Abstract
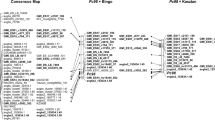
Crown rust, which is caused by Puccinia coronata f. sp. avenae, P. Syd. & Syd., is the most destructive disease of cultivated oats (Avena sativa L.) throughout the world. Resistance to the disease that is based on a single gene is often short-lived because of the extremely great genetic diversity of P. coronata, which suggests that there is a need to develop oat cultivars with several resistance genes. This study aimed to identify amplified fragment length polymorphism AFLP markers that are linked to the major resistance gene, Pc68, and to amplify the F6 genetic map from Pc68/5*Starter × UFRGS8. Seventy-eight markers with normal segregation were discovered and distributed in 12 linkage groups. The map covered 409.4 cM of the Avena sativa genome. Two AFLP markers were linked in repulsion to Pc68: U8PM22 and U8PM25, which flank the gene at 18.60 and 18.83 centiMorgans (cM), respectively. The marker U8PM25 is located in the linkage group 4_12 in the Kanota × Ogle reference oat population. These markers should be useful for transferring Pc68 to genotypes with good agronomic characteristics and for pyramiding crown rust resistance genes.


Similar content being viewed by others
References
Barzen E, Stahl R, Fuchs E, Borchardt DC, Salamini F (2004) Development of coupling-repulsion phase SCAR markers diagnostic for the sugar beet Rr1 allele conferring resistance to rhizomania. Mol Breed 3:231–238
Brogin RL (2005) Mapeamento de genes de resistência à ferrugem e de QTLs envolvidos à Septoriose em soja. Dissertation. Universidade de São Paulo, Piracicaba, 93 p
Bryan GJ, Stephenson P, Collins A, Kirby J, Smith JB, Gale MD (1999) Low levels of DNA sequence variation among adapted genotypes of hexaploid wheat. Theor Appl Genet 99:192–198
Bush AL, Wise RP (1996) Crown rust resistance loci on linkage groups 4 and 13 in cultivated oat. J Hered 87:427–432
Cervera MT, Ruiz-Garcia L, Martinez-Zapater JM (2002) Analysis of DNA methylation in Arabidopsis thaliana based on methylation-sensitive AFLP markers. Mol Genet Genomics 268:543–552
Chen CX, Wang ZL, Yang DE, Ye CJ, Zhao YB, Jin DM, Weng ML, Wang B (2004) Molecular tagging and genetic mapping of the disease resistance gene RppQ to southern corn rust. Theor Appl Genet 108:945–950
Chen G, Chong J, Gray M, Prashar S, Procunier JD (2006) Identification of single-nucleotide polymorphisms linked to resistance gene Pc68 to crown rust in cultivated oat. Can J Plant Pathol 28:214–222
Cheng DW, Armstrong KC, Tinker N, Wight C, He S, Lybaert A, Fedak G, Molnar SJ (2002) Genetic and physical mapping of Lrk10-like receptor kinase sequences in hexaploid oat (Avena sativa L.). Genome 45:100–109
Chong J (2000) Incidence and virulence of Puccinia coronata in Canada from 1996 to 1998. Can J Plant Pathol 22:99–109
Chong J, Howes NK, Brown PD, Harder DE (1994) Identification of the stem rust resistance gene Pg9 and its association with crown rust resistance and endosperm proteins in ‘Dumont’ oat. Genome 37:440–447
Chong J, Leonard KJ, Salmeron JJ (2000) A North American system of nomenclature for Puccinia coronata f.sp. avenae. Plant Dis 84:580–585
Cruz CD, Schuster I (2004) GQMOL: aplicativo computacional para análise de dados moleculares e de suas associações com caracteres quantitativos [Online]. Available at http://www.ufv.br/dbg/gqmol/gqmol.htm
Duble CM, Quint M, Melchinger AE (2003) Saturation of two chromosome regions conferring resistance to SCMV with SSR and AFLP markers by targeted BSA. Theor Appl Genet 106:485–493
Giovanni C, Dell’Orco P, Bruno A, Ciccarese F, Lotti C, Ricciardi L (2004) Identification of PCR-based markers (RAPD, AFLP) linked to a novel powdery mildew resistance gene (ol-2) in tomato. Plant Sci 166:41–48
Golas TM, Sikkem A, Gros J, Feron RMC, Van den Berg RG, Van der Weerden GM, Mariani C, Allefs J (2010) Identification of a resistance gene Rpi-dlc1 to Phytophthora infestans in European accessions of Solanum dulcamara. Theor Appl Genet 120:797–808
Haley SD (1994) Selection for monogenic pest resistance traits with coupling and repulsion-phase RAPD markers. Crop Sci 34:1061–1066
Jackson EJ, Avant JB, Overturf KE, Bonman JM (2006) A quantitative assay of Puccinia coronata f. sp. avenae DNA in Avena sativa. Plant Dis 90:629–636
Jackson EJ, Obert DE, Menz M, Hu G, Bonman JM (2007) Qualitative and quantitative trait loci conditioning resistance to Puccinia coronata pathotypes NQMG and LGCG in the oat (Avena sativa L.) cultivars Ogle and TAM O-301. Theor Appl Genet. doi:10.1007/s00122-007-0687-x
Johnston MR, Carsten LD, Douglas L, Sands DC (2000) Epidemic development and virulence in 1995–1998 of Puccinia coronata, a potential biocontrol agent of wild oats on San Clemente Island. Biol Control 17:250–257
Kamei A, Tsuro M, Kubo N, Hayashi T, Wang N, Fujimura T, Hirai M (2010) QTL mapping of clubroot resistance in radish (Raphanus sativus L.). Theor Appl Genet 120:1021–1027
Kaur S, Cogan NOI, Ye G et al (2009) Genetic map construction and QTL mapping of resistance to blackleg (Leptosphaeria maculans) disease in Australian canola (Brassica napus L.) cultivars. Theor Appl Genet 120:71–83
Koeyer D, Orr W, Lybaert A, Deyl J, Chenier C, Tinker N, Mcelroy A, Chong J, Molnar S (2000) Scar markers linked to the Pc68 resistance allele are an effective tool for selection. In: Cross RJ (ed) International oats conference. New Zealand Institute for Crop and Food Research Limited, Christchurch, New Zealand, pp 137–140
Leonard KJ (2002) Oat lines with effective adult plant resistance to crown rust. Plant Dis 86:593–598
Leonard KJ (2003) Regional frequencies of virulence of oat crown rust in the United States from 1990 through 2000. Plant Dis 87:1301–1310
Leonard KJ, Martinelli JA (2005) Virulence of oat crown rust in Brazil and Uruguay. Plant Dis 89:802–808
Li CD, Rossnagel BG, Scoles GJ (2000) The development of oat microsatellite markers and their use in identifying relationships among Avena species and oat cultivars. Theor Appl Genet 101:1259–1268
Liu XH, Tan ZB, Rong TZ (2009) Molecular mapping of a major QTL conferring resistance to SCMV based on immortal RIL population in maize. Euphytica 167:229–235
Locatelli AB, Federizzi LC, Milach SCK, Wight CP, Molnar SJ, Chapados JT, Tinker NA (2006) Loci affecting flowering time in oat under short-day conditions. Genome 49:1528–1538
Martinelli JA, Federizzi LC, Bennedeti AC (1994) Redução do rendimento de grãos de aveia em função da severidade da ferrugem da folha. Summa Phytopathologica 20:116–118
Matsuo K, Silke S, Georgiev O, Marti P, Giovannini N, Rungger D (1998) An embryonic demethylation mechanism involving binding of transcription factors to replicating DNA. EMBO J 17:1446–1453
McCallum BD, Fetch T, Chong J (2007) Cereal rust control in Canada. Aust J Agr Res 58:639–647
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis—a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Nat Acad Sci USA 88:9828–9832
Nikaido A, Yoshimaru H, Tsumura Y, Suyama Y, Murai M, Nagasaka K (1999) Segregation distortion for AFLP markers in Cryptomeria japonica. Genes Genetic Syst 74:55–59
O’Donoughue LS, Kianian SF, Rayapati PJ et al (1995) A molecular linkage map of cultivated oat. Genome 38:368–380
O’Donoughue LS, Chong J, Wight CP, Fedak G, Molnar SJ (1996) Localization of stem rust resistance genes and associated molecular markers in cultivates oat. Phytopathology 86:719–727
Orr W, Koeyer D, Chenier C, Tinker N, Molnar SJ (1999) SCAR markers for rust resistance genes Pc 68, Pg3, and Pg9 designed for marker assisted selection in oats. Plant & Animal Genome VII Conference, San Diego, CA, p 345
Penner GA, Chong J, Levesquelemay M, Molnar SJ, Fedak G (1993a) Identification of a Rapd marker linked to the oat stem rust gene-Pg3. Theoret Appl Genetics 85:702–705
Penner GA, Chong J, Wight CP, Molnar SJ, Fedak G (1993b) Identification of an Rapd marker for the crown rust resistance gene Pc68 in oats. Genome 36:818–820
Portyanko VA, Hoffman DL, Lee M, Holland JB (2001) A linkage map of hexaploid oat based on grass anchor DNA clones and its relationship to other oat maps. Genome 44:249–265
Randhawa HS, Popovic Z, Menzies J, Knox R, Fox S (2009) Genetics and identification of molecular markers linked to resistance to loose smut (Ustilago tritici) race T33 in durum wheat. Euphytica 169:151–157
Renganayaki K, Fritz AK, Sadasivam S et al (2002) Mapping and progress toward map-based cloning of Brown planthopper biotype-4 resistance gene introgressed from Oryza officinalis into cultivated Rice, O. sativa. Crop Sci 42:2112–2117
Roy A, Frascaria N, Mackay J, Bousquet J (1992) Segregating random amplified polymorphic DNAs (RAPDs) in Betula alleghaniensis. Theor Appl Genet 85:173–180
Rubiales D, Niks RE (2000) Combination of mechanisms of resistance to rust fungi as a strategy to increase durability. Options Mediterraneennes 40:333–339
Silva HD, Vencovsky R (2002) Testes múltiplos e associação de marcas associadas a QTL’s. Sci Agr 99:755–762
Tan YD, Wan CL, Zhu YF, Lu C, Xiang ZH, Deng HW (2001) An amplified fragment length polymorphism map of the silkworm. Genetics 157:1277–1284
Testolin R, Huang WG, Cipriani G, Retamales J (2001) Towards a linkage map in kiwifruit (Actinidia chinensis Planch.) based on microsatellites and satured with AFLP markers. Acta Horticultural 498:79–84
Tinker NA, Kilian A, Wight CP, Heller-Uszynska K, Wenzl P, Rines HW, Bjornstad A, Howarth CJ, Jannink J-L, Anderson JM, Rossnagel BG, Stuthman DD, Sorrells ME, Jackson EW, Tuvesson S, Kolb FL, Olsson O, Federizzi LC, Carson ML, Ohm HW, Molnar SJ, Scoles GJ, Eckstein PE, Bonman JM, Ceplitis A, Langdon T (2009) New DArT markers for oat provide enhanced map coverage and global germplasm characterization. BMC Genomics 10:39. doi:10.1186/1471-2164-10-39
Vieira EA, Carvalho FIF, Chaves MS, ACde Oliveira, Benin G, Hartwig I, Silva JAG, Bertan I, Martins AF, Martins LF (2007) Virulence variability of Puccinia coronata f. sp. avenae isolates collected in three counties from Rio Grande do Sul. Plant Dis 91:66–70
Vogl C, Xu SZ (2000) Multipoint mapping of viability and segregation distorting loci using molecular markers. Genetics 155:1439–1447
Vos P, Hogers R, Bleeker M et al (1995) Aflp—a new technique for DNA-fingerprinting. Nucleic Acids Res 23:4407–4414
Wang YH, Thomas CE, Dean RA (1997) A genetic map of melon (Cucumis melo L.) based on amplified fragment length polymorphism (AFLP) markers. Theor Appl Genet 95:791–798
Weng Y, Lazar MD (2002) Amplified fragment length polymorphism- and simple sequence repeat-based molecular tagging and mapping of greenbug resistance gene Gb3 in wheat. Plant Breed 121:218–223
Werner K, Friedt W, Ordon F (2005) Strategies for pyramiding resistance genes against the barley yellow mosaic virus complex (BaMMV, BaYMV, BaYMV-2). Mol Breed 16:45–55
Wight CP, Tinker NA, Kianian SF et al (2003) A molecular marker map in ‘Kanota’ × ‘Ogle’ hexaploid oat (Avena spp.) enhanced by additional markers and a robust framework. Genome 46:28–47
Wong LSL, Mckenzie RIH, Harder DE, Martens JW (1983) The Inheritance of Resistance to Puccinia-Coronata and of Floret Characters in Avena-Sterilis. Can J Genet Cytol 25:329–335
Yu J, Herrmann M (2006) Inheritance and mapping of a powdery mildew resistance gene introgressed from Avena macrostachya in cultivated oat. Theor Appl Genet 113:429–437
Zar JH (2009) Biostatistical analysis. Prentice Hall, New Jersey, USA
Zhu S, Kaeppler HF (2003) Identification of quantitative trait loci for resistance to crown rust in oat line MAM17-5. Crop Sci 43:358–366
Acknowledgment
The authors thank to the Brazilian Council for Scientific and Technological Development (CNPq) for the scholarship provided to the first author.
Author information
Authors and Affiliations
Corresponding author
Additional information
Part of the Master of Science dissertation of Franceli R. Kulcheski.
Rights and permissions
About this article
Cite this article
Kulcheski, F.R., Graichen, F.A.S., Martinelli, J.A. et al. Molecular mapping of Pc68, a crown rust resistance gene in Avena sativa . Euphytica 175, 423–432 (2010). https://doi.org/10.1007/s10681-010-0198-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-010-0198-8