rust surveillance, conducted at the University of Sydney since 1921. EP is funded by Spanish Ministry of Science and Innovation [PID2019-104518RB-100], (AEI/FEDER, UE) and regional government through the AGR-253 group, the European Regional and Social Development Funds.
Author information
Authors and Affiliations
Contributions
RFP conceived the review and wrote the first draft of the manuscript. All authors read, contributed content, commented on, and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Additional information
Communicated by Hermann Buerstmayr.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Park, R.F., Boshoff, W.H.P., Cabral, A.L. et al. Breeding oat for resistance to the crown rust pathogen Puccinia coronata f. sp. avenae: achievements and prospects. Theor Appl Genet 135, 3709–3734 (2022). https://doi.org/10.1007/s00122-022-04121-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-022-04121-z