Abstract
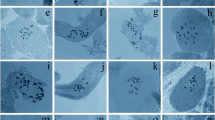
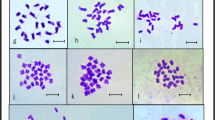
The present investigation systematically examined the distribution of 5S and 45S rDNA sites in six distinct Fritillaria species, all characterized as diploid with 2n = 2x = 24 chromosomes. Fritillaria assyriaca ecotypes displayed variable numbers of B chromosomes (Bs), ranging from one to four, while F. zagrica exhibited two additional B chromosomes. Terminal or sub-terminal chromosomal regions harbored one to two pairs of 5S rDNA sites. Regarding 45S rDNA sites, F. raddeana featured a singular pair, F. assyriaca, F. zagrica, and F. persica exhibited three pairs, F. avromanica presented four pairs, and F. chlorantha displayed eight pairs, predominantly situated distally, except for F. raddeana and F. chlorantha, which demonstrated interstitial and sub-terminal locations, respectively. Chromosome and karyotype indices facilitated the identification of, F. avromanica and F. chlorantha as species with the most symmetrical and asymmetrical chromosomes, respectively. Cluster analysis of the karyotype similarity matrix revealed incongruities between the observed number and distribution of rDNA sites and the established taxonomic classifications, particularly notable in F. chlorantha within the Fritillaria subgenus. The results provide significant insights into the genetic diversity and karyotype characteristics of Fritillaria, challenging conventional taxonomic frameworks. The observed variations in the numbers and locations of rDNA sites underscore the necessity for a nuanced understanding of genetic relationships within the Fritillaria subgenus.


Similar content being viewed by others
Data Availability
All chromosome slides is available in faculty of agriculture, cytogenetic lab. The datasets generated and analyzed during the current study is available.
References
Abbo S, Miller T, Reader S, Dunford R, King I (1994) Detection of ribosomal DNA sites in lentil and chickpea by fluorescent in situ hybridization. Genome 37(4):713–716
Adachi J, Watanabe K, Fukui K, Ohmido N, Kosuge K (1997) Chromosomal location and reorganization of the 45S and 5S rDNA in theBrachyscome lineariloba complex (Asteraceae). J Plant Res 110(3):371–377
Ahmadi-Roshan M, Karimzadeh G, Babaei A, Jafari H (2016) Karyological studies of Fritillaria (Liliaceae) species from Iran. Cytologia 81(2):133–141
Ambrožová K, Manda´kova T, Buresˇ P, Neumann P, Leitch IJ, Koblížková A et al (2011) Diverse retrotransposon families and an AT-rich satellite DNA revealed in giant genomes of Fritillaria lilies. Ann Botany 107(2):255–268
Arano H (1963) Cytological studied in subfamily Carduoideae (Compositae) of Japan IX. Botanic Magazine (Tokyo) 76:32–39
Badaeva E, Dedkova O, Gay G, Pukhalskyi V, Zelenin A, Bernard S et al (2007) Chromosomal rearrangements in wheat: their types and distribution. Genome 50(10):907–926
Bennett MD, Leitch IJ (2011) Nuclear DNA amounts in angiosperms: targets, trends and tomorrow. Ann Botany 107(3):467–590
Bennett MD, Smith J (1976) Nuclear DNA amounts in angiosperms. Philosophical Trans Royal Soc Lond B Biol Sci 274(933):227–274
Bharathan G, Lambert G, Galbraith D (1994) Nuclear DNA content of monocotyledons and related taxa. Am J Bot 81(3):381–386
Bogunić F, Siljak-Yakovlev S, Muratović E, Ballian D (2011) Different karyotype patterns among allopatric Pinus nigra (Pinaceae) populations revealed by molecular cytogenetics. Plant Biol 13(1):194–200
Chacón J, Sousa A, Baeza CM, Renner SS (2012) Ribosomal DNA distribution and a genus-wide phylogeny reveal patterns of chromosomal evolution in Alstroemeria (Alstroemeriaceae). Am J Bot 99(9):1501–1512
Chen J-F, Staub JE, Adelberg JW, Jiang J (1999) Physical mapping of 45S rRNA genes in Cucumis species by fluorescence in situ hybridization. Can J Bot 77(3):389–393
Chung M-C, Lee Y-I, Cheng Y-Y, Chou Y-J, Lu C-F (2008) Chromosomal polymorphism of ribosomal genes in the genus Oryza. Theor Appl Genet 116(6):745–753
Day PD, Berger M, Hill L, Fay MF, Leitch AR, Leitch IJ et al (2014) Evolutionary relationships in the medicinally important genus Fritillaria L.(Liliaceae). Mol Phylogenet Evol 80:11–19
Drouin G, De Sa MM (1995) The concerted evolution of 5S ribosomal genes linked to the repeat units of other multigene families. Mol Biol Evol 12(3):481–493
Dubcovsky J, Dvorák J (1995) Ribosomal RNA multigene loci: nomads of the Triticeae genomes. Genetics 140(4):1367–1377
Fedorov A (1969) Chromosome numbers of flowering plants Acad. Sci USSR, Leningrad (now St Petersburg)
Gerlach WL, Bedbrook J (1979) Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acids Res 7(7):1869–1885
Gerlach WL, Dyer TA (1980) Sequence organization of the repeating units in the nucleus of wheat which contain 5S rRNA genes. Nucleic Acids Res 8(21):4851–4865
Hamon P, Siljak-Yakovlev S, Srisuwan S, Robin O, Poncet Vr, Hamon S et al (2009) Physical mapping of rDNA and heterochromatin in chromosomes of 16 Coffea species: a revised view of species differentiation. Chromosome Res 17(3):291–304
Han Y, Zhang Z, Liu J, Lu J, Huang S, Jin W (2008) Distribution of the tandem repeat sequences and karyotyping in cucumber (Cucumis sativus L.) by fluorescence in situ hybridization. Cytogenet Genome Res 122(1):80–88
Hanson RE, Islam-Faridi N, Percival M, Crane EA, Ji CF, McKnight Y, T. D., et al (1996) Distribution of 5S and 18Sâ€28S rDNA loci in a tetraploid cotton (Gossypium hirsutum L.) and its putative diploid ancestors. Chromosoma 105(1):55–61
Hazbavi F, Hosseini S, Mirzaghaderi G, Advay M (2019) Karyotypic Variation in five species of the Genus Fritillaria (Liliaceae). Iran J Bot 25(2):127–134
Heslop-Harrison J (2000) Comparative genome organization in plants: from sequence and markers to chromatin and chromosomes. Plant Cell 12(5):617–635
Huziwara Y (1962) Karyotype analysis in some genera of Compositae. VIII. Further studies on the chromosomes of Aster. Am J Bot 49:116–119
Jafari H, Babaei A, Karimzadeh G, Ahmadi-Roshan M (2014) Cytogenetic study on some Fritillaria species of Iran. Plant Syst Evol 300(6):1373–1383
Khaniki GB (1997) Fritillaria Atrolineata (Liliaceae), a new species from Iran. Edinb J Bot 54(2):171–181
Khaniki G (2002a) Chromosome number of all Iranian species of Fritillaria Caucasica group (Liliaceae). Nucleus 45(6–11):103–108
Khaniki G (2002b) Chromosome number of Fritillaria Subgenera Petilium and Theresia (Liliaceae). Nucleus 45(1–2):6–11
Khaniki G (2005) Giemsa C-banding studies on interphase nuclei of Iranian species of Fritillaria and Rhinopetalum (Liliaceae). Proc Natl Acad Sci India Sect B 75(4):294
Khaniki GB, Persson K (1997) Nectary morphology in south west Asian Fritillaria (Liliaceae). Nord J Bot 17(6):579–611
Khourang M, Babaei A, Sefidkon F, Naghavi MR, Asgari D, Potter D (2014) Phylogenetic relationship in Fritillaria spp. of Iran inferred from ribosomal ITS and chloroplast trnl-trnf sequence data. Biochem Syst Ecol 57:451–457
Knight CA, Ackerly DD (2002) Variation in nuclear DNA content across environmental gradients: a quantile regression analysis. Ecol Lett 5(1):66–76
Knight CA, Molinari NA, Petrov DA (2005) The large genome constraint hypothesis: evolution, ecology and phenotype. Ann Botany 95(1):177–190
Leitch I, Heslop-Harrison J (1992) Physical mapping of the 18S–5.8 S–26S rRNA genes in barley by in situ hybridization. Genome 35(6):1013–1018
Levan A, Fredga K, Sandberg AA (1964) Nomenclature for centromeric position on chromosomes. Hereditas 52(2):201–220
Linde-Laursen I, Ibsen E, Bothmer RV, Giese H (1992) Physical localization of active and inactive rRNA gene loci in Hordeum marinum spp. Gussoneanum (4 x) by in situ hybridization. Genome 35(6):1032–1036
Liu ZL, Zhang D, Wang XQ, Ma XF, Wang XR (2003) Intragenomic and interspecific 5S rDNA sequence variation in five Asian pines. Am J Bot 90(1):17–24
Lou Q, Iovene M, Spooner DM, Buell CR, Jiang J (2010) Evolution of chromosome 6 of Solanum species revealed by comparative fluorescence in situ hybridization mapping. Chromosoma 119(4):435–442
Mantovani M, Abel S, Moreira-Filho O (2005) Conserved 5S and variable 45S rDNA chromosomal localisation revealed by FISH in Astyanax scabripinnis (Pisces, Characidae). Genetica 123(3):211–216
Mondin M, Aguiar-Perecin ML (2011) Heterochromatin patterns and ribosomal DNA loci distribution in diploid and polyploid Crotalaria species (Leguminosae, Papilionoideae), and inferences on karyotype evolution. Genome 54(9):718–726
Orosová M, Marec F, Oros M, Xi BW, Scholz T (2010) A chromosome study and localization of 18S rDNA in Khawia saurogobii (Cestoda: Caryophyllidea). Parasitol Res 106(3):587–593
Paszko B (2006) A critical review and a new proposal of karyotype asymmetry indices. Plant Syst Evol 258(1):39–48
Peruzzi L, Eroglu H (2013) Karyotype asymmetry: again, how to measure and what to measure? Comp Cytogenet 7(1):1–9
Peruzzi L, Leitch I, Caparelli K (2008) Chromosome diversity and evolution in Liliaceae. Ann Botany 103(3):459–475
Puttick MN, Clark J (1820) Donoghue PC (2015) Size is not everything: rates of genome size evolution, not C-value, correlate with speciation in angiosperms. Proc R Soc B: Biol Sci 282:20152289
Raina S, Mukai Y (1999) Detection of a variable number of 18S-5.8 S-26S and 5S ribosomal DNA loci by fluorescent in situ hybridization in diploid and tetraploid Arachis species. Genome 42(1):52–59
Rechinger KH, Browicz K, Persson K, Wendelbo P (1990) Liliaceae II. In: Rechinger KH (ed) Flora Iranica, vol 165. Graz Akademische Druk und Verlagsanstalt
Rix E (1977) Fritillaria in Iran. Iran J Bot 1:75–95
Rix E (2001) Fritillaria. A Revised Classification. The Fritillaria Group of the Alpine Garden Society. United Kingdom
Roa F, Guerra M (2012) Distribution of 45S rDNA sites in chromosomes of plants: structural and evolutionary implications. BMC Evol Biol 12(1):1–13
Robledo G, Lavia G, Seijo G (2009) Species relations among wild Arachis species with the A genome as revealed by FISH mapping of rDNA loci and heterochromatin detection. Theor Appl Genet 118(7):1295–1307
Ronsted N, Law S, Thornton H, Fay MF, Chase MW (2005) Molecular phylogenetic evidence for the monophyly of Fritillaria and Lilium (Liliaceae; Liliales) and the infrageneric classification of Fritillaria. Mol Phylogenet Evol 35(3):509–527
Rosato M, Castro M, Rosselló JA (2008) Relationships of the Woody Medicago species (section Dendrotelis) assessed by molecular cytogenetic analyses. Ann Botany 102(1):15–22
Roy V, Monti-Dedieu L, Chaminade N, Siljak-Yakovlev S, Aulard S, Lemeunier Fo et al (2005) Evolution of the chromosomal location of rDNA genes in two Drosophila species subgroups: ananassae and melanogaster. Heredity 94(4):388–395
Samaropoulou S, Bareka P, Kamari G (2016) Karyomorphometric analysis of Fritillaria Montana group in Greece. Comp Cytogenet 10(4):679–695
Shishido R, Sano Y, Fukui K (2000) Ribosomal DNAs: an exception to the conservation of gene order in rice genomes. Mol Gen Genet MGG 263(4):586–591
Singh M, Kumar R, Nagpure N, Kushwaha B, Mani I, Chauhan U et al (2009) Population distribution of 45S and 5S rDNA in golden mahseer, Tor putitora: population-specific FISH marker. J Genet 88(3):315–320
Stebbins GL (1971) Chromosomal evolution in higher plants. Edward Arnold, London
Taketa S, Harrison G, Heslop-Harrison J (1999) Comparative physical mapping of the 5S and 18S-25S rDNA in nine wild Hordeum species and cytotypes. Theor Appl Genet 98(1):1–9
Temsch EM, Temsch W, Ehrendorfer-Schratt L, Greilhuber J (2010) Heavy Metal Pollution, Selection, and genome size: the species of the Žerjav Study revisited with Flow Cytometry. J Bot 1–11
Tomović G, Vukojičić V, Snežana, Zlatković B, Stevanović V (2007) Fritillaria (Liliaceae) in Serbia: distribution, habitats and some taxonomic notes. Phytologia Balcanica 13(3):359–370
Vinogradov AE (2003) Selfish DNA is maladaptive: evidence from the plant Red List. Trends Genet 19(11):609–614
Watanabe K, Yahara T, Denda T, Kosuge K (1999) Chromosomal evolution in the genus Brachyscome (Asteraceae, Astereae): statistical tests regarding correlation between changes in karyotype and habit using phylogenetic information. J Plant Res 112(2):145–161
Zarco CR (1986) A new method for estimating karyotype asymmetry. Taxon 35(3):526–530
Zhao X, Lu J, Zhang Z, Hu J, Huang S, Jin W (2011) Comparison of the distribution of the repetitive DNA sequences in three variants of Cucumis sativus reveals their phylogenetic relationships. J Genet Genomics 38(1):39–45
Acknowledgements
We deeply thank M. Advay (University of Tehran) for providing the seeds of some Fritillaria samples.
Funding
The present research was carried out by the grant provided for MSc students by the University of Kurdistan.
Author information
Authors and Affiliations
Contributions
In this manuscript, Neda Seifoori has done the experimental work; Ghader Mirzaghaderi and shahla Hosseini analyzed, interpreted, wrote the results, and reviewed the manuscript.
Corresponding author
Ethics declarations
Ethical Approval
This declaration is not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Seifoori, N., Mirzaghaderi, G. & Hosseini, S. Chromosomal Positions of 5S and 45S rDNA in Some Iranian Fritillaria (Liliaceae) Species. Plant Mol Biol Rep (2024). https://doi.org/10.1007/s11105-024-01467-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11105-024-01467-0