Abstract
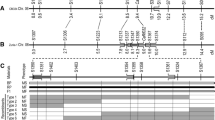
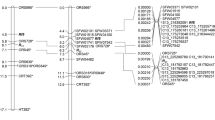
Cytoplasmic male-sterile line CMS ANN3 was discovered from a wild annual Helianthus annuus L. accession PI 413180. A single dominant fertility restoration (Rf) gene for CMS ANN3 was identified in PI 413180, P 21, RHA 801, RCMG 1, and RHA 280, designated as Rf9. This study reports the mapping of the Rf9 locus using an F2 population of 160 individuals. Bulked segregant analysis with SSR primers identified two polymorphic SSR markers in linkage group 3 of the sunflower genetic map. A linkage map of Rf9 was constructed with 16 markers including 11 SSR/STS and five SNP markers, covering a genetic distance of 87.3 cM. The Rf9 gene is located between a co-dominant marker ORS1114 and a dominant marker NSA002021, with a distance of 4.1 and 1.0 cM, respectively. The physical distance between the two closest flanking markers corresponds to 1.44 and 2.05 Mb on the HA 412-HO and XRQ assemblies, respectively. The closely linked marker for Rf9 will facilitate marker-assisted selection in breeding programs, which provides a basis for further characterization of this gene, and for studying the mechanism between cytoplasmic male sterility and Rf genes. The mapping of the Rf gene will also be useful for studying Rf gene suppression and nuclear–cytoplasm interactions.



Similar content being viewed by others
Abbreviations
- MAS:
-
Maker-assisted selection
- SSR:
-
Simple sequence repeats
- STS:
-
Sequence-tagged sites
- LOD:
-
Logarithm of odds
References
Aarts MG, Dirkse WG, Stiekema WJ, Pereira A (1993) Transposon tagging of a male sterility gene in Arabidopsis. Nature 363:715–717
Abratti G, Bazzalo ME, León A (2008) Mapping a novel fertility restoration gene in sunflower. In: Velasco L (ed) Proceedings of the 17th international sunflower conference, 8–12 June. Córdoba, Spain, pp 617–621
Bentolila S, Alfonso AA, Hanson MR (2002) A pentatricopeptide repeat-containing gene restores fertility to male-sterile plants. Proc Natl Acad Sci USA 99:10887–10892
Bowers JE, Bachlava E, Brunick RL, Rieseberg LH, Knapp SJ, Burke JM (2012) Development of a 10,000 locus genetic map of the sunflower genome based on multiple crosses. G3 Genes Genomes Genet 2:721–729
Brown GG, Formanova N, Jin H, Wargachuk R, Dendy C, PatilP LM, Zhang J, Cheung WY, Landry BS (2003) The radish Rfo restorer gene of Ogura cytoplasmic male sterility encodes a protein with multiple pentatricopeptide repeats. Plant J 35:262–272
Cui X, Wise RP, Schnable PS (1996) The rf2 nuclear restorer gene of male-sterile T-cytoplasm maize. Science 272:1334–1336
Eckardt NA (2006) Cytoplasmic male sterility and fertility restoration. Plant Cell 18:515–517. https://doi.org/10.1105/tpc.106.041830
Feng J, Jan CC (2008) Introgression and molecular tagging of Rf4, a new male fertility restoration gene from wild sunflower Helianthus maximiliani L. Theor Appl Genet 117:241–249
Fujii S, Toriyama K (2009) Suppressed expression of RETROGRADE-REGULATED MALE STERILITY restores pollen fertility in cytoplasmic male sterile rice plants. Proc Natl Acad Sci USA 106:9513–9518. https://doi.org/10.1073/pnas.0901860106
Gentzbittel L, Vear F, Zhang Y-X, Bervillé A, Nicolas P (1995) Development of a consensus linkage RFLP map of cultivated sunflower (Helianthus annuus L.). Theor Appl Genet 90:1079–1086
Gentzbittel L, Mestries E, Mouzeyar S, Mazeyrat F, Badaoui S, Vear F, Tourvieille de Labrouhe D, Nicolas P (1999) A composite map of expressed sequences and phenotypic traits of the sunflower (Helianthus annuus L.) genome. Theor Appl Genet 99:218–234. https://doi.org/10.1007/s001220051228
Horn R, Kusterer B, Lazarescu E, Prüfe M, Friedt W (2003) Molecular mapping of the Rf1 gene restoring pollen fertility in PET1-based F1 hybrids in sunflower (Helianthus annuus L.). Theor Appl Genet 106:599–606
Horn R, Radanovic A, Fuhrmann L, Sprycha Y, Hamrit S, Jockovic M, Miladinovic D, Jansen C (2019) Development and validation of markers for the fertility restorer gene Rf1 in sunflower. Int J Mol Sci 20:1260. https://doi.org/10.3390/ijms20061260
Hulke BS, Grassa CJ, Bowers JE, Burke JM, Qi L, Talukder ZI, Rieseberg LH (2015) A unified single nucleotide polymorphism map of sunflower (Helianthus annuus L.) derived from current genomic resources. Crop Sci 55:1696–1702
Jan CC (2000) Cytoplasmic male sterility in two wild Helianthus annuus L. accessions and their fertility restoration. Crop Sci 40:1535–1538
Jan CC (2003) Silencing of fertility restoration genes in sunflower. Helia 26:1–6
Jan CC, Rutger JN (1988) Mitomycin C- and streptomycin-induced male sterility in cultivated sunflower. Crop Sci 28:792–795
Jan CC, Vick BA, Miller JF, Kahler AL, Butler ET III (1998) Construction of an RFLP linkage map for cultivated sunflower. Theor Appl Genet 96:15–22
Josefsson C, Dilkes B, Comai L (2006) Parent-dependent loss of gene silencing during interspecies hybridization. Curr Biol 16:1322–1328
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Kusterer B, Horn R, Friedt W (2005) Molecular mapping of the fertility restoration locus Rf1 in sunflower and development of diagnostic markers for the restorer gene. Euphytica 143:35–42
Lai Z, Livingstone K, Zou Y, Church SA, Knapp SJ, Andrews J, Rieseberg LH (2005) Identification and mapping of SNPs from ESTs in sunflower. Theor Appl Genet 111:1532–1544
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newberg LA (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Leclercq P (1969) Une sterilite cytoplasmique chez le tournesol. Ann Amelior Plant 19:99–106
Liu F, Cui X, Horner HT, Weiner H, Schnabel PS (2001) Mitochondrial aldehyde dehydrogenase activity is required for male fertility in maize. Plant Cell 13:1063–1078
Liu Z, Mulpuri S, Feng J, Vick BA, Jan CC (2012) Molecular mapping of the Rf3 fertility restoration gene to facilitate its utilization in breeding confection sunflower. Mol Breed 29:275–284
Liu Z, Wang D, Feng J, Seiler GJ, Cai X, Jan CC (2013) Diversifying sunflower germplasm by integration and mapping of a novel male fertility restoration gene. Genetics 193:727–737
Liu Z, Cai X, Seiler GJ, Jan CC (2014) Interspecific amphiploid-derived alloplasmic male sterility with defective anthers, narrow disc florets, and small ray flowers in sunflower. Plant Breed. https://doi.org/10.1111/pbr.12216
Liu Z, Long Y, Xu SS, Seiler GJ, Jan CC (2019) Unique fertility restoration suppressor genes for male-sterile CMS ANN2 and CMS ANN3 cytoplasms in sunflower (Helianthus annuus L.). Mol Breed 39:22
Liu Z, Gu W, Seiler GJ, Jan CC (2020) A unique cytoplasmic-nuclear interaction in sunflower (Helianthus annuus L.) causing reduced-vigor plants and the genetics of vigor restoration. Front Plant Sci 11:1010. https://doi.org/10.3389/fpls.2020.01010
Liu Z, Wang H, Jan CC (2016) Additional Rf genes for CMS GIG2 and their molecular mapping. In: Proceeding of 38th sunflower research workshop, 12–13 Jan. National Sunflower Association, Fargo, ND. https://www.sunflowernsa.com/uploads/research/1298/2016NSA-Rfgenemappingposter-Liu-final.pdf
Long YM, Chao WS, Ma GJ, Xu SS, Qi LL (2017) An innovative SNP genotyping method adapting to multiple platforms and throughputs. Theor Appl Genet 130:597–607
Ma G, Long Y, Song Q, Talukder ZI, Shamimuzzaman M, Qi L (2021) Map and sequence-based chromosome walking towards cloning of the male fertility restoration gene Rf5 linked to R11 in sunflower. Sci Rep 11:777. https://doi.org/10.1038/s41598-020-80659-6
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci USA 88:9828–9832
Mulpuri S, Liu Z, Feng J, Gulya TJ, Jan CC (2009) Inheritance and molecular mapping of a downy mildew resistance gene, Pl13 in cultivated sunflower (Helianthus annuus L.). Theor Appl Genet 119:795–803
Owens GL, Baute GJ, Hubner A, Rieseberg LH (2018) Genomic sequences and copy number evolution during hybrid crop development in sunflowers. Evol Appl 11:1–12
Postel Z, Touzet P (2020) Cytonuclear genetic incompatibilities in plant speciation. Plants (basel) 9(4):E487. https://doi.org/10.3390/plants9040487
Qi LL, Seiler GJ, Vick BA, Gulya TJ (2012) Genetics and mapping of the R11 gene conferring resistance to recently emerged rust races, tightly linked to male fertility restoration, in sunflower (Helianthus annuus L.). Theor Appl Genet 125:921–932
Schnabel U, Engelmann U, Horn R (2008) Development of markers for the use of the PEF1 cytoplasm in sunflower hybrid breeding. Plant Breed 127:587–591
Seiler GJ, Qi LL, Marek LF (2017) Utilization of sunflower crop wild relatives for cultivated sunflower improvement. Crop Sci 57:1083–1101. https://doi.org/10.2135/cropsci2016.10.0856
Serieys H (1996) Identification, study and utilization in breeding programs of new CMS sources FAO Progress Report. Helia 19:144–158
Serieys H (2005) Identification, study and utilization in breeding programs of new CMS sources. Sunflower subnetwork progress report. FAO Subnetwork, Rome, pp 47–53
Sinha R, Rajam MV (2013) RNAi silencing of three homologues of S-adenosylmethionine decarboxylase gene in tapetal tissue of tomato results in male sterility. Plant Mol Biol 82:169–180. https://doi.org/10.1007/s11103-013-0051-2
Talukder ZI, Gong L, Hulke BS, Pegadaraju V, Song Q, Schultz Q, Qi L (2014) A high-density SNP map of sunflower derived from RAD-sequencing facilitating fine-mapping of the rust resistance gene R12. PLoS ONE 9:e98628
Talukder ZI, Ma G, Hulke BS, Jan CC, Qi L (2019) Linkage mapping and genome-wide association studies of the Rf gene cluster in sunflower (Helianthus annuus L.) and their distribution in world sunflower collections. Front Genet 10:216. https://doi.org/10.3389/fgene.2019.00216
Tang S, Yu JK, Slabaugh B, Shintani K, Knapp J (2002) Simple sequence repeat map of the sunflower genome. Theor Appl Genet 105:1124–1136
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Wang Z, Zou Y, Li X, Zhang Q, Chen L, Wu H, Su D, Chen Y, Guo J, Luo D, Long Y, Zhong Y, Liu YG (2006) Cytoplasmic male sterility of rice with boro II cytoplasm is caused by a cytotoxic peptide and is restored by two related PPR motif genes via distinct modes of mRNA silencing. Plant Cell 18:676–687
Wolf SL, Miller JF (1985) Fertility restoration response of various sunflower cytoplasms. In: Proceedings 11th international sunflower conference, Mar del Plata, Argentina, 10–13 Mar, pp 549–552
Yang Q, Nong X, Xu J, Huang F, Wang F, Wu J, Zhang C, Liu C (2021) Unraveling the genetic basis of fertility restoration for cytoplasmic male sterile line WNJ01A originated from Brassica juncea in Brassica napus. Front Plant Sci 12:721980. https://doi.org/10.3389/fpls.2021.721980
Yu JK, Tang S, Slabaugh MB, Heesacker A, Cole G, Herring M, Soper J, Han F, Chu WC, Webb DM, Thompson L, Edwards KJ, Berry S, Leon AJ, Grondona M, Olungu C, Maes N, Knapp SJ (2003) Towards a saturated molecular genetic linkage map for cultivated sunflower. Crop Sci 43:367–387
Yue B, Vick BA, Cai X, Hu J (2010) Genetic mapping for the Rf1 (fertility restoration) gene in sunflower (Helianthus annuus L.) by SSR and TRAP markers. Plant Breed 129:24–28
Acknowledgements
Technical assistance of Lisa A. Brown and Dr. Yunming Long is greatly appreciated. The authors thank Ming Zhang, Hongxia Wang, and Jichong Zhang for their great help in conducting this study.
Funding
This work was supported [in part] by the U.S. Department of Agriculture, Agricultural Research Service through the USDA-ARS National Sclerotinia Initiative Grant No. 3060-21220-028-00D, and the USDA-ARS, Current Research Information System Project No. 3060-21000-043-00D. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture. The USDA is an equal opportunity provider and employer.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: CCJ, ZL. Performed the experiment: ZL, LZ, CCJ. Analyzed the data: ZL, LZ, CCJ. Wrote the paper: ZL, CCJ, GJS, LZ. Commented on paper before submission: ZL, CCJ, LZ, GJS.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests, that are directly or indirectly related to the work submitted for publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10681_2023_3176_MOESM1_ESM.xlsx
Table S1. Gene annotation identified a total of 53 putative Rf-related genes on the target Rf9-containing region on chromosome 3 in the XRQ-v2.1 reference genome
10681_2023_3176_MOESM2_ESM.xlsx
Table S2. Blast HanXRQr2Chr03g0134161 to HA 412-HO-v2.0 assembly identified 22 sequences in the target Rf9-containing region on chromosome 3 in the HA 412-HO-v2.0 reference genome
Rights and permissions
About this article
Cite this article
Liu, Z., Zhang, L., Seiler, G.J. et al. Molecular mapping of the Rf9 gene from RCMG 1 for CMS ANN3 derived from wild sunflower (Helianthus annuus L.). Euphytica 219, 46 (2023). https://doi.org/10.1007/s10681-023-03176-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-023-03176-3