Abstract
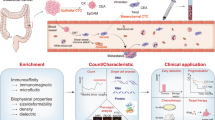
Colorectal cancer (CRC) is one of the main causes of cancer-related morbidity and mortality across the globe. Although serum biomarkers such as carcinoembryonic antigen (CEA) and carbohydrate antigen 19–9 (CA-199) have been prevalently used as biomarkers in various cancers, they are neither very sensitive nor highly specific. Repeated tissue biopsies at different times of the disease can be uncomfortable for cancer patients. Additionally, the existence of tumor heterogeneity and the results of local biopsy provide limited information about the overall tumor biology. Against this backdrop, it is necessary to look for reliable and noninvasive biomarkers of CRC. Circulating tumor cells (CTCs), which depart from a primary tumor, enter the bloodstream, and imitate metastasis, have a great potential for precision medicine in patients with CRC. Various efficient CTC isolation platforms have been developed to capture and identify CTCs. The count of CTCs, as well as their biological characteristics and genomic heterogeneity, can be used for the early diagnosis, prognosis, and prediction of treatment response in CRC. This study reviewed the existing CTC isolation techniques and their applications in the clinical diagnosis and treatment of CRC. The study also presented their limitations and provided future research directions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Despite recent advancements, colorectal cancer (CRC) is still one of the leading causes of cancer-related deaths worldwide. With 1.65 million new cases and about 835,000 deaths annually, CRC is ranked as the third most prevalent cancer globally and second in terms of mortality [1, 2]. The 5-year survival rate of CRC is as high as 90% when diagnosed at stage I or II, while it decreases to 13% when diagnosed at stage III or IV [1]. Although some patients can be cured by radical surgery, patients with heavy tumor burden, neurovascular invasion, and even distant metastasis exhibit an extremely poor prognosis. Carcinoembryonic antigen (CEA) is an established biomarker with reported efficacy for the treatment and monitoring of human cancers. It is the only biomarker recommended for monitoring and evaluating the prognosis of CRC in the current treatment guidelines [3]. Several blood tumor markers, including CA72-4 and CA19-9, are known to have limited sensitivity and specificity [4]. The identification of novel noninvasive biomarkers in plasma for diagnostic and prognostic purposes is thus of critical importance.
Tumor biopsies are now regarded as the “gold standard” in the diagnosis, prognosis, and prediction of therapy response in treating patients with cancers [5]. However, cancer cells are highly heterogeneous at the single-cell level, and biopsy specimens containing a small amount of tumor tissue may not represent entire cancer as a whole [6]. Besides, in the course of disease progression, cancers may have escape mutations and epigenetic alterations as a result of dynamic molecular changes [7, 8], which may also contribute to the cancer progression and resistance to therapy. Nevertheless, it is impractical to perform repeat tumor tissue biopsies during the patient’s illness. Just as colonoscopy, while tumor biopsy can be used to examine the entire colon and diagnose CRC, it is also an invasive technique that can cause complications such as bleeding, perforation, and cardiovascular accidents [9,10,11]. Not only are new biomarkers needed to evaluate the prognosis, treatment, and recurrence of CRC, but the search for these biomarkers with new minimally invasive techniques is also essential.
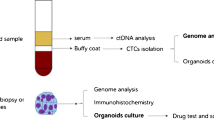
Compared with traditional tissue biopsy, liquid biopsy is a revolutionary technique that presents a static snapshot of the tumor and offers unique and enormous advantages. It provides information on the cancer burden in “real time” and unveils the evolution and heterogeneity of the disease over time [12]. The main targets of liquid biopsy include circulating tumor cells (CTCs), circulating tumor DNA (ctDNA), and extracellular vesicles. Tumor cells that enter the bloodstream from primary or metastatic cancers are called CTCs, the concept of which was first proposed by Ashworth at the end of the nineteenth century [13,14,15]. The abundance of CTCs in peripheral blood is incredibly low, with approximately 1 CTC per 106–107 white blood cells. Furthermore, the frequency of CTCs is even lower when solid tumors are confined to local growth [16], making the detection and isolation of CTCs extremely difficult and challenging. However, these rare cells can provide a wealth of tumor-related clinical information. Prompting the development of more sensitive platforms for isolating, enriching, and investigating CTCs can provide a noninvasive alternative for diagnosing, monitoring treatment, and evaluating the prognosis of malignant tumors. In-depth analysis of CTCs provides useful information for investigating the molecular characteristics of cancers, early detection of primary and metastatic lesions, and personalized treatment, mainly in terms of prognostic evaluation, stratified targeting of patients and real-time monitoring of treatment effects, identification of treatment targets, and drug resistance mechanisms [17]. This review presented the development of CTC isolation methods and their clinical applications in CRC and discussed their feasibility as potential biomarkers for the diagnosis and prognosis of CRC.
Overview of the biological and clinical significance of CTCs
During tumor growth, individual or clusters of tumor cells detach from the primary site and pass through the blood vessels, eventually entering the circulation and forming “circulating tumor cells.” Moreover, tumor cells may move through channels formed by the proteolysis of other tumor cells or pre-existing channels formed by tissue structures [15]. In fact, most CTCs released in the bloodstream die in an early stage due to the combined effects of mechanical and environmental factors such as shear stress, oxidative stress, and immune system response [18]. As a result, the half-life of CTCs in the cycle is noticeably short, usually between 1 and 2 h [19], and only a few drug-resistant cells can extravasate and spread. CTCs must undergo a series of adaptations to survive in the changing environment. The most important of these adaptations is the process of epithelial-to-mesenchymal transition (EMT), which allows tumor cells to reduce epithelial characteristics, limit polarity, and promote the transformation of mesenchymal phenotypes, characterized by higher migratory and invasive potential [20,21,22,23]. The activation of EMT in cancer cells has important functional consequences, given that mesenchymal tumor cells are more aggressive and resistant to treatment [24]. The proportion of mesenchymal CTCs increases with disease progression, eventually resulting in distant metastasis [25]. Over the recent years, it has been suggested that tumor cells must undergo a reverse mesenchymal-to-epithelial transition to acquire the ability to proliferate and thus develop metastatic tumors. Therefore, it is believed that tumor cells with an intermediate phenotype can spread to distant sites and grow most effectively [26]. Additionally, CTCs can form aggregates with leukocytes, endothelial cells, or platelets, called CTC clusters [27]. Initially, it was hypothesized that CTC clusters arose from the co-invasion and propagation of oligoclonal populations of tumor cells [27, 28]. Nonetheless, recent studies have shown that these clusters can be formed by tumor cells aggregating in the vascular system induced by the affinity interaction of the hyaluronan receptor CD44 [29]. Compared with a single tumor cell, CTC clusters are relatively rare in circulation but exhibit remarkably greater resistance to apoptosis and more metastatic potential [30, 31]. Moreover, studies on CTC clusters in the peripheral blood of patients with CRC have shown that CTC clusters are not malignant, but rather tumor-derived endothelial cells associated with tumor vascular characteristics; notably, the isolation and counting of these CTC clusters can distinguish between healthy individuals and patients with early-stage CRC having a high degree of accuracy (≤ IIa) [32].
Precision medicine remains a realistic vision in the field of oncology. However, tumor cell heterogeneity is a huge obstacle; many factors contribute to it and affect several biological processes associated with tumor progression [33]. Therefore, detecting CTCs in the peripheral blood of patients with CRC should be considered not only as a less invasive option than actual biopsy but also as a new approach for a comprehensive understanding of the whole tumor heterogeneity. The in-depth study of CTCs may enable the unrevealing of the molecular characteristics of tumors, identification of biomarkers for targeted therapies, prediction of the effectiveness of treatment, and precise assessment of tumor prognosis, thus enabling precision medicine.
CTC enrichment techniques
The techniques for isolating CTCs can be divided into two categories according to the inherent physical characteristics of the cells or the specific antigens expressed on the surface (Table 1). CTCs are usually identified by immunocytochemistry, immunofluorescence, or other techniques after being captured [34]. CTC separation and capture techniques have been constantly updated with increasing efficiency and purity in recent years.
Isolation based on physical properties
CTCs usually exhibit low density and tend to be comparatively larger in diameter compared with other blood cells [52]. A large number of platforms for CTC isolation based on physical properties have satisfying capture efficiency, which can be roughly divided into the following categories.
Based on size
Isolation by size of epithelial tumor cells (ISET) is the platform for capturing CTCs after blood filtration based on the difference in cell size; it can capture CTCs from 1 mL of peripheral blood samples for counting and immunomorphological analysis [35]. Su et al. [53] developed a new integrated microfluidic device to isolate CTCs based on the difference in size and deformability between tumor cells and normal blood cells. CTCs were successfully captured in the peripheral blood samples of all patients with advanced CRC within 3 h. Cohen et al. [36] proposed the Parsortix PR1 system, which allows CTC enrichment based on cell size and membrane deformability; also, the expression of more than 2000 cancer-related genes can be obtained in the downstream analysis of captured CTCs using the HTG EdgeSeq nuclease protection assay. Ribeiro Samy et al. [54] developed a size- and deformability-based microfluidic chip for capturing CTCs. Moreover, the droplet digital pathologic complete response (PCR) (ddPCR) was used for the CTC molecular characterization, revealing the existence of APC gene mutations in most patients with CRC. Likewise, ScreenCell [41], CellSieve [42], and FMSA (Flexible Micro Spring Array) [43] are all microfluidic platforms that capture CTCs based on the principle of cell size difference.
Based on a density gradient
This separation method takes advantage of the principle that cells differ in density from each other. It can cause stratification of cells, with CTCs remaining in the monocyte-enriched layer, by adding the separation medium to the blood and centrifuging it. Density gradient centrifugation is a simple, reliable, and inexpensive method. Nonetheless, the disadvantages of this method include the loss of large amounts of CTCs and the fact that leukocytes cannot be readily removed. This results in considerably low purity of obtained CTCs, which is not conducive for further downstream experimentation [55]. Ficoll-Paque was not initially used to isolate CTCs, but later, it was used to capture CTCs in the peripheral blood of cancer patients with satisfactory results [37]. Rosenberg et al. [38] proposed a device to achieve higher CTC purity, called OncoQuick, which uses porous membranes to reduce the number of blood cells with similar size to CTCs and has been shown to have a high CTC recovery rate.
Based on inertia
The differences in the size and density of CTCs from other blood cells reflect different cell dynamics. Di Carlo et al. [56] proposed an inertial microfluidic system to isolate CTCs without antibody labeling, and the capture efficiency of the device was as high as 90%; it could process 2.5 mL of blood every minute.
Based on electrophoresis
Gascoyne et al. [39] proposed dielectrophoretic field-flow fractionation (DepFFF) to isolate CTCs; it works on the principle that CTCs can be separated in the presence of an electric field due to the different sizes and charges of cells. Thus, the platform exploits the intrinsic properties of CTCs without adding biomarkers to the cell surface and can provide unmodified live CTCs for in vitro culture and downstream analysis [57, 58]. The shortcomings of this device include the low capture rate of approximately 70% and the low efficiency of processing the peripheral blood of approximately 1 mL per hour [59].
Based on photoacoustic effect
Bhattacharyya et al. [60] used photoacoustic flow cytometry (PAFC) to detect CTCs in the peripheral blood of patients with breast cancer. It works by the absorption of laser light through nanoparticles labeled with antibodies on target cells. They also found that this method could be used to study the response of patients with breast cancer to treatment as well as in vitro drug trials [60].
Isolation based on immune affinity
Besides the capture methods based on physical principles, the immunoaffinity-based capture methods take advantage of the expression of specific biomarkers on the surface of CTCs. CellSearch is a CTC capture platform developed by Veridex and approved by the Food and Drug Administration (FDA). It works by using immunomagnetic beads with EpCAM antibodies modified on the beads to capture CTCs through immunoaffinity with the EpCAM antigen expressed on the surface of CTCs and then subsequently generating a shunt under the action of a magnetic field. The captured CTCs can then be counted and immunoassayed [45]. Based on this, the CTC counts and dynamic changes may provide prognostic information in patients with cancers [45]. Recently, several studies used this platform to capture CTCs and found correlations with the poor prognosis of patients with CRC, thus demonstrating the potential of CTC as a prognostic biomarker [61,62,63,64,65,66,67]. Furthermore, the CTC count has been approved by the FDA as a standard for evaluating the prognosis of metastatic CRC [61], showing that liquid biopsy has a great potential for clinical application in the future. However, the CellSearch is limited by the amount of blood taken. The expression of EpCAM could be reduced due to the EMT process, resulting in the false-negative rate of this single antibody capture system. Therefore, other types of antibodies, such as N-cadherin, Vimentin, and combinations of multiple antibodies [68,69,70], which have immunoaffinity to cells with mesenchymal properties, have also been introduced into the CTC capture platform for capturing cells expressing low quantities of EpCAM. In addition, antibodies that do not depend on epithelial and mesenchymal properties have also been developed for CTC capture. A subunit of the VAR2CSA protein called rVAR2 was recombinantly expressed in E. coli by Agerbaek et al. and successfully used to capture CTCs from tumor patients [71, 72].
The development of microfluidic-based CTC capture platforms is also a hot research topic in recent years. It can use not only the difference in cell size, but also immunoaffinity, to separate CTCs, and thus has certain advantages over ordinary immunomagnetic beads. CTC-Chip [47], HB-Chip [48], and CTC-iChip [49] are a few representative capture platforms of such microfluidic systems that use immunoaffinity for CTC capture based on the principle that target cells can specifically bind to microchannels or structurally modified bioactive molecules in microfluidic devices. Although CTCs can be easily captured by combining cell surface − specific antigens with bioactive molecules via immunoaffinity, the purity of CTCs may be compromised because, in addition to white blood cells, the whole blood contains platelets, neutrophils, eosinophils, lymphocytes, and other cells, resulting in nonspecific capture [73]. To solve this problem, negative selection platforms were designed to remove non-target cells such as white blood cells and red blood cells [74]. Since the enrichment process is not affected by the expression of cell surface antigen, this type of device has a high capture efficiency and thus facilitates further downstream experiments. However, the limitation is that the purity of isolated CTCs is lower than that of positive enrichment and is prone to the loss of CTCs [75]. Although studies have suggested that CTC counting and differentiation of CTC subtypes by immunostaining may be effective in evaluating the prognosis of various cancers, a comprehensive molecular and functional analysis of CTC may better characterize the metastatic potential of tumors to develop precise treatment plans. Several capture platforms allow for downstream analysis after CTCs are isolated. For example, IsoFlux uses flow control and immunomagnetic capture to achieve high purity of CTCs for downstream analysis; also, the platform can effectively reduce the damage of CTCs during the capture process [50]. AdnaTest is a CTC capture platform that has been modified with three antibodies (EpCAM, EGFR, and MUC1), allowing for the identification of tumor-associated transcripts by RT-PCR [51]. As of now, many studies have used this category of platforms for CTC capture and downstream analysis to investigate the association between gene expression and cancer prognosis [76,77,78]. Nowadays, with the rapid development of nanotechnology, nanomaterials have been successfully introduced into the study of CTC isolation. The basic principle is based on the strong interaction between CTCs and nano-substrates, such as nanocolumns, nanofibers, nanoparticles, nanotubes, and nanorough surfaces, all of which exhibit satisfactory capture sensitivity and efficiency [79].
Clinical applications of CTCs
Traditional diagnostic and staging detection strategies for CRC have many limitations, resulting in an inadequate understanding of disease progression. Therefore, the exploration of representative biomarkers is urgently needed for early diagnosis, recurrence prediction, and prognostic evaluation of patients with CRC. A wealth of information on primary tumor prediction, disease progression, and prognostic follow-up can be obtained by detecting CTCs in peripheral blood (Table 2).
Early diagnosis
Tsai et al. [80] used the self-developed platform called CellMax to isolate and subsequently count CTCs from the peripheral blood of 667 Taiwanese adults who underwent enteroscopy. A predefined algorithm was then used for early diagnosis. The result showed that the platform had high specificity (86%) and sensitivity (95%) to distinguish between benign and malignant tumors. This also indicated that CTC detection had a good application prospect in the early diagnosis of CRC. Baek et al. [81] set the cutoff value of CTCs to 5/7.5 mL and used this as a criterion to differentiate between patients with CRC and healthy people with good sensitivity (75%) and specificity (100%). The results of these two studies showed the promising application of CTC detection in the early diagnosis of CRC. However, uniform criteria for differentiating benign from malignant tumors are still lacking.
Prognosis evaluation
Several relevant studies have demonstrated a strong correlation of CTC counts with cancer progression and prognosis [102,103,104,105]. Additionally, FDA has approved the use of CTC counting as a prognostic tool for metastatic prostate, colon, and breast cancers [61]. In a prospective multicenter study involving 430 patients with mCRC, the baseline CTCs were counted using immunomagnetic beads. Then, the patients were divided into positive and negative groups according to the cutoff value of 3/7.5 mL. The result showed that the median progression-free survival (PFS: 4.5 months vs 7.9 months; P = 0.0002) and the overall survival (OS: 9.4 months vs 18.5 months; P = 0.0001) was shorter in the positive group than in the negative group [61]. A recent study of 121 patients with CRC found that 71 patients were positive for baseline CTCs, and the positivity rate highly correlated with the depth of tumor invasion, lymphatic metastasis, distant metastasis, TNM stage, and serum CEA levels [51]. Moreover, the Kaplan–Meier survival curve showed that PFS (14 months vs 23 months, P = 0.001) and OS (18 months vs 25 months, P = 0.003) of baseline CTC-positive patients were significantly shorter than those of negative patients [82]. Yang et al. [83] proposed that the positive rate of preoperative CTCs in CRC positively correlated with the tumor stage and serum CEA level while the positive rate of postoperative CTCs positively correlated with the tumor stage and independently correlated with the tumor recurrence-free survival (RFS) rate. Additionally, the study demonstrated that postoperative CTC positivity was an independent indicator of poor prognosis in patients with CRC [76]. Moreover, the risk of recurrence was higher in postoperative CTC-positive patients than in preoperative CTC-positive patients [83]. Many studies have confirmed that the CTC count is an independent factor for PFS and OS in CRC [67, 84,85,86,87]. However, Le et al. [106] performed CTC isolation from the pulmonary venous blood of 24 patients with CRC treated with pulmonary metastasectomy. The results revealed no significant correlation of PFS and OS with CTC counts. Although CTC counts have attracted widespread interest as a useful marker for cancer prognosis, they are still not widely available for clinical application. Moreover, uniform cutoff values for assessing the prognosis and progression of cancer are still lacking. Therefore, studies have been conducted to investigate the genes expressed by CTCs and correlate their expression with the clinicopathological features of cancers [78, 107,108,109]. Chen et al. [82] reported that the epithelial cell transforming sequence 2 (ECT2) oncogene expressed by CTCs in peripheral blood had higher sensitivity than the serum CEA level; it was highly expressed in patients with advanced-stage CRC. Furthermore, the ECT2 gene expressed on the surface of CTCs can be used as a marker for the prognosis of patients with CRC [89]. Another study including 70 patients with CRC showed that the expression level of MAGEA1-6 or hTERT genes in CTCs was significantly higher in patients with T3 and T4 stages than in patients with T1 and T2 stages [90]. Ning et al. [110] showed that the expression of the Akt-2 gene in CTCs of patients with CRC predicted shorter PFS and OS; also, the median survival time of patients with CTCAkt−2+ was significantly shorter than that of patients with CTCAkt−2−. Nowadays, the expression of CK20 [92], CEACAM5 [93], COX-2 [94], and LGR-5 [95] has also been used to monitor the progression and prognosis in patients with CRC.
Treatment monitoring
CRC is overly aggressive and prone to metastasis. Hence, the selection of the most appropriate treatment and the best time for patients is highly challenging. Therefore, reliable biomarkers for accurate monitoring of the treatment effect are urgently required. Additionally, CTC subclones can reflect the heterogeneity of cancers in real time and show their different abilities to evade treatment; thus, CTCs can be monitored dynamically to understand the response to treatment in time [98]. Shi et al. [97] proposed that the number of CTCs in the peripheral blood of patients with liver metastases from CRC significantly reduced 7 and 30 days after cryotherapy. Moreover, the expression of serum CEA, EpCAM, CK18, and CK19 gradually decreased with time. Delgado-Urena et al. detected the CTCs of 77 patients with mCRC treated with FOLFOX and bevacizumab before treatment and 12 and 24 weeks after treatment. The patients were divided into good response group (complete response group, partial response group, or stable group) and poor response group. The final results revealed that the positive rates of CTCs and CTCVEGFR+ decreased in the group with good treatment response [98], suggesting that the dynamic detection of CTCs could monitor the treatment in a timely manner, allowing physicians to understand the treatment response and change the treatment regimen early. In another study of 35 patients with gastrointestinal cancer, Yue et al. [99] reported that CTCs with high expression of PD-L1 could be used as a biomarker to predict the PD-1/PD-L1 treatment response. In yet another study of preoperative neoadjuvant therapy for CRC, Troncarelli Flores et al. reported that the baseline CTCs TYMS+/ RAD23B+ were associated with poor reactions of neoadjuvant chemoradiation (NCRT) therapy. CTCs TYMS−/RAD23B− were detected in all PCR patients after neoadjuvant therapy, 83.5% of partial response (PR) patients, but not in no response (NR) patients; in comparison, CTCsTyms+/RAD23B+ were not detected in PCR or PR patients but could be detected in 83.5% of NR patients [100].
Culture of CTCs in vitro
The establishment of permanent CTC cell lines in vitro has been one of the challenging studies in recent times. Indeed, CTC cell lines can be used to identify proteins and pathways associated with cancer cell stemness and metastasis, as well as to test the sensitivity of cancers to anti-cancer drugs [111]. Cayrefourcq et al. [112] successfully cultivated a CTC cell line isolated from a patient with colon cancer named CTC-MCC-41, which had been cultured for more than 1 year by the time the study was published. The cell line has been characterized at the genome, transcriptome, proteome, and secretome levels. More importantly, functional studies revealed that the CTC cell line could quickly form tumors after transplantation in immunodeficient mice. Pantel et al. performed a follow-up study on the CTC-MCC-41 cell line and found that CTC-MCC-41 had the following characteristics besides the ability to expand in vitro for more than 2 years: (A) stem cell-like epithelial characteristics, (B) epithelial/mesenchymal phenotype, (C) potential for rapid initiation of angiogenesis in vitro, and (D) bone-like characteristics. Moreover, the cell line had characteristics similar to those of primary tumors and metastatic lymph nodes in patients with CRC [113]. De et al. [114] proposed a scheme for the 3D culture of CTCs in vitro, which used a polycaprolactone scaffold to deposit an extracellular matrix on it. The study also confirmed the existence of a CTC mixed phenotype, thus helping to better understand the relationship between CTC phenotype and CRC prognosis. Grillet et al. [115] conducted downstream experiments after culturing CTCs of patients with CRC in vitro. The results showed that the patients who successfully established CTC-MCC-411 died soon after receiving chemotherapy with the FOLFIRI regimen. The drug sensitivity test also revealed that the cell line established in vitro was resistant to the FOLFIRI regimen. Notably, the results revealed significant overexpression of genes associated with xenograft resistance in their cultured CTC lines. Cytotoxicity assays further confirmed the potential use of the model in predicting drug response in patients with CRC.
CTC single-cell analysis
TC single-cell analysis. Tumor cells have a high degree of heterogeneity at the single-cell level, which may be one of the factors responsible for increasing the incidence of cancer recurrence and metastasis [33, 116]. Single-cell technology can help unravel the heterogeneity of tumors. Therefore, an in-depth analysis of a single CTC allows the real-time monitoring of somatic mutation profiles and genomic expression of drug-resistant clones in patients with cancer, contributing to a deeper understanding of the treatment of human cancers such as CRC [33, 117]. Comparing a single CTC with primary and metastatic tumors may help identify the clones that cause metastasis and reveal the molecular biomarkers of metastasis [118]. Single-cell RNA sequencing (scRNA-seq) can be used as a powerful tool to discover new tumor progression markers in CTCs. Recent studies have reported the genomic and transcriptomic analyses of single CTC using next-generation sequencers [118, 119]. Abouleila et al. [6] reported a method of combining single-cell mass spectrometry with microfluidic cell enrichment and obtained a nontargeted molecular map of a single CTC from the peripheral blood of patients with gastrointestinal cancer. This revealed the differences in metabonomic characteristics among CTCs from different tumor populations. Pei et al. [120] reported an integrated microfluidic device for the phenotypic analysis of a single CTC. This device can allow the automatic isolation of CTCs from whole blood after sequential single-cell phenotyping with high-purity (92% ± 3%) cell sorting and high-throughput processing capacity (5 mL/3 h). This study reported the collection of a single CTC from xenografted mice and patients with different stages of CRC and obtained the correlations of CTC characteristics with clinical tumor stage and treatment response. Single-cell transcriptomics can detect the expression levels of a single cell in a given population, and hence the interest in this area of research has increased [121]. Bian et al. [122] established a platform called the single-cell triple omics sequencing (SCTrio-seq) technique. This platform is capable of detecting gene mutations, transcription, and methylation simultaneously at the single-cell level. Ten patients with CRC underwent SCTrio-seq. The results provided insight into tumor evolution and linked DNA methylation to genetic genealogy. Additionally, the results confirmed that DNA methylation levels were consistent within the genealogy but might differ significantly between clones [122]. Using single-cell analysis techniques, it is now possible to deconstruct cancers into their constituent cell types and thus enable the understanding of the biological characteristics of cancer. The analysis of CTC gene expression patterns of CRC at the single-cell level can help find key information for evaluating prognostic biomarkers and developing precise and personalized cancer therapies.
ctDNA analysis in CRC
Plasma ctDNA refers to DNA fragments of tumor origin and is a subset of plasma cell–free DNA (cfDNA). The cfDNA also includes DNA from other sources in the circulation, mainly germline DNA due to hematopoietic cell death [123, 124]. Hence, figuring out the origin of cfDNA may be a challenge. Besides, the shedding of ctDNA is extremely varied; it may be as little as 0.01%, or it might account for the majority of total plasma cfDNA [125]. Studies demonstrated that the amount of ctDNA was primarily associated with tumor burden and tumor type [126]. From a heterogeneous viewpoint, ctDNA offers a more comprehensive overview of the range of mutations present in a patient’s tumor [127]. Originally, the real-time quantitative PCR (qPCR) detection system was used to identify ctDNA-specific mutations [128]. However, limited by its sensitivity and specificity, PCR can only be performed in patients with advanced cancer having high ctDNA levels [129]. In contrast, digital PCR (dPCR) has a higher sensitivity in detecting and quantifying ctDNA and is currently widely used in bodily fluid biomarker studies [128, 130]. Still, one limitation of PCR-based techniques is that they cannot be used to investigate a wide range and diverse types of genomic alterations [131]. With the development of RNA sequencing technology, next-generation sequencing (NGS) has overcome these shortcomings and can detect copy number alterations, mutations, and other chromosomal rearrangements, including inversions, translocations, and reversals [132]. The recent growth in the studies demonstrates the value of ctDNA in the early diagnosis [133, 134], prognosis [135, 136], and response to treatment [137, 138] in CRC.
Compared with ctDNA, which is extensively fragmented and compounded by significant background, CTCs are intact cells providing more genetic information about the origin of tumors and potentially allowing for more specific diagnosis and more precise treatment plans for patients with CRC [139].
Summary and future prospects
Thanks to the continuous advancements in diagnostic and treatment technologies, the prognosis of patients with CRC has improved remarkably in recent decades. Nonetheless, the early diagnosis, treatment monitoring, and prognostic evaluation of CRC still have some limitations; therefore, a large number of patients with CRC still die every year. At present, a variety of sensitive, high-throughput, and efficient CTC isolation platforms have been successfully established, which can be used to count CTCs in the peripheral blood of patients with CRC dynamically. Additionally, genes and proteins expressed by CTCs can be detected to evaluate the prognosis and recurrence of CRC. However, a uniform CTC cutoff value for clinical assessment of CRC progression and prognosis still lacks due to the differences in sampling, storage time, and enrichment methods. The downstream analysis of CTCs in patients with CRC and the single-cell technology have also improved the understanding of tumor formation, development, metastasis, and heterogeneity, which is conducive to the development of a personalized drug detection platform. Despite several CTC detection methods, the application of CTCs in clinical practice is not yet widespread due to the lack of a standardized detection platform with uniform high sensitivity and specificity. Most of the studies are single-center studies with a small number of cases. This has eventually led to many studies with different results due to interindividual differences, such as pathogenic factors, tumor stage, ethnicity, and geographical factors, to name a few. Against this backdrop, an important research direction for the future is to include a large number of cases and conduct multicenter prospective studies. Besides the number of CTCs and expressed genes or proteins as the criteria for cancer diagnosis and prognosis, CTCs in combination with other indicators such as CA-199 or ctDNA are worth investigating because they can increase the accuracy of diagnostic and prognostic assessments [69, 140]. Tumor immunotherapy has recently attracted much attention and is one of the hot spots in the field of tumor treatment. The expression of immune checkpoints on the surface of CTCs is also associated with the effect of immunotherapy. Therefore, the blocking of these checkpoints can be used as an immunotherapeutic approach. Lian et al. successfully targeted PD-L1 and CD47 on the surface of CTCs, which eventually caused a significant inhibition of tumor growth and metastasis [141]. Although CTCs are not prevalently used and promoted in clinical practice, their role in tumor prognostic assessment cannot be ignored. Therefore, one of the future research directions should be to develop a CTC capture platform with higher efficiency, specificity, and sensitivity. Additionally, more comprehensive and in-depth studies of CTCs from a new perspective must be conducted to develop a more personalized and precise treatment plan for each cancer patient.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68(6):394–424. https://doi.org/10.3322/caac.21492
Global Burden of Disease Cancer C, Fitzmaurice C, Abate D, Abbasi N, Abbastabar H, Abd-Allah F, Abdel-Rahman O, Abdelalim A, Abdoli A, Abdollahpour I et al (2019) Global, regional, and national cancer incidence, mortality, years of life lost, years lived with disability, and disability-adjusted life-years for 29 cancer groups, 1990 to 2017: a systematic analysis for the Global Burden of Disease Study. JAMA Oncol. https://doi.org/10.1001/jamaoncol.2019.2996
Primrose JN, Perera R, Gray A, Rose P, Fuller A, Corkhill A, George S, Mant D (2014) Effect of 3 to 5 years of scheduled CEA and CT follow-up to detect recurrence of colorectal cancer: the FACS randomized clinical trial. JAMA 311(3):263–270. https://doi.org/10.1001/jama.2013.285718
Carpelan-Holmstrom M, Louhimo J, Stenman UH, Alfthan H, Haglund C (2002) CEA, CA 19–9 and CA 72–4 improve the diagnostic accuracy in gastrointestinal cancers. Anticancer Res 22(4):2311–2316
Khattak A (2016) Liquid biopsies: advancing cancer research through drops of blood. Intern Med J 46(3):376–377. https://doi.org/10.1111/imj.12996
Abouleila Y, Onidani K, Ali A, Shoji H, Kawai T, Lim CT, Kumar V, Okaya S, Kato K, Hiyama E et al (2019) Live single cell mass spectrometry reveals cancer-specific metabolic profiles of circulating tumor cells. Cancer Sci 110(2):697–706. https://doi.org/10.1111/cas.13915
Micevic G, Theodosakis N, Bosenberg M (2017) Aberrant DNA methylation in melanoma: biomarker and therapeutic opportunities. Clin Epigenetics 9:34. https://doi.org/10.1186/s13148-017-0332-8
Zaretsky JM, Garcia-Diaz A, Shin DS, Escuin-Ordinas H, Hugo W, Hu-Lieskovan S, Torrejon DY, Abril-Rodriguez G, Sandoval S, Barthly L et al (2016) Mutations Associated with acquired resistance to PD-1 blockade in melanoma. N Engl J Med 375(9):819–829. https://doi.org/10.1056/NEJMoa1604958
Chen C, Läcke E, Stock C, Hoffmeister M, Brenner H (2017) Colonoscopy and sigmoidoscopy use among older adults in different countries: a systematic review. Prev Med 103:33–42. https://doi.org/10.1016/j.ypmed.2017.07.021
Zhang J, Cheng Z, Ma Y, He C, Lu Y, Zhao Y, Chang X, Zhang Y, Bai Y, Cheng N (2017) Effectiveness of screening modalities in colorectal cancer: a network meta-analysis. Clin Colorectal Cancer 16(4):252–263. https://doi.org/10.1016/j.clcc.2017.03.018
Force USPST, Bibbins-Domingo K, Grossman DC, Curry SJ, Davidson KW, Epling JW Jr, Garcia FAR, Gillman MW, Harper DM, Kemper AR et al (2016) Screening for colorectal cancer: US Preventive Services Task Force recommendation statement. JAMA 315(23):2564–2575. https://doi.org/10.1001/jama.2016.5989
Gazzaniga P, Raimondi C, Nicolazzo C, Carletti R, di Gioia C, Gradilone A, Cortesi E (2015) The rationale for liquid biopsy in colorectal cancer: a focus on circulating tumor cells. Expert Rev Mol Diagn 15(7):925–932. https://doi.org/10.1586/14737159.2015.1045491
TR A, (1869) A case of cancer in which cells similar to those in the tumours were seen in the blood after death. Aust Med J 14:146–147
Liberko M, Kolostova K, Bobek V (2013) Essentials of circulating tumor cells for clinical research and practice. Crit Rev Oncol Hematol 88(2):338–356. https://doi.org/10.1016/j.critrevonc.2013.05.002
Joosse SA, Gorges TM, Pantel K (2015) Biology, detection, and clinical implications of circulating tumor cells. EMBO Mol Med 7(1):1–11. https://doi.org/10.15252/emmm.201303698
Torino F, Bonmassar E, Bonmassar L, De Vecchis L, Barnabei A, Zuppi C, Capoluongo E, Aquino A (2013) Circulating tumor cells in colorectal cancer patients. Cancer Treat Rev 39(7):759–772. https://doi.org/10.1016/j.ctrv.2012.12.007
Alix-Panabières C, Pantel K (2016) Clinical applications of circulating tumor cells and circulating tumor DNA as liquid biopsy. Cancer Discov 6(5):479–491. https://doi.org/10.1158/2159-8290.Cd-15-1483
Danese E, Montagnana M, Lippi G (2019) Circulating molecular biomarkers for screening or early diagnosis of colorectal cancer: which is ready for prime time? Ann Transl Med 7(21):610. https://doi.org/10.21037/atm.2019.08.97
Lim SH, Becker TM, Chua W, Ng WL, de Souza P, Spring KJ (2014) Circulating tumour cells and the epithelial mesenchymal transition in colorectal cancer. J Clin Pathol 67(10):848–853. https://doi.org/10.1136/jclinpath-2014-202499
Markiewicz A, Nagel A, Szade J, Majewska H, Skokowski J, Seroczynska B, Stokowy T, Welnicka-Jaskiewicz M, Zaczek AJ (2018) Aggressive phenotype of cells disseminated via hematogenous and lymphatic route in breast cancer patients. Transl Oncol 11(3):722–731. https://doi.org/10.1016/j.tranon.2018.03.006
Mani SA, Guo W, Liao MJ, Eaton EN, Ayyanan A, Zhou AY, Brooks M, Reinhard F, Zhang CC, Shipitsin M et al (2008) The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133(4):704–715. https://doi.org/10.1016/j.cell.2008.03.027
Ksiazkiewicz M, Markiewicz A, Zaczek AJ (2012) Epithelial-mesenchymal transition: a hallmark in metastasis formation linking circulating tumor cells and cancer stem cells. Pathobiology 79(4):195–208. https://doi.org/10.1159/000337106
Markiewicz A, Żaczek AJ (2017) The landscape of circulating tumor cell research in the context of epithelial-mesenchymal transition. Pathobiology 84(5):264–283. https://doi.org/10.1159/000477812
Yu M, Bardia A, Wittner BS, Stott SL, Smas ME, Ting DT, Isakoff SJ, Ciciliano JC, Wells MN, Shah AM et al (2013) Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 339(6119):580–584. https://doi.org/10.1126/science.1228522
Zheng X, Fan L, Zhou P, Ma H, Huang S, Yu D, Zhao L, Yang S, Liu J, Huang A et al (2017) Detection of circulating tumor cells and circulating tumor microemboli in gastric cancer. Transl Oncol 10(3):431–441. https://doi.org/10.1016/j.tranon.2017.02.007
Tam WL, Weinberg RA (2013) The epigenetics of epithelial-mesenchymal plasticity in cancer. Nat Med 19(11):1438–1449. https://doi.org/10.1038/nm.3336
Aceto N, Bardia A, Miyamoto DT, Donaldson MC, Wittner BS, Spencer JA, Yu M, Pely A, Engstrom A, Zhu H et al (2014) Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell 158(5):1110–1122. https://doi.org/10.1016/j.cell.2014.07.013
Cheung KJ, Padmanaban V, Silvestri V, Schipper K, Cohen JD, Fairchild AN, Gorin MA, Verdone JE, Pienta KJ, Bader JS et al (2016) Polyclonal breast cancer metastases arise from collective dissemination of keratin 14-expressing tumor cell clusters. Proc Natl Acad Sci U S A 113(7):E854-863. https://doi.org/10.1073/pnas.1508541113
Liu X, Taftaf R, Kawaguchi M, Chang YF, Chen W, Entenberg D, Zhang Y, Gerratana L, Huang S, Patel DB et al (2019) Homophilic CD44 interactions mediate tumor cell aggregation and polyclonal metastasis in patient-derived breast cancer models. Cancer Discov 9(1):96–113. https://doi.org/10.1158/2159-8290.CD-18-0065
Yousefi M, Ghaffari P, Nosrati R, Dehghani S, Salmaninejad A, Abarghan YJ, Ghaffari SH (2020) Prognostic and therapeutic significance of circulating tumor cells in patients with lung cancer. Cell Oncol (Dordr) 43(1):31–49. https://doi.org/10.1007/s13402-019-00470-y
Fabisiewicz A, Grzybowska E (2017) CTC clusters in cancer progression and metastasis. Med Oncol 34(1):12. https://doi.org/10.1007/s12032-016-0875-0
Cima I, Kong SL, Sengupta D, Tan IB, Phyo WM, Lee D, Hu M, Iliescu C, Alexander I, Goh WL et al (2016) Tumor-derived circulating endothelial cell clusters in colorectal cancer. Sci Transl Med 8(345):345ra389. https://doi.org/10.1126/scitranslmed.aad7369
Keller L, Pantel K (2019) Unravelling tumour heterogeneity by single-cell profiling of circulating tumour cells. Nat Rev Cancer 19(10):553–567. https://doi.org/10.1038/s41568-019-0180-2
Nagrath S, Jack RM, Sahai V, Simeone DM (2016) Opportunities and challenges for pancreatic circulating tumor cells. Gastroenterology 151(3):412–426. https://doi.org/10.1053/j.gastro.2016.05.052
Vona G, Sabile A, Louha M, Sitruk V, Romana S, Schutze K, Capron F, Franco D, Pazzagli M, Vekemans M et al (2000) Isolation by size of epithelial tumor cells: a new method for the immunomorphological and molecular characterization of circulatingtumor cells. Am J Pathol 156(1):57–63. https://doi.org/10.1016/s0002-9440(10)64706-2
Cohen EN, Jayachandran G, Hardy MR, Venkata Subramanian AM, Meng X, Reuben JM (2020) Antigen-agnostic microfluidics-based circulating tumor cell enrichment and downstream molecular characterization. PLoS ONE 15(10):e0241123. https://doi.org/10.1371/journal.pone.0241123
Weitz J, Kienle P, Lacroix J, Willeke F, Benner A, Lehnert T, Herfarth C, von Knebel DM (1998) Dissemination of tumor cells in patients undergoing surgery for colorectal cancer. Clin Cancer Res 4(2):343–348
Rosenberg R, Gertler R, Friederichs J, Fuehrer K, Dahm M, Phelps R, Thorban S, Nekarda H, Siewert JR (2002) Comparison of two density gradient centrifugation systems for the enrichment of disseminated tumor cells in blood. Cytometry 49(4):150–158. https://doi.org/10.1002/cyto.10161
Gascoyne PR, Noshari J, Anderson TJ, Becker FF (2009) Isolation of rare cells from cell mixtures by dielectrophoresis. Electrophoresis 30(8):1388–1398. https://doi.org/10.1002/elps.200800373
Galanzha EI, Zharov VP (2013) Circulating tumor cell detection and capture by photoacoustic flow cytometry in vivo and ex vivo. Cancers (Basel) 5(4):1691–1738. https://doi.org/10.3390/cancers5041691
Desitter I, Guerrouahen BS, Benali-Furet N, Wechsler J, Janne PA, Kuang Y, Yanagita M, Wang L, Berkowitz JA, Distel RJ et al (2011) A new device for rapid isolation by size and characterization of rare circulating tumor cells. Anticancer Res 31(2):427–441
Adams DL, Stefansson S, Haudenschild C, Martin SS, Charpentier M, Chumsri S, Cristofanilli M, Tang CM, Alpaugh RK (2015) Cytometric characterization of circulating tumor cells captured by microfiltration and their correlation to the Cell Search(®) CTC test. Cytometry A 87(2):137–144. https://doi.org/10.1002/cyto.a.22613
Harouaka RA, Zhou MD, Yeh YT, Khan WJ, Das A, Liu X, Christ CC, Dicker DT, Baney TS, Kaifi JT et al (2014) Flexible micro spring array device for high-throughput enrichment of viable circulating tumor cells. Clin Chem 60(2):323–333. https://doi.org/10.1373/clinchem.2013.206805
Qi LN, Xiang BD, Wu FX, Ye JZ, Zhong JH, Wang YY, Chen YY, Chen ZS, Ma L, Chen J et al (2018) Circulating tumor cells undergoing EMT provide a metric for diagnosis and prognosis of patients with hepatocellular carcinoma. Cancer Res 78(16):4731–4744. https://doi.org/10.1158/0008-5472.Can-17-2459
Allard WJ, Matera J, Miller MC, Repollet M, Connelly MC, Rao C, Tibbe AG, Uhr JW, Terstappen LW (2004) Tumor cells circulate in the peripheral blood of all major carcinomas but not in healthy subjects or patients with nonmalignant diseases. Clin Cancer Res 10(20):6897–6904. https://doi.org/10.1158/1078-0432.CCR-04-0378
Talasaz AH, Powell AA, Huber DE, Berbee JG, Roh KH, Yu W, Xiao W, Davis MM, Pease RF, Mindrinos MN et al (2009) Isolating highly enriched populations of circulating epithelial cells and other rare cells from blood using a magnetic sweeper device. Proc Natl Acad Sci U S A 106(10):3970–3975. https://doi.org/10.1073/pnas.0813188106
Nagrath S, Sequist LV, Maheswaran S, Bell DW, Irimia D, Ulkus L, Smith MR, Kwak EL, Digumarthy S, Muzikansky A et al (2007) Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 450(7173):1235–1239. https://doi.org/10.1038/nature06385
Stott SL, Hsu CH, Tsukrov DI, Yu M, Miyamoto DT, Waltman BA, Rothenberg SM, Shah AM, Smas ME, Korir GK et al (2010) Isolation of circulating tumor cells using a microvortex-generating herringbone-chip. Proc Natl Acad Sci U S A 107(43):18392–18397. https://doi.org/10.1073/pnas.1012539107
Karabacak NM, Spuhler PS, Fachin F, Lim EJ, Pai V, Ozkumur E, Martel JM, Kojic N, Smith K, Chen PI et al (2014) Microfluidic, marker-free isolation of circulating tumor cells from blood samples. Nat Protoc 9(3):694–710. https://doi.org/10.1038/nprot.2014.044
Harb W, Fan A, Tran T, Danila DC, Keys D, Schwartz M, Ionescu-Zanetti C (2013) Mutational analysis of circulating tumor cells using a novel microfluidic collection device and qPCR assay. Transl Oncol 6(5):528–538. https://doi.org/10.1593/tlo.13367
Danila DC, Samoila A, Patel C, Schreiber N, Herkal A, Anand A, Bastos D, Heller G, Fleisher M, Scher HI (2016) Clinical validity of detecting circulating tumor cells by AdnaTest assay compared with direct detection of tumor mRNA in stabilized whole blood, as a biomarker predicting overall survival for metastatic castration-resistant prostate cancer patients. Cancer J 22(5):315–320. https://doi.org/10.1097/ppo.0000000000000220
Lin HK, Zheng S, Williams AJ, Balic M, Groshen S, Scher HI, Fleisher M, Stadler W, Datar RH, Tai YC et al (2010) Portable filter-based microdevice for detection and characterization of circulating tumor cells. Clin Cancer Res 16(20):5011–5018. https://doi.org/10.1158/1078-0432.Ccr-10-1105
Su W, Yu H, Jiang L, Chen W, Li H, Qin J (2019) Integrated microfluidic device for enrichment and identification of circulating tumor cells from the blood of patients with colorectal cancer. Dis Markers 2019:8945974. https://doi.org/10.1155/2019/8945974
Ribeiro-Samy S, Oliveira MI, Pereira-Veiga T, Muinelo-Romay L, Carvalho S, Gaspar J, Freitas PP, López-López R, Costa C, Diéguez L (2019) Fast and efficient microfluidic cell filter for isolation of circulating tumor cells from unprocessed whole blood of colorectal cancer patients. Sci Rep 9(1):8032. https://doi.org/10.1038/s41598-019-44401-1
Gabriel MT, Calleja LR, Chalopin A, Ory B, Heymann D (2016) Circulating tumor cells: a review of non-EpCAM-based approaches for cell enrichment and isolation. Clin Chem 62(4):571–581. https://doi.org/10.1373/clinchem.2015.249706
Di Carlo D (2009) Inertial microfluidics. Lab Chip 9(21):3038–3046. https://doi.org/10.1039/b912547g
Moon HS, Kwon K, Kim SI, Han H, Sohn J, Lee S, Jung HI (2011) Continuous separation of breast cancer cells from blood samples using multi-orifice flow fractionation (MOFF) and dielectrophoresis (DEP). Lab Chip 11(6):1118–1125. https://doi.org/10.1039/c0lc00345j
Gupta V, Jafferji I, Garza M, Melnikova VO, Hasegawa DK, Pethig R, Davis DW (2012) ApoStream(TM), a new dielectrophoretic device for antibody independent isolation and recovery of viable cancer cells from blood. Biomicrofluidics 6(2):24133. https://doi.org/10.1063/1.4731647
Jiang J, Zhao H, Shu W, Tian J, Huang Y, Song Y, Wang R, Li E, Slamon D, Hou D et al (2017) An integrated microfluidic device for rapid and high-sensitivity analysis of circulating tumor cells. Sci Rep 7:42612. https://doi.org/10.1038/srep42612
Bhattacharyya K, Goldschmidt BS, Viator JA (2016) Detection and capture of breast cancer cells with photoacoustic flow cytometry. J Biomed Opt 21(8):87007. https://doi.org/10.1117/1.Jbo.21.8.087007
Cohen SJ, Punt CJ, Iannotti N, Saidman BH, Sabbath KD, Gabrail NY, Picus J, Morse M, Mitchell E, Miller MC et al (2008) Relationship of circulating tumor cells to tumor response, progression-free survival, and overall survival in patients with metastatic colorectal cancer. J Clin Oncol 26(19):3213–3221. https://doi.org/10.1200/JCO.2007.15.8923
Bork U, Rahbari NN, Scholch S, Reissfelder C, Kahlert C, Buchler MW, Weitz J, Koch M (2015) Circulating tumour cells and outcome in non-metastatic colorectal cancer: a prospective study. Br J Cancer 112(8):1306–1313. https://doi.org/10.1038/bjc.2015.88
Lalmahomed ZS, Mostert B, Onstenk W, Kraan J, Ayez N, Gratama JW, Grunhagen D, Verhoef C, Sleijfer S (2015) Prognostic value of circulating tumour cells for early recurrence after resection of colorectal liver metastases. Br J Cancer 112(3):556–561. https://doi.org/10.1038/bjc.2014.651
Sotelo MJ, Sastre J, Maestro ML, Veganzones S, Viéitez JM, Alonso V, Grávalos C, Escudero P, Vera R, Aranda E (2015) Role of circulating tumor cells as prognostic marker in resected stage III colorectal cancer. Annals Oncol Off J Eur Soc Med Oncol 26(3):535
van Dalum G, Stam GJ, Scholten LF, Mastboom WJ, Vermes I, Tibbe AG, De Groot MR, Terstappen LW (2015) Importance of circulating tumor cells in newly diagnosed colorectal cancer. Int J Oncol 46(3):1361–1368. https://doi.org/10.3892/ijo.2015.2824
Tsai WS, Chen JS, Shao HJ, Wu JC, Lai JM, Lu SH, Hung TF, Chiu YC, You JF, Hsieh PS et al (2016) Circulating tumor cell count correlates with colorectal neoplasm progression and is a prognostic marker for distant metastasis in non-metastatic patients. Sci Rep 6:24517. https://doi.org/10.1038/srep24517
Bahnassy AA, Salem SE, Mohanad M, Abulezz NZ, Abdellateif MS, Hussein M, Zekri CAN, Zekri AN, Allahloubi NMA (2019) Prognostic significance of circulating tumor cells (CTCs) in Egyptian non-metastatic colorectal cancer patients: a comparative study for four different techniques of detection (Flowcytometry, Cell Search, Quantitative Real-time PCR and Cytomorphology). Exp Mol Pathol 106:90–101. https://doi.org/10.1016/j.yexmp.2018.12.006
Liu H, Wang Z, Chen C, Ding P, Sun N, Pei R (2019) Dual-antibody modified PLGA nanofibers for specific capture of epithelial and mesenchymal CTCs. Colloids Surf B Biointerfaces 181:143–148. https://doi.org/10.1016/j.colsurfb.2019.05.031
Wei T, Zhang X, Zhang Q, Yang J, Chen Q, Wang J, Li X, Chen J, Ma T, Li G et al (2019) Vimentin-positive circulating tumor cells as a biomarker for diagnosis and treatment monitoring in patients with pancreatic cancer. Cancer Lett 452:237–243. https://doi.org/10.1016/j.canlet.2019.03.009
Takeda K, Yamada T, Takahashi G, Iwai T, Ueda K, Kuriyama S, Koizumi M, Matsuda A, Shinji S, Ohta R et al (2019) Analysis of colorectal cancer-related mutations by liquid biopsy: Utility of circulating cell-free DNA and circulating tumor cells. Cancer Sci 110(11):3497–3509. https://doi.org/10.1111/cas.14186
Bang-Christensen SR, Pedersen RS, Pereira MA, Clausen TM, Løppke C, Sand NT, Ahrens TD, Jørgensen AM, Lim YC, Goksøyr L et al. (2019) Capture and detection of circulating glioma cells using the recombinant VAR2CSA malaria protein. Cells 8 (9). https://doi.org/10.3390/cells8090998
Agerbaek MO, Bang-Christensen SR, Yang MH, Clausen TM, Pereira MA, Sharma S, Ditlev SB, Nielsen MA, Choudhary S, Gustavsson T et al (2018) The VAR2CSA malaria protein efficiently retrieves circulating tumor cells in an EpCAM-independent manner. Nat Commun 9(1):3279. https://doi.org/10.1038/s41467-018-05793-2
Ferreira MM, Ramani VC, Jeffrey SS (2016) Circulating tumor cell technologies. Mol Oncol 10(3):374–394. https://doi.org/10.1016/j.molonc.2016.01.007
Lara O, Tong X, Zborowski M, Chalmers JJ (2004) Enrichment of rare cancer cells through depletion of normal cells using density and flow-through, immunomagnetic cell separation. Exp Hematol 32(10):891–904
Pantel K, Alix-Panabieres C (2019) Liquid biopsy and minimal residual disease - latest advances and implications for cure. Nat Rev Clin Oncol 16(7):409–424. https://doi.org/10.1038/s41571-019-0187-3
Alva A, Friedlander T, Clark M, Huebner T, Daignault S, Hussain M, Lee C, Hafez K, Hollenbeck B, Weizer A et al (2015) Circulating tumor cells as potential biomarkers in bladder cancer. J Urol 194(3):790–798. https://doi.org/10.1016/j.juro.2015.02.2951
Ma Y, Luk A, Young FP, Lynch D, Chua W, Balakrishnar B, de Souza P, Becker TM (2016) Droplet digital PCR based androgen receptor variant 7 (AR-V7) detection from prostate cancer patient blood biopsies. Int J Mol Sci 17 (8). https://doi.org/10.3390/ijms17081264
Russo GI, Bier S, Hennenlotter J, Beger G, Pavlenco L, van de Flierdt J, Hauch S, Maas M, Walz S, Rausch S et al (2018) Expression of tumour progression-associated genes in circulating tumour cells of patients at different stages of prostate cancer. BJU Int 122(1):152–159. https://doi.org/10.1111/bju.14200
Ding P, Wang Z, Wu Z, Zhu W, Liu L, Sun N, Pei R (2020) Aptamer-based nanostructured interfaces for the detection and release of circulating tumor cells. J Mater Chem B 8(16):3408–3422. https://doi.org/10.1039/c9tb02457c
Tsai WS, You JF, Hung HY, Hsieh PS, Hsieh B, Lenz HJ, Idos G, Friedland S, Yi-Jiun Pan J, Shao HJ et al (2019) Novel circulating tumor cell assay for detection of colorectal adenomas and cancer. Clin Transl Gastroenterol 10(10):e00088. https://doi.org/10.14309/ctg.0000000000000088
Baek DH, Kim GH, Song GA, Han IS, Park EY, Kim HS, Jo HJ, Ko SH, Park DY, Cho YK (2019) Clinical potential of circulating tumor cells in colorectal cancer: a prospective study. Clin Transl Gastroenterol 10(7):e00055. https://doi.org/10.14309/ctg.0000000000000055
Wang L, Zhou S, Zhang W, Wang J, Wang M, Hu X, Liu F, Zhang Y, Jiang B, Yuan H (2019) Circulating tumor cells as an independent prognostic factor in advanced colorectal cancer: a retrospective study in 121 patients. International Journal of Colorectal Disease (1)
Yang C, Shi D, Wang S, Wei C, Zhang C, Xiong B (2018) Prognostic value of pre- and post-operative circulating tumor cells detection in colorectal cancer patients treated with curative resection: a prospective cohort study based on ISET device. Cancer Manag Res 10:4135–4144. https://doi.org/10.2147/CMAR.S176575
Dizdar L, Fluegen G, van Dalum G, Honisch E, Neves RP, Niederacher D, Neubauer H, Fehm T, Rehders A, Krieg A et al (2019) Detection of circulating tumor cells in colorectal cancer patients using the GILUPI Cell Collector: results from a prospective, single-center study. Mol Oncol 13(7):1548–1558. https://doi.org/10.1002/1878-0261.12507
Bidard FC, Kiavue N, Ychou M, Cabel L, Stern MH, Madic J, Saliou A, Rampanou A, Decraene C, Bouche O et al. (2019) Circulating tumor cells and circulating tumor DNA detection in potentially resectable metastatic colorectal cancer: a prospective ancillary study to the Unicancer Prodige-14 Trial. Cells 8 (6). https://doi.org/10.3390/cells8060516
Zhang D, Zhao L, Zhou P, Ma H, Huang F, Jin M, Dai X, Zheng X, Huang S, Zhang T (2017) Circulating tumor microemboli (CTM) and vimentin+ circulating tumor cells (CTCs) detected by a size-based platform predict worse prognosis in advanced colorectal cancer patients during chemotherapy. Cancer Cell Int 17:6. https://doi.org/10.1186/s12935-016-0373-7
Nicolazzo C, Raimondi C, Gradilone A, Emiliani A, Zeuner A, Francescangeli F, Belardinilli F, Seminara P, Loreni F, Magri V et al. (2019) Circulating tumor cells in right- and left-sided colorectal cancer. Cancers (Basel) 11 (8). https://doi.org/10.3390/cancers11081042
Arrazubi V (2019) Circulating tumor cells in patients undergoing resection of colorectal cancer liver metastases. Clinical Utility for LongTerm Outcome: A Prospective Trial. Ann Surg Oncol
Chen CJ, Sung WW, Chen HC, Chern YJ, Hsu HT, Lin YM, Lin SH, Peck K, Yeh KT (2017) Early assessment of colorectal cancer by quantifying circulating tumor cells in peripheral blood: ECT2 in diagnosis of colorectal cancer. Int J Mol Sci 18 (4). https://doi.org/10.3390/ijms18040743
Kim DD, Yang CS, Chae HD, Kwak SG, Jeon CH (2017) Melanoma antigen-encoding gene family member A1–6 and hTERT in the detection of circulating tumor cells following CD45(-) depletion and RNA extraction. Oncol Lett 14(1):837–843. https://doi.org/10.3892/ol.2017.6226
Ning Y, Zhang W, Hanna DL, Yang D, Okazaki S, Berger MD, Miyamoto Y, Suenaga M, Schirripa M, El-Khoueiry A et al (2018) Clinical relevance of EMT and stem-like gene expression in circulating tumor cells of metastatic colorectal cancer patients. Pharmacogenomics J 18(1):29–34. https://doi.org/10.1038/tpj.2016.62
Hinz S, Hendricks A, Wittig A, Schafmayer C, Tepel J, Kalthoff H, Becker T, Roder C (2017) Detection of circulating tumor cells with CK20 RT-PCR is an independent negative prognostic marker in colon cancer patients - a prospective study. BMC Cancer 17(1):53. https://doi.org/10.1186/s12885-016-3035-1
Messaritakis I, Sfakianaki M, Papadaki C, Koulouridi A, Vardakis N, Koinis F, Hatzidaki D, Georgoulia N, Kladi A, Kotsakis A et al (2018) Prognostic significance of CEACAM5mRNA-positive circulating tumor cells in patients with metastatic colorectal cancer. Cancer Chemother Pharmacol 82(5):767–775. https://doi.org/10.1007/s00280-018-3666-9
Cai J, Huang L, Huang J, Kang L, Xian CJ (2018) Associations between the cyclooxygenaseexpression in circulating tumor cells and the clinicopathological features of patients with colorectal cancer. J Cell Biochem 120 (4)
Wang WY, Wan L, Wu SY, Yang JG, Zhou Y, Liu F, Wu ZZ, Cheng Y (2018) Mesenchymal marker and LGR5 expression levels in circulating tumor cells correlate with colorectal cancer prognosis. Cell Oncol 41(5):495–504. https://doi.org/10.1007/s13402-018-0386-4
Satelli A, Mitra A, Brownlee Z, Xia X, Bellister S, Overman MJ, Kopetz S, Ellis LM, Meng QH, Li S (2015) Epithelial-mesenchymal transitioned circulating tumor cells capture for detecting tumor progression. Clin Cancer Res 21(4):899–906. https://doi.org/10.1158/1078-0432.CCR-14-0894
Shi J, Li Y, Liang S, Zeng J, Liu G, Mu F, Li H, Chen J, Liu T, Niu L (2016) Analysis of circulating tumor cells in colorectal cancer liver metastasis patients before and after cryosurgery. Cancer Biol Ther 17(9):935–942. https://doi.org/10.1080/15384047.2016.1210731
Delgado-Urena M, Ortega FG, de Miguel-Perez D, Rodriguez-Martinez A, Garcia-Puche JL, Ilyine H, Lorente JA, Exposito-Hernandez J, Garrido-Navas MC, Delgado-Ramirez M et al (2018) Circulating tumor cells criteria (CyCAR) versus standard RECIST criteria for treatment response assessment in metastatic colorectal cancer patients. J Transl Med 16(1):251. https://doi.org/10.1186/s12967-018-1624-2
Yue C, Jiang Y, Li P, Wang Y, Xue J, Li N, Li D, Wang R, Dang Y, Hu Z et al (2018) Dynamic change of PD-L1 expression on circulating tumor cells in advanced solid tumor patients undergoing PD-1 blockade therapy. Oncoimmunology 7(7):e1438111. https://doi.org/10.1080/2162402X.2018.1438111
Troncarelli Flores BC, Souza ESV, Ali Abdallah E, Mello CAL, Gobo Silva ML, Gomes Mendes G, Camila Braun A, Aguiar Junior S, Thome Domingos Chinen L (2019) Molecular and kinetic analyses of circulating tumor cells as predictive markers of treatment response in locally advanced rectal cancer patients. Cells 8 (7). https://doi.org/10.3390/cells8070641
Huang XZ, Gao P, Song YX, Sun JX, Chen XW, Zhao JH, Liu J, Xu HM, Wang ZN (2014) Relationship between circulating tumor cells and tumor response in colorectal cancer patients treated with chemotherapy: a meta-analysis. BMC Cancer 14(1):976. https://doi.org/10.1186/1471-2407-14-976
Yang C, Zou K, Yuan Z, Guo T, Xiong B (2018) Prognostic value of circulating tumor cells detected with the Cell Search System in patients with gastric cancer: evidence from a meta-analysis. Onco Targets Ther 11:1013–1023. https://doi.org/10.2147/OTT.S154114
Wang PP, Liu SH, Chen CT, Lv L, Li D, Liu QY, Liu GL, Wu Y (2020) Circulating tumor cells as a new predictive and prognostic factor in patients with small cell lung cancer. J Cancer 11(8):2113–2122. https://doi.org/10.7150/jca.35308
Lucci A, Hall CS, Patel SP, Narendran B, Bauldry JB, Royal RE, Karhade M, Upshaw JR, Wargo JA, Glitza IC et al (2020) Circulating tumor cells and early relapse in node-positive melanoma. Clin Cancer Res 26(8):1886–1895. https://doi.org/10.1158/1078-0432.CCR-19-2670
Xiaoyu L, Changyu L, Junhao L, Tianzhu Y, Guofeng Z, Jiemin C, Guoping L, Yang Z, Wenhui L, Xiaolin W (2018) Detection of CTCs in portal vein was associated with intrahepatic metastases and prognosis in patients with advanced pancreatic cancer. J Cancer 9(11):2038–2045
Le UT, Bronsert P, Picardo F, Riethdorf S, Haager B, Rylski B, Czerny M, Beyersdorf F, Wiesemann S, Pantel K et al (2018) Intraoperative detection of circulating tumor cells in pulmonary venous blood during metastasectomy for colorectal lung metastases. Sci Rep 8(1):8751. https://doi.org/10.1038/s41598-018-26410-8
Abreu M, Cabezas-Sainz P, Alonso-Alconada L, Ferreirós A, Mondelo-Macía P, Lago-Lestón RM, Abalo A, Díaz E, Palacios-Zambrano S, Rojo-Sebastian A et al. (2020) Circulating tumor cells characterization revealed TIMP1 as a potential therapeutic target in ovarian cancer. Cells 9 (5). https://doi.org/10.3390/cells9051218
Cai F, Cai L, Zhou Z, Pan X, Wang M, Chen S, Luis MAF, Cen C, Biskup E (2019) Prognostic role of Tif1gamma expression and circulating tumor cells in patients with breast cancer. Mol Med Rep 19(5):3685–3695. https://doi.org/10.3892/mmr.2019.10033
Cheng B, Tong G, Wu X, Cai W, Li Z, Tong Z, He L, Yu S, Wang S (2019) Enumeration and characterization of circulating tumor cells and its application in advanced gastric cancer. Onco Targets Ther 12:7887–7896. https://doi.org/10.2147/OTT.S223222
Welinder C, Jansson B, Lindell G, Wenner J (2015) Cytokeratin 20 improves the detection of circulating tumor cells in patients with colorectal cancer. Cancer Lett 358(1):43–46. https://doi.org/10.1016/j.canlet.2014.12.024
Soler A, Cayrefourcq L, Mazard T, Babayan A, Lamy PJ, Assou S, Assenat E, Pantel K, Alix-Panabières C (2018) Autologous cell lines from circulating colon cancer cells captured from sequential liquid biopsies as model to study therapy-driven tumor changes. Sci Rep 8(1):15931. https://doi.org/10.1038/s41598-018-34365-z
Cayrefourcq L, Mazard T, Joosse S, Solassol J, Ramos J, Assenat E, Schumacher U, Costes V, Maudelonde T, Pantel K et al (2015) Establishment and characterization of a cell line from human circulating colon cancer cells. Cancer Res 75(5):892–901. https://doi.org/10.1158/0008-5472.CAN-14-2613
Pantel K, Alix-Panabieres C (2016) Functional studies on viable circulating tumor cells. Clin Chem 62(2):328–334. https://doi.org/10.1373/clinchem.2015.242537
De T, Goyal S, Balachander G, Chatterjee K, Kumar P, Babu KG, Rangarajan A (2019) A novel ex vivo system using 3D polymer scaffold to culture circulating tumor cells from breast cancer patients exhibits dynamic E-M phenotypes. J Clin Med 8 (9). https://doi.org/10.3390/jcm8091473
Grillet F, Bayet E, Villeronce O, Zappia L, Lagerqvist EL, Lunke S, Charafe-Jauffret E, Pham K, Molck C, Rolland N et al (2017) Circulating tumour cells from patients with colorectal cancer have cancer stem cell hallmarks in culture. Gut 66(10):1802–1810. https://doi.org/10.1136/gutjnl-2016-311447
Dagogo-Jack I, Shaw AT (2018) Tumour heterogeneity and resistance to cancer therapies. Nat Rev Clin Oncol 15(2):81–94. https://doi.org/10.1038/nrclinonc.2017.166
Economopoulou P, Georgoulias V, Kotsakis A (2017) Classifying circulating tumor cells to monitor cancer progression. Expert Rev Mol Diagn 17(2):153–165. https://doi.org/10.1080/14737159.2017.1275572
Ortiz V, Yu M (2018) Analyzing circulating tumor cells one at a time. Trends Cell Biol 28(10):764–775. https://doi.org/10.1016/j.tcb.2018.05.004
Zhu Z, Qiu S, Shao K, Hou Y (2018) Progress and challenges of sequencing and analyzing circulating tumor cells. Cell Biol Toxicol 34(5):405–415. https://doi.org/10.1007/s10565-017-9418-5
Pei H, Li L, Wang Y, Sheng R, Wang Y, Xie S, Shui L, Si H, Tang B (2019) Single-cell phenotypic profiling of CTCs in whole blood using an integrated microfluidic device. Anal Chem 91(17):11078–11084. https://doi.org/10.1021/acs.analchem.9b01647
Tieng FYF, Baharudin R, Abu N, Mohd Yunos RI, Lee LH, Ab Mutalib NS (2020) Single cell transcriptome in colorectal cancer-current updates on its application in metastasis, chemoresistance and the roles of circulating tumor cells. Front Pharmacol 11:135. https://doi.org/10.3389/fphar.2020.00135
Bian S, Hou Y, Zhou X, Li X, Yong J, Wang Y, Wang W, Yan J, Hu B, Guo H et al (2018) Single-cell multiomics sequencing and analyses of human colorectal cancer. Science 362(6418):1060–1063. https://doi.org/10.1126/science.aao3791
Snyder MW, Kircher M, Hill AJ, Daza RM, Shendure J (2016) Cell-free DNA comprises an in vivo nucleosome footprint that informs its tissues-of-origin. Cell 164(1–2):57–68. https://doi.org/10.1016/j.cell.2015.11.050
Aucamp J, Bronkhorst AJ, Badenhorst CPS, Pretorius PJ (2018) The diverse origins of circulating cell-free DNA in the human body: a critical re-evaluation of the literature. Biol Rev Camb Philos Soc 93(3):1649–1683. https://doi.org/10.1111/brv.12413
Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N, Bartlett BR, Wang H, Luber B, Alani RM et al. (2014) Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med 6 (224):224ra224. https://doi.org/10.1126/scitranslmed.3007094
Ye Q, Ling S, Zheng S, Xu X (2019) Liquid biopsy in hepatocellular carcinoma: circulating tumor cells and circulating tumor DNA. Mol Cancer 18(1):114. https://doi.org/10.1186/s12943-019-1043-x
Cheng ML, Pectasides E, Hanna GJ, Parsons HA, Choudhury AD, Oxnard GR (2021) Circulating tumor DNA in advanced solid tumors: clinical relevance and future directions. CA Cancer J Clin 71(2):176–190. https://doi.org/10.3322/caac.21650
Diaz LA Jr, Bardelli A (2014) Liquid biopsies: genotyping circulating tumor DNA. J Clin Oncol 32(6):579–586. https://doi.org/10.1200/JCO.2012.45.2011
Marcuello M, Vymetalkova V, Neves RPL, Duran-Sanchon S, Vedeld HM, Tham E, van Dalum G, Flugen G, Garcia-Barberan V, Fijneman RJ et al (2019) Circulating biomarkers for early detection and clinical management of colorectal cancer. Mol Aspects Med 69:107–122. https://doi.org/10.1016/j.mam.2019.06.002
Wan N, Weinberg D, Liu TY, Niehaus K, Ariazi EA, Delubac D, Kannan A, White B, Bailey M, Bertin M et al (2019) Machine learning enables detection of early-stage colorectal cancer by whole-genome sequencing of plasma cell-free DNA. BMC Cancer 19(1):832. https://doi.org/10.1186/s12885-019-6003-8
Yi X, Ma J, Guan Y, Chen R, Yang L, Xia X (2017) The feasibility of using mutation detection in ctDNA to assess tumor dynamics. Int J Cancer 140(12):2642–2647. https://doi.org/10.1002/ijc.30620
Dawson SJ, Tsui DW, Murtaza M, Biggs H, Rueda OM, Chin SF, Dunning MJ, Gale D, Forshew T, Mahler-Araujo B et al (2013) Analysis of circulating tumor DNA to monitor metastatic breast cancer. N Engl J Med 368(13):1199–1209. https://doi.org/10.1056/NEJMoa1213261
Phallen J, Sausen M, Adleff V, Leal A, Hruban C, White J, Anagnostou V, Fiksel J, Cristiano S, Papp E et al. (2017) Direct detection of early-stage cancers using circulating tumor DNA. Sci Transl Med 9 (403). doi:https://doi.org/10.1126/scitranslmed.aan2415
Wu X, Zhang Y, Hu T, He X, Zou Y, Deng Q, Ke J, Lian L, He X, Zhao D et al (2021) A novel cell-free DNA methylation-based model improves the early detection of colorectal cancer. Mol Oncol 15(10):2702–2714. https://doi.org/10.1002/1878-0261.12942
Bidard FC, Kiavue N, Ychou M, Cabel L, Stern MH, Madic J, Saliou A, Rampanou A, Decraene C, Bouché O et al. (2019) Circulating tumor cells and circulating tumor DNA detection in potentially resectable metastatic colorectal cancer: a prospective ancillary study to the Unicancer Prodige-14 Trial. Cells 8 (6). doi:https://doi.org/10.3390/cells8060516
Tie J, Cohen JD, Lo SN, Wang Y, Li L, Christie M, Lee M, Wong R, Kosmider S, Skinner I et al (2021) Prognostic significance of postsurgery circulating tumor DNA in nonmetastatic colorectal cancer: Individual patient pooled analysis of three cohort studies. Int J Cancer 148(4):1014–1026. https://doi.org/10.1002/ijc.33312
Cremolini C, Rossini D, Dell’Aquila E, Lonardi S, Conca E, Del Re M, Busico A, Pietrantonio F, Danesi R, Aprile G et al (2019) Rechallenge for patients With RAS and BRAF wild-type metastatic colorectal cancer with acquired resistance to first-line cetuximab and irinotecan: a phase 2 single-arm clinical trial. JAMA Oncol 5(3):343–350. https://doi.org/10.1001/jamaoncol.2018.5080
Khakoo S, Carter PD, Brown G, Valeri N, Picchia S, Bali MA, Shaikh R, Jones T, Begum R, Rana I et al (2020) MRI tumor regression grade and circulating tumor DNA as complementary tools to assess response and guide therapy adaptation in rectal cancer. Clin Cancer Res 26(1):183–192. https://doi.org/10.1158/1078-0432.Ccr-19-1996
Dong J, Jan YJ, Cheng J, Zhang RY, Meng M, Smalley M, Chen PJ, Tang X, Tseng P, Bao L et al. (2019) Covalent chemistry on nanostructured substrates enables noninvasive quantification of gene rearrangements in circulating tumor cells. Sci Adv 5 (7):eaav9186. doi:https://doi.org/10.1126/sciadv.aav9186
Radovich M, Jiang G, Hancock BA, Chitambar C, Nanda R, Falkson C, Lynce FC, Gallagher C, Isaacs C, Blaya M et al (2020) Association of circulating tumor DNA and circulating tumor cells after neoadjuvant chemotherapy with disease recurrence in patients with triple-negative breast cancer: preplanned secondary analysis of the BRE12-158 Randomized Clinical Trial. JAMA Oncol. https://doi.org/10.1001/jamaoncol.2020.2295
Lian S, Xie R, Ye Y, Lu Y, Cheng Y, Xie X, Li S, Jia L (2019) Dual blockage of both PD-L1 and CD47 enhances immunotherapy against circulating tumor cells. Sci Rep 9(1):4532. https://doi.org/10.1038/s41598-019-40241-1
Funding
This study was supported by the Chinese Natural Science Foundation (Grant number: 81672970), the State Key Laboratory Open Subjects (Grant number: GZK1202010), the Jiangsu Provincial Health Planning Commission Fund Project (Grant number: CXTDA2017016), the Suzhou Industrial Innovation Project (Grant number: SS202088), the Suzhou Key Medical Center (Grant number: LCZX201505), and the Wujiang Science and Education Project (Grant number: wwk202009).
Author information
Authors and Affiliations
Contributions
Mingchao Hu and Chungen Xing contributed to the idea for the study. Zhili Wang, Zeen Wu, and Pi Ding contributed to the literature search. Renjun Pei and Qiang Wang contributed to the manuscript review.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hu, M., Wang, Z., Wu, Z. et al. Circulating tumor cells in colorectal cancer in the era of precision medicine. J Mol Med 100, 197–213 (2022). https://doi.org/10.1007/s00109-021-02162-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-021-02162-3