Abstract
MYB transcription factors exist in a large copy number and control various plant phenotypes. We cloned R2R3 MYB-type transcription factors that determine the coloration of basal sheaths in barley and wheat coleoptiles. These genes are highly homologous to maize C1 and rice OsC1, regulators for anthocyanin biosynthesis, but they control seed pigmentation in maize and rice. On the basis of high homology, barley and wheat counterparts are designated HvC1 and TaC1, respectively. HvC1 gene is located on the short arm of chromosome 7H, and TaC1 genes are located on the short arms of chromosomes 7A, 7B, and 7D (TaC1-A1, B1, and D1, respectively). HvC1 is a strong candidate for Ant1 because of (1) complete co-segregation of anthocyanin pigmentation phenotype of the basal sheath with the HvC1 genotype in genetic mapping, and (2) complete deletion of the HvCl gene in two anthocyanin-decreased allelic mutants (ant1.1 and ant1.2) that were induced by irradiation. In contrast, colorless coleoptile wheat lines had lesions in all three genomes consisting of a single-nucleotide substitution or a 1-bp deletion of TaC1-A1, a 1.7-kb insertion of TaC1-B1, and a 2.0-kb insertion of TaC1-D1. At least one normal TaC1 gene appears to be sufficient to produce anthocyanin pigments in wheat coleoptiles. Previous crossing experiments localized Rc (red coleoptile) genes to homoeologous group 7 chromosomes and deduced Rc genotypes of several wheat lines. Their TaC1 gene sequence variation coincided with deduced Rc genotypes; therefore, the present molecular genetic study demonstrates that TaC1 is a strong candidate for Rc in wheat.







Similar content being viewed by others
References
Cockram J, White J, Zuluaga DL, Smith D, Comadran J, Macaulay M, Luo Z, Kearsey MJ, Werner P, Harrap D, Tapsell C, Liu H, Hedley PE, Stein N, Schulte D, Steuernagel B, Marshall DF, Thomas WT, Ramsay L, Mackay I, Balding DJ, Waugh R, O’Sullivan DM (2010) Genome-wide association mapping to candidate polymorphism resolution in the unsequenced barley genome. Proc Natl Acad Sci USA 107:21611–21616
Cone KC, Cocciolone SM, Burr FA, Burr B (1993) Maize anthocyanin regulatory gene Pl is a duplicate of C1 that functions in the plant. Plant Cell 5:1795–1805
Devos KM, Dubcovsky J, Dvorak J, Chinoy CN, Gale MD (1995) Structural evolustion of wheat chromosomes 4A, 5A, and 7B and its impact on recombination. Theor Appl Genet 91:282–288
Druka A, Kudrna D, Rostoks N, Brueggeman R, von Wettstein D, Kleinhofs A (2003) Chalcone isomerase gene from rice (Oryza sativa) and barley (Hordeum vulgare): physical, genetic and mutation mapping. Gene 302:171–178
Endo TR, Gill BS (1996) The deletion stocks of common wheat. J Heredity 87:295–307
Franckowiak J, Lundqvist U (2012) Descriptions of barley genetic stocks for 2012. Barley Genet Newsl 42:36–792
Himi E, Noda K (2005) Red grain colour gene (R) of wheat is a Myb-type transcription factor. Euphytica 143:239–242
Himi E, Maekawa M, Miura H, Noda K (2011) Development of PCR markers for Tamyb10 related to R-1, red grain color gene in wheat. Theor Appl Genet 122:1561–1576
Himi E, Yamashita Y, Haruyama N, Yanagisawa T, Maekawa M, Taketa S (2012) Ant28 gene for proanthocyanidin synthesis encoding the R2R3 MYB domain protein (Hvmyb10) highly affects grain dormancy in barley. Euphytica 188:141–151
Hu J, Anderson B, Wessler SR (1996) Isolation and characterization of rice R genes: evidence for distinct evolutionary paths in rice and maize. Genetics 142:1021–1031
Islam AKMR, Shepherd KW, Sparrow DHB (1981) Isolation and characterization of euplasmic wheat–barley chromosome addition lines. Heredity 46:161–174
Jende-Strid B (1984) Coordinator’s report: anthocyanin genes. Barley Genet Newsl 14:76–79
Jende-Strid B, Lundqvist U (1978) Diallelic tests of anthocyanin-deficient mutants. Barley Genet Newsl 8:57–59
Khlestkina EK, Pestsova EG, Roder MS, Borner A (2002) Molecular mapping, phenotypic expression and geographical distribution of genes determining anthocyanin pigmentation of coleoptiles in wheat (Triticum aestivum L.). Theor Appl Genet 104:632–637
Khlestkina EK, Roder MS, Salina EA (2008) Relationship between homoeologous regulatory and structural genes in allopolyploid genome—a case study in bread wheat. BMC Plant Biol 8:88
Khlestkina EK, Roder MS, Pshenichnikova TA, Borner A (2010) Functional diversity at the Rc (rec coleoptile) gene in bread wheat. Mol Breed 25:125–132
Khlestkina EK, Antonova EV, Pershina LA, Soloviev AA, Badaeva ED, Borner A, Salina EA (2011) Variability of Rc (red coleoptile) alleles in wheat and wheat-alien genetic stock collections. Cereal Res Commun 39:465–474
Kristiansen K, Rohde W (1991) Structure of the Hordeum vulgare gene encoding dihydroflavonol-4-reductase and molecular analysis of ant18 mutants blocked in flavonoid synthesis. Mol Gen Genet 230:49–59
Li S (2014) Transcriptional control of flavonoid biosynthesis: fine-tuning of the MYB-bHLH-WD40 (MBW) complex. Plant Signal Behav 9(1):e27522
Li WL, Faris JD, Chittoor JM, Leach JE, Hulbert SH, Liu DJ, Chen PD, Gill BS (1999) Genomic mapping of defense response genes in wheat. Thero Appl Genet 98:226–233
Li JZ, Sjakste TG, Roder MS, Ganal MW (2003) Development and genetic mapping of 127 new microsatellite markers in barley. Theor Appl Genet 107:1021–1027
Ling HQ, Zhao S, Liu D, Wang J, Sun H, Zhang C, Fan H, Li D, Dong L, Tao Y, Gao C, Wu H, Li Y, Cui Y, Guo X, Zheng S, Wang B, Yu K, Liang Q, Yang W, Lou X, Chen J, Feng M, Jian J, Zhang X, Luo G, Jiang Y, Liu J, Wang Z, Sha Y, Zhang B, Wu H, Tang D, Shen Q, Xue P, Zou S, Wang X, Liu X, Wang F, Yang Y, An X, Dong Z, Zhang K, Zhang X, Luo MC, Dvorak J, Tong Y, Wang J, Yang H, Li Z, Wang D, Zhang A, Wang J (2013) Draft genome of the wheat A-genome progenitor Triticum urartu. Nature 496:87–90
McIntosh RA, Yamazaki Y, Dubcovsky J, Rogers J, Morris C, Somers DJ, Appels R, Devos KM (2010) Catalogue of gene symbols for wheat National BioResource Project (NBRP)/KOMUGI
Neff MM, Chory J (1998) Genetic interactions between phytochrome A, phytochrome B, and cryptochrome 1 during Arabidopsis development. Plant Physiol 118:27–35
Ouyang S, Zhu W, Hamilton J, Lin H, Campbell M, Childs K, Thibaud-Nissen F, Malek RL, Lee Y, Zheng L, Orvis J, Haas B, Wortman J, Buell CR (2007) The TIGR rice genome annotation resource: improvements and new features. Nucleic Acids Res 35:D883–D887
Petroni K, Tonelli C (2011) Recent advances on the regulation of anthocyanin synthesis in reproductive organs. Plant Sci 181:219–229
Rostoks N, Mudie S, Cardle L, Russell J, Ramsay L, Booth A, Svensson JT, Wanamaker SI, Walia H, Rodriguez EM, Hedley PE, Liu H, Morris J, Close TJ, Marshall DF, Waugh R (2005) Genome-wide SNP discovery and linkage analysis in barley based on genes responsive to abiotic stress. Mol Genet Genomics 274:515–527
Saitoh K, Onishi K, Mikami I, Thidar K, Sano Y (2004) Allelic diversification at the C (OsC1) locus of wild and cultivated rice: nucleotide changes associated with phenotypes. Genetics 168:997–1007
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Sorrells ME, La Rota M, Bermudez-Kandianis CE, Greene RA, Kantety R, Munkvold JD, Miftahudin MA, Ma X, Gustafson PJ, Qi LL, Echalier B, Gill BS, Matthews DE, Lazo GR, Chao S, Anderson OD, Edwards H, Linkiewicz AM, Dubcovsky J, Akhunov ED, Dvorak J, Zhang D, Nguyen HT, Peng J, Lapitan NL, Gonzalez-Hernandez JL, Anderson JA, Hossain K, Kalavacharla V, Kianian SF, Choi DW, Close TJ, Dilbirligi M, Gill KS, Steber C, Walker-Simmons MK, McGuire PE, Qualset CO (2003) Comparative DNA sequence analysis of wheat and rice genomes. Genome Res 13:1818–1827
Taketa S, Yuo T, Sakurai Y, Miyake S, Ichii M (2011) Molecular mapping of the short awn 2 (lks2) and dense spike 1 (dsp1) genes on barley chromosome 7H. Breeding Science 61:80–85
Varshney RK, Marcel TC, Ramsay L, Russell J, Roder MS, Stein N, Waugh R, Langridge P, Niks RE, Graner A (2007) A high density barley microsatellite consensus map with 775 SSR loci. Theor Appl Genet 114:1091–1103
Winkel-Shirley B (2001) Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol 126:485–493
Acknowledgments
This work was supported by the Wesco Scientific Promotion Foundation and the Sapporo Bioscience Foundation. The authors thank Ms. Fumiko Katayama, Mr. Yuichi Kikuchi, and Ms. Noriko Hirose for their technical assistance and Dr. M. Röder (IPK) for barley SSR primer information. CS deletion lines for the short arms of 7A, 7B, and 7D were provided by the National BioResource Project.
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
438_2015_991_MOESM1_ESM.pptx
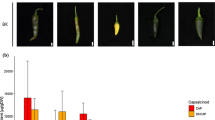
Supplementary material 1 (PPTX 1553 kb) Supplementary Fig. S1 a Base-color comparison of the leaf sheath of Bonus, ant1.1, ant1.2, ant1.4, and ant1.56. Awns of the heading stages of Bonus (b) and ant1.1 (c), and auricles of the flag leaves of Bonus (d) and ant1.1 (e). f Relative anthocyanin content of the auricles (black) of the flag leaves and awns (white) at the heading stage. Error bars represent standard error. *P < 0.05Supplementary Fig. S2 a Schematic diagrams of chromosomes 7BS and 7DS. Chromosomal breaking positions of partial deletion lines are shown with arrows, and numbers in parentheses indicate fraction lengths. b Amplification of TaC1-B1 in CS and its partial deletion lines of 7BS (upper) and TaC1-D1 in CS and its partial deletion lines of 7DS (lower).Supplementary Fig. S3 Alignment of genomic nucleotide sequences of three wheat C1 genes (TaC1-A1 of Hope, TaC1-B1 of Hope, and TaC1-D1 of NS67). Gray-boxed regions represent exons. Putative initiation codons (ATG) and stop codons (TAA) are underlined. White arrowheads indicate the insertion in TaC1-B1 and -D1. White arrows show mutation sites in TaC1-A1. Recognition sites of restriction enzyme (PmlI and ClaI) are boxed.Supplementary Fig. S4 TaC1 genotyping of one colorless coleoptile line (CS) and nine red coleoptile lines (8019R1, Hope, NS67, Douson1, M808, Cornell595-W, Ardito, AUS1408, and OW104). a PCR products of TaC1-A1 using TaC1-LP15 and TaC1-RP4 with (+) or without (-) ApaI treatment. Undigested lines (CS, M808, Cornell595-W, Ardito, AUS1408, and OW104) are genotyped to Rc-A1a. b Partial alignments of TaC1-A1 sequences of Hope (Rc-A1b) and NS67 (Rc-A1a). A primer for PCR is shown with a black arrow in alignment. The TaC1-RP15 primer generates a recognition site of EcoRV (GATATC) with a mismatched base printed in lower case and indicated by a gray arrow. c PCR products of TaC1-A1 using TaC1-LP6 and TaC1-RP15 with (+) or without (-) EcoRV treatment. Digested lines (NS67 and Douson1) are genotyped to Rc-A1a. d PCR product of TaC1-B1 with pairs of TaC1-LP11 and TaC1-RP8 (upper), and the PCR product of TaC1-B1 with pairs of TaC1-LP10 and TaC1-RP8 (lower). e PCR product of TaC1-D1 with pairs of TaC1-LP3 and TaC1-RP11 (upper), and the PCR product of TaC1-D1 with pairs of TaC1-LP7 and TaC1-RP11 (lower)
Rights and permissions
About this article
Cite this article
Himi, E., Taketa, S. Isolation of candidate genes for the barley Ant1 and wheat Rc genes controlling anthocyanin pigmentation in different vegetative tissues. Mol Genet Genomics 290, 1287–1298 (2015). https://doi.org/10.1007/s00438-015-0991-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-015-0991-0