Abstract
The mitochondrial genomes of D. melacanthus and D. furcatus were sequenced and used to investigate the phylogenetic relationships with 54 species of Pentatomidae. Their mitogenomes are 17,197 and 15,444 bp-long, respectively, including 13 protein-coding genes (PCGs), 2 ribosomal RNA genes, and 22/21 transfer RNA genes, with conserved gene arrangement. Leu, Lys, and Ser were the most common amino acids in their PCGs. PCGs evolutionary analysis indicated their mitogenomes are under purifying selection, and the most conserved genes are from the cytochrome complex, reinforcing their suitability as markers for molecular taxonomy. We identified 490 mtSSRs in 56 Pentatomidae species, with large variation and a positive correlation between mtSSR number and genome size. Three mtSSRs were identified in each Diceraeus species. Only the mtSSR in the nad6 (D. melacanthus) and nad4 (D. furcatus) appear to have application as molecular markers for species characterization. Phylogenetic analysis confirmed the monophyly of Pentatomidae. However, our analysis challenged the monophyly of Pentatominae and Podopinae. We also detected unexpected relationships among some tribes and genera, highlighting the complexity of the internal taxonomic structure of Pentatomidae. Both Diceraeus species were grouped in the same clade with the remaining Carpocorini analyzed.
Similar content being viewed by others
Introduction
The green-belly bugs Diceraeus furcatus and D. melacanthus (Hemiptera: Pentatomidae) are important pests of soybean, winter cereal, and especially maize in Brazil1,2,3,4. Their increased abundance is attributed to the agricultural system in practice in Brazil, with the continuous cultivation of maize following soybean harvest. This system provides year-round food and shelter for green-belly bugs and other pests5,6,7. Despite their relevance as agricultural pests, few molecular studies have been conducted on these species8,9, especially concerning their genetic delimitation, given their morphological similarity and considerable phenotypic plasticity10.
The identification of some insects is challenging and it requires additional tools, such as the use of integrative diagnostic methods based on molecular, morphological, and paleontological data11. In recent decades, the use of DNA barcode markers based on a short region of the mitochondrial cytochrome oxidase I gene (cox1) has been the most used molecular tool. The cox1 became widely used for species recognition and delineation in various taxonomic groups12, although it has shown limited application for species delineation in taxonomic groups carrying low intraspecific variability13.
The mitochondrial genome (mtDNA) has emerged as a powerful tool for insect diversity and phylogenetic studies, expanding the potential pool of informative marker genes13. Its unique features, such as small size14, stable genetic composition15, maternal inheritance16, orthologous genes and low intermolecular recombination rates17, make it suitable for evolutionary studies. mtDNA serves as a source of molecular markers for population genetics, comparative evolution, divergence time analysis18,19,20, and gene rearrangements21,22. mtDNA markers are also instrumental in species recognition and delineation23, resolving conflicts of species complexes and in studying the biodiversity of taxonomic groups lacking experts for identification.
The mitochondrial genome of insects is circular, compact and up to 18 kb. It typically comprises 37 genes, including 13 protein-coding genes (PCGs), 2 ribosomal RNA genes (rRNA), and 22 transfer RNA genes (tRNA)14,24,25. The gene arrangement is often conserved26 and even small changes can provide valuable insights into phylogenetic relationships among organisms27. In addition, insect mitogenomes usually have a large non-coding region (A + T-rich region), containing essential regulatory elements for replication and transcription28,29.
Microsatellites—repetitive sequences of simple tandem repeats (SSRs)—have been extensively used as genetic markers30,31. SSRs are abundant and carry high mutation rates32. They are present in coding and non-coding regions of eukaryotic and prokaryotic genomes33,34,35,36,37. SSRs can occur in the nuclear36,38, cytoplasmic, and mitochondrial genomes39,40. Their location in the mitogenome determines their functional role in gene regulation, development and evolution36,41. Although evidence suggests non-random SSR distribution in genomes, especially in plants42, information on the identification and characterization of mitochondrial microsatellites (mtSSRs) and their roles in insects is scarce and poorly understood.
In this study, we sequenced and described the mitogenomes of D. furcatus and D. melacanthus. We analyzed the structure and nucleotide composition of their mitochondrial genomes and compared them with that of other Pentatomidae species, providing insights for the development of additional molecular markers. Additionally, we conducted phylogenetic analyses to determine the systematic placement of D. furcatus and D. melacanthus relative to other Pentatomidae species.
Material and methods
Sample collection and mtDNA extraction
Males and females from a laboratory-reared population of D. melacanthus maintained under controlled conditions were individualized shortly after emergence. At the end of the pre-reproductive period, males and females were paired to form individual couples. Females with the highest number of fertile eggs were kept with their progeny until reaching the 3rd instar, when nymphs were subjected to mitochondrial DNA (mtDNA) extraction. Specimens of D. furcatus were provided by Dr. Antonio Ricardo Panizzi (Embrapa Trigo, Passo Fundo/RS). The obtained specimens underwent mtDNA extraction using the commercial Mitochondrial DNA Isolation Kit (Abcam®), following the manufacturer’s instructions. The samples were stored at − 20 °C and submitted for sequencing at the Multiuser Center for Agricultural Biotechnology, USP/Esalq.
Sequencing, assembly, and annotation of the mitochondrial genome
Libraries for sequencing the mtDNA of D. melacanthus were prepared using the Nextera XT DNA Library Prep system (Illumina), while those for D. furcatus were prepared using the TruSeq Nano DNA Library Prep system (Illumina), given the total amount of mtDNA obtained for each of the studied species. The mitochondrial genome sequencing of D. melacanthus was conducted on the Illumina MiSeq V3 platform (2 × 250 bp), while that of D. furcatus was performed on the NextSeq platform (2 × 100 bp).
The quality of raw reads was assessed using FastQC version 0.11.943. Adapters and low-quality bases were removed using Trimmomatic44, applying the following parameters: TruSeq2-PE.fa:0:30:10:8:true SLIDINGWINDOW:4:24 LEADING:3 TRAILING:3 HEADCROP:3 MINLEN:25.
The de novo assembly of the mitogenomes was performed using the NovoPlasty algorithm45, using a partial sequence of the cytochrome oxidase I gene from D. melacanthus (SRX2888372) deposited in the National Center for Biotechnology Information (NCBI) database (https://www.ncbi.nlm.nih.gov/) as a seed. Annotation of the assembled mtDNA was conducted using the MITOS Web Server for mitochondrial annotation (http://mitos.bioinf.uni-leipzig.de/index.py)46. The same software was employed to determine gene, rRNA, and tRNA boundaries, as well as to predict RNA secondary structures. GenomeVX (http://wolfe.ucd.ie/GenomeVx/)47 was used to visualize the complete mitogenomes.
Nucleotide and amino acid composition, the relative synonymous codon usage (RSCU), the number of non-synonymous substitutions per non-synonymous site (ka), the number of synonymous substitutions per synonymous site (ks), and the ratio (ka/ks) for each protein-coding gene (PCG) were calculated in Mega X48. Ka and Ks values were obtained using trimmed PCG sequences. AT-GC content and skew statistics were calculated using CGView49.
The complete mitogenomes of D. melacanthus and D. furcatus were also utilized to identify short mitochondrial DNA sequence repeats (mtSSRs) with the MIcroSAtellite tool (https://webblast.ipk-gatersleben.de/misa/)50. The minimum repeat length used was ≥ 12 for mononucleotides, ≥ 6 for dinucleotides, ≥ 4 for trinucleotides, and ≥ 3 for tetra-, penta-, and hexanucleotides. The interruption between two microsatellites was considered as zero. To facilitate comparisons of genomes of different sizes, SSRs were standardized per 1 kilobase (kb) of genome, calculating the relative abundance (RA) and relative density (RD) for the identified mtSSRs. RA provides the total number of SSR classes per kb of genome, and RD offers the total length (in nucleotides) contributing to 1 kb of genome nucleotides51.
The mitochondrial genomes of D. melacanthus and D. furcatus have been deposited in GenBank under accession numbers PP235949 e PP235950.
Phylogenetic analyses
Phylogenetic analyses were conducted using the mitogenomes of D. melacanthus and D. furcatus, and of 54 species of Pentatomidae available at the NCBI GenBank database (Table 1). Two additional mitogenomes from Aleyrodidae and Reduviidae were used as outgroups (Table 1). We pre-processed the dataset using the MitoPhAST V3.0 pipeline52. Nucleotide sequences of PCGs were extracted, translated into amino acids, and aligned using Clustal Omega53. Poorly aligned regions were removed with trimAI54, and a supermatrix was constructed with the concatenated alignments of PCGs in the following order: nad1, nad4, nad3, cox3, cox1, nad4L, cytb, atp8, nad6, nad2, cox2, atp6, and nad5. Phylogenetic analyses were performed using Maximum Likelihood (ML) and Bayesian Inference (BI) methods. ML analyses were conducted in IQ-TREE v1.6.1055, using the mtInv + F + R5 model, as determined by ModelFinder56 through the Akaike Information Criterion (AIC), with 100,000 bootstrap iterations. Bayesian analysis of site-homogeneous models was performed in MrBayes v. 3.2.757. Two independent runs with four chains (three heated and one cold) were conducted simultaneously for 10 million generations, with sampling every 1000 generations and 25% of burn-in. The remaining samples were used to construct the consensus tree and Bayesian posterior probabilities. Another Bayesian analysis under a site-heterogeneous model was implemented using PhyloBayes MPI 1.5a on the CIPRES Science Gateway 3.358. After removing constant sites from the alignment, two independent chains starting from a random tree were run under the CAT + GTR + C4 (GTR) and C6 models. The final trees were visualized using FigTree v.1.4.4 software (http://tree.bio.ed.ac.uk/software/figtree/). Tree topologies obtained with each phylogenetic analytical method used were compared using the IQ-TREE web server (http://iqtree.cibiv.univie.ac.at/)59. The data sets of P123 and AA were subjected to Kishino–Hasegawa (KH)60, Shimodaira–Hasegawa (SH)61, weighted Kishino–Hasegawa test (WKH)60, weighted SH test (WSH)61, expected Likelihood Weight (ELW)62, and the approximately unbiased (AU) tests63 with 1000 replicates.
Results and discussion
Organization and characteristics of mitogenomes
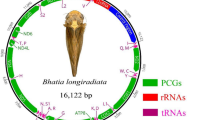
The mitochondrial genomes of D. melacanthus and D. furcatus are 17,197 and 15,444 bp—long circular, double-stranded DNA molecules, respectively. They contain 13 protein-coding genes (PCGs), 2 ribosomal RNA genes (rRNAs), and 22 (D. melacanthus) or 21 (D. furcatus) transfer RNA genes (tRNAs). The mitogenome of D. furcatus lacks the isoleucine tRNA gene (trnI). Thus, the mitogenome of D. melacanthus carries a total of 37 genes, while that of D. furcatus 36 (Table 2). The pseudogene nad5-1 was identified in both mitogenomes. These mitogenomes have a conserved gene arrangement, with 23 genes located on the sense strand and 14 (13 in D. furcatus) on the antisense strand (Fig. 1). The intergenic spacers ranged from 2 to 1281 bp in D. melacanthus, and from 1 to 591 bp in D. furcatus. Gene overlaps in their mitogenomes varied between 1 and 11 bp (Fig. 1).
The A + T content (AT%) in the mitogenome of D. melacanthus is 74%, with a lower AT% in PCGs (71.3%) than in rRNAs (74.2%) and tRNAs (75.7%) (Table 2), resulting in a mitogenome with a positive AT skew (0.149) and negative GC skew (− 0.154). The PCGs exhibit a slightly negative AT skew due to the biased AT% in some of the nad genes (nad1, nad4, nad4L, nad5) (Table 2). The tRNAs and rRNAs also have a negative AT skew. The AT% of the mitogenome of D. furcatus was 72.4%, with PCGs carrying lower AT% (70.5%) than rRNAs (73.8%) and tRNAs (75.8%). The AT skews of tRNAs (0.003) and of the complete mitochondrial genome (0.174) of D. furcatus are positive, while those of PCGs (− 0.188) and rRNAs (− 0.163) are negative. The negative AT skew of PCGs is also likely due to the nad genes (Table 2).
The biased AT% of the whole mitogenomes, PCGs, rRNAs, and tRNAs genes of Diceraeus is similar to that of other Pentatomidae species76,91. Small variations in AT% among different genes have been attributed to DNA repair, selection, and mutations that occur during the evolutionary process81,92.
Mitochondrial genomes exhibit the characteristic strand asymmetry, usually reflected by AT and GC skews. In this study, AT (positive) and GC (negative) skews resemble the patterns normally reported for insects29,91,92,93. The length of concatenated PCGs of D. melacanthus (10,533 bp) was close to that of D. furcatus (10,351 bp), corresponding to approximately 61 and 67% of their mitogenomes, similarly to various Pentatomidae14,29,70,82,94. Nine PCGs (nad2, cox1, cox2, atp8, atp6, cox3, nad3, nad6, cytb) are encoded on the main strand and four on the complementary strand (nad5, nad4, nad4L, nad1) (Fig. 1). nad5 is the longest PCG (1,677 bp for both species), while nad4L the shortest (264 bp for both species). ATD (ATA, ATT, ATG) and TTG are the most common initiation codons used in PCGs as reported for other Pentatomidae64,79,94,95,96.
Fourteen tRNA genes of D. melacanthus (trnM, trnW, trnL2, trnK, trnD, trnG, trnA, trnR, trnN, trnS1, trnE, trnT, trnS2, and trnI) and 13 of D. furcatus (except trnI) are encoded on the main strand, with eight on the secondary strand (trnQ, trnC, trnY, trnF, trnH, trnP, trnL1, and trnV) (Fig. 1), all with the typical cloverleaf secondary structure (Fig. 2)64,76,94,95,97. The D. melacanthus mitogenome carries 20 tRNAs common to most metazoans, contrary to information from another mitogenome of the same species, which reports the absence of trnI69. In our study, the mitogenome of D. furcatus presented 19 out of the 20 commonly reported tRNAs.
Secondary structure of tRNAs from (a) D. melacanthus and (b) D. furcatus. trnA—alanine; trnC—cysteine; trnD—aspartate; trnE—glutamate; trnF—phenylalanine; trnG—glycine; trnH—histidine; trnI—isoleucine; trnK—lysine; trnL—leucine; trnM—methionine; trnN—asparagine; trnP—proline; trnQ—glutamine; trnR—arginine; trnS—serine; trnT—threonine; trnV—valine; trnW—tryptophan; trnY – tyrosine.
The loss of tRNA is considered rare in insect mitogenomes98, although many reports can be found. The gene cluster trnT, trnQ, and trnM was not found in the mitogenome of the hemipteran Nabis apicalis99; trnI was not detected in Aleurochiton aceris89; trnS1 and trnQ were lost by Aleurodicus dugesii89; Aleurocanthus camelliae lacks trnI100; Neomaskellia andropogonis does not have trnA, trnR, trnN, trnI, and trnS289. The loss of mitochondrial genes can be tolerated by the cell when their function becomes unnecessary or when it is taken over by a substitute gene with a similar role, or yet because it has been functionally transferred to the nucleus. The transference of gene functions from the mitogenome to the nuclear genome has many advantages101,102,103,104,105, including effects on species differentiation and the functional evolution of mitogenomes106. Genes encoding tRNAs in mitogenomes exhibit a mutation rate five times higher than nuclear genes, which undergo intense purifying selection107. However, there is no evidence that this occurred with the absent tRNA in this study. Instead, its function may have been replaced by nuclear tRNAs, given the frequent use of the isoleucine amino acid. trnL and trnS were duplicated in both mitogenomes. trnS copies were located in the same strand (sense), while trnL in different strands. Gene duplications also contribute to tRNA gene number variation among taxa108, and the loss of mitochondrial tRNAs in eukaryotes is not uncommon101.
The 16S and 12S rRNA genes are in the antisense strand of the mitogenomes of D. melacanthus and D. furcatus (Figs. 1 and 1S). The larger rRNA subunit (16S) is located between trnL1 and a spacer region, and has identical sizes in both species (657 bp). The smaller rRNA subunit (12S) is located between the trnV gene and another spacer region, with of 791 bp in length for D. melacanthus and 792 bp for D. furcatus.
We demonstrated that the mitogenomes of D. melacanthus and D. furcatus share the same base composition pattern and conservation of content and gene arrangement observed after comparative analyses with other Pentatomidae mitogenomes14,29,70,82,94.
Leucine (Leu), lysine (Lys), serine (Ser), methionine (Met), and asparagine (Asp) are the most frequently used amino acids in the mitogenome of Diceraeus spp. (Fig. 3). Both Diceraeus species showed similar patterns of relative synonymous codon usage (RSCU). The most frequent codon (UUA–Leu2) used in D. melacanthus is the same of other Pentatomidae91, differing from the preferred AGA (Ser1) and UUA (Leu2) in the mitochondrial genome of D. furcatus. The preferred codons in PCGs showed a strong AT bias due to the predominance of A or T nucleotides in the third position of codons, following a pattern observed in Pentatomidae69,91,94,109.
There are 27 codons preferentially used (RSCU value > 1) for the 20 amino acids in PCGs in D. melacanthus and 26 in D. furcatus mitogenomes, representing less than half of the total set of codons (62 in D. melacanthus; 61 in D. furcatus) (Fig. 4). In D. melacanthus, all of them end in AT nucleotides, but only 84.6% in D. furcatus. Yet, in D. furcatus a large portion of synonymous codons are degenerate at the third nucleotide position, indicating greater selection pressure at this position from GC to AT110. The top five most used codons in D. melacanthus mitogenome are UUA (Leu2), GGA (Gly), UCA (Ser2), GUA (Val), and CGA (Arg); and AGA (Ser1), UUA (Leu2), UCA (Ser2), GUA (Val), GGA (Gly) in D. furcatus. Both Diceraeus species shared the preferential codon usage (Fig. 4).
The evolutionary patterns of the subfamily Pentatominae were observed by calculating the synonymous substitution rate (ks), nonsynonymous substitution rate (ka), and the ka/ks ratio (ω), pairwise for each PCG (Fig. 5)109,111. All PCGs had ω < 1 (< 0.500), indicating evolution under purifying selection29,109. The most conserved genes with the lowest ω values are cytb (ω = 0.078), cox3 (ω = 0.083), cox2 (ω = 0.091), and cox1 (ω = 0.159), are typically used as markers for molecular taxonomy, reinforcing their suitability for this purpose12,29,95,109. However, we were unable to efficiently amplify the barcode region of the cox1 gene for Diceraeus species using the universal set of primers available (LCO1490/jgLCO1490 and HCO2198/jgHCO2198)112,113. Regarding the specific substitution rates of each gene, Ks ranged from 0.59 (nad4) to 0.94 (nad3), while Ka varied from 0.07 (cox2, cox3, cytb) to 0.40 (atp8).
Mitochondrial microsatellites (mtSSRs)
The mitogenomes of D. melacanthus and D. furcatus presented 3 mtSSRs each, with two shared tetranucleotide mtSSRs equal in size (Table 3). The shared mtSSRs were located in the same coding gene (nad4) and intergenic region. The 18 bp-long trinucleotide of D. melacanthus was located in nad6 gene, while the 12 bp-long tetranucleotide of D. furcatus was located in nad5 gene (Table 3).
The number of mtSSRs identified in D. melacanthus and D. furcatus was within the range observed in the mitogenome of the 56 pentatomid species analyzed (3–20 mtSSRs). A total of 490 mtSSRs were found, representing less than 1% of the mitochondrial genomes analyzed and within the range of microsatellites identified in the nuclear genome of insects (0.02–3.1%)114. The number of mtSSRs was positively related with the size of the mitochondrial genome (r2 = 0.83, p < 0.05), but with a wide variation in abundance among insect species (Fig. 6a), as reported elsewhere39,51. The lowest abundance (0.17 mtSSR/kb) and relative density (0.64 bp/kb) of mtSSRs were observed for D. melacanthus, while the highest values occurred in Plautia fimbriata (abundance = 1.26 mtSSR/kb; relative density = 4.15 bp/kb) (Fig. 6b). The abundance and relative density of mtSSRs in the mitogenome of pentatomids did not correlate with mitogenome size as previously reported in studies of insect genomes114. The GC content (%) of mtSSRs among species was very similar, ranging from 21.9 to 28.3%. The existence of a relationship between the number of microsatellites, abundance, and relative density with the size and GC content of mitochondrial genome in plants51 was not observed in pentatomids.
(a) Relationship between the number of mtSSRs and the mitogenome size (kb) from 56 Pentatomidae species; (b) Genome size (kb) and genomic density (bp/kb) of mtSSRs from 56 Pentatomidae species. The code names are based on the first two letters of the genus and species name (see Table 1).
The majority of mtSSRs of pentatomids were found in coding regions (70%), especially tri- and tetranucleotides (Fig. 7a). No mononucleotides were found in non-coding regions. Studies on mitogenomes of lower eukaryotes115 and animals116 also detected a higher abundance of mtSSRs in genic regions, possibly due to the prokaryotic origin of mitochondria (endosymbiont hypothesis)116. This is different from nuclear genomes in which SSRs are predominantly located in non-coding regions, including introns117. A significant portion of mtSSRs of pentatomids located in coding regions (Fig. 7b) is found in NADH dehydrogenase genes involved in the electron transport chain. nad6 had the highest abundance of microsatellites (18%), followed by nad4 (14%) and nad2 (11%). Additionally, 13% of mtSSRs were located in 16S rRNA.
Tetranucleotides (43.1%) and trinucleotides (40.4%) were the most abundant types of mtSSRs, with all other types of mtSSRs representing less than 7% of abundance (Fig. 8). Studies on SSRs in insects are relatively uncommon but indicate a predominance of di- and trinucleotides114. In plants, where the majority of such studies are conducted, mono-, di-, and trinucleotides often predominate39,118,119,120,121,122. Among all mtSSRs in the mitogenomes of pentatomids, 98.8% of them had only A and T nucleotides in their composition. Dinucleotides were represented only by 6–9 repeats of AT/TA. Motifs TAT (27.1%), TAA (21%), and ATA (17.4%) with 4–6 repeats prevailed in trinucleotides. In tetranucleotides, 3–4 repeats of motifs AAAT (26.2%), TAAA (18.3%), and TTAT (14%) were the most common. Pentanucleotides were predominantly composed of motifs AAATA (30%), AAAAT (16.6%), AAATT (10.0%), and AATAA (10.0%), all with 3 repeats. Mono- and hexanucleotides represented only 1.4% and 1.6% of the total mtSSRs of pentatomids, respectively.
Phylogenetic relationships in Pentatomidae
Four phylogenetic trees were obtained using different analytical methods (ML, CAT + GTR + C4, C60, and BI) of concatenated amino acid sequences of PCGs from 56 species of Pentatomidae (+ 2 outgroup species) (Fig. 9 and 2S). Topology tests (Table 1S) indicated tree topologies produced using CAT + GTR + C4, C60, and BI methods resulted in significant different log-likelihoods (p < 0.05). We chose to explore the BI tree due to the highest posterior probability value obtained (Fig. 9). The recovered phylogeny demonstrated that the Pentatomidae family is a monophyletic group with maximum posterior probability support (PP = 1), reinforcing previous reports68,69,123,124,125,126. However, our data challenge the monophyly hypothesis of Pentatominae and Podopinae, as representatives from both subfamilies intermingle, corroborating published findings69. Asopinae and Phyllocephalinae formed independent clades (PP = 1), although within Pentatominae. The monophyly and position of Asopinae within Pentatominae have been previously proposed based on morphology124 and molecular taxonomy81,127, highlighting the need for more detailed internal classification, especially because Asopinae is not divided into tribes126,128,129. Neojurtina typica (Pentatominae: Pentatomini) is placed as the oldest representative within the family94,95. Other Pentatominae species are also dispersed on the tree, such as Placosternum urus and Piezodorus guildinii, forming sister group relationships with representatives of Phyllocephalinae and Podopinae, respectively (Fig. 9).
Phylogenetic relationships of tribes within Pentatomidae reconstructed from mtDNA sequences of 13 PCGs using the BI method. The numbers on the branches are the posterior probabilities (PP). The length of the branches is proportional to the genetic distance. Outgroup branches that don’t follow the scale are dashed. The subfamilies of Pentatomidae are shown by the colors in each species, and the tribes by vertical bars.
The internal taxonomic structure of Pentatomidae has generally been considered stable. However, Pentatominae has been the subject of continuous questioning, revisions, and adjustments125,128,129. These questions arise because the studies that formed the basis for taxonomic classifications lacked high-quality phylogenetic methodology, had incomplete sampling, and poorly defined characters, limiting the impact of their findings126. Pentatominae comprises more than 40 tribes, including many genera that are challenging to accommodate in other subfamilies129,130.
The Pentatomini tribe is recognized as having less precise taxonomic definition76. The Nezarini and Antestini tribes are positioned within the same clade, suggesting a possible relationship due to their significant morphological similarity126,131,132. However, they are not considered monophyletic, although combined, they form a monophyletic group94.
Representatives of Halyini, also within Pentatominae, resolved into a single clade with high support probability, although positioned separately from most Pentatominae representatives (Fig. 9). The tribes Eysarcorini, Strachiini, Halyini, and Caystrini, and the genera Pentatoma, Plautia, and Diceraeus proved to be monophyletic. The two available mitochondrial genomes for D. melacanthus resolved into parallel branches forming a sister group with D. furcatus, in a clade with high posterior probability, which also includes another Carpocorini, Euschistus heros, in a slightly more external position. However, the Carpocorini tribe is not monophyletic, a result supported by recent analyses involving morphological and molecular data126.
Eysarcoris gibbosus was the first to branch off into the Eysarcorini clade, however, with Carbula sinica separating it from the other representatives of its genus. The positioning of E. gibbosus out of the Eysarcoris group supports the proposed reclassification of E. gibbosus to the genus Stagnomus (tribe Eysarcorini)133, which supports the closer systematically positioning of E. gibbosus to C. sinica76,133,134.
Members of the Piezodorini tribe have often been associated with members of the Menidini131, Nezarini132, Eurysaspini129, and the genus Rhaphigaster126 because of some common external morphological characteristics. However, molecular data based on mitogenome analysis did not support a relationship of Piezodorini with these groups but rather with the Podopinae subfamily (Scotinophara lurida and Graphosoma rubrolineatum), although with low support. The position of Graphosoma and Scotinophora raises some questions because they group with a clade of pentatomines, and positioned themselves as sister species. Others also agree with the monophyly of Podopinae, even though there are several apomorphies that disagree with this classification126.
Our data highlights the conservation of gene content and arrangement between D. melacanthus and D. furcatus, enhancing our understanding of the mitochondrial diversity in Pentatomidae. The identification and characterization of mtSSRs in coding regions offer new perspectives for phylogenetic and genetic studies within the family, suggesting potential molecular markers for population genetics and evolutionary analyses. The phylogenetic analysis underscores the need for revisions in internal taxonomic classification, challenging the monophyly of Pentatominae and Podopinae. In summary, this study not only contributes to the specific understanding of D. melacanthus and D. furcatus but also sheds light on broader issues of the taxonomy and evolution of Pentatomidae, providing insights for future research and refinements in taxonomic classifications.
Data availability
The new mitogenomes have been deposited in GenBank at https://www.ncbi.nlm.nih.gov/ under accession numbers PP235949 e PP235950. The other datasets analysed during the current study are available in GenBank, as referenced in the article.
References
Ávila, C. J. & Panizzi, A. R. Occurrence and damage by Dichelops (Neodichelops) melacanthus (Dallas) (Heteroptera: Pentatomidae) on corn. An. Soc. Entomol. Bras. 24, 193–194 (1995).
Chocorosqui, V. R. & Panizzi, A. R. Impact of cultivation systems on Dichelops melacanthus (Dallas) (Heteroptera: Pentatomidae) population and damage and its chemical control on wheat. Neotrop. Entomol. 33, 487–492 (2004).
Manfredi-Coimbra, S., Silva, J. J., Chocorosqui, V. R. & Panizzi, A. R. Danos do percevejo barriga-verde Dichelops melacanthus (Dallas) (Heteroptera: Pentatomidae) em trigo. Cien. Rural 35, 1243–1247 (2005).
Salvadori, J. R., Pereira, P. R. V. S. & Corrêa-Ferreira, B. S. Pragas ocasionais em lavouras de soja no Rio Grande do Sul. Embrapa Trigo: Documentos Online 91, 34 (2007).
Chiesa, A. C. M., Sismeiro, M. N. S., Pasini, A. & Roggia, S. Tratamento de sementes para manejo do percevejo-barriga-verde na cultura de soja e milho em sucessão. Pesqui. Agropecu. Bras. 51, 301–308 (2016).
Gomes, E. C., Hayashida, R. & Bueno, A. F. Dichelops melacanthus and Euschistus heros injury on maize: Basis for re-evaluating stink bug thresholds for IPM decisions. Crop Prot. 130, 105050 (2020).
Smaniotto, L. F. & Panizzi, A. R. Interactions of selected species of stink bugs (Hemiptera: Heteroptera: Pentatomidae) from leguminous crops with plants in the Neotropics. Fla. Entomol. 98, 7–17 (2015).
Pinheiro, D. H. et al. Suitable reference genes for RT-qPCR analysis in Dichelops melacanthus (Hemiptera: Pentatomidae). Mol. Biol. Rep. 47, 4989–5000 (2020).
Silva, J. J., Ventura, M. U., Silva, F. A. C. & Panizzi, A. R. Population dynamics of Dichelops melacanthus (Dallas) (Heteroptera: Pentatomidae) on host plants. Neotrop. Entomol. 42, 141–145 (2013).
Grazia, J. Revisão do gênero Dichelops Spinola, 1837 (Heteroptera, Pentatomidae, Pentatomini). Iheringia. Serie Zoologia 53, 3–119 (1978).
Trautwein, M. D., Wiegmann, B. M., Beutel, R., Kjer, K. M. & Yeates, D. K. Advances in insect phylogeny at the dawn of the postgenomic era. Annu. Rev. Entomol. 57, 449–468 (2012).
Hebert, P. D. N. & Gregory, T. R. The promise of dna barcoding for taxonomy. Syst. Biol. 54, 852–859 (2005).
Nelson, L. A. et al. Beyond barcoding: A mitochondrial genomics approach to molecular phylogenetics and diagnostics of blowflies (Diptera: Calliphoridae). Gene 511, 131–142 (2012).
Cameron, S. L. Insect mitochondrial genomics: Implications for evolution and phylogeny. Annu. Rev. Entomol. 59, 95–117 (2014).
Wolstenholme, D. Genetic novelties in mitochondrial genomes of multicellular animals. Curr. Opin. Genet. Dev. 2, 918–925 (1992).
Barr, C. M., Neiman, M. & Taylor, D. R. Inheritance and recombination of mitochondrial genomes in plants, fungi and animals. New Phytol. 168, 39–50 (2005).
Dellaporta, S. L. et al. Mitochondrial genome of Trichoplax adhaerens supports Placozoa as the basal lower metazoan phylum. Proc. Natl. Acad. Sci. USA 103, 8751–8756 (2006).
Hao, Y. J. et al. Complete mitochondrial genomes of Anopheles stephensi and A. dirus and comparative evolutionary mitochondriomics of 50 mosquitoes. Sci. Rep. 7, 7666 (2017).
Pakrashi, A., Kumar, V., Banerjee, D., Tyagi, K. & Kalleshwaraswamy, C. M. Gene rearrangements in the mitochondrial genomes of whiteflies (Hemiptera: Aleyrodinae): plesiomorphies, synapomorphies and autapomorphies. Mol. Biol. Rep. (PrePrint) (2021)
Yang, L. et al. Characterization of the complete mitochondrial genome of Orthaga olivacea Warre (Lepidoptera Pyralidae) and comparison with other lepidopteran insects. PloS One 15, e0227831. https://doi.org/10.1371/journal.pone.0227831 (2020).
Tyagi, K. et al. Rearrangement and evolution of mitochondrial genomes in Thysanoptera (Insecta). Sci. Rep. 10, 695 (2020).
Tyagi, K. et al. The gene arrangement and phylogeny using mitochondrial genomes in spiders (Arachnida: Araneae). Int. J. Biol. Macromol. 146, 488–496 (2020).
Higgs, P. G., Jameson, D., Jow, H. & Rattray, M. The evolution of tRNA-Leu genes in animal mitochondrial genomes. J. Mol. Evol. 57, 435–445 (2003).
Song, N., Liang, A. P. & Bu, C. P. A molecular phylogeny of Hemiptera inferred from mitochondrial genome sequences. PloS One 7, e48778. https://doi.org/10.1371/journal.pone.0048778 (2012).
Zhang, D. X. & Hewitt, G. M. Insect mitochondrial control region: A review of its structure, evolution and usefulness in evolutionary studies. Biochem. Syst. Ecol. 25, 99–120 (1997).
Curole, J. P. & Kocher, T. D. Mitogenomics: Digging deeper with complete mitochondrial genomes. Trends Ecol. Evol. 14, 394–398 (1999).
Boore, J. L., Lavrov, D. V. & Brown, W. M. Gene translocation links insects and crustaceans. Nature 392, 667–668 (1998).
Boore, J. L. Animal mitochondrial genomes. Nucleic Acids Res. 27, 1767–1780 (1999).
Yuan, M. L., Zhang, Q. L., Guo, Z. L., Wang, J. & Shen, Y. Y. Comparative mitogenomic analysis of the superfamily Pentatomoidea (Insecta: Hemiptera: Heteroptera) and phylogenetic implications. BMC Genom. 16, 460. https://doi.org/10.1186/s12864-015-1679-x (2015).
Kumar, S., Singh, A. & Shanker, A. pSATdb: A database of mitochondrial common, polymorphic, and unique microsatellites. Life Sci. Alliance 5, e202101307. https://doi.org/10.2650/lsa.202101307 (2022).
Parida, S. K. et al. Informative genomic microsatellite markers for efficient genotyping applications in sugarcane. Theor. Appl. Genet. 118, 327–338 (2009).
Gemayel, R., Cho, J., Boeynaems, S. & Verstrepen, K. J. Beyond junk-variable tandem repeats as facilitators of rapid evolution of regulatory and coding sequences. Genes 3, 461–480. https://doi.org/10.3390/genes3030461 (2012).
Christians, J. K. & Watt, C. A. Mononucleotide repeats represent an important source of polymorphic microsatellite markers in Aspergillus nidulans. Mol. Ecol. Resour. 9, 572–578 (2009).
Tautz, D. & Renz, M. Simple sequences are ubiquitous repetitive components of eukaryotic genomes. Nucleic Acids Res. 12, 4127–4138 (1984).
Zane, L., Bargelloni, L. & Patarnello, T. Strategies for microsatellite isolation: A review. Mol. Ecol. 11, 1–16 (2002).
Kalia, R. K., Rai, M. K., Kalia, S., Singh, R. & Dhawan, A. K. Microsatellite markers: An overview of the recent progress in plants. Euphytica 177, 309–334 (2011).
Phumichai, C., Phumichai, T. & Wongkaew, A. Novel chloroplast microsatellite (cpSSR) markers for genetic diversity assessment of cultivated and wild Hevea rubber. Plant Mol. Biol. 33, 1486–1498 (2015).
Zhao, Z. et al. Genome-wide analysis of tandem repeats in plants and green algae. G3: Genes Genomes Genet. 4, 67–78. https://doi.org/10.1534/g3.113.008524 (2014).
Rajendrakumar, P. et al. Simple sequence repeats in organellar genomes of rice: Frequency and distribution in genic and intergenic regions. Bioinformatics 23, 1–4 (2007).
Soranzo, N., Provan, J. & Powell, W. An example of microsatellite length variation in the mitochondrial genome of conifers. Genome 42, 158–161 (1999).
Lawson, M. J. & Zhang, L. Distinct patterns of SSR distribution in the Arabidopsis thaliana and rice genomes. Genome Biol. 7, R14 (2006).
Vieira, M. L. C., Santini, L., Diniz, A. L. & Munhoz, C. F. Microsatellite markers: what they mean and why they are so useful. Genet. Mol. Biol. 39, 312–328 (2016).
Andrews, S. FastQC: A quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2010)
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dierckxsens, N., Mardulyn, P. & Smits, G. NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res. 45, e18 (2017).
Bernt, M. et al. MITOS: Improved de novo metazoan mitochondrial genome annotation. Mol. Phylogenet. Evol. 69, 313–319 (2013).
Conant, G. C. & Wolfe, K. H. GenomeVx: Simple web-based creation of editable circular chromosome maps. Bioinformatics 24, 861–862 (2008).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Grant, J. R. & Stothard, P. The CGView server: A comparative genomics tool for circular genomes. Nucleic Acids Res. 36, W181–W184 (2008).
Beier, S., Thiel, T., Münch, T., Scholz, U. & Mascher, M. MISA-web: A web server for microsatellite prediction. Bioinformatics 33, 2583–2585. https://doi.org/10.1093/bioinformatics/btx198 (2017).
Freitas, K. E. J. et al. An empirical analysis of mtSSRs: Could microsatellite distribution patterns explain the evolution of mitogenomes in plants?. Funct. Integr. Genom. 22, 35–53 (2022).
Tan, M. H., Gan, H. M., Schultz, M. B. & Austin, C. M. MitoPhAST, a new automated mitogenomic phylogeny tool in the post-genomic era with a case study of 89 decapod mitogenomes including eight new freshwater crayfish mitogenomes. Mol. Phylogenet. Evol. 85, 180–188 (2015).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal omega. Mol. Syst. Biol. 7, 539. https://doi.org/10.1038/msb.2011.75 (2011).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973. https://doi.org/10.1093/bioinformatics/btp348 (2009).
Nguyen, T. T. T., Meewan, M., Ryan, S. & Austin, C. M. Genetic diversity and translocation in the marron, Cherax tenuimanus (Smith): Implications for management and conservation. Fish. Manag. Ecol. 9, 163–173 (2002).
Kalyaanamoorthy, S. et al. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Ronquist, F. et al. MrBayes 3.2: Efficient bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Miller, M., Pfeiffer, W. & Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. Gateway Computing Environments Workshop (GCE), 1–8. https://doi.org/10.1109/GCE.2010.5676129 (2010)
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Kishino, H. & Hasegawa, M. Evaluation of the maximum likelihood estimate of the evolutionary tree topologies from DNA sequence data, and the branching order in hominoidea. J. Mol. Evol. 29, 170–179 (1989).
Shimodaira, H. & Hasegawa, M. Multiple comparisons of log-likelihoods with applications to phylogenetic inference. Mol. Biol. Evol. 16, 1114 (1999).
Strimmer, K. & Rambaut, A. Inferring confidence sets of possibly misspecified gene trees. Proc. R. Soc. B. 269, 137–142 (2002).
Shimodaira, H. An approximately unbiased test of phylogenetic tree selection. Syst. Biol. 51, 492–508 (2002).
Mu, Y. L., Zhang, C. H., Zhang, Y. J., Yang, L. & Chen, X. S. Characterizing the complete mitochondrial genome of Arma custos and Picromerus lewisi (Hemiptera: Pentatomidae: Asopinae) and conducting phylogenetic analysis. J. Insect Sci. 22, 6 (2022).
Wu, Y. et al. Characterization of the complete mitochondrial genome of Arma custos (Hemiptera: Pentatomidae). Mitochondrial DNA Part B 5, 2624–2626 (2020).
Liu, Y. et al. Higher-level phylogeny and evolutionary history of Pentatomomorpha (Hemiptera: Heteroptera) inferred from mitochondrial genome sequences. Syst. Entomol. 44, 810–819 (2019).
Jiang, P. Studies on the comparative mitochondrial genomics and phylogeny of Heteroptera (Insecta: Hemiptera). PhD, China Agricultural University, Beijing (2017)
Xu, S. et al. Comparative mitogenomics and phylogenetic analyses of Pentatomoidea (Hemiptera: Heteroptera). Genes 12, 1306. https://doi.org/10.3390/genes12091306 (2021).
Gonçalves, L. T., Pezzi, P. H. & Bianchi, F. M. Four new stink bug mitogenomes corroborate the internal inconsistencies in the classification of Pentatomidae (Hemiptera). Zootaxa 5120, 128–142 (2022).
Zhao, Q. et al. Complete mitochondrial genome of Dinorhynchus dybowskyi (Hemiptera: Pentatomidae: Asopinae) and phylogenetic analysis of Pentatomomorpha species. J. Insect Sci. 18, 44 (2018).
Zhang, Q. L. et al. Adaptive evidence of mitochondrial genomes in Dolycoris baccarum (Hemiptera: Pentatomidae) to divergent altitude environments. Mitochondrial DNA Part A 30, 9–15 (2019).
Guo, Y., Xiao, J., Li, D. & Wang, J. The complete mitochondrial genome of the stink bug Eocanthecona furcellata (Hemiptera: Pentatomidae). Mitochondrial DNA Part B 6, 3085–3086 (2021).
Gao, S. et al. PacBio full-length transcriptome profiling of insect mitochondrial gene expression. RNA Biol. 13, 820–825 (2016).
Zhao, W. et al. Comparative mitogenomic analysis of the Eurydema genus in the context of representative Pentatomidae (Hemiptera: Heteroptera) taxa. J. Insect Sci. 19, 1–12 (2019).
Zhao, Q., Chen, C., Liu, J. & Wei, J. Characterization of the complete mitochondrial genome of Eysarcoris aeneus (Heteroptera: Pentatomidae), with its phylogenetic analysis. Mitochondrial DNA Part B 4, 2096–2097 (2019).
Li, R., Li, M., Yan, J., Bai, M. & Zhang, H. Five mitochondrial genomes of the genus Eysarcoris Hahn, 1834 with phylogenetic implications for the Pentatominae (Hemiptera: Pentatomidae). Insects 12, 597. https://doi.org/10.3390/insects12070597 (2021).
Chen, Q., Niu, X., Fang, Z. & Weng, Q. The complete mitochondrial genome of Eysarcoris guttigerus (Hemiptera: Pentatomidae). Mitochondrial DNA Part B 5, 687–688 (2020).
Chen, C., Wei, J., Ji, W. & Zhao, Q. The first complete mitochondrial genome from the subfamily Phyllocephalinae (Heteroptera: Pentatomidae) and its phylogenetic analysis. Mitochondrial DNA Part B. 2, 938–939 (2017).
Wang, J., Zhang, L., Yang, X. Z., Zhou, M. Q. & Yuan, M. L. The first mitochondrial genome for the subfamily Podopinae (Hemiptera: Pentatomidae) and its phylogenetic implications. Mitochondrial DNA Part B 2, 219–220 (2017).
Lee, Y. et al. A mitochondrial genome phylogeny of Mytilidae (Bivalvia: Mytilida). Mol. Phylogenetics Evol. 139, 106533 (2019).
Li, Q. et al. Characterization and comparative analysis of six complete mitochondrial genomes from ectomycorrhizal fungi of the Lactarius genus and phylogenetic analysis of the Agaricomycetes. Int. J. Biol. Macromol. 121, 249–260 (2019).
Zhao, L. et al. The complete mitochondrial genome of Menida violacea (Hemiptera: Pentatomidae) and its phylogenetic implication. Mitochondrial DNA Part B 4, 1953–1954 (2019).
Hua, J. et al. Comparative and phylogenomic studies on the mitochondrial genomes of Pentatomomorpha (Insecta: Hemiptera: Heteroptera). BMC Genom. 9, 610. https://doi.org/10.1186/1471-2164-9-610 (2008).
Chen, W. T., Zhang, L. J., Cao, Y. & Yuan, M. L. The complete mitochondrial genome of Palomena viridissima (Hemiptera: Pentatomidae) and phylogenetic analysis. Mitochondrial DNA Part B 6, 1326–1327 (2021).
Zhao, L. et al. The complete mitochondrial genome of Pentatoma rufipes (Hemiptera, pentatomidae) and its phylogenetic implications. ZooKeys 1042, 51–72 (2021).
Zhao, Q. et al. The first mitochondrial genome for the subfamily Asopinae (Heteroptera: Pentatomidae) and its phylogenetic implications. Mitochondrial DNA Part B 2, 804–805 (2017).
Wang, Y., Duan, Y. & Yang, X. The complete mitochondrial genome of Plautia crossota (Hemiptera: Pentatomidae). Mitochondrial DNA Part B 4, 2281–2282 (2019).
Zhao, Q. et al. The complete mitochondrial genome of Zicrona caerulea (Linnaeus) (Hemiptera: Pentatomidae: Asopinae) and its phylogenetic implications. Zootaxa 4747, 547–561 (2020).
Thao, M. L. L., Baumann, L. & Baumann, P. Organization of the mitochondrial genomes of whiteflies, aphids, and psyllids (Hemiptera, Sternorrhyncha). BMC Evol. Biol. 4, 25 (2004).
Dotson, E. M. & Beard, C. B. Sequence and organization of the mitochondrial genome of the Chagas disease vector, Triatoma dimidiata. Insect Mol. Biol. 10, 205–215 (2001).
Guo, Z. & Yuan, M. Research progress of mitochondrial genomes of Hemiptera insects. Sci. Sin. 46, 151–166 (2016).
Hassanin, A., Léger, N. & Deutsch, J. Evidence for multiple reversals of asymmetric mutational constraints during the evolution of the mitochondrial genome of Metazoa, and consequences for phylogenetic inferences. Syst. Biol. 54, 277–298 (2005).
Wei, S. J. et al. New views on strand asymmetry in insect mitochondrial genomes. PloS One 5, 1–10. https://doi.org/10.1371/journal.pone.0012708 (2010).
Ding, X. et al. Comparative mitogenomics and phylogenetic analyses of the genus Menida (Hemiptera, Heteroptera, Pentatomidae). ZooKeys 1138, 29–48 (2023).
Lian, D. et al. Comparative analysis and phylogeny of mitochondrial genomes of Pentatomidae (Hemiptera: Pentatomoidea). Front. Genet. https://doi.org/10.3389/fgene.2022.1045193 (2022).
Zhao, W. et al. Characterization of the complete mitochondrial genome and phylogenetic implications for Eurydema maracandica (Hemiptera: Pentatomidae). Mitochondrial DNA Part B 2, 550–551 (2017).
Jühling, F. et al. Improved systematic tRNA gene annotation allows new insights into the evolution of mitochondrial tRNA structures and into the mechanisms of mitochondrial genome rearrangements. Nucleic Acids Res. 40, 2833–2845 (2012).
Lu, C., Huang, X. & Deng, J. The challenge of Coccidae (Hemiptera: Coccoidea) mitochondrial genomes: The case of Saissetia coffeae with novel truncated tRNAs and gene rearrangements. Int. J. Biol. Macromol. 158, 854–864 (2020).
Li, H. et al. Comparative mitogenomic analysis of damsel bugs representing three tribes in the family Nabidae (Insecta: Hemiptera). PloS One 7, e45925. https://doi.org/10.1371/journal.pone.0045925 (2012).
Chen, S. C. et al. The complete mitochondrial genome of Aleurocanthus camelliae: Insights into gene arrangement and genome organization within the family Aleyrodidae. Int. J. Mol. Sci. 17, 1843 (2016).
Adams, K. & Palmer, J. D. Evolution of mitochondrial gene content: gene loss and transfer to the nucleus. Mol. Phylogenetics Evol. 29, 380–395 (2003).
Rogalski, M., Karcher, D. & Bock, R. Superwobbling facilitates translation with reduced tRNA sets. Nat. Struct. Mol. Biol. 15, 192–198 (2008).
Salinas, T., Duchêne, A. M. & Maréchal-Drouard, L. Recent advances in tRNA mitochondrial import. Trends Biochem. Sci. 33, 320–329 (2008).
Saunier, A. et al. Mitochondrial genomes of the Baltic clam Macoma balthica (Bivalvia: Tellinidae): Setting the stage for studying mito-nuclear incompatibilities. BMC Evol. Biol. 14, 259 (2014).
Warren, J. M. & Sloan, D. B. Interchangeable parts: The evolutionarily dynamic tRNA population in plant mitochondria. Mitochondrion 52, 144–156 (2020).
Adams, K. L., Qiu, Y. L., Stoutemyer, M. & Palmer, J. D. Punctuated evolution of mitochondrial gene content: high and variable rates of mitochondrial gene loss and transfer to the nucleus during angiosperm evolution. Proc. Natl. Acad. Sci. USA 99, 9905–9912 (2002).
Lynch, M. Mutation accumulation in nuclear, organelle, and prokaryotic transfer RNA genes. Mol. Biol. Evol. 14, 914–925 (1997).
Domes, K., Maraun, M., Scheu, S. & Cameron, S. L. The complete mitochondrial genome of the sexual oribatid mite Steganacarus magnus: Genome rearrangements and loss of tRNAs. BMC Genom. 9, 532. https://doi.org/10.1186/1471-2164-9-532 (2008).
Li, H. & Li, J. Comparative analysis of four complete mitogenomes from hoverfly genus Eristalinus with phylogenetic implications. Sci. Rep. 12, 4164 (2022).
Fourdrilis, S., Frias Martins, A. M. & Backeljau, T. Relation between mitochondrial DNA hyperdiversity, mutation rate and mitochondrial genome evolution in Melarhaphe neritoides (Gastropoda: Littorinidae) and other Caenogastropoda. Sci. Rep. 8, 17964 (2018).
Yang, Z. & Bielawski, J. P. Statistical methods for detecting molecular adaptation. Trends Ecol. Evol. 15, 496–503 (2000).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Marine Biol. Biotechnol. 3, 294–299 (1994).
Geller, J., Meyer, C., Parker, M. & Hawk, H. Redesign of PCR primers for mitochondrial cytochrome c oxidase subunit I for marine invertebrates and application in all-taxa biotic surveys. Mol. Ecol. Resour. 13, 851–861 (2013).
Ding, S., Wang, S., He, K., Jiang, M. & Li, F. Large-scale analysis reveals that the genome features of simple sequence repeats are generally conserved at the family level in insects. BMC Genom. 18, 848. https://doi.org/10.1186/s12864-017-4234-0 (2017).
Levdansky, E. et al. Coding tandem repeats generate diversity in Aspergillus fumigatus genes. Eukaryot. Cell 6, 1380–1391 (2007).
Shakyawar, S. K., Joshi, B. K. & Kumar, D. SSR repeat dynamics in mitochondrial genomes of five domestic animal species. Bioinformation 4, 158–163 (2009).
Hancock, J. M. The contribution of slippage-like processes to genome evolution. J. Mol. Evol. 41, 1038–1047 (1995).
Anand, K., Kumar, S., Alam, A. & Shankar, A. Mining of microsatellites in mitochondrial genomes of order Hypnales (Bryopsida). Plant Sci. Today 6, 635–638. https://doi.org/10.14719/pst.2019.6.sp1.697 (2019).
Filiz, E. SSRs mining of Brassica species in mitochondrial genomes: Bioinformatic approaches. Hortic. Environ. Biotechnol. 54, 548–553 (2013).
George, B., Bhatt, B. S., Awasthi, M., George, B. & Singh, A. K. Comparative analysis of microsatellites in chloroplast genomes of lower and higher plants. Curr. Genet. 61, 665–677 (2015).
Khera, P., Saxena, R., Sameerkumar, C. V., Saxena, K. & Varshney, R. K. Mitochondrial SSRs and their utility in distinguishing wild species, CMS lines and maintainer lines in pigeonpea (Cajanus cajan L.). Euphytica 206, 737–746 (2015).
Kuntal, H., Sharma, V. & Daniell, H. Microsatellite analysis in organelle genomes of Chlorophyta. Bioinformation 8, 255–259 (2012).
Bianchi, F. M. & Gonçalves, L. T. Borrowing the Pentatomomorpha tome from the DNA barcode library: Scanning the overall performance of cox1 as a tool. J. Zoolog. Syst. Evol. Res. 59, 992–1012 (2021).
Gapud, V. D. L. P. A generic revision of the subfamily Asopinae, with consideration of its phylogenetic position in the family Pentatomidae and superfamily Pentatomoidea (Hemiptera Heteroptera). Philipp. Entomol. 8, 865–961 (1991).
Grazia, J., Schuh, R. T. & Wheeler, W. C. Phylogenetic relationships of family groups in Pentatomoidea based on morphology and DNA sequences (Insecta: Heteroptera). Cladistics 24, 932–976 (2008).
Roca-Cusachs, M. et al. Opening Pandora’s box: molecular phylogeny of the stink bugs (Hemiptera: Heteroptera: Pentatomidae) reveals great incongruences in the current classification. Syst. Entomol. 47, 36–51 (2022).
Wu, Y. Z. et al. The evolutionary position of Lestoniidae revealed by molecular autapomorphies in the secondary structure of rRNA besides phylogenetic reconstruction (Insecta: Hemiptera: Heteroptera). Zool. J. Linn. Soc. 177, 750–763 (2016).
Rider, D. A. Stirotarsinae, new subfamily for Stirotarsus abnormis Bergroth (Heteroptera: Pentatomidae). Ann. Entomol. Soc. Am. 93, 802–806 (2000).
Rider, D. A. et al. Higher Systematics of the Pentatomoidea. In Invasive Stink Bugs and Related Species (Pentatomoidea) (ed. McPherson, J. E.) 178 (CRC Press, 2018).
Genevcius, B. C. et al. Phylogeny of the stink bug tribe Chlorocorini (Heteroptera, Pentatomidae) based on DNA and morphological data: The evolution of key phenotypic traits. Syst. Entomol. 46, 327–338 (2021).
Gross, G. F. Plant-Feeding and Other Bugs (Hemiptera) of South Australia, Heteroptera–Part 1. In Handbook of the Flora and Fauna of South Australia (eds Jessop, J. P. & Toelken, H. R.) 1–250 (Handbooks Committee, 1975).
Linnavuori, R. E. Pentatomidae and Acanthosomatidae (Heteroptera) of Nigeria and the Ivory Coast, with remarks on species of the adjacent countries in West and Central Africa. Acta Zool. Fenn. 163, 1–176 (1982).
Roca-Cusachs, M. & Jung, S. Redefining Stagonomus Gorski based on morphological and molecular data (Pentatomidae: Eysarcorini). Zootaxa 4658, 368–374 (2019).
Zhang, H. F. Studies on the cytotaxonomy of five species of the family Pentatomidae (Hemiptera: Heteroptera). Entomotaxonomia 23, 265–276 (2001).
Acknowledgements
The authors acknowledge the Coordination of Higher Education Personnel Improvement (CAPES–CODE 1) for providing a PhD. fellowship to support LCD.
Author information
Authors and Affiliations
Contributions
Conceptualization: L. C. D. and F. L. C.; data curation: L. C. D.; formal analysis: L. C. D. and F. L. C.; funding acquisition: F. L. C.; investigation: L. C. D. and F. L. C.; methodology: L. C. D. and F. L. C.; project administration: F. L. C.; supervision: F. L. C.; writing original draft: L. C. D. and F. L. C.; writing, review and editing: all authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dallagnol, L.C., Cônsoli, F.L. Evolutionary and phylogenetic insights from the mitochondrial genomic analysis of Diceraeus melacanthus and D. furcatus (Hemiptera: Pentatomidae). Sci Rep 14, 12861 (2024). https://doi.org/10.1038/s41598-024-63584-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-63584-w
- Springer Nature Limited