Abstract
Coffea arabica is an allotetraploid of high economic importance. C. arabica transcriptome is a combination of the transcripts of two parental genomes (C. eugenioides and C. canephora) that gave rise to the homeologous genes of the species. Previous studies have reported the transcriptional dynamics of C. arabica. In these reports, the ancestry of homeologous genes was identified and the overall regulation of homeologous differential expression (HDE) was explored. One of these genes is part of the FRIGIDA-like family (FRL), which includes the Arabidopsis thaliana flowering-time regulation protein, FRIGIDA (FRI). As nonfunctional FRI proteins give rise to rapid-cycling summer annual ecotypes instead of vernalization-responsive winter-annuals, allelic variation in FRI can modulate flowering time in A. thaliana. Using bioinformatics, genomic analysis, and the evaluation of gene expression of homeologs, we characterized the FRL gene family in C. arabica. Our findings indicate that C. arabica expresses 10 FRL homeologs, and that, throughout flower and fruit development, these genes are differentially transcribed. Strikingly, in addition to confirming the expression of FRL genes during zygotic embryogenesis, we detected FRL expression during direct somatic embryogenesis, a novel finding regarding the FRL gene family. The HDE profile of FRL genes suggests an intertwined homeologous gene regulation. Furthermore, we observed that FLC gene of C. arabica has an expression profile similar to that of CaFRL genes.
Similar content being viewed by others
Introduction
Coffea arabica and C. canephora are the species responsible for the production of all coffee beans worldwide. As an allotetraploid (2n = 4x = 44), the C. arabica genome is composed of the diploid genomes (2n = 2x = 22) of its ancestors, C. canephora and C. eugenioides, which became subgenomes within this species (CaCc and CaCe, respectively)1,2,3. Coffea eugenioides is a bush-like plant that inhabits mild-temperature highlands and produces low caffeine-containing small fruits4. Coffea canephora trees inhabit warm tropical-equatorial lowlands and produce high caffeine-containing seeds5. The two parental species are closely related, and the two subgenomes in C. arabica have low sequence divergence (i.e., 1.3% average difference in the genes)1, which is also correlated with the autogamous reproductive strategy of C. arabica.
Several studies have found that the transcriptional set of C. arabica is a combination of the homeologous gene expression of the CaCc and CaCe subgenomes3,6,7,8,9,10,11,12. It is extremely likely that the homeologous differential expression (HDE) in C. arabica is responsible for the plasticity in phenotype modulation in different tissues and under different biological conditions. In fact, allopolyploidization has been considered a contributor to speciation and plant adaptation to broader habitats6,13,14,15,16. Although homeolog loss and silencing were found to be common in the CaCc subgenome, which suggested CaCe dominance, neither of the two subgenomes were preferentially expressed in C. arabica8. Therefore, it appears that each gene has its own homeologous expression coordination, providing global intertwined homeolog regulation in C. arabica.
In Arabidopsis thaliana, FRIGIDA (FRI) is a key protein that regulates flowering transition by activating the flowering locus C (FLC), which encodes a central flowering repressor that controls the plant response to vernalization17,18,19. FRI acts as a scaffold protein that interacts with other proteins to assemble a complex that binds to the FLC promoter region, thereby triggering its expression, and consequently, inhibiting flowering20. On the contrary, vernalization has no effect on FRI expression, and instead promotes flowering by causing the epigenetic repression of FLC expression19.
FRLs (FRIGIDA-Like genes) have been found in all sequenced plant genomes, regardless of whether the species displays vernalization. Even though FRI is connected to flowering regulation, members of this gene family are also associated with other biological processes connected with reproduction, such as embryonic development21 and seed maturation22. Based on single-nucleotide polymorphism (SNP)–based detection of homeologous genes, two C. arabica FRLs were suggested to display HDE3 (more details in the Methods section). Given the advances in genome and transcriptome sequencing of C. arabica, C. canephora, and C. eugenioides23,24,25, we further characterized the Coffea FRLs by evaluating their sequence features, phylogenetics, and cis-regulatory elements, and by characterizing the C. arabica FRL transcription and HDE in tissues such as flowers and fruits, and during direct somatic embryogenesis. In addition, C. arabica FLC expression was evaluated, which indicated an expression profile similar to that of FRL genes. Our results provide strong support to the hypothesis that FRLs are active in diverse stages of plant reproduction.
Results
Characterization of FRL genes in Coffea
Eight sequences of A. thaliana FRLs were used to search the BlastP database against the C. canephora genome sequence23. Five genes were found to be similar to the corresponding genes in A. thaliana (Table 1). Next, C. canephora FRL sequences were used in the BlastP search against the C. arabica genome sequence (http://www.phvtozome.net) and the C. eugenioides EST databank25. Five sequences were found in C. eugenioides and 10 in C. arabica. After aligning all the FRLs from the abovementioned Coffea species, it was possible to assign C. arabica homeologous FRLs using the same SNP alignment–based strategy3. Genes considered as present in C. canephora subgenome were designated as x.1, and the genes considered as present in C. eugenioides subgenome were designated as x.2 (CaCe; Table 1). It should be mentioned that, because we did not have access to the C. eugenioides genome yet, the C. eugenioides FRL genes were not completely described (e.g., complete gene annotation, presence of genes, and orthologs).
FRIGIDA domain PF07899 (https://pfam.xfam.org) was detected in all the analyzed sequences as well as those used for protein alignment (Fig. 1). In addition, the N-terminus of the FRLs was evaluated to classify Coffea FRLs within gene families based on protein sequence analyses described by Risk et al.22. Supplementary Fig. S1 (supplementary note) shows the five FRL families and the presence of members of C. canephora and C. arabica in each A. thaliana FRL family.
Sequence alignment of Frigida domain in Coffea (Ca, Cc) and Arabdopsis (At) Frigida-like proteins. Black background, more than 90% of conservation between amino acids; Dark gray background and white letters, conservation between amino acids 89–80%; Light gray background and black letters, conservation between amino acids 79–60%.
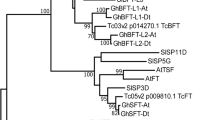
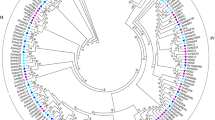
To gain insight into the evolutionary relationships of FRL genes in Coffea and other plant genomes, the neighbor-joining method was used to construct a phylogenetic tree (Fig. 2). Sequences of C. canephora FRL and its respective A. thaliana orthologs were grouped within the same clade (Fig. 2). Mixed sequences from monocotyledons and dicotyledons were found within different clades, suggesting an ancestral FRL origin before the divergence of the plant clades. In addition, we observed that each C. canephora FRL was allocated to an FRL subfamily, as described by Risk et al.22, which is not the case for tomato and potato (Solanaceae), which lack AtFRL1 homeologs, and rice and sorghum (Poaceae), which lack AtFRI homeologs (Fig. 2).
Phylogenetic analysis of CcFRL proteins with orthologous proteins of A. thaliana (At), S. bicolor (Sorbi), O. sativa (ORYSA), S. lycopersicum (SOLLY), S. tuberosum (SOLTU) and V. vinifera (VITVI). The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) is shown next to the branches. Bootstrap values >50 are shown on the tree. The evolutionary distances were computed using the p-distance method.
Coffea arabica homeologous FRL assignments
SNP alignment-based strategy was used to assign the FRL homeologous genes in C. arabica. Briefly, C. arabica (2), C. canephora (1), and C. eugenioides (1) sequences similar to each A. thaliana FRL were aligned based on the SNP profile. C. arabica genes were assigned as derived from the C. canephora subgenome (CaCc; FRL x.1) or C. eugenioides subgenome (CaCe; FRL x.2). The C. arabica, C. eugenioides, and C. canephora FRL genes were aligned to construct a dendogram, which confirmed the subgenome assignment (Supplementary Fig. S2, supplementary note). SNPs observed in silico allowed for designing of a homeolog-specific primer in C. arabica according to the TaqMAMA method26 (Supplementary Table S1, supplementary note), or containing an indel of at least three nucleotides (Supplementary Fig. S3, supplementary note). The subgenome specificity of each homeolog-specific primer was tested with quantitative real-time (qRT-PCR) using cDNA from the leaves of C. canephora, C. eugenioides, and C. arabica. As expected, the primers designed from the CaCe subgenome amplified only the C. eugenioides cDNA, and primers that matched the CaCc subgenome amplified only the C. canephora cDNA (Fig. 3, left and middle columns). In contrast, both the primers were effective in amplifying the C. arabica homeologs (CaCe and CaCc) in each FRL, indicating that these genes and, consequently, both the subgenomes, are transcriptionally active in C. arabica (Fig. 3, right column).
Expression profiles of CaFRL homeologous genes (CaCc and CaCe) in leaves of C. arabica, C. canephora, C. eugenioides and a 50:50 mix of the parental cDNAs (C. canephora and C. eugenioides). Gray bars refer to CaCc and white bars refer to CaCe. Values of three technical replicates are presented as mean ± SD (error bars). Transcript abundances were normalized using the expression of UBI (ubiquitin) as reference gene. Asterisks indicate significant differences (P < 0.05) between homeologous genes.
Interestingly, the FRL homeologs displayed different expression profiles in C. arabica leaves. For example, CaFRL-1 and CaFRL-2 CaCe homeologs (FRL x.2) were expressed more than were the CaCc homeologs (FRL x.1). Inversely, for CaFRL-4 and CaFRL-5, the homeologous expression of CaCc was greater than that of CaCe (Fig. 3). The CaFRL-3 homeologous expression was balanced (Fig. 3).
To further verify the homeolog-specific findings, qPCR experiments using a 50:50 mix of the parental cDNAs were performed to ensure that the primers, when being amplified from the tetraploid, were indeed behaving in a homeolog-specific manner. Briefly, we made a 50:50 mix of cDNA from C. eugenioides and C. canephora, and carried out qPCR with homeolog-specific primers for each gene. We did not perform multiplex analysis but included the cDNA mix and each homeolog primer in separate wells. We confirmed the amplification of each homeolog in the mix, and interestingly, noted a similar expression rate as seen in ancestral samples, when comparing the expression scales of ancestral amplifications with that of the 50:50 mix (Fig. 3).
The homeologous promoters of CaFRLs were also assigned and evaluated with an aim to find the putative differential cis-elements among them. The results can be found in the Supplementary file (Figs S4–S7, supplementary note).
Coffea arabica FRL expression during flower development
CaFRL expression was assessed during four stages of C. arabica floral development, depicted in Fig. 4A: green floral buds, white floral buds <10 mm (white 1), white floral buds >10 mm (white 2), and open flowers (anthesis). All the CaFRLs showed higher expression levels in the white 1 stage than in other floral stages, and in particular, CaFRL-5 (Fig. 5A) showed a decay in transcription during advanced floral development. The evaluation of HDE (Fig. 5B) revealed that the expressions of CaFRL-2, CaFRL-4, and CaFRL-5 were subgenome biased (CaFRL-2 and CaFRL-5 toward CaCe and CaFRL-4 toward CaCc), whereas CaFRL-1 and CaFRL-3 homeologs tended to be similarly expressed throughout flower development (Fig. 5B).
Anatomical view of C. arabica organs and tissues at which CaFRL gene expression was evaluated. (A) Flowers at different stages (green cluster, white cluster, white candle and anthesis). Scale: 5 mm. (B) Fruits (From top to bottom: whole fruit, fruit cross section, fruit longitudinal section, embryos; from left to right: days after flowering). Perisperm (pe), embryo (eb), endosperm (end), pericarp (pe). Scale: fruits = 2 mm, embryos = 1 mm. (C) Foliar explants collected throughout DSE. 0 days (0d), 8 days (8d), 16 days (16d), 28 days (28d), 60 days (60d), embryo formation (Embryos). Adaxial epidermis (AD), palisade parenchyma (Pp), spongy parenchyma (Sp),vascular bundle (Vb), abaxial epidermis (AB), stomata (St); asterisk indicates intense mitosis in spongy parenchyma; Arrow indicates the beginning of cellular division at spongy parenchyma; Pró-embryonic mass (Pm); Meristematic cells (Mc) Scale: 0d-28d = 50 µm, 60d and Embryos = 200 µm.
Gene expression analysis of CaFRL genes and CaFLC gene in C. arabica flowers. (A) Heat map visualization of CaFRL expression in flowers at different stages (see material and methods). The sum of relative homeolog expressions was used as numerical input for creating the heat map scale from light green (weakly expressed) to red (strongly expressed). ‘Green cluster’ sample was used as internal calibrator. (B) Expression profiles of homeologous genes (CaCc and CaCe) of CaFRL family in flowers at different stages (green cluster, white 1 floral bud, white 2 floral bud and anthesis). Values of three technical replicates are presented as mean ± SD (error bars). Transcript abundances were normalized using the expression of UBI (ubiquitin) as reference gene. Asterisks indicate significant differences (P < 0.05) between homeologous genes.
Coffea arabica FRL expression during fruit development
Coffee fruit development is a long process that can be evaluated using the cross and longitudinal sections of the fruit. Between 60 and 90 DAF, the perisperm (inner fruit) and pericarp (outer fruit) develop. Perisperm, a prevalent inner tissue, gradually disappears and is replaced by the endosperm, surrounded by a thin tissue of silver skin membrane (Fig. 4B). By 120 DAF, the embryo can be visualized, and by 180 DAF, it achieves its final length and morphology (Fig. 4B). Based on these macroscopic parameters, 60 and 90 DAF contained perisperm and pericarp samples, while the subsequent harvest days contained pericarp, endosperm, and embryo (Fig. 4B).
The expression analysis heat map showed that all CaFRLs were expressed in fruits, especially in the embryo and endosperm, with different transcriptional profiles (Fig. 6). CaFRL-1 had a nearly specific expression that manifests during the late-endosperm stage (Fig. 6A). CaFRL-2 had the highest expression in the embryo (120–240 DAF), followed by the endosperm (210–240 DAF). CaFRL-3 had the highest expression in the endosperm (120–240 DAF) and embryo (150 DAF). CaFRL-4 had the highest expression in the perisperm (90 DAF) and embryo (150–240 DAF). Finally, CaFRL-5 had the highest expression in the endosperm (240 DAF). As for the role of HDE in fruit development, the expression profile of CaFRL-1 was intertwined; CaCe homeolog was prevalent in the embryo, whereas a more balanced pattern was seen in the other tissues (Fig. 6B). CaFRL-2 differential expression had a bias toward a CaCe homeolog in all the analyzed tissues (Fig. 6B). In contrast, CaFRL-3 did not present an expression bias, a very different pattern than that of CaFRL-4, for which CaCc homeolog was expressed more than the CaCe homeolog (Fig. 6B). The evaluation of CaFRL-5 HDE in the fruits revealed an intertwined profile, with some bias toward the CaCe homeolog (Fig. 6B).
Gene expression analysis of CaFRL genes and CaFLC gene during C. arabica fruit development (A) Heat map visualization of CaFRL expression in fruits at different stages of fruit development. The sum of relative homeolog expressions was used as numerical input for creating the heat map scale, from light green (weakly expressed) to red (strongly expressed). ‘60 daf pe’ sample was used as internal calibrator (B) Expression profiles of homeologous genes (CaCc and CaCe) of CaFRL family in fruits at different tissues. Perisperm (pe), embryo (eb), endosperm (end) and pericarp (po) and stages of ripening (60–240 daf, days after flowering). Values of three technical replicates are presented as mean ± SD (error bars). Transcript abundances were normalized using the expression of UBI (ubiquitin) as reference gene. Asterisks indicate significant differences (P < 0.05) between homeologous genes.
Coffea arabica FRL expression during direct somatic embryogenesis
The evidence that CaFRLs were expressed in C. arabica embryos during fruit development prompted a hypothesis that these genes could also be expressed in “artificial” in vitro direct somatic embryogenesis (DSE). Detailed histological analyses of C. arabica DSE were used for evaluating the origin of early tissue embryogenesis and establish the most appropriate timing for harvesting an embryo (Fig. 4C). Eight days after inoculation, rapid cell division begins in the mesophyll, particularly in the spongy parenchyma cells (Fig. 4C). Such division intensifies 60 d after inoculation, as the first evidence of proembryogenic mass (PM) development (Fig. 4C). At this stage, mesophyll cells show an evident nucleus, dense cytoplasm, and small intercellular space. Sixty days after explant inoculation, PM appears with meristematic cells (MC) along its border. From this moment, during the different stages of development, different morphologies of the embryos (e.g., globular, heart, torpedo) begin to form (Fig. 4C).
Using heat-map analyses, the CaFRL gene expression was evaluated throughout DSE (Fig. 7). In general, all five CaFRL genes increased their expression at the 60-day stage, had the highest expression at the globular-embryo stage, and maintained high transcriptional levels in all the other samples of embryo shapes (Fig. 7A). In terms of HDE, CaFRL-1 did not have an expression bias toward one subgenome (Fig. 7B). The CaCe subgenome homeolog from CaFRL-2 appeared to be slightly more expressed than the CaCc homeolog (Fig. 7B). In the globular, heart, and torpedo stages, the C. eugenioides CaFRL-3 homeolog was more expressed. The same pattern as observed for CaFRL-4 and CaFRL-5, with the C. eugenioides homeolog being preferentially expressed (Fig. 7B).
Gene expression analysis of CaFRL genes and CaFLC gene in C. arabica direct somatic embryogenesis (A) Heat map visualization of CaFRL expression in DSE at different stages. The sum of relative homeolog expressions was used as numerical input for creating the heat map scale from light green (weakly expressed) to red (strongly expressed). ‘8d’ sample was used as internal calibrator (B) Expression profiles of homeologous genes (CaCc and CaCe) of CaFRL family during DSE at different stages: 8 days (8d), 16 days (16d), 28 days (28d), 60 days (60d), globular embryos (gl), heart embryos (he) and torpedo embryos (to). Values of three technical replicates are presented as mean ± SD (error bars). Transcript abundances were normalized using the expression of UBI (ubiquitin) as reference gene. Asterisks indicate significant differences (P < 0.05) between homeologous genes.
Coffea arabica FLC gene expression is similar to FRL gene transcription
As mentioned above, FRI regulates FLC expression. To check whether C. arabica FLC follows C. arabica FRL genes, we examined its expression in flowers, fruits, and somatic embryos. Arabidopsis thaliana (FLC NP_196576.1) was used as bait for Blast analysis against C. arabica, C. eugenioides, and C. canephora genome databases. Sequences were retrieved and aligned, indicating that C. arabica FLC homeologous genes and their homeologs in C. eugenioides and C. canephora have very similar sequences (Supplementary Fig. S8, supplementary note). Primers designed for HDE failed to discriminate the CaFLC homeologs (data not shown). Using primers that aligned in both homeologous (full primer), FLC was more expressed in the floral white 1 stage (Figs 5A, 8), similar to the FRL genes. In fruits, FLC have prevalent expression in embryo and endosperm, mainly in the final stages of fruit development (Figs 6A, 8), also coinciding with FRL expression, especially that of CaFRL3 and CaFRL4. In DSE, CaFLC showed the highest expression in the 60-day stage and globular embryo stage (Figs 7A, 8).
Overview of CaFLC gene expression. From left to right (clockwise direction): flower, direct somatic embryogenesis (DSE), fruit perisperm (Pe), embryo (Eb), endosperm (end) and fruit pericarp (po) in stages of ripening (60–240 daf, days after flowering). Values of three technical replicates are presented as mean ± SD (error bars). Transcript abundances were normalized using the expression of UBI (ubiquitin) as reference gene.
Discussion
FRIGIDA-like proteins (FRLs) are required for regulating the flowering time in A. thaliana. In general, Arabidopsis accessions have two different flowering-time-related phenotypes. The first requires cold winters for flowering in spring (vernalization-responsive winter annuals), the second is a rapid-cycling summer annual. Differences in the expression of MADS-Box protein FLC, a key repressor of flowering and activator of vegetative development in Arabidopsis17,27, discriminate between the two phenotypes28. The FRI gene is known to increase the FLC RNA levels in winter-annual accessions, thereby delaying flowering until the FLC is silenced by vernalization18,29. In contrast, rapid-cycling accessions have low FLC levels because the FRI is inactive due to FRI allelic variation18. In addition, FRI forms the FRI-C complex with transcriptional activators FRIGIDA ESSENTIAL1 (FES1), FLC EXPRESSOR (FLX), and SUPPRESSOR OF FRI4 (SUF4)30. Moreover, the SWR complex, which acts as a chromatin remodeler to FLC, is recruited by FRI20.
Allelic sequence variation of FRI modulates the flowering time in A. thaliana, and FRI loss-of-function explains most of the variation in flowering time in early-flowering ecotypes. Nevertheless, in this study, we did not focus on the FRI allelic variation in C. arabica accessions, but on the homeolog variation in the species and the differential expression levels of these homeologs, particularly within reproduction-related organs. Such analyses evaluated the variation in FRL gene expression from an ancestry-spatiotemporal viewpoint instead of a population viewpoint, thus, connecting the FRL polymorphism between the C. arabica parental genomes (C. canephora and C. eugenioides) to developmental processes.
CaFRL homeolog sequence analysis
Five FRLs were found in the C. canephora genome, and their putative orthologs were identified in the genome of C. arabica and the RNAseq assembly of C. eugenioides (Table 1). Ten FRLs were found in C. arabica, which was in agreement with the hypothesis that this species is an allotetraploid that is most likely derived from the hybridization of the unreduced gametes from C. canephora (or a canephoroid species group) and C. eugenioides, both apparently containing five FRLs. Interestingly, all the homeologs were expressed in at least one of the conditions analyzed (see below). The C. arabica and C. canephora FRL orthologs showed structural differences in the genes, including differences in gene size, number of introns, and protein size (Table 1), which could be the result of recombination, transposon action, or other molecular events during the evolution of both species.
The number of FRLs found in C. canephora was lower than that found in A. thaliana (8 sequences), S. lycopersicum (12 sequences), S. tuberosum (11 sequences), V. vinifera (9 sequences), S. bicolor (10 sequences), and O. sativa (11 sequences). The presence of least number of FRLs in C. canephora (and likely C. eugenioides) might indicate either a gene family retraction (gene loss) in these species or an expansion of FRLs in the other annotated species. In fact, the second hypothesis appears to be more plausible because the phylogenetic tree (Fig. 2) indicates a series of paralogs in S. tuberosum, V. vinifera, O. sativa, and S. lycopersicum. Risk et al.22 showed that Solanaceae (tomato and potato) species lack AtFRL-1 homeologs, and Poaceae monocots rice and sorghum do not have genes homeologous to AtFRI (Fig. 2), suggesting that the FRL sequence identity, together with the FRL gene family width, might be important for species-, family-, or even clade-specific developmental processes (i.e., flowering and embryogenesis) that could respond to diverse environmental adaptations.
Risk et al.22 classified FRIGIDA-like genes based on evident differences on the N-terminus of A. thaliana genes containing an FRI-like domain. According to the authors, the contribution of the AtFRI N-terminus appears to be limited to promoting FLC expression, whereas the C-terminus is necessary for protein-protein interactions and the promotion of consecutive FLC transcription. Interestingly, both CaFRLs homeologous to AtFRI (CaFRL-3.1 and CaFRL-3.2) contain a C-terminus extension compared with the A. thaliana gene (Fig. 1).
CaFRLs display homeologous differential expression
It was possible to discriminate homeologous genes based on the alignment among C. canephora, C. eugenioides, and C. arabica FRL sequences. This inference based on sequence alignment was confirmed by expression analysis using Taq-MAMA primer design (Fig. S3) on the leaves of the three species, which showed that CaCc FRLs were expressed only in C. canephora and C. arabica, and that CaCe FRLs were expressed only in C. eugenioides and C. arabica (Fig. 3). In addition, these results confirmed the effectiveness of this alignment-based strategy.
In general, both homeologs from each CaFRL gene were expressed under at least one condition in our analyses (Figs 5–7); therefore, we could not detect gene silencing in C. arabica FLRs. Instead, these results indicated a more sophisticated regulation of gene expression. This result differs from those of homeolog analyses in other allopolyploid species such as cotton, for which the genes from one subgenome have been silenced or lost during the evolution of polyploidy31,32. When two or more different genomes are combined within a single cell, they must respond to the consequences of genome duplication, especially with respect to duplicate copies of genes with similar or redundant functions33. There are some possibilities for the regulation of homeologous genes in polyploids, such as (i) retention of original or similar function for the new homeologs, (ii) functional diversification of one of the homeologs, or (iii) silencing of one of these genes34. However, homeologous genes could also exhibit unequal expression patterns (i.e., levels of ancestral dominance)35, and might vary according to different types of stress8,36 and among different organs13, as case described here. The differential expressions of these homeologs, which implicitly present sequence differences, might result in myriad combinations of protein-protein interaction that could regulate a series of developmental processes.
The presence of cis-elements that were connected to an environmental response (i.e., heat stress, MEJA and gibberellin response, light response; Supplementary note) is in accordance with the idea that FRI genes are a part of the bridge that connects environmental conditions to development. Nevertheless, there is no direct connection between cis-element presence/absence and gene expression of the homeologs, because most differential homeologous cis-elements are present in CaCe FRL promoters and genes from CaCc are expressed (Figs 5–7). With a more specific set of genes, the same entangled gene expression regulation described by previous authors might occur with CaFRLs, most likely with trans-factors from one subgenome acting in the other subgenome, or by epigenetic factors such as histone modification, DNA methylation, or regulatory RNAs. It is not surprising that FRLs could be epigenetically modulated, given that several genes involved in flowering and embryogenesis exhibit this kind of regulation37,38. One of the most interesting expression profiles of HDE that suggests trans-action was from CaFRL-4. During flower and fruit development, CaCc homeolog CaFRL-4.1 was notably the most expressed (Figs 5, 6); however, during DSE, the expression profile changed completely with the CaCe homeolog CaFRL-4.2 being more expressed (Fig. 7). One possibility is that MS medium used for DSE contains molecules that could activate the transcription of CaCe homeolog instead of CaCc. This is an example of puzzling homeologous gene regulation, which appears to rely on specific trans-factors from a tissue or developmental process (i.e., somatic embryogenesis vs zygotic embryogenesis; see below).
CaFRLs might exert functions in late flower development
Flowering in Coffea plants usually occurs after a period of drought, when the onset of rain triggers flowering and anthesis. Flowering time in Coffea is a complex feature that is partially dependent on environmental factors, such as photoperiod and vernalization, but also on rain39. These external signals modulate a regulatory network to prevent the plant from blossoming too soon or too late in the season. In Coffea, these signals include drought, which triggers the reproductive differentiation of vegetative buds, and a rainy season, which allows flower and fruit development39.
A detailed morphological analysis of the C. arabica flowering mechanism had been provided by de Oliveira et al.40, indicating that, together with environmental cues, floral meristem ontogenesis is also an important factor that affects asynchronous flowering events. The same authors assessed MADS-box expression along floral development and discovered important differences between the spatiotemporal expression of classical Arabidopsis MADS-box and their orthologs in C. arabica40. In this sense, MADS-box sub- or neo-functionalization could be the cause of morphological idiosyncrasies in C. arabica flower development, such as mucilage secretion and formation of epipetalous stamens. In addition, the authors pointed out that innovative spatiotemporal coexpression of MADS-box (i.e., FLC) with its partners (i.e., FRI) might be related to these new functions.
Choi et al.21 reported that Arabidopsis FLC and FRI are expressed in flower buds/meristems in open flowers, and more specifically in ovules of nonvernalized plants, indicating that these genes are involved in female gametogenesis. We also found that C. arabica FLC is expressed in flowers. Barreto et al.41 detected FLC expression in organs exposed to abiotic and biotic stresses, We identified a quite similar expression pattern across CaFRLs, with high transcription at the white 1 stage and lower transcription during the later stages, except for CaFRL-3 (AtFRI ortholog), the expression of which increased later during anthesis (Fig. 5A). Despite its putative importance in ovule development, it was hypothesized that during flower development, FRI might activate FLC to act as a repressor of SOC1, thus, stimulating the SEP3 expression, and consequently, final floral organ development42.
CaFRLs appear to be involved in embryogenesis and endosperm development
Choi et al.21 provided a comprehensive analysis of FLC and FLC regulator expression during reproductive development, including fruit development and embryogenesis. As mentioned above, the authors found that FLC was expressed in open flowers. Furthermore, the gene is transcribed in nonvernalized ovules, but not in pollen or vernalized ovules21. Nevertheless, the FLC expression is reactivated after fertilization in embryos but not in the endosperm. FRI is expressed in ovules, independent of vernalization, but not in the pollen. The gene is then reactivated in embryos following the FLC expression pattern21. CaFLCs have prevalent expression in embryo, similar to the A. thaliana FLC gene21. In our analysis, all CaFRLs were expressed during fruit development, although each one displaying a different expression profile (Fig. 6). Overall, the genes were expressed in the perisperm, endosperm, and embryo in diverse profiles, with much lower expression in the pericarp. One outstanding difference between AtFRI and CaFRL-3 is that, although the former is conspicuously expressed only in embryos21, the latter is also expressed in endosperms and at a much higher level than in embryos (Fig. 6). An inspection of the expression of other A. thaliana FRLs could reveal expression patterns similar to those found in C. arabica FRLs, possibly pointing out that some AtFRIs are expressed in endosperm; however, differences in fruit tissue ontogenesis between A. thaliana and C. arabica can explain the discrepancy in our data.
After fecundation, C. arabica fruit contains mainly the pericarp, which is composed of the exocarp (peel), mesocarp, and endocarp, as well as perisperm, which develops from the nucleus of the ovule soon after the fertilization43. Perisperm is an aqueous tissue with intense cell division and expansion. At approximately 100 DAF, perisperm is progressively replaced by triploid endosperm44. As storage tissue, mature endosperm accumulates nutrients that are mobilized by the embryo during seed germination. The evidence that CaFRLs have an increased expression during the final stages of fruit development suggests that these genes could be engaged in the regulation of the physiological deposition and storage of endosperm compounds, which are quantitatively and qualitatively responsible for coffee beverage quality45.
CaFRLs are expressed during somatic embryogenesis
One outstanding result in our data was the expression of FRL genes, especially that of CaFRL-3, CaFRL-4, and CaFRL-5, during C. arabica DSE, which is responsible for the formation of somatic embryos or embryogenic tissue directly from the explant without the development of an intermediate callus phase46. Interestingly, CaFRLs were expressed in the initial stages of embryo development during DSE, suggesting its participation in embryo maturation. Another important result was that CaFLC was also expressed during DSE, strongly suggesting that FRL genes can trigger FLC expression in artificial embryogenesis. Another MADS-box gene, AGL15, was found to play an essential role in somatic embryogenesis in both soybean and Arabidopsis47. Somatic embryogenesis (SE) is an interesting process during which plants regenerate a new plant from a single cell or a group of somatic cells44. Many studies have investigated the relationship between SE and zygotic embryogenesis (ZE). Nic-Can et al.48 studied SE in C. canephora and found that the genes involved in zygotic embryogenesis—LEC1, BABY BOOM1, and WOX4—are expressed during SE development in this plant. The fact that FRLs, FLC, and the ZE-related genes mentioned above are expressed in both ZE and SE clearly indicates that both embryogenesis processes share common developmental pathways, and thus, suggests that FRLs and FLC are embryogenesis-related genes.
By evaluating FRL gene expression in reproduction-related organs/tissues, we confirmed previous genome-wide homeologous gene expression pattern that indicated intertwined regulation of C. arabica homeologs. Furthermore, we found that FRL genes are expressed in C. arabica late flowering stages, endosperm, and embryo during ZE, and most importantly, during SE. Our study provides insights for the study of FRL genes, with a new perspective of FRIGIDA gene action in allopolyploids.
Methods
Biological material
Leaf samples were collected from C. arabica (Catuaí Amarelo IAC62), C. canephora and C. eugenioides from the germplasm of IAC (Campinas Agronomic Institute) located in Campinas, São Paulo, 54′21″S/47°03′39″W). Flowers and fruits were collected from C. arabica (Catuaí amarelo IAC62). The collection of the flowers was carried out in September 2016 according to the development of the bud flowers. The fruits were collected monthly from November 2015 to May 2016, following 60 days after flowering (DAF), 90 DAF, 120 DAF, 150 DAF, 180 DAF, 210 DAF and 240 DAF. The samples were collected in biological triplicates (Plants L7P9, L7P14 and L8P7), each plant consisting in a replica. The tissues perisperm, endosperm, pericarp and embryo were separated, frozen in liquid nitrogen and stored in a freezer at −80 °C. Direct Somatic Embryogenesis (DSE) was performed according to the methodology described by Ramos et al.49, using C. arabica leaves (Catuaí Amarelo IAC 62) as primary explant. Briefly,leaves were cut in laminar flow cabinet, removing the midrib and edges, obtaining explants of 1 cm2, which were inoculated with the adaxial side in contact with the culture medium, then kept in dark in a temperature of 25 °C ± 2 °C. For DSE, Murashige-Skoog (MS) medium was used with half the concentration of macronutrients and micronutrients, added with 20 g L-1 sucrose and 10 µM of isopentenyl adenine (2 iP). Samples were collected from the moment of inoculation (day 0) and throughout embryogenesis (8, 16, 28 and 60 days) until the shapes of the developing embryos at each stage could be detected (e.g., globular, heart, torpedo).
Morphoanatomical analyses
Morphological analyses were performed from embryos obtained by Direct Somatic embryogenesis (DSE). Tissues were maintained on MS half medium, collected at the time of in vitro inoculation (0 days) and at different stages of development of the somatic embryos (8, 16, 28, 60 days after inoculation in the culture medium, globular embryo, heart embryo and torpedo embryo). For the anatomical analysis, the samples were fixed in FAA 50 solution (formaldehyde, acetic acid and ethanol 50%, 5: 5: 90), dehydrated in ethanol series and infiltrated in plastic resin (Leica Historesin®) according to the manufacturer’s instructions. The samples were sectioned using a manual rotary microtome (Leica®) with type C razor, in the thickness of 5 μm. Sections were stained with 0.05% toluidine blue in phosphate and citrate buffer pH 4.5 and mounted on “Entellan®” synthetic resin (Merck®). Documentation of results was performed by capturing images using the Olympus DP71 camcorder coupled to the Olympus BX 51 microscope.
Genomic data and in silico analyses
Single-nucleotide polymorphism (SNP)–based detection of homeologous genes in C. arabica was previously described by Vidal et al.3. Biefly, the authors have used the alignment of EST sequences from C. canephora and C. arabica to infer that the sequences in C. arabica that have a SNP pattern similar to those in C. canephora originated from the CaCc subgenome, and that the ones that did not have a similar pattern were from the CaCe subgenome. These inferences were confirmed by polymerase chain reaction (PCR) using the ancestors’ DNA. Based on the expression levels, determined by counting the number of reads per tissue in each homeologous haplotype, the authors could assign genes that could hypothetically display homeologous gene expression3.
Identification of orthologs of the FRIGIDA gene family was performed using eight Arabidopsis thaliana FLPs as baits in BlastP searches. Their orthologs in Coffea canephora, Solanum lycopersicum, Solanum tuberosum, Vitis vinifera, Sorghum bicolor and Oryza sativa were identified in the following databases: Coffee Genome Hub50, NCBI (http://www.ncbi.nlm.nih.gov), TAIR (http://www.arabidopsis.org), AtGDB (http://www.plantgdb.org/AtGDB) Phytozome (http://www.phvtozome.net), Sol Genomics Network (http://solgenomics.net), SIGDB (http://www.plantgdb.org/SIGDB), Grape Genome Database (http://www.genoscope.cns.fr/externe), Gramene Database (http://www.gramene.org), and Rice Genome Annotation (http://rice.plantbiology.msu.edu). A second search was performed to identify the orthologs of the selected genes in C. arabica and C. eugenioides. The complete transcribed sequences (CDS) of the FRL genes of C. canephora were used as search queries in UC Davis C. arabica sequencing initiative (https://phytozome.jgi.doe.gov/pz/portal.html#!info?alias=Org_Carabica_er) and in the RNAseq reads of leaves and fruits of C. eugenioides25 (SRA sequence read alignment; https://www.ncbi.nlm.nih.gov/sra). Alignments were performed using the CLUSTALW tool and edited in the GeneDoc program (http://www.nrbsc.org/gfx/genedocA). Genes that did not contain specific domains were removed. Phylogenetic analysis was performed using the MEGA software51. The search for cis regulatory elements was performed using PlantCare platform (http://bioinformatics.psb.ugent.be/webtools/plantcare/html).
RNA extraction and real-time qPCR assays
RNA was extracted using the Concert™ Plant RNA Purification Reagent (Invitrogen). RNA (1 µg) was previously treated with 1 U/µL DNAseI (Invitrogen). cDNA samples were synthesized according to according SuperScript® III Reverse Transcriptase kit protocol (Invitrogen) and used for qPCR reaction. For each reaction, 1 μl of the appropriate cDNA dilutions, 0.2 μL of the primer forward, 0.2 μL of the reverse primer at 10 mM each and 5 μL of Platinum® SYBR® Green qPCR SuperMix-UDG with ROX (Invitrogen). The reaction was supplemented with 3.6 μL Milli-Q water to a final volume of 10 μL per reaction. For each condition, the same reaction was performed three times to overlap and confirm the results in the apparatus. The data were analyzed in the program 7500 Fast Software (software v2.1.1). The samples were processed in triplicates, always accompanied by the negative controls (NTC: “in the template control”) that did not contain cDNA. The negative control in the reactions is used to verify the absence of exogenous cDNA contamination in the SYBR, primers or water mixtures. Gene expression levels were normalized to expression level of ubiquitin (UBQ10) as a constitutive reference52. Expression was expressed as relative quantification by applying the formula (1 + E) − ΔCt, where ΔCtartget = Cttarget gene − Ctference gene, as previously described53. Relative expression was The LinReg software54 was used to calculate the efficiency of each pair of primers per reaction. The statistical analyses (ANOVA and Tukey tests) were performed using STATISTICA software (StatSoft). The expression data was formatted by R3.4.3 software for representation. The sum of relative homeolog expressions was used as numerical input for creating the heat map. Primers were designed according to qPCR TaqMAMA method3,26.
References
Cenci, A., Combes, M. C. & Lashermes, P. Genome evolution in diploid and tetraploid Coffea species as revealed by comparative analysis of orthologous genome segments. Plant Mol. Biol. 78, 135–145 (2012).
Lashermes, P. et al. Molecular characterisation and origin of the Coffea arabica L. genome. Mol. Gen. Genet. 261, 259–266 (1999).
Vidal, R. O. et al. A high-throughput data mining of single nucleotide polymorphisms in Coffea species expressed Sequence tags suggests Differential Homeologous Gene Expression in the Allotetraploid Coffea arabica. Plant Physiol. 154, 1053–1066 (2010).
Maurin, O. et al. Towards a phylogeny for Coffea (Rubiaceae): Identifying well-supported lineages based on nuclear and plastid DNA sequences. Ann. Bot. 100, 1565–1583 (2007).
Leroy, T. et al. Construction and charcterization of a Coffea canephora BAC library to study the organization of sucrose biosynthesis genes. Theor. Appl. Genet. 111, 1032–1041 (2005).
Bardil, A., de Almeida, J. D., Combes, M. C., Lashermes, P. & Bertrand, B. Genomic expression dominance in the natural allopolyploid Coffea arabica is massively affected by growth temperature. New Phytol. 192, 760–774 (2011).
Combes, M. C., Cenci, A., Baraille, H., Bertrand, B. & Lashermes, P. Homeologous gene expression in response to growing temperature in a recent allopolyploid (Coffea arabica L.). J. Hered. 103, 36–46 (2012).
Combes, M. C., Dereeper, A., Severac, D., Bertrand, B. & Lashermes, P. Contribution of subgenomes to the transcriptome and their intertwined regulation in the allopolyploid Coffea arabica grown at contrasted temperatures. New Phytol. 200, 251–260 (2013).
Cotta, M. G. et al. Lipid transfer proteins in coffee: isolation of Coffea orthologs, Coffea arabica homeologs, expression during coffee fruit development and promoter analysis in transgenic tobacco plants. Plant Mol. Biol. 85, 11–31 (2014).
de Carvalho, K. et al. Homeologous genes involved in mannitol synthesis reveal unequal contributions in response to abiotic stress in Coffea arabica. Mol. Genet. Genomics 289, 951–963 (2014).
Lashermes, P., Hueber, Y., Combes, M. C., Severac, D. & Dereeper, A. Inter-genomic DNA exchanges and homeologous gene silencing shaped the nascent allopolyploid coffee genome (Coffea arabica L.). G3 (Bethesda). 6, 2937–2948 (2016).
Marraccini, P. et al. RBCS1 expression in coffee: Coffea orthologs, Coffea arabica homeologs, and expression variability between genotypes and under drought stress. BMC Plant Biology 11, 85 (2011).
Adams, K. L., Cronn, R., Percifield, R. & Wendel, J. F. Genes duplicated by polyploidy show unequal contributions to the transcriptome and organ-specific reciprocal silencing. Proc. Natl. Acad. Sci. USA 100, 4649–4654 (2003).
Comai, L. The advantages and disadvantages of being polyploid. Nat. Rev. Genet. 6, 836–846 (2005).
Jackson, S. & Chen, Z. J. Genomic and expression plasticity of polyploidy. Curr. Opin. Plant Biol. 13, 153–159 (2010).
Soltis, D. E., Visger, C. J. & Soltis, P. S. The polyploidy revolution then…and now: Stebbins revisited. Am. J. Bot. 101, 1057–1078 (2014).
Michaels, S. D. & Amasino, R. M. FLOWERING LOCUS C encodes a novel MADS domain protein that acts as a repressor of flowering. Plant Cell 11, 949–956 (1999).
Johanson, U. et al. Molecular analysis of FRIGIDA, a major determinant of natural variation in Arabidopsis flowering time. Science. 290, 344–347 (2000).
Wang, J., Tian, L., Lee, H. S. & Chen, Z. J. Nonadditive regulation of FRI and FLC loci mediates flowering-time variation in Arabidopsis allopolyploids. Genetics. 173, 965–974 (2006).
Choi, K. et al. The FRIGIDA complex activates transcription of FLC, a strong flowering repressor in Arabidopsis, by recruiting chromatin modification factors. Plant Cell 23, 289–303 (2011).
Choi, J. et al. Resetting and regulation of Flowering Locus C expression during Arabidopsis reproductive development. Plant J. 57, 918–931 (2009).
Risk, J. M., Laurie, R. E., Macknight, R. C. & Day, C. L. FRIGIDA and related proteins have a conserved central domain and family specific N- and C- terminal regions that are functionally important. Plant Mol. Biol. 73, 493–505 (2010).
Denoeud, F. et al. The coffee genome provides insight into the convergent evolution of caffeine biosynthesis. Science 345, 1181–1184 (2014).
Mondego, J. M. C. et al. An EST-based analysis identifies new genes and reveals distinctive gene expression features of Coffea arabica and Coffea canephora. BMC Plant Biology 11, 30 (2011).
Yuyama, P. M. et al. Transcriptome analysis in Coffea eugenioides, an Arabica coffee ancestor, reveals differentially expressed genes in leaves and fruits. Mol. Genet. Genomics. 291, 323–336 (2016).
Li, B., Kadura, I., Fu, D. J. & Watson, D. E. Genotyping with TaqMAMA. Genomics 83, 311–320 (2004).
Li, D. et al. A repressor complex governs the integration of flowering signals in Arabidopsis. Dev. Cell 15, 110–120 (2008).
Gazzani, S., Gendall, A. R., Lister, C. & Dean, C. Analysis of the molecular basis of flowering time variation in Arabidopsis accessions. Plant Physiol. 132, 1107–1114 (2003).
Shindo, C. et al. Role of FRIGIDA and FLOWERING LOCUS C in determining variation in flowering time of Arabidopsis. Plant Physiol. 138, 1163–1173 (2005).
Andersson, C. R. et al. The FLX gene of Arabidopsis is required for FRI-dependent activation of FLC expression. Plant Cell Physiol. 49, 191–200 (2008).
Adams, K. L., Percifield, R. & Wendel, J. F. Organ-specific silencing of duplicated genes in a newly synthesized cotton allotetraploid. Genetics 168, 2217–2226 (2004).
Flagel, L. E., Wendel, J. F. & Udall, J. A. Duplicate gene evolution, homoeologous recombination, and transcriptome characterization in allopolyploid cotton. BMC Genomics. 13, 302 (2012).
Flagel, L. E. & Wendel, J. F. Gene duplication and evolutionary novelty in plants. New Phytol. 183, 557–564 (2009).
Wendel, J. F., Lisch, D., Hu, G. & Mason, A. S. The long and short of doubling down: polyploidy, epigenetics, and the temporal dynamics of genome fractionation. Curr. Opin. Genet. Dev. 49, 1–7 (2018).
Grover, C. E. et al. Homoeolog expression bias and expression level dominance in allopolyploids. New Phytol. 196, 966–971 (2012).
Liu, Z. & Adams, K. L. Expression partitioning between genes duplicated by polyploidy under abiotic stress and during organ development. Curr. Biol. 17, 1669–1674 (2007).
Sung, S. & Amasino, R. M. Vernalization and epigenetics: how plants remember winter. Curr. Opin. Plant Biol. 7, 4–10 (2004).
De-la-Peña, C., Nic-Can, G. I., Galaz-Ávalos, R. M., Avilez-Montalvo, R. & Loyola-Vargas, V. M. The role of chromatin modifications in somatic embryogenesis in plants. Front. Plant Sci. 6, 635 (2015).
Majerovicz, N. & Söndahl, M. R. Induction and differentiation of reproductive buds in Coffea arabica L. Braz. J. Plant Physiol. 17, 247–254 (2005).
de Oliveira, R. R., Cesarino, I., Mazzafera, P. & Dornelas, M. C. Flower development in Coffea arabica L.: new insights into MADS-box genes. Plant Reprod. 27, 79–94 (2014).
Barreto, H. G. et al. In Silico and quantitative analyses of the putative FLC-like homologue in coffee (Coffea arabica L.). Plant Mol. Biol. Rep. 30, 29–35 (2012).
Lee, J. & Lee, I. Regulation and function of SOC1, a flowering pathway integrator. J. Exp. Bot. 61, 2247–2254 (2010).
de Castro, R. D. & Marraccini, P. Cytology, biochemistry and molecular changes during coffee fruit development. Brazil. J. Plant Physiol. 18, 175–199 (2006).
Zimmerman, J. L. Somatic embryogenesis: A model for early development in higher plants. Plant Cell. 5, 1411–1423 (1993).
Privat, I. et al. Differential regulation of grain sucrose accumulation and metabolism in Coffea arabica (Arabica) and Coffea canephora (Robusta) revealed through gene expression and enzyme activity analysis. New Phytol. 178, 781–797 (2008).
Yang, X. & Zhang, X. Regulation of somatic embryogenesis in higher plants. Crit. Rev. Plant Sci. 29, 36–57 (2010).
Thakare, D. et al. The MADS-domain transcriptional regulator AGAMOUS-LIKE 15 promotes somatic embryo development in Arabidopsis and soybean. Plant Physiol. 146, 1663–1672 (2008).
Nic-Can, G. I. et al. New insights into somatic embryogenesis: leafy cotyledon1, baby boom1 and WUSCHEL-related homeobox 4 are epigenetically regulated in Coffea canephora. PLoS One. 8, e72160 (2013).
Ramos, L., Yokoo, E. & Gonçalves, W. Direct somatic embryogenesis is genotype specific in coffee. In: COLLOQUE Scientifique International sur le Café (ASIC), 763–766 (1993).
Dereeper, A. et al. The coffee genome hub: a resource for coffee genomes. Nucleic Acids Res. 43(Database issue), D1028–35 (2015).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Barsalobres-Cavallari, C. F., Severino, F. E., Maluf, M. P. & Maia, I. G. Identification of suitable internal control genes for expression studies in Coffea arabica under different experimental conditions. BMC Mol. Biol. 10, 1 (2009).
Bertrand, B. et al. The greater phenotypic homeostasis of the allopolyploid Coffea arabica improved the transcriptional homeostasis over that of both diploid parents. Plant Cell Physiol. 56, 2035–2051 (2015).
Ramakers, C., Ruijter, J. M., Deprez, R. H. & Moorman, A. F. Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci. Lett. 339, 62–66 (2003).
Acknowledgements
We thank Dr. Carlos Colombo (IAC) for logistics support and Centro de Café Alcides Carvalho (IAC) for maintenance of coffee germplasm used in this study. This work was supported by São Paulo State Research Foundation (FAPESP; 2013/17544-3).
Author information
Authors and Affiliations
Contributions
N.G.V. performed and conceived all experiments and wrote the paper. I.F.F. and J.L.S.M. performed somatic embryogenesis. J.C.R. help in embryo extraction and in fruit development evaluation. J.M.C.M. conceived the experiments and wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vieira, N.G., Ferrari, I.F., Rezende, J.C.d. et al. Homeologous regulation of Frigida-like genes provides insights on reproductive development and somatic embryogenesis in the allotetraploid Coffea arabica. Sci Rep 9, 8446 (2019). https://doi.org/10.1038/s41598-019-44666-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-44666-6
- Springer Nature Limited
This article is cited by
-
Genetic and epigenetic modes of the regulation of somatic embryogenesis: a review
Biologia Futura (2022)
-
Global identification of long non-coding RNAs involved in the induction of spinach flowering
BMC Genomics (2021)
-
Leaf transcriptomic signatures for somatic embryogenesis potential of Elaeis guineensis
Plant Cell Reports (2021)
-
Genome-wide identification and expression analysis of the frigida domain gene family in Prunus mume (Prunus mume Sieb. et Zucc.)
Horticulture, Environment, and Biotechnology (2021)