Abstract
A sensitive method for diagnosing coronavirus disease 2019 (COVID-19) is highly required to fight the current and future global health threats due to severe acute respiratory syndrome coronavirus 2 (SARS-CoV 2). However, most of the current methods exhibited high false‐negative rates, resulting in patient misdiagnosis and impeding early treatment. Nanoparticles show promising performance and great potential to serve as a platform for diagnosing viral infection in a short time and with high sensitivity. This review highlighted the potential of nanoparticles as platforms for the diagnosis of COVID-19. Nanoparticles such as gold nanoparticles, magnetic nanoparticles, and graphene (G) were applied to detect SARS-CoV 2. They have been used for molecular-based diagnosis methods and serological methods. Nanoparticles improved specificity and shorten the time required for the diagnosis. They may be implemented into small devices that facilitate the self-diagnosis at home or in places such as airports and shops. Nanoparticles-based methods can be used for the analysis of virus-contaminated samples from a patient, surface, and air. The advantages and challenges were discussed to introduce useful information for designing a sensitive, fast, and low-cost diagnostic method. This review aims to present a helpful survey for the lesson learned from handling this outbreak to prepare ourself for future pandemic.
Similar content being viewed by others
Introduction
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV 2) is associated with a pandemic of the acute respiratory disease called coronavirus disease 2019 (COVID-19) [1,2,3,4,5,6,7]. To date (March 2021), there are more than 118 million reported infections and over 2.6 million deaths. Most countries faced mandatory quarantines and lockdowns fighting this threat, leading to an economic crisis [8]. An infected person suffers from fever or chilis, dry cough, sore throat, diarrhea, headache, nausea or vomiting, fatigue, and breathing difficulty. All these symptoms or some of them can be observed. The infection of SARS-COV was also related to the cardiovascular system (CVS) [9], central nervous system (CNS), gastrointestinal tract (GIT) [10], and female reproductive strategies [11]. To date, there is no medicine/drug that has been proved to be effective in treating COVID-19. The only hope now is in the success of the emergency-approved vaccines [12, 13]. The early diagnosis of COVID-19 may help control the spread of the pandemic [14,15,16].
Nanotechnology describes the technology for using particles (nanoparticles) with at least one dimension in the nanometer range (1 nm = 10−9 m, Fig. 1). Nanoparticles exhibit distinct properties such as large surfaces, many active sites, and high adsorption capacities compared to bulk materials. Thus, they have been applied for several applications such as analytical chemistry [17,18,19,20,21,22,23,24,25,26], proteomics [27], sensing/biosensing [28,29,30,31,32,33,34,35], biotechnology [36,37,38,39,40,41,42,43,44,45], nanomedicine [46,47,48,49,50,51,52,53], drug delivery [54,55,56], gene transfer [57,58,59,60], wound healing [61], energy-based applications [62,63,64,65,66,67], and environmental applications [68,69,70,71,72,73,74,75]. Nanoparticles improved these applications by showing high performance [76,77,78,79,80,81,82,83,84] with a great potential for implementation into a miniaturized device, including wearable electronics [85,86,87]. Thus, they exhibit the great potential to improve the quality of life via controlling the viral spread through early detection of infection. Therefore, this review article summarized the current findings and future prospective of nanotechnology for the diagnosis of COVID-19.
Corona virus: structure and infection
Coronavirus (CoVs) refers to virus strain with club-shaped protein spikes on their surface, i.e., crown-like appearance. There are four classes of CoVs, such as alpha, beta, gamma, and delta. All these classes consist of a single-stranded positive-sense ribonucleic acid (RNA) genome. There are two types of alpha-coronaviruses (229E and NL63) and two types of beta-corona viruses (OC43 and HKU1), which can circulate in humans, causing common cold [88]. The human infection by beta coronavirus class (β-CoVs) was previously reported for the severe acute respiratory syndrome (SARS) and the Middle East respiratory syndrome (MERS) [89]. The new coronavirus, i.e., SARS-CoV 2, belongs to the beta class [90]. The infection with SARS-CoV 2 showed human-to-human transmission leading to the spread of infection in more than 122 countries worldwide [91,92,93,94]. The World Health Organization (WHO) and the Centers for Disease Control and Prevention (CDC) warned the people from the spread of the infection via cough and touch [91,92,93].
SARS-CoV 2 virus is an enveloped and non-segmented viruses of a single-stranded RNA genome (4 kilobases) with a nucleocapsid (Fig. 2). It contains two main compartments; protein and non-protein structures [95]. The nucleotide genome of COVID-19 is 80% and 96% identical to the SARS-CoV and the BatCoV RaTG13, respectively [96, 97]. SARS-CoV 2 contains four proteins: (i) S-protein (Spike glycoprotein, PDB: 5XL3), (ii) M-protein (Membrane or Matrix), (iii) E-protein (Envelope), and N- protein (Nucleocapsid phosphoprotein) [98,99,100,101,102,103]. These proteins play a vital role for the infection.
S-protein enables the attachment of the virus to host cells. M-protein promotes the entry of the SARS-CoV 2 virus into the host cells and maintains SARS-CoV 2 viral particles' membrane integrity. The E-protein is the smallest protein and plays a structural role in the assembly of the virus. The N-protein binds to the RNA and supports nucleocapsid formation [98,99,100,101,102,103]. Thus, they can be classified into core protein, e.g., N-protein, and (ii) envelope protein, e.g., S, M, and E proteins. The assembly of these constituents produces a particle size of 100 nm (Fig. 2).
SARS-CoV 2 infects the respiratory system and then spreads systemically to the heart, liver, and kidney [104]. A significant number of patients with SARS-CoV 2 suffers from mild to moderate symptoms. However, 15% of patients with SARS-CoV 2 exhibit severe pneumonia and approximately 5% progress acute respiratory distress syndrome (ARDS), leading to septic shock and multiple organ failure [105, 106]. A study showed that patients with severe SARS-CoV 2 exhibited substantially elevated serum levels of pro-inflammatory cytokines including Interleukin 6 (IL-6) and IL-1β, as well as IL-2, IL-8, IL-17, Granulocyte colony-stimulating factor (G-CSF), Granulocyte–macrophage colony-stimulating factor (GM-CSF), Interferon gamma-induced protein 10 (IP-10), monocyte chemoattractant protein 1 (MCP1), macrophage inflammatory protein (MIP1 α), and tumor necrosis factors (TNF), characterized as cytokine storm [107]. The infected person's analysis showed high expression of angiotensin-converting enzyme 2 (ACE2) in the respiratory tract and other organs [108, 109].
Data analysis (245 surface samples) and air places from hospital rooms of COVID-19 patients for SARS-CoV 2 RNA were reported [110]. The study revealed that 56.7% of the investigated rooms had at least one environmental surface contaminated with the virus (Fig. 3). Air sampling (from airborne infection isolation rooms (AIIRs) in the intensive care unit (ICU) and the general ward) is performed in three of the 27 AIIRs in the general ward. It detects SARS-CoV 2 PCR-positive aerosol particles of sizes > 4 µm and 1–4 µm in two rooms [110]. The analysis of wastewater is critical to monitor the emergence and spread of infectious COVID-19 disease at a population level [111]. There are several protocols for the extraction of the virus or its antigens from wastewater. The detection of SARS-CoV 2 in wastewater is paramount for monitoring public health [112].
Percentage of contaminated swabs from surface samples in hospital rooms. The figure was reprinted from Ref. [110]. This is an Open Access article distributed under the terms of the Creative Commons CC BY license
Nanotechnology
Nanotechnology is the technology of nanoparticles with a size of 1–200 nm (Fig. 2). Nanoparticles can be classified to:
-
1.
Metallic nanoparticles, e.g., gold nanoparticles (Au NPs), silver nanoparticles (Ag NPs);
-
2.
Metal oxide nanoparticles, e.g., iron oxide magnetic nanoparticle (Fe3O4 NPs);
-
3.
Carbon nanomaterials including 0-dimensional (0D, e.g., fullerenes (C60), carbon dots (C-dots)), 1D (carbon nanotunes (CNTs), 2D (e.g., graphene (G), graphene oxide (GO), and 3D (e.g., graphite);
-
4.
Quantum dots (QDs): CdS QDs, CdTe QDs, carbon QDs.
-
5.
Porous materials: metal–organic frameworks (MOFs), covalent organic frameworks (COFs)[113], silica;
-
6.
Polymers: natural polymers (e.g., chitosan, cellulose), and synthetic (e.g., polythiophene, polypyrrole);
-
7.
Lipid nanoparticles (LNPs): triglycerides, fatty acids, steroids, and waxes.
Nanoparticles exhibit several unique properties such as large surface area and simple modification with inorganic [114, 115], organic [116], and biomolecules such as enzymes and protein [117]. They exhibit properties such as catalytic activity [118,119,120,121,122,123], nanozyme [124,125,126], and antimicrobial [127, 128]. The large surface areas of nanoparticles ensure simple modification of their surface via chemical engineering. The particle size of nanomaterials (1–200 nm) is close to the viral particle size of SARS-CoV 2 (Fig. 2). Thus, they interact strongly, leading to significant changes in the electronic properties of nanoparticles. These interactions offer high selectivity and better sensitivity. They can be used for diagnosis, protection, and prevention [129,130,131,132].
Nanoparticles offer distinct properties compared to bulk materials. Metallic nanoparticles such as Au NPs, Ag NPs provide unique optical and electronic properties such as surface plasmonic resonance (SPR) [133,134,135], and localized surface plasmon resonance (LSPR) [136]. Thus, they can be used as a probe for conventional SPR, nanoplasmonic sensors, surface-enhanced Raman spectroscopy (SERS), plasmonic-enhanced fluorescence, and colorimetric method. Some metal oxides or chalcogenides such as molybdenum oxide (MoO3−x) [71] and molybdenum disulfide (MoS2)[137] also exhibited plasmonic properties. Carbon nanomaterials such as graphene show unique electronic, optical, and electrochemical properties [138]. They are promising for point-of-care tests (POCT).
Laboratory diagnosis of COVID-19
The early detection of SARS-COV-2 is necessary to control the widespread infection [139,140,141,142,143,144,145,146,147,148,149,150,151,152,153,154,155,156,157,158,159,160]. Several methods can be used for the diagnosis of COVID-19 [161,162,163,164,165,166,167,168,169,170,171,172,173,174,175,176,177,178,179,180,181,182,183]. The diagnosis of COVID-19 depends on the analysis of the patient's response due to the infection or the analysis of virus contents, e.g., RNA or their protein (Fig. 4). The patient's temperature (elevated temperature), feeling fatigued, and difficulty in breathing indicate infection (Fig. 4). However, these symptoms are lack specificity and may be observed due to the infection with other pathogens. The patient's pathological changes in organs such as the chest can be monitored via computerized tomography (CT) scan (Fig. 4). Like other pneumonia types, a CT scan may be a reliable test for screening SARS-COV 2 cases [184, 185]. However, the analysis required specialized equipment and failed to meet a large scale of requirement, and it may not provide benefit for point-of-care (POC) diagnosis of COVID-19. COVID-19 can be diagnosed via laboratory measurements [106] such as (1) hematologic (the increase in lymphocyte and white cell counts); (2) biochemical change due to the rise of liver function damage biomarker (Lactate dehydrogenase (LDH), α-hydroxybutyric dehydrogenase (α-HBDH), creatine phosphokinase (CPK), AST/ALT ratio (the concentration ratio between enzymes aspirate transaminase (AST) and aka alanine transaminase (ALT)); (3) kidney dysfunction (Creatinine levels in the blood); (4) increase in the inflammation biomarkers (Erythrocyte sedimentation rate (ESR), C-reactive protein (CRP), Procalcitonin (PCT)); (5) changes in the blood properties such as the increase in the time blood takes to clot (Prothrombin time, PT), fibrin degradation fragment (D-dimer) and plasma viscosity (PV). The analysis of markers such as pro-inflammatory cytokines [107] and ACE2 [90, 186] can also be used. These methods are usually used for the analysis of patients. They cannot be used for the analysis of contaminated samples such as surface and air. They are universal and can only be used as an indicator for any infection, i.e., lack of specificity. The laboratory screening is a qualitative analysis method and an indication of an illness related to COVID-19 or other diseases.
Several analytical methods can be used for the diagnosis of COVID-19. There are three main tests called molecular tests (e.g., genetic-based tests or nucleic acid tests (NATs)), antigen tests, and antibodies tests (e.g., serological tests). These methods can be classified to:-
-
I.
Genetic tests (viral nucleic acid tests): analysis of viral genome using techniques such as real-time-quantitative reverse transcription-polymerase chain reaction (RT-qPCR), isothermal amplification (e.g., Loop-mediated isothermal amplification (LAMP), nucleic acid sequence-based amplification (NASBA), transcription-mediated amplification (TMA), rolling circle amplification (RCA), Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)), and nanopore targeted sequencing (NTS) [187].
-
II.
Antigen tests: analysis of the viral proteins (membrane-bound spike proteins or the nucleocapsid proteins) using techniques such as colorimetric, field-effect transistor (FET), enzyme-linked immunosorbent assay (ELISA), and mass spectrometry (MS).
-
III.
Serological tests: analysis of the antibodies (Immunoglobulin M (IgM) and Immunoglobulin G (IgG)) against the virus [188,189,190]. The study of patient's antibodies can be achieved using methods such as electrical (EC) biosensors, localized surface plasmon resonance (LSPR), surface-enhanced Raman scattering (SERS), quartz crystal microbalance (QCM), fluorescence-based biosensor, colorimetric biosensor, gold immunochromatography, ELISA, chemiluminescence immunoassay, and piezoelectric microcantilever sensors (PEMS).
SARS-CoV 2 can be detected in various samples such as feces (1 × 107 copies/mL)[191], urine (1 × 102 copies/mL) [192, 193], saliva (5 × 104 copies/mL) [194], and respiratory tract (103–107 copies/mL)) [195,196,197]. The commonplace for sample collection is respiratory tract (upper part (pharyngeal swabs, nasal swabs, and nasal discharges), and lower part samples (sputum, airway secretions, and bronchoalveolar lavage fluid)). Nasopharyngeal samples are widely used due to the ease of collection, high viral load (103–107 copies/mL), and high stability during transportation or storage [198]. Sample can be self-collected at home via anterior nares swabs. However, the sample collection is painful because it requires the deep insertion of cotton-tipped plastic swabs. Other invasive places such as saliva, feces, and urine can also be used [199,200,201]. However, they contain low viral load and contain interfering species make the use of this specimen challenging for the diagnosis of COVID-19. The concentration of antibodies (IgG and IgM) generated in response to infection is found in blood with a concentration of 0.43–187.82 and 0.26–24.02 (chemiluminescence values divided by the cutoff), respectively.
Nanoparticles offer several advantages for bioanalytical methods that can be applied for the diagnosis of COVID-19. The large surface area of nanoparticles offers high sensitivity. Nanoparticles can be used for the preconcentration and enrichment of the low SARS-COV2 load. The surface can be modified to ensure high specificity.
Thermal scanning
Measuring a patient's body temperature using a thermometer or thermal scanner has been vastly used as one of the detection techniques for monitoring the infection of COVID-19 in public places such as airports, schools, and universities. This method can only detect elevated skin temperatures, which is different from the core body's temperature. Therefore, it lacks high precision to determine whether the individual possesses a fever or not. The body temperature can also be elevated due to other reasons such as exercise, walking, etc. Furthermore, fever is not one of the common symptoms in almost 75% of COVID-19’s patients. Nanoparticles are useful for wearable electronic temperature sensors temperature‐sensitive artificial skin [202].
Genetic-based analysis: PCR, RT-qPCR, LAMP, and CRISPR
The virus analysis, including SARS-CoV 2 using their genetic materials, is based on their nucleic acid analysis via amplification tests (NAATs). However, the content of the gene is usually low. Thus, there are several amplification methods, including PCR, strand displacement assay (SDA), or transcription-mediated assay (TMA). All these methods are based on Watson–Crick base pairing based on a primer molecule (single-stranded probe) capture DNA/ RNA target molecules.
Polymerase chain reaction (PCR) is a standard method for gene analysis. It aims to amplify DNA samples to several million to billions of copies using a tiny DNA sample. The technique uses three reagents: (1) primers (a short single-strand DNA fragments (oligonucleotides) that are a complementary sequence to the target DNA region), (2) a DNA polymerase (heat-stable DNA polymerase, such as Taq polymerase), and (3) deoxynucleoside triphosphates (dNTPs, the building blocks from which the DNA polymerase synthesizes a new DNA strand). The first step of the process includes: (1) denaturation (physical separation of DNA), (2) annealing (the primers attach to each of the single-stranded DNA templates), (3) extension/elongation in which DNA polymerase synthesizes a new DNA strand via adding free dNTPs in the 5′-to-3′ direction. The process is repeated nth cycles to produce 2n copies. The whole process uses 10–200 μL in small reaction tubes (0.2–0.5 mL volumes) in a thermal cycler based on the Peltier effect. It requires a series of 20–40 repeated temperature changes, with each cycle commonly consisting of 2–3 discrete temperature steps.
Real-time quantitative reverse transcription-polymerase chain reaction (RT-qPCR) is a genetic-based method for detecting and quantifying the virus. The procedure is based on converting viral RNA to complementary DNA (cDNA) using the reverse transcription method. In real-time RT-PCR, DNA amplification is monitored in real time as the PCR progresses using a fluorescent dye, a specific DNA probe labeled with a fluorescent molecule, and a quencher molecule (such as TaqMan assays). The process is a repeated amplification process for about 40 cycles until the viral cDNA can be detected, usually by a fluorescent or electrical signal.
The diagnosis of SARS-CoV 2 using RT-qPCR involved several steps: (1) Nasopharyngeal swab (15 min): cotton swab is inserted into the nostril to absorb secretions; (2) Collecting specimen is stored at 2–8 °C for up to 72 h or proceed to; (3) RNA extraction (requires 45 min); (5) The purified RNA is reverse transcribed to cDNA and amplified by qPCR. Positive SARS-CoV 2 patients cross the threshold line within 40 cycles. The specimen for diagnosis of early infection is usually collected via a nasopharyngeal (NP) swab or an oropharyngeal (OP) swab [203, 204]. Collecting combined NP and OP specimens seems to be the most effective approach [204]. A cotton swab must be inserted deeply into the nasal cavity for 10 s. This procedure is painful. RT-qPCR was also used for the detection of SARS-COV 2 in wastewater [205].
Isothermal amplification methods (e.g., recombinase polymerase amplification (RPA) and loop-mediated isothermal amplification (LAMP) were used for a nucleic acid amplification technique. The procedure takes place at one temperature, i.e., isothermal (no need of a thermocycler), and the amplification is continuous. The sample preparation is simple and requires no complicated steps. The method offers high specificity, efficiency, and rapidity under isothermal conditions. Several point-of-care RNA detection technologies that do not require special instruments exist, including reverse transcription–RPA (RT–RPA) and reverse transcription-LAMP (RT-LAMP). The stringency of detection by these isothermal amplification methods can be improved by incorporating an additional sequence-specific detection module, such as hybridization-based fluorescent oligonucleotide probes [206].
A rapid POC diagnostic test (< 20 min) based on RT-LAMP was reported using semiconductor technology (Fig. 5) [207]. The method depends on the detection of SARS-CoV 2 from an extracted RNA samples. The developed LAMP assay was tested on a real-time benchtop instrument (RT-qLAMP), showing a lower limit of detection of 10 RNA copies per reaction [207]. The results showed sensitivity and specificity of 91% and 100%, respectively, compared to RT-qPCR and average positive detection times of 15.45 ± 4.43 min (Fig. 5) [207]. Another POC diagnostic test based on microfluidic platforms was reported to detect viruses using the rolling circle amplification (RCA) method [208]. Viral samples can be detected via DNA hydrogel formation utilizing a platform of isothermal amplification of complementary targets (DhITACT) in microfluidic channels [208]. Self-assembled DNA hydrogel was briefly formed on the surface of microfluidic channels using single-stranded RCA via the isothermal amplification process [208]. These methods are promising for POC diagnosis. They can be used for public services in places such as airports, universities, and shops.
Diagnosis workflow of COVID-19. Figure reprinted with permission Ref. [207].
Clustered regularly interspaced short palindromic repeats (CRISPR)-based diagnostic systems were also proposed for COVID-19 diagnosis. CRISPR is a family of DNA sequences found in the genomes of prokaryotic organisms such as bacteria [209,210,211]. They are used to detect and destroy DNA similarly to bacteriophages during infections. Hence, these sequences play a vital role in the antiviral (i.e., anti-phage) defense system of prokaryotes. The CRISPR-Cas system is a prokaryotic immune system that confers resistance to foreign genetic elements. The CRISPR-Cas9 (CRISPR-associated) genome editing technique awards Nobel Prize in Chemistry 2020. It uses collateral cleavage activity of bystander nucleic acid probes of RNA-guided CRISPR-associated 12/13 (Cas12/13) nucleases [212]. A report incorporated RT-LAMP with CRISPR-Cas12a to detect SARS-CoV 2 in respiratory swab RNA extracts in a colorimetric lateral flow assay [213]. This method is promising and can be conjugated with well-established methods such as the colorimetric method.
Genetic-based methods exhibit several advantages. It offers high specificity. The technique is the standard protocol for the diagnosis of COVID-19. A single-tube assay RT-LAMP, CRISPRs, and the CRISPR-associated (Cas) enzyme Cas12a was recently proposed [214]. The method is simple and can be developed for POC. The technique offered high specificity, although sensitivity depends on the timing of disease presentation, sampling location, and severity of illness [203]. It usually takes about 4–24 h.
The genetic-based method is the standard method for virus analysis. However, it was reported that 20–50% of patients with SARS could not be confirmed by RT-PCR diagnostic [215]. The technique shows only sensitivities of 61–68%, 65–72%, 50–54%, and 58–63% for nasopharyngeal aspirate specimens, throat swab specimens, urine specimens, and stool specimens, respectively [215]. qPCR assay did not produce a detectable target signal in samples containing less than five copies of its amplified DNA target. Real-time RT-PCR may lead to false-negative results due to variations in several possible factors, such as the quality of the collected specimen, the viral load, the source of the reagents, and RNA preparation steps, and fluctuations in the viral load during different phases of the process. RT-qPCR requires sample concentration and preparation. It requires thermal cycling and multiple heating and cooling. It requires various protocols with labor work. The testing procedure requires special laboratory equipment that is often located at a central laboratory. However, portable approaches based on nanopore sequencers are promising for POC detection. The analysis requires at least up to 3 days. Thus, it is a time-consuming method. The sampling is painful. However, other sample places such as tears [216] and saliva [217] were reported. The genetic-based process requires a well-known target sequence to generate the primers that will allow its selective amplification. The small amount of contaminating DNA can also be amplified, resulting in misleading or ambiguous results. The PCR amplification can also be inhibited due to environmental samples such as humic acids. RT–RPA and RT-LAMP methods suffer from nonspecific amplification under isothermal conditions, leading to false-positive results. The false-positive results are exacerbated in the presence of non-sequence-specific probes, such as pH-sensitive dyes [218].
Lateral flow assay based on CRISPR-based diagnostic systems was reported [209]. The method is rapid (< 40 min), easy to implement, and showed high accuracy for the detection of SARS-CoV 2 from respiratory swab RNA extracts [213]. The process extracted the patient's sample RNA via DNA endonuclease-targeted CRISPR trans reporter (DETECTR) [219]. It depends on simultaneous extraction of RNA and detection of predefined coronavirus sequences. This method relies on a custom CRISPR Cas12a/gRNA complex that can be detected via a fluorescent probe (CRISPR-based fluorescent detection system, CRISPR-FDS) [220]. This assay showed a limit of detection (LOD) of 2 copies per sample [220]. Another method using specific high-sensitivity enzymatic reporter unlocking (SHERLOCK) assay was reported [221]. The process showed a LOD of 42 RNA copies per reaction with high specificity and sensitivity of 100% with a fluorescence readout, and 100% specific and 97% sensitive with a lateral flow readout [221].
Analysis based on nucleic acid sequencing such as nanopore sequencing was implemented for the detection of COVID-19. These techniques provide base-pair level information essential to mutation tracing and COVID-19 strain recognition [222, 223]. The method relies on electrophoretic force to translocate DNA, RNA, or protein molecules through an orifice. Nanopore sequencing is commercially available through the Oxford MiniION sequencer [224]. The two-dimensional gold nanoislands (Au NIs) functionalized with complementary DNA receptors can perform a sensitive detection of the selected sequences from SARS-CoV 2 through nucleic acid hybridization. The thermoplasmonic heat is generated on the same Au NIs chip for better sensing performance when illuminated at their plasmonic resonance frequency. The localized PPT heat is capable of elevating the in situ hybridization temperature and facilitating the accurate discrimination of two similar gene sequences. Our dual-functional LSPR biosensor exhibits a high sensitivity toward the selected SARS-CoV 2 sequences with a lower detection limit down to the concentration of 0.22 pM. It allows precise detection of the specific target in a multi-gene mixture [225].
Analysis of nucleic acid using other bioanalytical methods was also reported. Nucleic acid-based electrochemical biosensors were developed to detect the SARS-CoV 2 via the N protein gene's detection using gold nanoparticle-modified electrodes [226]. This method showed good sensitivity. However, it requires sample preparation, such as the RNA extraction step. The plasmonic photothermal (PPT) effect and LSPR were integrated into a dual-functional plasmonic biosensor to detect nucleic acid from SARS-CoV 2 [225]. The device is a chip-based application. The method is based on the use of two-dimensional gold nanoislands (Au NIs) that can form Au–S bonds with the thiol-cDNA receptor of RdRp, ORF1ab, or the E gene sequence. The plasmonic Au NIs generated local PPT heat that promoted fast and sensitive detection of nucleic acids. Au NIs improved the hybridization kinetics of fully matching strands. The dual-functional LSPR sensing system offered selective hybridization detection toward SARS-CoV 2 and SARS-CoV 1. The method provided sensitive detection of SARS-CoV 2 sequences with a LOD of 0.22 pM [225].
An electrochemical-based method using gold nanoparticles (Au NPs) was reported for the analysis of viral nucleocapsid phosphoprotein (N-gene, Fig. 6) [226]. The surface of Au NPs was capped with highly specific antisense oligonucleotides (ssDNA) targeting to ensure high selectivity toward N-gene. The sensing probes, e.g., ssDNA@Au NPs, were immobilized on a graphene paper-based electrochemical platform. The output signals can be recorded with a simple handheld reader [226]. This method offered rapid analysis (< 5 min), showed low cost, and can be easy to implement. It provided quantitative analysis with a broad linear detection range of 585.4–5.854 × 107 copies/μL and sensitivity of 231 copies/μL. It exhibited a LOD of 6.9 copies/μL without the need for nucleic acid amplification [226].
Operation steps for the COVID-19 electrochemical sensing platform: (A) sample collection via the nasal swab or saliva, (B) RNA extraction, (C) immobilization of RNA extract on the top of the graphene-ssDNA-AuNP platform, (D) incubation of 5 min, and (E) record the digital electrochemical output. Figure reprinted with permission from Ref. [226]
An electrochemical biosensor based on RNA amplification using isothermal RCA and differential pulse voltammetry (DPV) was reported (Fig. 7) [227]. It involves 4 simple steps: (1) sample collection, (2) extraction of RNA/DNA, (3) hybridization using RCA reaction, and (4) detection (Fig. 7) [227]. The extracted RNA/DNA was purified and separated using magnetic beads coated with streptavidin. Silica nanoparticles (SiNPs) were coated with two dyes redox system of methylene blue (MB) and acridine orange (AO), producing silica-methylene blue (SiMB) and silica-acridine orange (SiAO). SiMB and SiAO were used for the electrochemical measurements using a portable potentiostat called PalmSens4 connected to a laptop. This assay offers a LOD of 1 copy/μL of N and S genes (Fig. 7) [227]. This method is a one-step sandwich hybridization assay, rapid (2 h), sensitive, and accurate (100% concordance result with qRT-PCR). Furthermore, the potentiostat (PalmSen4) is portable, offering on-site, real-time diagnostic tests for COVID-19 [227].
a Schematic representation showing the detection workflow of SARS-CoV2 and b the detection setup using a portable PalmSens4 potentiostat device connected to a laptop. Figure reprinted with permission from Ref. [227]. This article is licensed under a Creative Commons Attribution 4.0 International License
Nucleic acid analysis using the colorimetric method was also reported. The plasmonic properties of nanoparticles such as Au NPs offer a simple colorimetric analysis of E gene of SARS-CoV 2 [228]. The method depends on the change of the plasmonic peak of Au NPs coated with nucleic acids for visual detection of PCR products of SARS-CoV 2 template [228]. The changes can also be observed via naked eyes [228]. This method is simple, sensitive, and can be implemented into strip-based technologies.
Antigen-bases methods
The SARS-CoV 2 antigen analysis, such as S-protein, N-protein, has been reported using different bioanalytical methods such as electrochemical bases sensors, field-effect transistors (FET). These types of procedures are well stabilized in the literature. They offered high sensitivity and selectivity. They can be easily implemented into portable devices. They did not require high experience for operation.
Electrochemical immunosensor (EC) can be used for viral analysis [229]. An electrochemical immunosensor based on an array of carbon electrodes (DEP)/Au NPs was reported to detect the recombinant on the surface of microfluidic channels [230]. Au NPs were electrodeposited into the glass carbon electrode. The layer of Au NPs immobilized S1 protein for a fixed concentration of antibody (10 µg/mL) that recognized the MERS-CoV antigen. The detection method of MERSA-CoV was based on measuring the current changes for the reduction peak of the Ferro/ferricyanide redox. The reduction in the peak current was due to the electron transfer efficiency reduction due to antibody–antigen binding. The immunosensor required 20 min for the analysis and offered a LOD of 1.0 pg/mL for MERS-CoV protein with a high selectivity degree. The LOD resulted lower than the LOD of ELISA (1 ng/mL). It was successfully applied to spiked nasal samples. Electrochemical sensors offered high sensitivity compared to commercial biosensors such as SPR-based chip (BIAcore systems) for viral analysis such as Influenza virus H1N1 and Dengue virus with LOD of 1 µg/mL, and 2.125 pM, respectively [231, 232]. An electrochemical-based method using electrode-tethered sensors bearing an analyte-binding antibody was reported (Fig. 8) [233]. The surface of the electrode was modified with a negatively charged DNA linker containing a redox system (ferrocene) and antibody (anti-spike) (Fig. 8). The antibody moiety binds to the virus's S-protein, causing changes in the kinetic of transport for a DNA–antibody complex (Fig. 8). This method is reagent-free viral sensing within 5 min (Fig. 8) [233].
An electrochemical approach for monitoring the kinetics of transport for a DNA–antibody complex. The figure was reprinted with permission from Ref. [233]. Copyrights belong to ACS, 2020
A cotton-tipped electrochemical immunosensor was reported for the detection of SARS-CoV 2 virus antigen (Fig. 9) [234]. The carbon nanofiber (CNF) screen-printed electrode was fabricated via diazonium electrografting (Fig. 9). The electrode was further modified with virus nucleocapsid (N) protein using cross-linking chemistry. The changes in square wave voltammetric (SWV) technique before and after interaction can be correlated with the SARS-CoV 2 load. This method offered a LOD of 0.8 pg/mL for SARS-CoV 2 (Fig. 9) [234]. It is a sensitive, selective, and fast analysis procedure. It can also be further modified to ensure high producibility and throughput analysis. The portable PalmSen4 potentiostat was used for S-protein or N-protein detection using magnetic beads to support immunological chain and secondary antibody with alkaline phosphatase [235]. A screen-printed electrode modified with carbon black nanomaterial was used for monitoring the enzymatic byproduct 1-naphthol. This method offered LODs of 19 ng/mL and 8 ng/mL for S and N-protein, respectively [235]. This method is sensitive and required a short analysis time (30 min).
Schematic of the Cotton-Tipped Electrochemical Immunosensor for COVID-19 virus. Figure reprinted with permission from Ref. [234].
Antigen analysis can be achieved using graphene [236], and gold (Au) nanoparticles [237] via FET, and Terahertz plasmonic metasensors, respectively. A novel antibody-based biosensor using FET was reported to detect the S-protein from SARS-CoV 2 (Fig. 10) [236]. The method is based on graphene sheets coated with a specific SARS-CoV 2 antibody (Fig. 10). The technique offered a LOD of 1 fg/mL, 100 fg/mL, and 16 pfu/mL (pfu refers to plaque-forming unit) in phosphate buffer saline, universal transport medium, and culture medium, respectively. The SARS-CoV 2 FET sensor discriminated between infected and non-infected individuals with a LOD of 242 copies/mL [236]. Terahertz plasmonic metasensors using Au NPs were used for the quantitative and qualitative detection of S-protein from SARS-CoV 2 [237]. It offered a LOD of 4.2 fM [237].
(a) Detection of SARS-CoV 2 virus from clinical samples using FET sensor for detection of SAR-CoV-2 virus from COVID-19 patients, (b) Real-time response of COVID-19 FET toward COVID-19 clinical sample and (c) scatter plot and error bar graph of normalized response, (d) Real-time response of FET toward the clinical sample and (e) related dose-dependent response curve. Figure reprinted with permission from Ref. [236]. Copyrights belong to ACS, 2020
Serological-based tests (SB-T)
Serological-based tests (SB-T) measure the body's immune response to the infection. They aim to the analysis of antibodies and proteins in the blood caused due to the infection by pathogens such as SARS-CoV 2. The SB-T measurements aren't related to the virus itself [238]. It is reliable to study the immune response and can be used to identify the recovery or post-infected people. It is also an invaluable test to determine the level of immunity reached in the population. The Food and Drug Administration (FDA) has approved several hundred tests, and most of them are based on molecular technology of immunoglobulin M (IgM) and IgG (Table 1) [239].
The detection of IgG and IgM in serum or whole blood has been demonstrated to be a reliable method for diagnosing COVID-19 with high specificity and sensitivity [240]. A point‐of‐care lateral flow immunoassay (LFIA) test using Au nanoparticles was used for the detection of IgM and IgG antibodies simultaneously against SARS‐CoV 2 virus in human blood (Fig. 11) [241]. The test strip's main body consists of five parts, including plastic backing, sample pad, conjugate pad, absorbent pad, and NC membrane (Fig. 11). The analysis using this method can be achieved within 15 min at 88.66% with a specificity of 90.63% [241]. A colorimetric method using Au NPs was also reported [242]. The analysis takes place in a solution. Au NPs were functionalized with antibodies for targeting three surface proteins of SARS-CoV 2, e.g., spike, envelope, and membrane. The detection of these proteins depends on the red-shifted SPR peak of Au NPs. The analysis requires few minutes [242]. These methods offer high sensitivity, selectivity and needed short analysis time (Table 1).
a Schematic illustration of a point‐of‐care lateral flow immunoassay (LFIA) for SARS‐CoV 2 IgM‐IgG combined antibody test, and b an illustration of different testing results; C refers to control line; G refers to IgG line; M refers to IgM line. Figure reprinted with permission from Ref. [241]. Reference [241] is an Open Access article distributed under the terms of the Creative Commons CC BY license
Magnetic chemiluminescence enzyme immunoassay (MCLIA) was reported for the detection of SARS-COV 2 via the analysis of IgG and IgM [240]. MCLIA showed high specificity and sensitivity in detecting serum IgG and IgM [240]. The positive rate of IgG and IgM was 71.4% and 57.2%, respectively [240]. The method exhibited high specificity compared to other respiratory pathogens, including influenza A virus, influenza B virus, parainfluenza virus, with high stability (coefficient of variation (CV) was below 6%) [240]. The serological testing using MCLIA may be helpful for the diagnosis of suspected patients with negative RT–PCR results and for the identification of asymptomatic infections [243].
A portable microfluidic immunoassay system was proposed to detect SARS-CoV 2 (Fig. 12) [244]. This method was easy to use, sensitive, rapid (< 15 min) and offered on-site detection of IgG/IgM/Antigen of SARS-CoV 2 simultaneously [244]. This method's cost is only about 5 yuan (0.71 dollars) and required only 10 min for analysis. The analysis required a spot of 10 μL of the specimen (blood, serum, plasma, pharyngeal swabs, alveolar lavage fluid, or fecal suspension) into the loading chamber of the microchip (Fig. 12). This was followed by the addition of sample dilution buffer (70 μL). The biomarkers of SARS-CoV 2 (IgG/IgM/antigen) are specifically bound to the fluorescent microspheres (FMS) labeled capture antibody [244]. The method showed the detection of the virus on different days for patients. It was also approved by the Center for Medical Device Evaluation (CMDE) in China and obtained European CE certification (Fig. 12) [244]. Groltex and Sanford Burnham Prebys Medical Discovery Institute developed a graphene-based biosensor platform consisting of deposited gold nanoislands, handheld reader units, and disposable plastic testing chips [245]. The "graphene sensor chip on plastic" platform required a small biological sample and can perform up to 4 to 12 viral tests [245]. Gold nanospikes were fabricated using electrodeposition and applied for pto-microfluidic sensing platform (localized surface plasmon resonance (LSPR) and microfluidic device)[246]. The antibody concentration was correlated with the LSPR wavelength peak shift of Au nanospikes caused due to the local refractive index change because of the antigen–antibody binding [246]. This method is a label-free microfluidic platform with a LOD of 0.08 ng/mL (0.5 pM) [246].
(A) Photograph of the portable home-made fluorescence detection equipment; (B) photograph of the immunoassay microchip ready to use; (C) schematic illustration of the microfluidic fluorescence immunoassay for IgG/IgM/antigen detection of SARS CoV 2. Figure reprinted with permission from Ref. [244].
Other methods
A functionalized magnetic nanoparticle (MNPs) using S-protein was proposed for the detection of SARS-CoV 2 using a mimic SARS-CoV 2 consisting of spike proteins and polystyrene beads (Fig. 13) [247]. The method depends on measuring the magnetic response of MNPs in an ac magnetic field with and without the virus (Fig. 13). This method offered rapid and sensitive detection of SARS-CoV 2 with a LOD of 0.084 nM (5.9 fmoles) [247].
Principle of magnetic field changes using (a) functionalized MNPs with and without (b) mimic SARS-CoV 2, and (c) the MPS signals response with and without mimic virus. Figure reprinted with permission from Ref. [247].
Mass spectrometry (MS) has advanced the analysis of virus [248]. MS depends on the ionization of the viral contents, e.g., protein or RNA. Soft ionization techniques such as matrix-assisted laser desorption/ionization mass spectrometry (MALDI-MS) are promising for direct detection of the intact virus [25, 41, 76, 249,250,251,252,253,254,255,256]. Nanoparticles are essential as the surface for MALDI-MS, i.e., surface-assisted laser desorption/ionization mass spectrometry (SALDI-MS). The method is label-free, offering high sensitivity. The presence of proteins such as S, E, N, and M enables simple analysis for SARS-CoV 2 using mass spectrometry [257]. Blood analysis using advanced MS-based proteomics approaches will help identify the essential protein patterns of COVID-19′s patients [258].
Concluding remarks
The past investments to discover viruses, detection, and antiviral treatment have rarely been made in the past, except for very few viruses. Thus, coronavirus has several accessory proteins that seem to be essential for the virus [100]; however, their functions aren't entirely characterized. Our knowledge about this novel virus remains very limited [94]. There are not enough nanotechnology approaches being explored to tackle the current outbreak using a sensitive diagnostic method. This task requires network coordination among pharmaceutical companies, scientists, governments, and WHO. The virus's analysis inside the body via analytical techniques such as imaging and tracking their places is highly required and essential for future threats.
The surface contamination by SARS-CoV 2 has been found to be more significant in the spread of COVID-19. The contamination of latex/nitrile gloves, N95 respirators, hospital scrubs, overshoes, and floors in a nosocomial environment has been considered a severe issue [259, 260]. It causes uncontrolled spreading of the disease and passes the infection to other patients and their families [261]. The detection of SARS-CoV 2 contamination using a simple bioanalytical method for these necessary tools is essential for protecting people in contact with infected persons. It is supposed to control the virus's spread and alert people by the infected places and surfaces.
Serological-based tests may help diagnose suspected patients with negative RT–PCR results and identify asymptomatic infections [243]. The test is susceptible, specific, and required a short time. However, the results of these methods may be varied significantly with interferences [262]. These methods lack high accuracy for the analysis within the first days after the onset of disease [263]. Computed tomography imaging, whole-genome sequencing, and electron microscopy are more precise [264]. A study indicated that an initial chest CT has a higher detection rate (98%) compared to reverse transcriptase‐polymerase chain reaction (RT‐PCR) (70%) in infected patients. Analysis using genetic-based methods such as PCR demonstrated that about 3% of patients have no primary positive RT‐PCR but have a positive chest CT; therefore, both tests are recommended for COVID‐19 patients [265]. Electrochemical-based methods conjugated nanotechnology is promising for several reasons [266]. They can be used for gene detection of (1) gene such as nucleocapsid phosphoprotein (N-gene) [226], RdRp gene [267]; (2) antigen [234]; and (3) antibodies. The method can be easily implemented into portable devices offering simple operation and on-site detection [227].
Nanotechnology may help to improve diagnosis [268, 269] using techniques such as advanced POCT (point of care tests) approaches [139]. The future development of portable microfluidic-based cartridges and lyophilized reagents to run the assay could enable POCT outside of the clinical diagnostic laboratory, such as airports, local emergency departments, and other clinic locations [270]. US Food and Drug Administration (FDA) approved several handheld methods for detecting COVID-19 (Table 2). The approved devices depend on the analysis of nucleic acid [271], biomarker IL-6 and CRP [272]. However, they are still costly and lack to quantify the total viral load in the infected person. There is no standard value to represent the viral load.
The sample collection and data analysis require careful investigation. There is no test with complete accuracy (100%) all of the time. Several things may affect the test’s accuracy. A person may have the virus infection, but the swab might not collect the virus from the nose or throat due to short contact time or low viral load. The virus may be present due to accidental contamination during sample collection, storage, or analysis. The storage conditions may also destroy the virus due to incorrect temperature or contamination with a disinfectant such as alcohol. The process and chemicals used during the analysis or sample preparation, such as RNA extraction, may affect the test result. These points should be considered during sample collection, storage, and analysis.
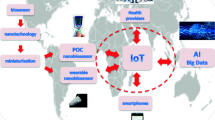
Nanotechnology can advance the diagnosis of COVID-19 and offer an advanced diagnostic approach based on a POC sensing technology [294]. It can be interfaced with the Internet of things artificial intelligence (AI) techniques, [295], and internet of medical things (IoMT)-integrated biosensors [296] for investigating practical informatics via data storage, sharing, and analytics. They can circumvent conventional techniques such as low sensitivity, low selectivity, high cost, and long diagnostic time [297]. They can be used for no pain sample analysis, such as the analysis of patient's saliva using graphene oxide (GO)/Au/Fiber Bragg grating (FBG) probe [298]. Nanotechnology can be used to advance technologies such as paper lateral flow assays [299], label-free biosensors [300,301,302], optical technologies [160, 303], and digital technologies [304].
References
WHO (2020) WHO. https://www.who.int/emergencies/diseases/novel-coronavirus-2019
Cao X (2020) COVID-19: immunopathology and its implications for therapy. Nat Rev Immunol 20:269–270. https://doi.org/10.1038/s41577-020-0308-3
Le Thanh T, Andreadakis Z, Kumar A et al (2020) The COVID-19 vaccine development landscape. Nat Rev Drug Discov 19:305–306. https://doi.org/10.1038/d41573-020-00073-5
Anderson RM, Heesterbeek H, Klinkenberg D, Hollingsworth TD (2020) How will country-based mitigation measures influence the course of the COVID-19 epidemic? Lancet 395:931–934. https://doi.org/10.1016/S0140-6736(20)30567-5
Fontanet A, Cauchemez S (2020) COVID-19 herd immunity: where are we? Nat Rev Immunol 20:583–584. https://doi.org/10.1038/s41577-020-00451-5
Zhou F, Yu T, Du R et al (2020) Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet 395:1054–1062. https://doi.org/10.1016/S0140-6736(20)30566-3
Clinical and virologic characteristics of the first 12 patients with coronavirus disease (2019) (COVID-19) in the United States. Nat Med 26:861–868. https://doi.org/10.1038/s41591-020-0877-5
Callaway E (2020) Time to use the p-word? Coronavirus enters dangerous new phase. Nature. https://doi.org/10.1038/d41586-020-00551-1
Zheng Y-Y, Ma Y-T, Zhang J-Y, Xie X (2020) COVID-19 and the cardiovascular system. Nat Rev Cardiol 17:259–260. https://doi.org/10.1038/s41569-020-0360-5
Xiao F, Tang M, Zheng X et al (2020) Evidence for gastrointestinal infection of SARS-CoV-2. Gastroenterology 158:1831-1833.e3. https://doi.org/10.1053/j.gastro.2020.02.055
Li M, Chen L, Zhang J et al (2020) The SARS-CoV-2 receptor ACE2 expression of maternal-fetal interface and fetal organs by single-cell transcriptome study. PLoS ONE 15:e0230295. https://doi.org/10.1371/journal.pone.0230295
Powell AE, Zhang K, Sanyal M et al (2021) A single immunization with spike-functionalized ferritin vaccines elicits neutralizing antibody responses against SARS-CoV-2 in mice. ACS Cent Sci 7:183–199. https://doi.org/10.1021/acscentsci.0c01405
Chaudhary V, Royal A, Chavali M, Yadav SK (2021) Advancements in research and development to combat COVID-19 using nanotechnology. Nanotechnol Environ Eng 6:8. https://doi.org/10.1007/s41204-021-00102-7
Hosseini A, Pandey R, Osman E et al (2020) Roadmap to the bioanalytical testing of COVID-19: from sample collection to disease surveillance. ACS Sensors 5:3328–3345. https://doi.org/10.1021/acssensors.0c01377
Aziz A, Asif M, Ashraf G et al (2021) Trends in biosensing platforms for SARS-CoV-2 detection: a critical appraisal against standard detection tools. Curr Opin Colloid Interface Sci. https://doi.org/10.1016/j.cocis.2021.101418
Kotru S, Klimuntowski M, Ridha H et al (2021) Electrochemical sensing: a prognostic tool in the fight against COVID-19. TrAC Trends Anal Chem. https://doi.org/10.1016/j.trac.2021.116198
Wu H-F, Gopal J, Abdelhamid HN, Hasan N (2012) Quantum dot applications endowing novelty to analytical proteomics. Proteomics 12:2949–2961. https://doi.org/10.1002/pmic.201200295
Abdelhamid HN, Wu H-F (2012) A method to detect metal-drug complexes and their interactions with pathogenic bacteria via graphene nanosheet assist laser desorption/ionization mass spectrometry and biosensors. Anal Chim Acta 751:94–104. https://doi.org/10.1016/j.aca.2012.09.012
Abdelhamid HN, Wu H-F (2015) Proteomics analysis of the mode of antibacterial action of nanoparticles and their interactions with proteins. TrAC Trends Anal Chem 65:30–46. https://doi.org/10.1016/j.trac.2014.09.010
Abdelhamid HN (2015) Ionic liquids for mass spectrometry: matrices, separation and microextraction. TrAC Trends Anal Chem 77:122–138. https://doi.org/10.1016/j.trac.2015.12.007
Abdelhamid HN, Gopal J, Wu H-FF (2013) Synthesis and application of ionic liquid matrices (ILMs) for effective pathogenic bacteria analysis in matrix assisted laser desorption/ionization (MALDI-MS). Anal Chim Acta 767:104–111. https://doi.org/10.1016/j.aca.2012.12.054
Yang Y, Shen K, Lin J et al (2016) A Zn-MOF constructed from electron-rich π-conjugated ligands with an interpenetrated graphene-like net as an efficient nitroaromatic sensor. RSC Adv 6:45475–45481. https://doi.org/10.1039/C6RA00524A
Nasser Abdelhamid H, Wu B-S, Wu H-F (2014) Graphene coated silica applied for high ionization matrix assisted laser desorption/ionization mass spectrometry: a novel approach for environmental and biomolecule analysis. Talanta 126:27–37. https://doi.org/10.1016/j.talanta.2014.03.016
Abdelhamid HN (2017) Organic matrices, ionic liquids, and organic matrices@nanoparticles assisted laser desorption/ionization mass spectrometry. TrAC Trends Anal Chem 89:68–98. https://doi.org/10.1016/j.trac.2017.01.012
Abdelhamid HN (2018) Nanoparticle assisted laser desorption/ionization mass spectrometry for small molecule analytes. Microchim Acta 185:200. https://doi.org/10.1007/s00604-018-2687-8
Abdelhamid HN, Bermejo-Gómez A, Martín-Matute B, Zou X (2017) A water-stable lanthanide metal–organic framework for fluorimetric detection of ferric ions and tryptophan. Microchim Acta 184:3363–3371. https://doi.org/10.1007/s00604-017-2306-0
Abdelhamid HN, Kumaran S, Wu H-F (2016) One-pot synthesis of CuFeO2 nanoparticles capped with glycerol and proteomic analysis of their nanocytotoxicity against fungi. RSC Adv 6:97629–97635. https://doi.org/10.1039/C6RA13396G
Abdelhamid HN, Wu H-F (2018) Selective biosensing of Staphylococcus aureus using chitosan quantum dots. Spectrochim Acta Part A Mol Biomol Spectrosc 188:50–56. https://doi.org/10.1016/j.saa.2017.06.047
Abdelhamid HN (2017) Lanthanide metal–organic frameworks and hierarchical porous zeolitic imidazolate frameworks: synthesis, properties, and applications. Stockholm University, Faculty of Science. http://urn.kb.se/resolve?urn=urn:nbn:se:su:diva-146398
Abdelhamid HN, Wu H-F (2015) Synthesis and multifunctional applications of quantum nanobeads for label-free and selective metal chemosensing. RSC Adv 5:50494–50504. https://doi.org/10.1039/C5RA07069D
Farghaly AA, Lam M, Freeman CJ et al (2016) Potentiometric measurements in biofouling solutions: comparison of nanoporous gold to planar gold. J Electrochem Soc 163:H3083–H3087. https://doi.org/10.1149/2.0101604jes
Freeman CJ, Farghaly AA, Choudhary H et al (2016) Microdroplet-based potentiometric redox measurements on gold nanoporous electrodes. Anal Chem 88:3768–3774. https://doi.org/10.1021/acs.analchem.5b04668
Khan RK, Farghaly AA, Silva TA et al (2019) Gold-nanoparticle-decorated titanium nitride electrodes prepared by glancing-angle deposition for sensing applications. ACS Appl Nano Mater 2:1562–1569. https://doi.org/10.1021/acsanm.8b02354
Farghaly AA, Khan RK, Collinson MM (2018) Biofouling-resistant platinum bimetallic alloys. ACS Appl Mater Interfaces 10:21103–21112. https://doi.org/10.1021/acsami.8b02900
Zulfajri M, Abdelhamid HN, Sudewi S et al (2020) Plant part-derived carbon dots for biosensing. Biosensors 10:68. https://doi.org/10.3390/bios10060068
Abdelhamid HN, Wu H-F (2013) Multifunctional graphene magnetic nanosheet decorated with chitosan for highly sensitive detection of pathogenic bacteria. J Mater Chem B 1:3950–3961. https://doi.org/10.1039/c3tb20413h
Abdelhamid HN, Wu H-F (2013) Probing the interactions of chitosan capped CdS quantum dots with pathogenic bacteria and their biosensing application. J Mater Chem B 1:6094–6106. https://doi.org/10.1039/c3tb21020k
Manikandan M, Nasser Abdelhamid H, Talib A, Wu H-F (2014) Facile synthesis of gold nanohexagons on graphene templates in Raman spectroscopy for biosensing cancer and cancer stem cells. Biosens Bioelectron 55:180–186. https://doi.org/10.1016/j.bios.2013.11.037
Gopal J, Abdelhamid HN, Hua P-Y, Wu H-F (2013) Chitosan nanomagnets for effective extraction and sensitive mass spectrometric detection of pathogenic bacterial endotoxin from human urine. J Mater Chem B 1:2463. https://doi.org/10.1039/c3tb20079e
Abdelhamid HN, Wu H-F (2016) Gold nanoparticles assisted laser desorption/ionization mass spectrometry and applications: from simple molecules to intact cells. Anal Bioanal Chem 408:4485–4502. https://doi.org/10.1007/s00216-016-9374-6
Bhaisare ML, Abdelhamid HN, Wu B-S, Wu H-F (2014) Rapid and direct MALDI-MS identification of pathogenic bacteria from blood using ionic liquid-modified magnetic nanoparticles (Fe3O4@SiO2). J Mater Chem B 2:4671–4683. https://doi.org/10.1039/C4TB00528G
Abdelhamid HN, Wu H-F (2014) Facile synthesis of nano silver ferrite (AgFeO2) modified with chitosan applied for biothiol separation. Mater Sci Eng C 45:438–445. https://doi.org/10.1016/j.msec.2014.08.071
Hua P-Y, Manikandan M, Abdelhamid HN, Wu H-F (2014) Graphene nanoflakes as an efficient ionizing matrix for MALDI-MS based lipidomics of cancer cells and cancer stem cells. J Mater Chem B 2:7334–7343
Abdelhamid HN, Wu H-F (2014) Polymer dots for quantifying the total hydrophobic pathogenic lysates in a single drop. Colloids Surf B Biointerfaces 115:51–60. https://doi.org/10.1016/j.colsurfb.2013.11.013
Sekar R, Kailasa SK, Abdelhamid HN et al (2013) Electrospray ionization tandem mass spectrometric studies of copper and iron complexes with tobramycin. Int J Mass Spectrom 338:23–29. https://doi.org/10.1016/j.ijms.2012.12.001
Shahnawaz Khan M, Abdelhamid HN, Wu H-F (2015) Near infrared (NIR) laser mediated surface activation of graphene oxide nanoflakes for efficient antibacterial, antifungal and wound healing treatment. Colloids Surf B Biointerfaces 127C:281–291. https://doi.org/10.1016/j.colsurfb.2014.12.049
Abdelhamid HN, Khan MS, Wu HF (2014) Graphene oxide as a nanocarrier for gramicidin (GOGD) for high antibacterial performance. RSC Adv 4:50035–50046. https://doi.org/10.1039/c4ra07250b
Wu B-S, Abdelhamid HN, Wu H-F (2014) Synthesis and antibacterial activities of graphene decorated with stannous dioxide. RSC Adv 4:3722. https://doi.org/10.1039/c3ra43992e
Abdelhamid HN, Talib A, Wu H-F (2015) Facile synthesis of water soluble silver ferrite (AgFeO2) nanoparticles and their biological application as antibacterial agents. RSC Adv 5:34594–34602. https://doi.org/10.1039/C4RA14461A
Dowaidar M, Abdelhamid HN, Hällbrink M et al (2017) Graphene oxide nanosheets in complex with cell penetrating peptides for oligonucleotides delivery. Biochim Biophys Acta - Gen Subj 1861:2334–2341. https://doi.org/10.1016/j.bbagen.2017.07.002
Dowaidar M, Abdelhamid HN, Hällbrink M et al (2017) Magnetic nanoparticle assisted self-assembly of cell penetrating peptides-oligonucleotides complexes for gene delivery. Sci Rep 7:9159. https://doi.org/10.1038/s41598-017-09803-z
Abdelhamid HN, Talib A, Wu HF (2017) One pot synthesis of gold–carbon dots nanocomposite and its application for cytosensing of metals for cancer cells. Talanta 166:357–363. https://doi.org/10.1016/j.talanta.2016.11.030
Abdelhamid HN (2016) Laser assisted synthesis, imaging and cancer therapy of magnetic nanoparticles. Mater Focus 5:305–323. https://doi.org/10.1166/mat.2016.1336
Abdelhamid HN, El-Bery HM, Metwally AA et al (2019) Synthesis of CdS-modified chitosan quantum dots for the drug delivery of Sesamol. Carbohydr Polym 214:90–99. https://doi.org/10.1016/j.carbpol.2019.03.024
Keservani R, Sharma A, Abdelhamid H (2019) Nanoparticulate drug delivery systems. CRC Press, Boca Raton
Abdelhamid HN, Wu H-F (2019) Nanoparticles advance drug delivery for cancer cells. In: Keservani RK, Sharma AK (eds) Nanoparticulate drug delivery systems. Apple Academic Press, Palm Bay, pp 121–150
Dowaidar M, Nasser Abdelhamid H, Hällbrink M et al (2018) Chitosan enhances gene delivery of oligonucleotide complexes with magnetic nanoparticles–cell-penetrating peptide. J Biomater Appl 33:392–401. https://doi.org/10.1177/0885328218796623
Abdelhamid HN, Dowaidar M, Hällbrink M, Langel Ü (2019) Cell penetrating peptides-hierarchical porous zeolitic imidazolate frameworks nanoparticles: an efficient gene delivery platform. SSRN Electron J. https://doi.org/10.2139/ssrn.3435895
Abdelhamid HN, Dowaidar M, Langel Ü (2020) Carbonized chitosan encapsulated hierarchical porous zeolitic imidazolate frameworks nanoparticles for gene delivery. Microporous Mesoporous Mater 302:110200. https://doi.org/10.1016/j.micromeso.2020.110200
Abdelhamid HN, Dowaidar M, Hällbrink M, Langel Ü (2020) Gene delivery using cell penetrating peptides-zeolitic imidazolate frameworks. Microporous Mesoporous Mater 300:110173. https://doi.org/10.1016/j.micromeso.2020.110173
Hussein KH, Abdelhamid HN, Zou X, Woo H-M (2019) Ultrasonicated graphene oxide enhances bone and skin wound regeneration. Mater Sci Eng C 94:484–492. https://doi.org/10.1016/j.msec.2018.09.051
Abdelhamid HN, El-Zohry AM, Cong J et al (2019) Towards implementing hierarchical porous zeolitic imidazolate frameworks in dye-sensitized solar cells. R Soc Open Sci 6:190723. https://doi.org/10.1098/rsos.190723
Goda MN, Abdelhamid HN, Said AE-AA (2020) Zirconium Oxide Sulfate-Carbon (ZrOSO4@C) Derived from Carbonized UiO-66 for Selective Production of Dimethyl Ether. ACS Appl Mater Interfaces 12:646–653. https://doi.org/10.1021/acsami.9b17520
Kassem AA, Abdelhamid HN, Fouad DM, Ibrahim SA (2019) Metal–organic frameworks (MOFs) and MOFs-derived CuO@C for hydrogen generation from sodium borohydride. Int J Hydrogen Energy 44:31230–31238. https://doi.org/10.1016/j.ijhydene.2019.10.047
Nahar L, Farghaly AA, Esteves RJA, Arachchige IU (2017) Shape controlled synthesis of Au/Ag/Pd nanoalloys and their oxidation-induced self-assembly into electrocatalytically active aerogel monoliths. Chem Mater 29:7704–7715. https://doi.org/10.1021/acs.chemmater.7b01731
Van Cleve T, Wang G, Mooney M et al (2021) Tailoring electrode microstructure via ink content to enable improved rated power performance for platinum cobalt/high surface area carbon based polymer electrolyte fuel cells. J Power Sources 482:228889. https://doi.org/10.1016/j.jpowsour.2020.228889
Abdelhamid HN (2020) Salts induced formation of hierarchical porous ZIF-8 and their applications for CO2 sorption and hydrogen generation via NaBH4 hydrolysis. Macromol Chem Phys 221:2000031. https://doi.org/10.1002/macp.202000031
Abdelhamid HN (2020) Zinc hydroxide nitrate nanosheets conversion into hierarchical zeolitic imidazolate frameworks nanocomposite and their application for CO2 sorption. Mater Today Chem 15:100222. https://doi.org/10.1016/j.mtchem.2019.100222
Yao Q, Bermejo Gómez A, Su J et al (2015) Series of highly stable isoreticular lanthanide metal–organic frameworks with expanding pore size and tunable luminescent properties. Chem Mater 27:5332–5339. https://doi.org/10.1021/acs.chemmater.5b01711
Abdelhamid HN, Bhaisare ML, Wu H-F (2014) Ceria nanocubic-ultrasonication assisted dispersive liquid-liquid microextraction coupled with matrix assisted laser desorption/ionization mass spectrometry for pathogenic bacteria analysis. Talanta 120:208–217. https://doi.org/10.1016/j.talanta.2013.11.078
Etman AS, Abdelhamid HN, Yuan Y et al (2018) Facile water-based strategy for synthesizing MoO3–x nanosheets: efficient visible light photocatalysts for dye degradation. ACS Omega 3:2193–2201. https://doi.org/10.1021/acsomega.8b00012
Ashour RM, Abdelhamid HN, Abdel-Magied AF et al (2017) Rare earth ions adsorption onto graphene oxide nanosheets. Solvent Extr Ion Exch 35:91–103. https://doi.org/10.1080/07366299.2017.1287509
Abdelhamid HN, Zou X (2018) Template-free and room temperature synthesis of hierarchical porous zeolitic imidazolate framework nanoparticles and their dye and CO 2 sorption. Green Chem 20:1074–1084. https://doi.org/10.1039/C7GC03805D
Abdelhamid HN, Wu H-F (2015) Synthesis of a highly dispersive sinapinic acid@graphene oxide (SA@GO) and its applications as a novel surface assisted laser desorption/ionization mass spectrometry for proteomics and pathogenic bacteria biosensing. Analyst 140:1555–1565
Abdelhamid HN (2015) Delafossite nanoparticle as new functional materials: advances in energy, nanomedicine and environmental applications. Mater Sci Forum 832:28–53. https://doi.org/10.4028/www.scientific.net/MSF.832.28
Nasser Abdelhamid H, Wu HF (2013) Furoic and mefenamic acids as new matrices for matrix assisted laser desorption/ionization-(MALDI)-mass spectrometry. Talanta 115:442–450. https://doi.org/10.1016/j.talanta.2013.05.050
Abdelhamid HN, Khan MS, Wu H-FF (2014) Design, characterization and applications of new ionic liquid matrices for multifunctional analysis of biomolecules: a novel strategy for pathogenic bacteria biosensing. Anal Chim Acta 823:51–60. https://doi.org/10.1016/j.aca.2014.03.026
Gopal J, Abdelhamid HN, Huang JH, Wu HF (2016) Nondestructive detection of the freshness of fruits and vegetables using gold and silver nanoparticle mediated graphene enhanced Raman spectroscopy. Sens Actuators B Chem 224:413–424. https://doi.org/10.1016/j.snb.2015.08.123
Valencia L, Abdelhamid HN (2019) Nanocellulose leaf-like zeolitic imidazolate framework (ZIF-L) foams for selective capture of carbon dioxide. Carbohydr Polym 213:338–345. https://doi.org/10.1016/j.carbpol.2019.03.011
Abdel-Magied AF, Abdelhamid HN, Ashour RM et al (2019) Hierarchical porous zeolitic imidazolate frameworks nanoparticles for efficient adsorption of rare-earth elements. Microporous Mesoporous Mater 278:175–184. https://doi.org/10.1016/j.micromeso.2018.11.022
Abdelhamid HN (2015) Ionic liquids matrices for laser assisted desorption/ionization mass spectrometry. Mass Spectrom Purif Tech. https://doi.org/10.4172/2469-9861.1000109
Iqbal MN, Abdel-Magied AF, Abdelhamid HN et al (2017) Mesoporous ruthenium oxide: a heterogeneous catalyst for water oxidation. ACS Sustain Chem Eng 5:9651–9656. https://doi.org/10.1021/acssuschemeng.7b02845
Abdelhamid HN, Wilk-Kozubek M, El-Zohry AM et al (2019) Luminescence properties of a family of lanthanide metal–organic frameworks. Microporous Mesoporous Mater 279:400–406. https://doi.org/10.1016/j.micromeso.2019.01.024
Abdelhamid HN (2016) Physicochemical properties of proteomic ionic liquids matrices for MALDI-MS. J Data Min Genom Proteom 7(2153–0602):1000
Tricoli A, Nasiri N, De S (2017) Wearable and miniaturized sensor technologies for personalized and preventive medicine. Adv Funct Mater 27:1605271. https://doi.org/10.1002/adfm.201605271
Pal RK, Farghaly AA, Collinson MM et al (2016) Photolithographic micropatterning of conducting polymers on flexible silk matrices. Adv Mater 28:1406–1412. https://doi.org/10.1002/adma.201504736
Pal RK, Farghaly AA, Wang C et al (2016) Conducting polymer-silk biocomposites for flexible and biodegradable electrochemical sensors. Biosens Bioelectron 81:294–302. https://doi.org/10.1016/j.bios.2016.03.010
Amanat F, Krammer F (2020) SARS-CoV-2 vaccines: status report. Immunity 52:583–589. https://doi.org/10.1016/j.immuni.2020.03.007
Song Z, Xu Y, Bao L et al (2019) From SARS to MERS, thrusting coronaviruses into the spotlight. Viruses 11:59. https://doi.org/10.3390/v11010059
Zhou P, Yang X-L, Wang X-G et al (2020) A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579:270–273. https://doi.org/10.1038/s41586-020-2012-7
Holshue ML, DeBolt C, Lindquist S et al (2020) First case of 2019 novel coronavirus in the United States. N Engl J Med 382:929–936. https://doi.org/10.1056/NEJMoa2001191
Rothe C, Schunk M, Sothmann P et al (2020) Transmission of 2019-nCoV infection from an asymptomatic contact in Germany. N Engl J Med 382:970–971. https://doi.org/10.1056/NEJMc2001468
Phan LT, Nguyen TV, Luong QC et al (2020) Importation and human-to-human transmission of a novel coronavirus in Vietnam. N Engl J Med 382:872–874. https://doi.org/10.1056/NEJMc2001272
Lai C-C, Shih T-P, Ko W-C et al (2020) Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and coronavirus disease-2019 (COVID-19): the epidemic and the challenges. Int J Antimicrob Agents 55:105924. https://doi.org/10.1016/j.ijantimicag.2020.105924
Schoeman D, Fielding BC (2019) Coronavirus envelope protein: current knowledge. Virol J 16:69. https://doi.org/10.1186/s12985-019-1182-0
Chan JF-W, Yuan S, Kok K-H et al (2020) A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet 395:514–523. https://doi.org/10.1016/S0140-6736(20)30154-9
Paraskevis D, Kostaki EG, Magiorkinis G et al (2020) Full-genome evolutionary analysis of the novel corona virus (2019-nCoV) rejects the hypothesis of emergence as a result of a recent recombination event. Infect Genet Evol 79:104212. https://doi.org/10.1016/j.meegid.2020.104212
Masters PS (2006) The molecular biology of coronaviruses. Adv Virus Res 66:193–292. https://doi.org/10.1016/S0065-3527(06)66005-3
Wang C, Zheng X, Gai W et al (2017) MERS-CoV virus-like particles produced in insect cells induce specific humoural and cellular imminity in rhesus macaques. Oncotarget 8:12686–12694. https://doi.org/10.18632/oncotarget.8475
Liu DX, Fung TS, Chong KK-L et al (2014) Accessory proteins of SARS-CoV and other coronaviruses. Antiviral Res 109:97–109. https://doi.org/10.1016/j.antiviral.2014.06.013
Mortola E, Roy P (2004) Efficient assembly and release of SARS coronavirus-like particles by a heterologous expression system. FEBS Lett 576:174–178. https://doi.org/10.1016/j.febslet.2004.09.009
de Haan CAM, Rottier PJM (2005) Molecular interactions in the assembly of coronaviruses. Adv Virus Res 64:165–230. https://doi.org/10.1016/S0065-3527(05)64006-7
Neuman BW, Kiss G, Kunding AH et al (2011) A structural analysis of M protein in coronavirus assembly and morphology. J Struct Biol 174:11–22. https://doi.org/10.1016/j.jsb.2010.11.021
Li X, Wang L, Yan S et al (2020) Clinical characteristics of 25 death cases with COVID-19: a retrospective review of medical records in a single medical center, Wuhan, China. Int J Infect Dis 94:128–132. https://doi.org/10.1016/j.ijid.2020.03.053
Huang C, Wang Y, Li X et al (2020) Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395:497–506. https://doi.org/10.1016/S0140-6736(20)30183-5
Xu Z, Shi L, Wang Y et al (2020) Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir Med 8:420–422. https://doi.org/10.1016/S2213-2600(20)30076-X
Qin C, Zhou L, Hu Z et al (2020) Dysregulation of immune response in patients with coronavirus 2019 (COVID-19) in Wuhan, China. Clin Infect Dis 71:762–768. https://doi.org/10.1093/cid/ciaa248
Du L, He Y, Zhou Y et al (2009) The spike protein of SARS-CoV—a target for vaccine and therapeutic development. Nat Rev Microbiol 7:226–236. https://doi.org/10.1038/nrmicro2090
Chen I-Y, Chang SC, Wu H-Y et al (2010) Upregulation of the chemokine (C-C Motif) ligand 2 via a severe acute respiratory syndrome coronavirus Spike-ACE2 signaling pathway. J Virol 84:7703–7712. https://doi.org/10.1128/JVI.02560-09
Chia PY, Coleman KK, Tan YK et al (2020) Detection of air and surface contamination by SARS-CoV-2 in hospital rooms of infected patients. Nat Commun 11:2800. https://doi.org/10.1038/s41467-020-16670-2
Zahedi A, Monis P, Deere D, Ryan U (2021) Wastewater-based epidemiology—surveillance and early detection of waterborne pathogens with a focus on SARS-CoV-2, Cryptosporidium and Giardia. Parasitol Res. https://doi.org/10.1007/s00436-020-07023-5
Lahrich S, Laghrib F, Farahi A et al (2021) Review on the contamination of wastewater by COVID-19 virus: Impact and treatment. Sci Total Environ 751:142325. https://doi.org/10.1016/j.scitotenv.2020.142325
Abdellah AR, Abdelhamid HN, El-Adasy A-BAAM et al (2020) One-pot synthesis of hierarchical porous covalent organic frameworks and two-dimensional nanomaterials for selective removal of anionic dyes. J Environ Chem Eng 8:104054. https://doi.org/10.1016/j.jece.2020.104054
Abdelhamid HN (2021) Dehydrogenation of sodium borohydride using cobalt embedded zeolitic imidazolate frameworks. J Solid State Chem 297:122034. https://doi.org/10.1016/j.jssc.2021.122034
Abdelhamid HN (2021) A review on hydrogen generation from the hydrolysis of sodium borohydride. Int J Hydrogen Energy 46:726–765. https://doi.org/10.1016/j.ijhydene.2020.09.186
Abdelhamid HN (2020) Dye encapsulated hierarchical porous zeolitic imidazolate frameworks for carbon dioxide adsorption. J Environ Chem Eng 8:104008. https://doi.org/10.1016/j.jece.2020.104008
Abdelhamid HN (2021) Biointerface between ZIF-8 and biomolecules and their applications. Biointerface Res Appl Chem 11:8283–8297. https://doi.org/10.33263/BRIAC111.82838297
Abdelhamid HN, Goda MN, Said AE-AA (2020) Selective dehydrogenation of isopropanol on carbonized metal–organic frameworks. Nano-Struct Nano-Objects 24:100605. https://doi.org/10.1016/j.nanoso.2020.100605
Kassem AA, Abdelhamid HN, Fouad DM, Ibrahim SA (2020) Hydrogenation reduction of dyes using metal–organic framework-derived CuO@C. Microporous Mesoporous Mater 305:110340. https://doi.org/10.1016/j.micromeso.2020.110340
Kassem AA, Abdelhamid HN, Fouad DM, Ibrahim SA (2020) Catalytic reduction of 4-nitrophenol using copper terephthalate frameworks and CuO@C composite. J Environ Chem Eng. https://doi.org/10.1016/j.jece.2020.104401
Abdelhamid HN (2020) High performance and ultrafast reduction of 4-nitrophenol using metal–organic frameworks. J Environ Chem Eng. https://doi.org/10.1016/j.jece.2020.104404
Abdelhamid HN (2020) UiO-66 as a catalyst for hydrogen production via the hydrolysis of sodium borohydride. Dalt Trans 49:10851–10857. https://doi.org/10.1039/D0DT01688H
Abdelhamid HN (2020) Hierarchical porous ZIF-8 for hydrogen production via the hydrolysis of sodium borohydride. Dalt Trans 49:4416–4424. https://doi.org/10.1039/D0DT00145G
Abdelhamid HN, Sharmoukh W (2021) Intrinsic catalase-mimicking MOFzyme for sensitive detection of hydrogen peroxide and ferric ions. Microchem J 163:105873. https://doi.org/10.1016/j.microc.2020.105873
Abdelhamid HN, Mahmoud GA-E, Sharmoukh W (2020) Correction: a cerium-based MOFzyme with multi-enzyme-like activity for the disruption and inhibition of fungal recolonization. J Mater Chem B 8:7557–7557. https://doi.org/10.1039/D0TB90139C
Abdelhamid HN, Mahmoud GA-E, Sharmouk W (2020) A cerium-based MOFzyme with multi-enzyme-like activity for the disruption and inhibition of fungal recolonization. J Mater Chem B 8:7548–7556. https://doi.org/10.1039/D0TB00894J
Yousef MS, Abdelhamid HN, Hidalgo M et al (2021) Antimicrobial activity of silver-carbon nanoparticles on the bacterial flora of bull semen. Theriogenology 161:219–227. https://doi.org/10.1016/j.theriogenology.2020.12.006
Kumaran S, Abdelhamid HN, Hasan N, Wu H-F (2020) Cytotoxicity of palladium nanoparticles against Aspergillus niger. Nanosci Nanotechnol-Asia 10:80–85. https://doi.org/10.2174/2210681208666180904113754
Talebian S, Wallace GG, Schroeder A et al (2020) Nanotechnology-based disinfectants and sensors for SARS-CoV-2. Nat Nanotechnol 15:618–621. https://doi.org/10.1038/s41565-020-0751-0
Yu L, Peel GK, Cheema FH et al (2020) Catching and killing of airborne SARS-CoV-2 to control spread of COVID-19 by a heated air disinfection system. Mater Today Phys 15:100249. https://doi.org/10.1016/j.mtphys.2020.100249
Pandey A, Nikam AN, Mutalik SP et al (2021) Architectured therapeutic and diagnostic nanoplatforms for combating SARS-CoV-2: role of inorganic, organic, and radioactive materials. ACS Biomater Sci Eng 7:31–54. https://doi.org/10.1021/acsbiomaterials.0c01243
Medhi R, Srinoi P, Ngo N et al (2020) Nanoparticle-based strategies to combat COVID-19. ACS Appl Nano Mater 3:8557–8580. https://doi.org/10.1021/acsanm.0c01978
Chin LK, Son T, Hong J-S et al (2020) Plasmonic sensors for extracellular vesicle analysis: from scientific development to translational research. ACS Nano 14:14528–14548. https://doi.org/10.1021/acsnano.0c07581
Wang X, Yuan X, Fu K et al (2021) Colorimetric analysis of extracellular vesicle surface proteins based on controlled growth of Au aptasensors. Analyst. https://doi.org/10.1039/D0AN02080J
Deng J, Zhao S, Liu Y et al (2020) Nanosensors for diagnosis of infectious diseases. ACS Appl Bio Mater. https://doi.org/10.1021/acsabm.0c01247
Badilescu S, Raju D, Bathini S, Packirisamy M (2020) Gold nano-island platforms for localized surface plasmon resonance sensing: a short review. Molecules 25:4661. https://doi.org/10.3390/molecules25204661
Peng X, Zhou Y, Nie K et al (2020) Promising near-infrared plasmonic biosensor employed for specific detection of SARS-CoV-2 and its spike glycoprotein. New J Phys 22:103046. https://doi.org/10.1088/1367-2630/abbe53
Prattis I, Hui E, Gubeljak P et al (2021) Graphene for biosensing applications in point-of-care testing. Trends Biotechnol. https://doi.org/10.1016/j.tibtech.2021.01.005
Tang Y-W, Schmitz JE, Persing DH, Stratton CW (2020) Laboratory diagnosis of COVID-19: current issues and challenges. J Clin Microbiol. https://doi.org/10.1128/JCM.00512-20
Ong DSY, de Man SJ, Lindeboom FA, Koeleman JGM (2020) Comparison of diagnostic accuracies of rapid serological tests and ELISA to molecular diagnostics in patients with suspected coronavirus disease 2019 presenting to the hospital. Clin Microbiol Infect 26:1094.e7-1094.e10. https://doi.org/10.1016/j.cmi.2020.05.028
Castillo-Henríquez L, Brenes-Acuña M, Castro-Rojas A et al (2020) Biosensors for the detection of bacterial and viral clinical pathogens. Sensors 20:6926. https://doi.org/10.3390/s20236926
Paliwal P, Sargolzaei S, Bhardwaj SK et al (2020) Grand Challenges in bio-nanotechnology to manage the COVID-19 pandemic. Front Nanotechnol. https://doi.org/10.3389/fnano.2020.571284
Alhalaili B, Popescu IN, Kamoun O et al (2020) Nanobiosensors for the detection of novel coronavirus 2019-nCoV and other pandemic/epidemic respiratory viruses: a review. Sensors 20:6591. https://doi.org/10.3390/s20226591
Chakraborty D, Kumar S, Chandrasekaran N, Mukherjee A (2020) Viral diagnostics and preventive techniques in the era of COVID-19: role of nanoparticles. Front Nanotechnol. https://doi.org/10.3389/fnano.2020.588795
Fani M, Zandi M, Soltani S, Abbasi S (2020) Future developments in biosensors for field-ready SARS-CoV-2 virus diagnostics. Biotechnol Appl Biochem. https://doi.org/10.1002/bab.2033
Cui F, Zhou HS (2020) Diagnostic methods and potential portable biosensors for coronavirus disease 2019. Biosens Bioelectron 165:112349. https://doi.org/10.1016/j.bios.2020.112349
Mahapatra S, Chandra P (2020) Clinically practiced and commercially viable nanobio engineered analytical methods for COVID-19 diagnosis. Biosens Bioelectron 165:112361. https://doi.org/10.1016/j.bios.2020.112361
Ji T, Liu Z, Wang G et al (2020) Detection of COVID-19: A review of the current literature and future perspectives. Biosens Bioelectron 166:112455. https://doi.org/10.1016/j.bios.2020.112455
Asif M, Ajmal M, Ashraf G et al (2020) The role of biosensors in coronavirus disease-2019 outbreak. Curr Opin Electrochem 23:174–184. https://doi.org/10.1016/j.coelec.2020.08.011
Zamora-Ledezma C, Medina E et al (2020) Biomedical science to tackle the COVID-19 pandemic: current status and future perspectives. Molecules 25:4620. https://doi.org/10.3390/molecules25204620
Wölfel R, Corman VM, Guggemos W et al (2020) Virological assessment of hospitalized patients with COVID-2019. Nature 581:465–469. https://doi.org/10.1038/s41586-020-2196-x
Kilic T, Weissleder R, Lee H (2020) Molecular and immunological diagnostic tests of COVID-19: current status and challenges. iScience 23:101406. https://doi.org/10.1016/j.isci.2020.101406
Zhang L, Guo H (2020) Biomarkers of COVID-19 and technologies to combat SARS-CoV-2. Adv Biomark Sci Technol 2:1–23. https://doi.org/10.1016/j.abst.2020.08.001
Kailasa SK, Mehta VN, Koduru JR et al (2021) An overview of molecular biology and nanotechnology based analytical methods for the detection of SARS-CoV-2: promising biotools for the rapid diagnosis of COVID-19. Analyst. https://doi.org/10.1039/D0AN01528H
Yuan X, Yang C, He Q et al (2020) Current and perspective diagnostic techniques for COVID-19. ACS Infect Dis 6:1998–2016. https://doi.org/10.1021/acsinfecdis.0c00365
Bhalla N, Pan Y, Yang Z, Payam AF (2020) Opportunities and challenges for biosensors and nanoscale analytical tools for pandemics: COVID-19. ACS Nano 14:7783–7807. https://doi.org/10.1021/acsnano.0c04421
Weiss C, Carriere M, Fusco L et al (2020) Toward nanotechnology-enabled approaches against the COVID-19 pandemic. ACS Nano 14:6383–6406. https://doi.org/10.1021/acsnano.0c03697
Suleman S, Shukla SK, Malhotra N et al (2021) Point of care detection of COVID-19: advancement in biosensing and diagnostic methods. Chem Eng J 414:128759. https://doi.org/10.1016/j.cej.2021.128759
Chen L, Zhang G, Liu L, Li Z (2021) Emerging biosensing technologies for improved diagnostics of COVID-19 and future pandemics. Talanta 225:121986. https://doi.org/10.1016/j.talanta.2020.121986
Lukose J, Chidangil S, George SD (2021) Optical technologies for the detection of viruses like COVID-19: progress and prospects. Biosens Bioelectron 178:113004. https://doi.org/10.1016/j.bios.2021.113004
Shabani E, Dowlatshahi S, Abdekhodaie MJ (2021) Laboratory detection methods for the human coronaviruses. Eur J Clin Microbiol Infect Dis 40:225–246. https://doi.org/10.1007/s10096-020-04001-8
Sharma S, Saini S, Khangembam M, Singh V (2021) Nanomaterials-based biosensors for COVID-19 detection—a review. IEEE Sens J 21:5598–5611. https://doi.org/10.1109/JSEN.2020.3036748
Jayamohan H, Lambert CJ, Sant HJ et al (2021) SARS-CoV-2 pandemic: a review of molecular diagnostic tools including sample collection and commercial response with associated advantages and limitations. Anal Bioanal Chem 413:49–71. https://doi.org/10.1007/s00216-020-02958-1
Yüce M, Filiztekin E, Özkaya KG (2021) COVID-19 diagnosis—a review of current methods. Biosens Bioelectron 172:112752. https://doi.org/10.1016/j.bios.2020.112752
Laghrib F, Saqrane S, El Bouabi Y et al (2021) Current progress on COVID-19 related to biosensing technologies: new opportunity for detection and monitoring of viruses. Microchem J 160:105606. https://doi.org/10.1016/j.microc.2020.105606
Gowri A, Kumar NA, Anand S (2021) Recent advances in nanomaterials based biosensors for point of care (PoC) diagnosis of Covid-19—a minireview. TrAC Trends Anal Chem. https://doi.org/10.1016/j.trac.2021.116205
Dave P, Rojas-Cessa R, Dong Z, Umpaichitra V (2020) Survey of saliva components and virus sensors for prevention of COVID-19 and infectious diseases. Biosensors 11:14. https://doi.org/10.3390/bios11010014
Gupta R, Sagar P, Priyadarshi N et al (2020) Nanotechnology-based approaches for the detection of SARS-CoV-2. Front Nanotechnol. https://doi.org/10.3389/fnano.2020.589832
Iravani S (2020) Nano- and biosensors for the detection of SARS-CoV-2: challenges and opportunities. Mater Adv 1:3092–3103. https://doi.org/10.1039/D0MA00702A
Antiochia R (2020) Nanobiosensors as new diagnostic tools for SARS, MERS and COVID-19: from past to perspectives. Microchim Acta 187:639. https://doi.org/10.1007/s00604-020-04615-x
Xu L, Li D, Ramadan S et al (2020) Facile biosensors for rapid detection of COVID-19. Biosens Bioelectron 170:112673. https://doi.org/10.1016/j.bios.2020.112673
Sheikhzadeh E, Eissa S, Ismail A, Zourob M (2020) Diagnostic techniques for COVID-19 and new developments. Talanta 220:121392. https://doi.org/10.1016/j.talanta.2020.121392
Khan H, Kushwah KK, Singh S et al (2021) Smart technologies driven approaches to tackle COVID-19 pandemic: a review. 3 Biotech 11:50. https://doi.org/10.1007/s13205-020-02581-y
Chellasamy G, Arumugasamy SK, Govindaraju S, Yun K (2020) Analytical insights of COVID-19 pandemic. TrAC Trends Anal Chem 133:116072. https://doi.org/10.1016/j.trac.2020.116072
Taha BA, Al Mashhadany Y, Hafiz Mokhtar MH et al (2020) An analysis review of detection coronavirus disease 2019 (COVID-19) based on biosensor application. Sensors 20:6764. https://doi.org/10.3390/s20236764
Mattioli IA, Hassan A, Oliveira ON, Crespilho FN (2020) On the challenges for the diagnosis of SARS-CoV-2 based on a review of current methodologies. ACS Sens 5:3655–3677. https://doi.org/10.1021/acssensors.0c01382
Taleghani N, Taghipour F (2021) Diagnosis of COVID-19 for controlling the pandemic: a review of the state-of-the-art. Biosens Bioelectron 174:112830. https://doi.org/10.1016/j.bios.2020.112830
Sharifi M, Hasan A, Haghighat S et al (2021) Rapid diagnostics of coronavirus disease 2019 in early stages using nanobiosensors: challenges and opportunities. Talanta 223:121704. https://doi.org/10.1016/j.talanta.2020.121704
Srivastava M, Srivastava N, Mishra PK, Malhotra BD (2021) Prospects of nanomaterials-enabled biosensors for COVID-19 detection. Sci Total Environ 754:142363. https://doi.org/10.1016/j.scitotenv.2020.142363
Manmana Y, Kubo T, Otsuka K (2021) Recent developments of point-of-care (POC) testing platform for biomolecules. TrAC Trends Anal Chem 135:116160. https://doi.org/10.1016/j.trac.2020.116160
Datta M, Singh DD, Naqvi AR (2021) Molecular diagnostic tools for the detection of SARS-CoV-2. Int Rev Immunol. https://doi.org/10.1080/08830185.2020.1871477
Ghaffari M, Mollazadeh-Bajestani M, Moztarzadeh F et al (2021) An overview of the use of biomaterials, nanotechnology, and stem cells for detection and treatment of COVID-19: towards a framework to address future global pandemics. Emergent Mater. https://doi.org/10.1007/s42247-020-00143-9
Giri B, Pandey S, Shrestha R et al (2021) Review of analytical performance of COVID-19 detection methods. Anal Bioanal Chem 413:35–48. https://doi.org/10.1007/s00216-020-02889-x
Zhao D, Yao F, Wang L et al (2020) A comparative study on the clinical features of coronavirus 2019 (COVID-19) pneumonia with other pneumonias. Clin Infect Dis 71:756–761. https://doi.org/10.1093/cid/ciaa247
Jain R, Gupta M, Taneja S, Hemanth DJ (2021) Deep learning based detection and analysis of COVID-19 on chest X-ray images. Appl Intell 51:1690–1700. https://doi.org/10.1007/s10489-020-01902-1
Hamming I, Timens W, Bulthuis M et al (2004) Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. a first step in understanding SARS pathogenesis. J Pathol 203:631–637. https://doi.org/10.1002/path.1570
Wang M, Fu A, Hu B et al (2020) Nanopore targeted sequencing for the accurate and comprehensive detection of SARS-CoV-2 and other respiratory viruses. Small 16:2002169. https://doi.org/10.1002/smll.202002169
Santiago I (2020) Trends and innovations in biosensors for COVID-19 mass testing. ChemBioChem. https://doi.org/10.1002/cbic.202000250
Ejazi SA, Ghosh S, Ali N (2021) Antibody detection assays for COVID-19 diagnosis: an early overview. Immunol Cell Biol 99:21–33. https://doi.org/10.1111/imcb.12397
Chong Y, Ikematsu H, Tani N et al (2021) Clinical significance of SARS-CoV-2-specific IgG detection with a rapid antibody kit for COVID-19 patients. Influenza Other Respir Viruses 15:13–18. https://doi.org/10.1111/irv.12802
Ng SC, Chan FKL, Chan PKS (2020) Screening FMT donors during the COVID-19 pandemic: a protocol for stool SARS-CoV-2 viral quantification. Lancet Gastroenterol Hepatol 5:642–643. https://doi.org/10.1016/S2468-1253(20)30124-2
Han MS, Seong M-W, Heo EY et al (2020) Sequential analysis of viral load in a neonate and her mother infected with severe acute respiratory syndrome coronavirus 2. Clin Infect Dis 71:2236–2239. https://doi.org/10.1093/cid/ciaa447
Lu X, Zhang L, Du H et al (2020) SARS-CoV-2 infection in children. N Engl J Med 382:1663–1665. https://doi.org/10.1056/NEJMc2005073
To KK-W, Tsang OT-Y, Leung W-S et al (2020) Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. Lancet Infect Dis 20:565–574. https://doi.org/10.1016/S1473-3099(20)30196-1
Pan Y, Zhang D, Yang P et al (2020) Viral load of SARS-CoV-2 in clinical samples. Lancet Infect Dis 20:411–412. https://doi.org/10.1016/S1473-3099(20)30113-4
Bedford J, Enria D, Giesecke J et al (2020) COVID-19: towards controlling of a pandemic. Lancet 395:1015–1018. https://doi.org/10.1016/S0140-6736(20)30673-5
Lescure F-X, Bouadma L, Nguyen D et al (2020) Clinical and virological data of the first cases of COVID-19 in Europe: a case series. Lancet Infect Dis 20:697–706. https://doi.org/10.1016/S1473-3099(20)30200-0
Wang X, Tan L, Wang X et al (2020) Comparison of nasopharyngeal and oropharyngeal swabs for SARS-CoV-2 detection in 353 patients received tests with both specimens simultaneously. Int J Infect Dis 94:107–109. https://doi.org/10.1016/j.ijid.2020.04.023
To KKW, Yip CCY, Lai CYW et al (2019) Saliva as a diagnostic specimen for testing respiratory virus by a point-of-care molecular assay: a diagnostic validity study. Clin Microbiol Infect 25:372–378. https://doi.org/10.1016/j.cmi.2018.06.009
Wang W-K, Chen S-Y, Liu I-J et al (2004) Detection of SARS-associated coronavirus in throat wash and saliva in early diagnosis. Emerg Infect Dis 10:1213–1219. https://doi.org/10.3201/eid1007.031113
To KK-W, Tsang OT-Y, Yip CC-Y et al (2020) Consistent detection of 2019 novel coronavirus in saliva. Clin Infect Dis 71:841–843. https://doi.org/10.1093/cid/ciaa149
Shin J, Jeong B, Kim J et al (2020) Sensitive wearable temperature sensor with seamless monolithic integration. Adv Mater 32:1905527. https://doi.org/10.1002/adma.201905527
Zou L, Ruan F, Huang M et al (2020) SARS-CoV-2 viral load in upper respiratory specimens of infected patients. N Engl J Med 382:1177–1179. https://doi.org/10.1056/NEJMc2001737
Kim C, Ahmed JA, Eidex RB et al (2011) Comparison of nasopharyngeal and oropharyngeal swabs for the diagnosis of eight respiratory viruses by real-time reverse transcription-PCR assays. PLoS ONE 6:e21610. https://doi.org/10.1371/journal.pone.0021610
Ahmed W, Angel N, Edson J et al (2020) First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: a proof of concept for the wastewater surveillance of COVID-19 in the community. Sci Total Environ 728:138764. https://doi.org/10.1016/j.scitotenv.2020.138764
Bhadra S, Riedel TE, Lakhotia S et al (2020) High-surety isothermal amplification and detection of SARS-CoV-2, including with crude enzymes. bioRxiv. https://doi.org/10.1101/2020.04.13.039941
Rodriguez-Manzano J, Malpartida-Cardenas K, Moser N et al (2021) Handheld point-of-care system for rapid detection of SARS-CoV-2 extracted RNA in under 20 min. ACS Cent Sci. https://doi.org/10.1021/acscentsci.0c01288
Lee HY, Jeong H, Jung IY et al (2015) DhITACT: DNA hydrogel formation by isothermal amplification of complementary target in fluidic channels. Adv Mater 27:3513–3517. https://doi.org/10.1002/adma.201500414
Rahimi H, Salehiabar M, Barsbay M et al (2021) CRISPR Systems for COVID-19 Diagnosis. ACS Sens. https://doi.org/10.1021/acssensors.0c02312
Harrington LB, Burstein D, Chen JS et al (2018) Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science (80-) 362:839–842. https://doi.org/10.1126/science.aav4294
Gootenberg JS, Abudayyeh OO, Lee JW et al (2017) Nucleic acid detection with CRISPR-Cas13a/C2c2. Science (80-) 356:438–442. https://doi.org/10.1126/science.aam9321
Gootenberg JS, Abudayyeh OO, Kellner MJ et al (2018) Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science (80-) 360:439–444. https://doi.org/10.1126/science.aaq0179
Broughton JP, Deng X, Yu G et al (2020) CRISPR-Cas12-based detection of SARS-CoV-2. Nat Biotechnol 38:870–874. https://doi.org/10.1038/s41587-020-0513-4
Pang B, Xu J, Liu Y et al (2020) Isothermal amplification and ambient visualization in a single tube for the detection of SARS-CoV-2 using loop-mediated amplification and CRISPR technology. Anal Chem 92:16204–16212. https://doi.org/10.1021/acs.analchem.0c04047
Yam WC, Chan KH, Poon LLM et al (2003) Evaluation of reverse transcription-PCR assays for rapid diagnosis of severe acute respiratory syndrome associated with a novel coronavirus. J Clin Microbiol 41:4521–4524. https://doi.org/10.1128/JCM.41.10.4521-4524.2003
Xia J, Tong J, Liu M et al (2020) Evaluation of coronavirus in tears and conjunctival secretions of patients with SARS-CoV-2 infection. J Med Virol 92:589–594. https://doi.org/10.1002/jmv.25725
Rutgers Clinical Genomics Laboratory TaqPath SARS-CoV-2 Assay EUA Summary. https://www.fda.gov/media/136875/download. Accessed 5 Feb 2021
Becherer L, Borst N, Bakheit M et al (2020) Loop-mediated isothermal amplification (LAMP)—review and classification of methods for sequence-specific detection. Anal Methods 12:717–746. https://doi.org/10.1039/C9AY02246E
Chen JS, Ma E, Harrington LB et al (2018) CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science (80-) 360:436–439. https://doi.org/10.1126/science.aar6245
Huang Z, Tian D, Liu Y et al (2020) Ultra-sensitive and high-throughput CRISPR-p owered COVID-19 diagnosis. Biosens Bioelectron 164:112316. https://doi.org/10.1016/j.bios.2020.112316
Patchsung M, Jantarug K, Pattama A et al (2020) Clinical validation of a Cas13-based assay for the detection of SARS-CoV-2 RNA. Nat Biomed Eng 4:1140–1149. https://doi.org/10.1038/s41551-020-00603-x
Esbin MN, Whitney ON, Chong S et al (2020) Overcoming the bottleneck to widespread testing: a rapid review of nucleic acid testing approaches for COVID-19 detection. RNA 26:771–783. https://doi.org/10.1261/rna.076232.120
Venkatesan BM, Bashir R (2011) Nanopore sensors for nucleic acid analysis. Nat Nanotechnol 6:615–624. https://doi.org/10.1038/nnano.2011.129
Lu H, Giordano F, Ning Z (2016) Oxford nanopore MinION sequencing and genome assembly. Genom Proteom Bioinform 14:265–279. https://doi.org/10.1016/j.gpb.2016.05.004
Qiu G, Gai Z, Tao Y et al (2020) Dual-functional plasmonic photothermal biosensors for highly accurate severe acute respiratory syndrome coronavirus 2 detection. ACS Nano 14:5268–5277. https://doi.org/10.1021/acsnano.0c02439
Alafeef M, Dighe K, Moitra P, Pan D (2020) Rapid, ultrasensitive, and quantitative detection of SARS-CoV-2 using antisense oligonucleotides directed electrochemical biosensor chip. ACS Nano 14:17028–17045. https://doi.org/10.1021/acsnano.0c06392
Chaibun T, Puenpa J, Ngamdee T et al (2021) Rapid electrochemical detection of coronavirus SARS-CoV-2. Nat Commun 12:802. https://doi.org/10.1038/s41467-021-21121-7
Karami A, Hasani M, Azizi Jalilian F, Ezati R (2021) Conventional PCR assisted single-component assembly of spherical nucleic acids for simple colorimetric detection of SARS-CoV-2. Sens Actuators B Chem 328:128971. https://doi.org/10.1016/j.snb.2020.128971
Ranjan P, Singhal A, Yadav S et al (2021) Rapid diagnosis of SARS-CoV-2 using potential point-of-care electrochemical immunosensor: toward the future prospects. Int Rev Immunol. https://doi.org/10.1080/08830185.2021.1872566
Layqah LA, Eissa S (2019) An electrochemical immunosensor for the corona virus associated with the Middle East respiratory syndrome using an array of gold nanoparticle-modified carbon electrodes. Microchim Acta 186:224. https://doi.org/10.1007/s00604-019-3345-5
Park TJ, Lee SJ, Kim D-K et al (2012) Development of label-free optical diagnosis for sensitive detection of influenza virus with genetically engineered fusion protein. Talanta 89:246–252. https://doi.org/10.1016/j.talanta.2011.12.021
Jahanshahi P, Wei Q, Jie Z et al (2017) Kinetic analysis of IgM monoclonal antibodies for determination of dengue sample concentration using SPR technique. Bioengineered 8:239–247. https://doi.org/10.1080/21655979.2016.1223413
Yousefi H, Mahmud A, Chang D et al (2021) Detection of SARS-CoV-2 viral particles using direct, reagent-free electrochemical sensing. J Am Chem Soc. https://doi.org/10.1021/jacs.0c10810
Eissa S, Zourob M (2021) Development of a low-cost cotton-tipped electrochemical immunosensor for the detection of SARS-CoV-2. Anal Chem 93:1826–1833. https://doi.org/10.1021/acs.analchem.0c04719
Fabiani L, Saroglia M, Galatà G et al (2021) Magnetic beads combined with carbon black-based screen-printed electrodes for COVID-19: a reliable and miniaturized electrochemical immunosensor for SARS-CoV-2 detection in saliva. Biosens Bioelectron 171:112686. https://doi.org/10.1016/j.bios.2020.112686
Seo G, Lee G, Kim MJ et al (2020) Rapid detection of COVID-19 causative virus (SARS-CoV-2) in human nasopharyngeal swab specimens using field-effect transistor-based biosensor. ACS Nano 14:5135–5142. https://doi.org/10.1021/acsnano.0c02823
Ahmadivand A, Gerislioglu B, Ramezani Z et al (2021) Functionalized terahertz plasmonic metasensors: femtomolar-level detection of SARS-CoV-2 spike proteins. Biosens Bioelectron 177:112971. https://doi.org/10.1016/j.bios.2021.112971
Coronavirus (COVID-19) Update: Serological Tests|FDA. https://www.fda.gov/news-events/press-announcements/coronavirus-covid-19-update-serological-tests. Accessed 26 Dec 2020
Nguyen T, Duong Bang D, Wolff A (2020) 2019 novel coronavirus disease (COVID-19): paving the road for rapid detection and point-of-care diagnostics. Micromachines 11:306. https://doi.org/10.3390/mi11030306
Cai X, Chen J, li Hu J-, et al (2020) A peptide-based magnetic chemiluminescence enzyme immunoassay for serological diagnosis of coronavirus disease 2019. J Infect Dis 222:189–193. https://doi.org/10.1093/infdis/jiaa243
Li Z, Yi Y, Luo X et al (2020) Development and clinical application of a rapid IgM-IgG combined antibody test for SARS-CoV-2 infection diagnosis. J Med Virol 92:1518–1524. https://doi.org/10.1002/jmv.25727
Della VB, Cennamo M, Minopoli A et al (2020) Colorimetric test for fast detection of SARS-CoV-2 in nasal and throat swabs. ACS Sens 5:3043–3048. https://doi.org/10.1021/acssensors.0c01742
Long Q-X, Liu B-Z, Deng H-J et al (2020) Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat Med 26:845–848. https://doi.org/10.1038/s41591-020-0897-1
Lin Q, Wen D, Wu J et al (2020) Microfluidic immunoassays for sensitive and simultaneous detection of IgG/IgM/Antigen of SARS-CoV-2 within 15 min. Anal Chem 92:9454–9458. https://doi.org/10.1021/acs.analchem.0c01635
Grolltex develops graphene-based sensor for Covid-19 detection|Graphene-Info. https://www.graphene-info.com/grolltex-develops-graphene-based-sensor-covid-19-detection. Accessed 30 Jan 2021
Funari R, Chu K-Y, Shen AQ (2020) Detection of antibodies against SARS-CoV-2 spike protein by gold nanospikes in an opto-microfluidic chip. Biosens Bioelectron 169:112578. https://doi.org/10.1016/j.bios.2020.112578
Zhong J, Rösch EL, Viereck T et al (2021) Toward rapid and sensitive detection of SARS-CoV-2 with functionalized magnetic nanoparticles. ACS Sens. https://doi.org/10.1021/acssensors.0c02160
Bertzbach LD, Kaufer BB, Karger A (2020) Applications of mass spectrometry imaging in virus research. Adv Virus Res. https://doi.org/10.1016/bs.aivir.2020.10.002
Abdelhamid HN, Wu H-F (2014) Ultrasensitive, rapid, and selective detection of mercury using graphene assisted laser desorption/ionization mass spectrometry. J Am Soc Mass Spectrom 25:861–868. https://doi.org/10.1007/s13361-014-0825-z
Abdelhamid HN (2020) Nanocytotoxicity using matrix-assisted laser desorption ionization mass spectrometry. Future Microbiol 15:385–387. https://doi.org/10.2217/fmb-2019-0260
Abdelhamid HN (2020) General methods for detection and evaluation of nanotoxicity. Nanotoxicity. https://doi.org/10.1016/B978-0-12-819943-5.00009-9
Abdelhamid HN, Wu H-F (2019) A new binary matrix for specific detection of mercury(II) using matrix-assisted laser desorption ionization mass spectrometry. J Am Soc Mass Spectrom 30:2617–2622. https://doi.org/10.1007/s13361-019-02324-1
Abdelhamid HN (2019) Nanoparticle-based surface assisted laser desorption ionization mass spectrometry: a review. Microchim Acta 186:682. https://doi.org/10.1007/s00604-019-3770-5
Abdelhamid HN (2019) Nanoparticles assisted laser desorption/ionization mass spectrometry. In: de la Guardia M, Esteve-Turrillas FA (eds) Handbook of smart materials in analytical chemistry. Wiley, Chichester, pp 729–755
Abdelhamid HN (2018) Ionic liquid-assisted laser desorption/ionization–mass spectrometry: matrices, microextraction, and separation. Methods Protoc 1:23. https://doi.org/10.3390/mps1020023
Abdelhamid HN, Chen Z-Y, Wu H-F (2017) Surface tuning laser desorption/ionization mass spectrometry (STLDI-MS) for the analysis of small molecules using quantum dots. Anal Bioanal Chem 409:4943–4950. https://doi.org/10.1007/s00216-017-0433-4
Zhang Y, Tang LV (2020) Overview of targets and potential drugs of SARS-CoV-2 according to the viral replication. J Proteome Res. https://doi.org/10.1021/acs.jproteome.0c00526
Saei AA, Sharifi S, Mahmoudi M (2020) COVID-19: nanomedicine uncovers blood-clot mystery. J Proteome Res 19:4364–4373. https://doi.org/10.1021/acs.jproteome.0c00425
Otter JA, Donskey C, Yezli S et al (2016) Transmission of SARS and MERS coronaviruses and influenza virus in healthcare settings: the possible role of dry surface contamination. J Hosp Infect 92:235–250. https://doi.org/10.1016/j.jhin.2015.08.027
Dowell SF, Simmerman JM, Erdman DD et al (2004) Severe acute respiratory syndrome coronavirus on hospital surfaces. Clin Infect Dis 39:652–657. https://doi.org/10.1086/422652
Lancet T (2020) COVID-19: protecting health-care workers. Lancet 395:922. https://doi.org/10.1016/S0140-6736(20)30644-9
Van Elslande J, Houben E, Depypere M et al (2020) Diagnostic performance of seven rapid IgG/IgM antibody tests and the Euroimmun IgA/IgG ELISA in COVID-19 patients. Clin Microbiol Infect 26:1082–1087. https://doi.org/10.1016/j.cmi.2020.05.023
Traugott M, Aberle SW, Aberle JH et al (2020) Performance of severe acute respiratory syndrome coronavirus 2 antibody assays in different stages of infection: comparison of commercial enzyme-linked immunosorbent assays and rapid tests. J Infect Dis 222:362–366. https://doi.org/10.1093/infdis/jiaa305
Udugama B, Kadhiresan P, Kozlowski HN et al (2020) Diagnosing COVID-19: the disease and tools for detection. ACS Nano 14:3822–3835. https://doi.org/10.1021/acsnano.0c02624
Xie X, Zhong Z, Zhao W et al (2020) Chest CT for typical coronavirus disease 2019 (COVID-19) pneumonia: relationship to negative RT-PCR testing. Radiology 296:E41–E45. https://doi.org/10.1148/radiol.2020200343
Khan MZH, Hasan MR, Hossain SI et al (2020) Ultrasensitive detection of pathogenic viruses with electrochemical biosensor: State of the art. Biosens Bioelectron 166:112431. https://doi.org/10.1016/j.bios.2020.112431
Fan Z, Yao B, Ding Y et al (2021) Entropy-driven amplified electrochemiluminescence biosensor for RdRp gene of SARS-CoV-2 detection with self-assembled DNA tetrahedron scaffolds. Biosens Bioelectron 178:113015. https://doi.org/10.1016/j.bios.2021.113015
Kalantar-Zadeh K, Ward SA, Kalantar-Zadeh K, El-Omar EM (2020) Considering the effects of microbiome and diet on SARS-CoV-2 infection: nanotechnology roles. ACS Nano 14:5179–5182. https://doi.org/10.1021/acsnano.0c03402
Chan WCW (2020) Nano research for COVID-19. ACS Nano 14:3719–3720. https://doi.org/10.1021/acsnano.0c02540
Xing Y, Zhao L, Cheng Z et al (2020) Microfluidics-based sensing of biospecies. ACS Appl Bio Mater. https://doi.org/10.1021/acsabm.0c01271
Carter LJ, Garner LV, Smoot JW et al (2020) Assay techniques and test development for COVID-19 diagnosis. ACS Cent Sci 6:591–605. https://doi.org/10.1021/acscentsci.0c00501
Herold T, Jurinovic V, Arnreich C et al (2020) Elevated levels of IL-6 and CRP predict the need for mechanical ventilation in COVID-19. J Allergy Clin Immunol 146:128-136.e4. https://doi.org/10.1016/j.jaci.2020.05.008
PerkinElmer New Coronavirus Nucleic Acid Detection Kit—Instructions for Use. https://www.fda.gov/media/136410/download. Accessed 30 Jan 2021
SARS-CoV-2 RNA, Qualitative Real-Time RT-PCR (Test Code 39433) Package Insert For Emergency Use Only For In-vitro Diagnostic Use-Rx Only. https://www.fda.gov/media/136231/download. Accessed 30 Jan 2021
1copyTM COVID-19 qPCR Multi Kit (Cat no. M22MD100M) Instructions for Use For in vitro diagnostic use For Emergency Use Authorization Only Prescription Use Only
Genesing. Primerdesign Ltd COVID-19 genesig Real-Time PCR assay. https://www.fda.gov/media/136823/download
Cepheid|Xpert® Xpress SARS-CoV-2 has received FDA Emergency Use Authorization. https://www.cepheid.com/en/about/SARS-CoV-2-Test-Development-Information. Accessed 30 Jan 2021
BioFire® COVID-19 Test—BioFire Defense. https://www.biofiredefense.com/covid-19test/. Accessed 30 Jan 2021
AvellinoCoV2. AvellioCoV2 test EUA Summary. https://www.fda.gov/media/136453/download. Accessed 30 Jan 2021
SimplexaTM COVID-19 Direct-Page 2
Abbott RealTime SARS-CoV-2 Assay (EUA). https://www.molecular.abbott/int/en/products/infectious-disease/RealTime-SARS-CoV-2-Assay. Accessed 30 Jan 2021
NeoPlex TM COVID-19 Detection Kit Multiplex RT-Real-time PCR Reagents for SARS-CoV-2 Detection
COVID-19 RT-qPCR Detection Kit Instructions for Use For Emergency Use Authorization Only. https://www.fda.gov/media/137895/download. Accessed 30 Jan 2021
Quick SARS-CoV-2 rRT-PCR Kit—Instructions for Use. https://www.fda.gov/media/137780/download. Accessed 30 Jan 2021
OPTI SARS-CoV-2 Test. https://www.fda.gov/media/137739/download. Accessed 31 Jan 2021
FTD SARS-CoV-2—Instructions for Use. https://www.fda.gov/media/137690/download. Accessed 31 Jan 2021
PI1109-C ePlex ® SARS-CoV-2 Test Assay Manual. https://www.fda.gov/media/136282/download. Accessed 31 Jan 2021
Coronavirus Update|Hologic Molecular Test Receives FDA Emergency Use Authorization. https://www.hologic.com/coronavirus-test. Accessed 31 Jan 2021
Hologic, Inc Aptima ® SARS-CoV-2 Assay (Panther ® System) Aptima ® SARS-CoV-2 Aptima SARS-CoV-2-Panther System
Rapid Diagnostics|Abbott Point of Care Testing. https://www.globalpointofcare.abbott/en/index.html. Accessed 31 Jan 2021
Sherlock Biosciences Receives FDA Emergency Use Authorization for CRISPR SARS-CoV-2 Rapid Diagnostic • Sherlock Biosciences. https://sherlock.bio/sherlock-biosciences-receives-fda-emergency-use-authorization-for-crispr-sars-cov-2-rapid-diagnostic/. Accessed 31 Jan 2021
Sofia SARS Antigen FIA|Quidel. https://www.quidel.com/immunoassays/rapid-sars-tests/sofia-sars-antigen-fia. Accessed 31 Jan 2021
Elecsys® IL-6. https://diagnostics.roche.com/us/en/products/params/elecsys-il-6.html. Accessed 31 Jan 2021
Campos EVR, Pereira AES, de Oliveira JL et al (2020) How can nanotechnology help to combat COVID-19? Opportunities and urgent need. J Nanobiotechnol 18:125. https://doi.org/10.1186/s12951-020-00685-4
Kaushik AK, Dhau JS, Gohel H et al (2020) Electrochemical SARS-CoV-2 sensing at point-of-care and artificial intelligence for intelligent COVID-19 management. ACS Appl Bio Mater 3:7306–7325. https://doi.org/10.1021/acsabm.0c01004
Jain S, Nehra M, Kumar R et al (2021) Internet of medical things (IoMT)-integrated biosensors for point-of-care testing of infectious diseases. Biosens Bioelectron. https://doi.org/10.1016/j.bios.2021.113074
Parihar A, Ranjan P, Sanghi SK et al (2020) Point-of-care biosensor-based diagnosis of COVID-19 holds promise to combat current and future pandemics. ACS Appl Bio Mater 3:7326–7343. https://doi.org/10.1021/acsabm.0c01083
Samavati A, Samavati Z, Velashjerdi M et al (2020) Sustainable and fast saliva-based COVID-19 virus diagnosis kit using a novel GO-decorated Au/FBG sensor. Chem Eng J. https://doi.org/10.1016/j.cej.2020.127655
Shirshahi V, Liu G (2021) Enhancing the analytical performance of paper lateral flow assays: from chemistry to engineering. TrAC Trends Anal Chem 136:116200. https://doi.org/10.1016/j.trac.2021.116200
Soler M, Estevez MC, Cardenosa-Rubio M et al (2020) How nanophotonic label-free biosensors can contribute to rapid and massive diagnostics of respiratory virus infections: COVID-19 Case. ACS Sens 5:2663–2678. https://doi.org/10.1021/acssensors.0c01180
Dhanabalan SS, Sriram S, Walia S et al (2021) Wearable Label-free optical biodetectors: progress and perspectives. Adv Photonics Res 2:2000076. https://doi.org/10.1002/adpr.202000076
Ruiz-Vega G, Soler M, Lechuga LM (2021) Nanophotonic biosensors for point-of-care COVID-19 diagnostics and coronavirus surveillance. J Phys Photonics 3:011002. https://doi.org/10.1088/2515-7647/abd4ee
Song M, Yang M, Hao J (2021) Pathogenic virus detection by optical nanobiosensors. Cell Rep Phys Sci 2:100288. https://doi.org/10.1016/j.xcrp.2020.100288
Das Mukhopadhyay C, Sharma P, Sinha K, Rajarshi K (2021) Recent trends in analytical and digital techniques for the detection of the SARS-Cov-2. Biophys Chem 270:106538. https://doi.org/10.1016/j.bpc.2020.106538
Acknowledgements
Funding was provided by Science and Technology Development Fund (EG) (Grant No. 35969).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no competing financial interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Abdelhamid, H.N., Badr, G. Nanobiotechnology as a platform for the diagnosis of COVID-19: a review. Nanotechnol. Environ. Eng. 6, 19 (2021). https://doi.org/10.1007/s41204-021-00109-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s41204-021-00109-0