Abstract
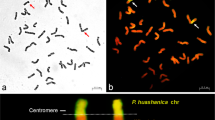
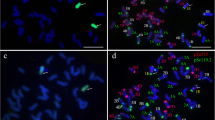
Psathyrostachys huashanica Keng (2n = 2x = 14, NsNs) is a valuable genetic resource for improved wheat breeding, because of its high fecundity, vigorous growth, and resistance to diseases. In this study, a novel wheat–P. huashanica 5Ns (5D) disomic substitution line DH66 was isolated from the F6 progeny of the heptaploid hybrid H8911 (2n = 7x = 49, AABBDDNs) and Triticum durum line Trs-372. Mitotic and meiotic observations showed that the chromosome karyotype of DH66 was 2n = 42 = 21II. Genomic in situ hybridization indicated that DH66 contained 40 wheat chromosomes and two P. huashanica chromosomes, which paired stably and were transmitted to the offspring. Fluorescence in situ hybridization showed that chromosome 5D was absent from DH66. Analysis using a 15K wheat chip demonstrated that the genotype of DH66 had strong matches with P. huashanica in terms of many single-nucleotide polymorphism (SNP) loci on the 5D chromosome, but few with line 7182 at the same SNPs. Verification using markers confirmed that a pair of wheat 5D chromosomes in DH66 were substituted by a pair of P. huashanica 5Ns chromosomes, and thus DH66 was characterized as a wheat–P. huashanica 5Ns (5D) disomic substitution line. Agronomic trait evaluations showed that, compared with its wheat parents, DH66 exhibited significant improvements, with reduced plant height and dough stability time, and superior resistance to stripe rust in the adult stage. Therefore, the novel cytogenetically stable substitution line DH66 can be used in wheat disease-resistant and high-quality breeding programs.








Similar content being viewed by others
References
Anamthawat-Jónsson K (1999) Variable genome composition in Triticum×Leymus amphiploids. Theor Appl Genet 99:1087–1093. https://doi.org/10.1007/s001220051313
Barrantes W, Fernández-del-Carmen A, López-Casado G, González-Sánchez MÁ, Fernández-Muñoz R, Granell A, Monforte AJ (2014) Highly efficient genomics-assisted development of a library of introgression lines of Solanum pimpinellifolium. Mol Breed 34:1817–1831. https://doi.org/10.1007/s11032-014-0141-0
Chen LG, Jun WU, Zhao JX, Liu SH, Yang QH, Wan-Li DU, Pang YH, Chen XH (2010) Screening, cloning and southern blotting analysis of specific repetitive DNA sequences of Psathyrostachys huashanica. Journal of Triticeae Crops 30:23–28. https://doi.org/10.1080/00949651003724790
Chen SY (1991) The hybridization between Triticum aestivum and Psathyrostachys huashanica. Acta Genet Sin 18:508–512
Chen XM (2005) Epidemiology and control of stripe rust [Puccinia striiformis f. sp. tritici] on wheat. Can J Plant Pathol 27:314–337. https://doi.org/10.1080/07060660509507230
Ciferri R (1955) The first interspecific wheat hybrids. J Heredity 46:81–83. https://doi.org/10.1093/oxfordjournals.jhered.a106529
Cifuentes M, Benavente E (2009) Wheat-alien metaphase I pairing of individual wheat genomes and D genome chromosomes in interspecific hybrids between Triticum aestivum L. and Aegilops geniculata Roth. Theor Appl Genet 119:805–813. https://doi.org/10.1007/s00122-009-1090-6
Danilova TV, Friebe B, Gill BS, Poland J, Jackson E (2018) Chromosome rearrangements caused by double monosomy in wheat-barley group-7 substitution lines. Cytogenet Genome Res 154:45–55. https://doi.org/10.1159/000487183
Do N, Akkaya M, Basit B, Tekrar D, Optimizasyonu Ç (2001) Optimization of PCR amplification of wheat simple sequence repeat DNA markers. Turk J Biol 25:153–158
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin 19:11–15
Du WL, Wang J, Lu M, Sun SG, Chen XH, Zhao JX, Yang QH, Wu J (2013a) Molecular cytogenetic identification of a wheat–Psathyrostachys huashanica Keng 5Ns disomic addition line with stripe rust resistance. Mol Breed 31:879–888. https://doi.org/10.1007/s11032-013-9841-0
Du WL, Wang J, Lu M, Sun SG, Chen XH, Zhao JX, Yang QH, Wu J (2014a) Characterization of a wheat-Psathyrostachys huashanica Keng 4Ns disomic addition line for enhanced tiller numbers and stripe rust resistance. Planta 239:97–105. https://doi.org/10.1007/s00425-013-1957-2
Du WL, Wang J, Pang YH, Li YL, Chen XH, Zhao J, Yang QH, Wu J (2013b) Isolation and characterization of a Psathyrostachys huashanica Keng 6Ns chromosome addition in common wheat. PLoS One 8:e53921. https://doi.org/10.1371/journal.pone.0053921
Du WL, Wang J, Pang YH, Wang LM, Wu J, Zhao JX, Yang QH, Chen XH (2014b) Isolation and characterization of a wheat-Psathyrostachys huashanica ‘Keng’ 3Ns disomic addition line with resistance to stripe rust. Genome 57:37–44. https://doi.org/10.1139/gen-2013-0199
Du WL, Wang J, Pang YH, Wu J, Zhao JX, Liu SH, Yang QH, Chen XH (2014c) Development and application of PCR markers specific to the 1Ns chromosome of Psathyrostachys huashanica Keng with leaf rust resistance. Euphytica 200:207–220. https://doi.org/10.1007/s10681-014-1145-x
Du WL, Wang J, Wang LM, Wu J, Zhao JX, Liu SH, Yang QH, Chen XH (2014d) Molecular characterization of a wheat–Psathyrostachys huashanica Keng 2Ns disomic addition line with resistance to stripe rust. Mol Gen Genomics 289:735–743. https://doi.org/10.1007/s00438-014-0844-2
Du WL, Wang J, Wang LM, Zhang J, Chen XH, Zhao JX, Yang QH, Wu J (2013c) Development and Characterization of a Psathyrostachys huashanica Keng 7Ns Chromosome Addition Line with Leaf Rust Resistance. PLoS One 8:e70879. https://doi.org/10.1371/journal.pone.0070879
Du WL, Zhao JX, Wang J, Wang LM, Wu J, Yang QH, Liu SH, Chen XH (2015) Cytogenetic and molecular marker-based characterization of a wheat-Psathyrostachys huashanica Keng 2Ns(2D) substitution line. Plant Mol Biol Report 33:414–423. https://doi.org/10.1007/s11105-014-0761-x
Du XY, Ma X, Min JZ, Zhang XC, Jia ZZ (2018) Development of a wheat-Aegilops searsii substitution line with positively affecting Chinese steamed bread quality. Breed Sci 68:289–293. https://doi.org/10.1270/jsbbs.17044
Fu J, Wang MN, Zhao JX, Chen SY, Hou WS, Yang QH (2003) Studies on cytogenetics and utilization of wheat-Psathyrostachys huashanica medium material H8911 with resistance to wheat take-all fungus. Acta Botan Boreali-Occiden Sin 23:2157–2162
Gale MD, Devos KM (1998) Plant comparative genetics after 10 years. Science 282:656–659. https://doi.org/10.1126/science.282.5389.656
Gupta P, Balyan H, Edwards K, Isaac P, Korzun V, Röder M, Gautier MF, Joudrier P, Schlatter A, Dubcovsky J, De la Pena R, Khairallah M, Penner G, Hayden M, Sharp P, Keller B, Wang R, Hardouin J, Jack P, Leroy P (2002) Genetic mapping of 66 new microsatellite (SSR) loci in bread wheat. Theor Appl Genet 105:413–422. https://doi.org/10.1007/s00122-002-0865-9
Han FP, Liu B, Fedak G, Liu ZH (2004) Genomic constitution and variation in five partial amphiploids of wheat–Thinopyrum intermedium as revealed by GISH, multicolor GISH and seed storage protein analysis. Theor Appl Genet 109:1070–1076. https://doi.org/10.1007/s00122-004-1720-y
Jing JX, Fu J, Yuan HX, Wang MN, Shang HS, Li ZQ (1999) A preliminary study on heredity of the resistance to stripe rust in three wild relatives of wheat. Acta Phytopathol Sin 29:147–150
Kang HY, Zhang ZJ, Xu LL, Qi WL, Tang Y, Wang H, Zhu W, Li DY, Zeng J, Wang Y, Fan X, Sha LN, Zhang HQ, Zhou YH (2016) Characterization of wheat-Psathyrostachys huashanica small segment translocation line with enhanced kernels per spike and stripe rust resistance. Genome 59:221–229. https://doi.org/10.1139/gen-2015-0138
King JL, Grewal S, Yang CY, Hubbart S, Scholefield D, Ashling S, Edwards KJ, Allen AM, Burridge A, Bloor C, Davassi A, da Silva GJ, Chalmers K, King IP (2017) A step change in the transfer of interspecific variation into wheat from Amblyopyrum muticum. Plant Biotechnol J 15:217–226. https://doi.org/10.1111/pbi.12606
Lang T, Li GR, Wang HJ, Yu ZH, Chen QH, Yang EN, Fu SL, Tang ZX, Yang ZJ (2018) Physical location of tandem repeats in the wheat genome and application for chromosome identification. https://doi.org/10.1007/s00425-018-3033-4
Liu SB, Wang HG (2005) Characterization of a wheat–Thinopyron intermedium substitution line with resistance to powdery mildew. Euphytica 143:229–233. https://doi.org/10.1007/s10681-005-3862-7
Liu WX, Jin Y, Rouse M, Friebe B, Gill B, Pumphrey MO (2011) Development and characterization of wheat-Ae. searsii Robertsonian translocations and a recombinant chromosome conferring resistance to stem rust. Theor Appl Genet 122:1537–1545. https://doi.org/10.1007/s00122-011-1553-4
Liu ZG, Yao WY, Shen XX, Chao KX, Fan Y, Li MZ, Wang BT, Li Q, Jing JX (2014) Molecular mapping of a stripe rust resistance gene YrH9020a transferred from Psathyrostachys huashanica Keng on wheat chromosome 6D. J Integr Agric 13:2577–2583. https://doi.org/10.1016/S2095-3119(14)60755-3
Ma DF, Fang ZW, Yin JL, Chao KX, Jing JX, Li Q, Wang BT (2016) Molecular mapping of stripe rust resistance gene YrHu derived from Psathyrostachys huashanica. Mol Breed 36:64. https://doi.org/10.1007/s11032-016-0487-6
Ma DF, Zhou XL, Hou L, Bai YB, Li Q, Wang HG, Tang MS, Jing JX (2013) Genetic analysis and molecular mapping of a stripe rust resistance gene derived from Psathynrostachys huashanica Keng in wheat line H9014-121-5-5-9. Mol Breed 32:365–372. https://doi.org/10.1007/s11032-013-9876-2
Ma H, Singh RP, Mujeeb KA (1995) Suppression/expression of resistance to stripe rust in synthetic hexaploid wheat (Triticum turgidum×T. tauschii). Euphytica 83:87–93. https://doi.org/10.1007/bf01678034
McIntosh R, Mu JM, Han DJ, Kang ZS (2018) Wheat stripe rust resistance gene Yr24/Yr26: A retrospective review. The Crop Journal 6:321–329. https://doi.org/10.1016/j.cj.2018.02.001
Pang YH, Chen XH, Zhao JX, Du WL, Cheng XN, Wu J, Li YL, Wang LM, Wang J, Yang QH (2014) Molecular cytogenetic characterization of a wheat-Leymus mollis 3D(3Ns) substitution line with resistance to leaf rust. Journal of Genetics and Genomics 41:205–214. https://doi.org/10.1016/j.jgg.2013.11.008
Patokar C, Sepsi A, Schwarzacher T, Kishii M, Heslop-Harrison JS (2016) Molecular cytogenetic characterization of novel wheat-Thinopyrum bessarabicum recombinant lines carrying intercalary translocations. Chromosoma 125:163–172. https://doi.org/10.1007/s00412-015-0537-6
Pestsova E, Ganal MW, Röder MS (2000) Isolation and mapping of microsatellite markers specific for the D genome of bread wheat. Genome 43:689–697. https://doi.org/10.1139/g00-042
Qi WL, Tang Y, Zhu W, Li DY, Diao CD, Xu LL, Zeng J, Wang Y, Fan X, Sha LN, Zhang HQ, Zheng YL, Zhou YH, Kang HY (2016) Molecular cytogenetic characterization of a new wheat-rye 1BL•1RS translocation line expressing superior stripe rust resistance and enhanced grain yield. Planta 244:405–416. https://doi.org/10.1007/s00425-016-2517-3
Röder MS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A Microsatellite Map of Wheat. Genetics 149:2007–2023
Ren TH, Chen F, Yan BJ, Zhang HQ, Ren ZL (2012) Genetic diversity of wheat–rye 1BL.1RS translocation lines derived from different wheat and rye sources. Euphytica 183:133–146. https://doi.org/10.1007/s10681-011-0412-3
Ren TH, Tang ZX, Fu SL, Yan BJ, Tan FQ, Ren ZL, Li Z (2017) Molecular cytogenetic characterization of novel wheat-rye T1RS.1BL translocation lines with high resistance to diseases and great agronomic traits. Front Plant Sci 8:799. https://doi.org/10.3389/fpls.2017.00799
Gill BS, Huang L, Kuraparthy V, Raupp W, Wilson DL, Friebe B (2008) Alien genetic resources for wheat leaf rust resistance, cytogenetic transfer, and molecular analysis. Aust J Agric Res 59:197–205. https://doi.org/10.1071/AR07315
Schneider LG, Molnar-Lang M, Graner A (2003) Fluorescence in situ hybridization polymorphism using two repetitive DNA clones in different cultivars of wheat. Plant Breed 122:396–400. https://doi.org/10.1046/j.1439-0523.2003.00891.x
Sepsi A, Molnár I, Szalay D, Molnár-Láng M (2008) Characterization of a leaf rust-resistant wheat–Thinopyrum ponticum partial amphiploid BE-1, using sequential multicolor GISH and FISH. Theor Appl Genet 116:825–834. https://doi.org/10.1007/s00122-008-0716-4
Sharma HC, Gill BS (1983) Current status of wide hybridization in wheat. Euphytica 32:17–31. https://doi.org/10.1007/bf00036860
Shirabe K, Ryota E, Kaori S, Akio K (2013) Genomic and chromosomal distribution patterns of various repeated DNA. Genome 56:131–137. https://doi.org/10.1139/gen-2013-0003
Su JN, Guo J, Wang CJ, Jing F, Zhao JX, Yang QH, Chen XH, Jun W (2015) Specific SCAR markers on chromosome 3Ns of Psathyrostachys huashanica Keng. Journal of Triticeae Crops 35:1–6
Sun G (1993) Intermeiocyte connections and cytomixis in intergeneric hybrids II. Triticum aestivumxPsathyrostachys huashanica. Wheat Information Service 77:13–18
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. https://doi.org/10.1093/molbev/msr121
Tang ZX, Yang ZJ, Fu SL (2014) Oligonucleotides replacing the roles of repetitive sequences pAs1, pSc119.2, pTa-535, pTa71, CCS1, and pAWRC.1 for FISH analysis. J Appl Genet 55:313–318. https://doi.org/10.1007/s13353-014-0215-z
Vaillancourt A, Nkongolo KK, Michael P, Mehes M (2008) Identification, characterisation, and chromosome locations of rye and wheat specific ISSR and SCAR markers useful for breeding purposes. Euphytica 159:297–306. https://doi.org/10.1007/s10681-007-9492-5
Wang J, Liu YL, Su HD, Guo XR, Han FP (2017) Centromere structure and function analysis in wheat-rye translocation lines. Plant J 91:199–207. https://doi.org/10.1111/tpj.13554
Wang J, Wang LM, Du WL, Chen LG, Liu SH, Wu J, Zhao JX, Yang QH, Chen XH (2014a) Development of 5Ns chromosome-specific SCAR markers for utilization in future wheat breeding programs. Russ J Genet 50:606–612. https://doi.org/10.1134/s1022795414060131
Wang LM, Liu Y, Du WL, Jing F, Wang ZH, Wu J, Chen XH (2015) Anatomy and cytogenetic identification of a wheat-Psathyrostachys huashanica Keng line with early maturation. PLoS One 10:e0131841. https://doi.org/10.1371/journal.pone.0131841
Wang S, Wong D, Forrest K, Allen A, Chao S, Huang BE, Maccaferri M, Salvi S, Milner SG, Cattivelli L (2014b) Characterization of polyploid wheat genomic diversity using a high-density 90,000 single nucleotide polymorphism array. Plant Biotechnol J 12:787–796. https://doi.org/10.1111/pbi.12183
Wang YJ, Quan W, Peng NN, Wang CY, Yang XF, Liu XL, Zhang H, Chen CH, Ji WQ (2016) Molecular cytogenetic identification of a wheat–Aegilops geniculata Roth 7Mg disomic addition line with powdery mildew resistance. Mol Breed 36:40. https://doi.org/10.1007/s11032-016-0463-1
Wu JH, Zeng QD, Wang QL, Liu SJ, Yu SZ, Mu JM, Huang S, Sela HN, Distelfeld A, Huang LL, Han DJ, Kang ZS (2018) SNP-based pool genotyping and haplotype analysis accelerate fine-mapping of the wheat genomic region containing stripe rust resistance gene Yr26. Theor Appl Genet 131:1481–1496. https://doi.org/10.1007/s00122-018-3092-8
Yang XF, Wang CY, Chen CH, Zhang H, Tian ZR, Li X, Wang YJ, Ji WQ (2015a) Chromosome constitution and origin analysis in three derivatives of Triticum aestivum–Leymus mollis by molecular cytogenetic identification. Genome 57:1–9. https://doi.org/10.1139/gen-2014-0161
Yang XF, Wang CY, Li X, Chen CH, Tian ZR, Wang YJ, Ji WQ (2015b) Development and molecular cytogenetic identification of a novel wheat-Leymus mollis Lm#7Ns (7D) disomic substitution line with stripe rust resistance. PLoS One 10:e0140227–e0140227. https://doi.org/10.1371/journal.pone.0140227
Yu SY, Long H, Yang H, Zhang J, Deng GB, Pan ZF, Zhang EL, Yu MQ (2012) Molecular detection of rye (Secale cereale L.) chromatin in wheat line 07jian126 (Triticum aestivum L.) and its association to wheat powdery mildew resistance. Euphytica 186:247–255. https://doi.org/10.1007/s10681-012-0649-5
Yuan FP, Zeng QD, Wu JH, Wang QL, Yang ZJ, Liang BP, Kang ZS, Chen XH, Han DJ (2018) QTL mapping and validation of adult plant resistance to stripe rust in Chinese wheat landrace humai 15. Front Plant Sci 9:968. https://doi.org/10.3389/fpls.2018.00968
Zhang W, Chao S, Manthey F, Chicaiza O, Brevis JC, Echenique V, Dubcovsky J (2008) QTL analysis of pasta quality using a composite microsatellite and SNP map of durum wheat. Theor Appl Genet 117:1361–1377. https://doi.org/10.1007/s00122-008-0869-1
Zhao JX, Du WL, Wu J, Cheng XN, Gao Y, Pang YH, Chen XH, Liu SH, Yang QH, Fu J (2013) Development and identification of a wheat-Leymus mollis multiple alien substitution line. Euphytica 190:45–52. https://doi.org/10.1007/s10681-012-0772-3
Zhao JX, Ji WQ, Wu J, Chen XH, Cheng XN, Wang JW, Pang YH, Liu SH, Yang QH (2010) Development and identification of a wheat-Psathyrostachys huashanica addition line carrying HMW-GS, LMW-GS and gliadin genes. Genet Resour Crop Evol 57:387–394. https://doi.org/10.1007/s10722-009-9477-4
Zhou SH, Zhang JP, Che YH, Liu WH, Lu YQ, Yang XM, Li XQ, Jia JZ, Liu X, Li LH (2018) Construction of Agropyron Gaertn. genetic linkage maps using a wheat 660K SNP array reveals a homoeologous relationship with the wheat genome. Plant Biotechnol J 16:818–827. https://doi.org/10.1111/pbi.12831
Zhu C, Wang YZ, Chen CH, Wang CY, Zhang A, Peng NN, Wang YJ, Zhang H, Liu XL, Ji WQ (2017) Molecular cytogenetic identification of a wheat-Thinopyrum ponticum substitution line with stripe rust resistance. Genome 60:860–867. https://doi.org/10.1139/gen-2017-0099
Acknowledgements
Many thanks to Dr. Duncan E. Jackson for his language editing and checking of scientific consistency.
Funding
Much appreciated financial support was provided by the National Science Foundation of China (31571650 and 31771785), National Key Research and Development Program of China (2017YFD0100701), and Key Projects in Shaanxi Provincial Agricultural Field (2018ZDXM-NY-006).
Author information
Authors and Affiliations
Contributions
JC Li wrote the paper. JC Li, JX Zhao and ZJ Yang conducted experiments. XN Cheng and FP Yuan contributed new reagents or analytical tools. XN Yao analyzed data. J Wu and QH Yang contributed new methods or models. JX Zhao and XH Chen conceived and designed the study.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, J., Yao, X., Yang, Z. et al. Molecular cytogenetic characterization of a novel wheat–Psathyrostachys huashanica Keng 5Ns (5D) disomic substitution line with stripe rust resistance. Mol Breeding 39, 109 (2019). https://doi.org/10.1007/s11032-019-1014-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-019-1014-3