Abstract
Compared to the major histocompatibility complex (MHC) of typical mammals, the chicken BF/BL region is small and simple, with most of the genes playing central roles in the adaptive immune response. However, some genes of the chicken MHC are almost certainly involved in innate immunity, such as the complement component C4 and the lectin-like receptor/ligand gene pair BNK and Blec. The poorly expressed classical class I molecule BF1 is known to be recognised by natural killer (NK) cells and, analogous to mammalian immune responses, the classical class I molecules BF1 and BF2, the CD1 homologs and the butyrophilin homologs called BG may be recognised by adaptive immune lymphocytes with semi-invariant receptors in a so-called adaptate manner. Moreover, the TRIM and BG regions next to the chicken MHC, along with the genetically unlinked Y and olfactory/scavenger receptor regions on the same chromosome, have multigene families almost certainly involved in innate and adaptate responses. On this chicken microchromosome, the simplicity of the adaptive immune gene systems contrasts with the complexity of the gene systems potentially involved in innate immunity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The major histocompatibility complex (MHC) is the genetic region that determines rapid graft rejection, due to the highly polymorphic classical class I and class II molecules that play central roles in the adaptive immune system by presentation of peptides to T cells, as well as being important in innate immunity as ligands for natural killer (NK) cells (Kaufman 2018a). The MHC of typical mammals is a large genomic region with hundreds of genes, among which a few encode classical MHC molecules along with many genes that have other functions (Knight and Trowsdale 2013). By contrast, the chicken MHC is a small and simple region, expressing a single class I molecule and a single class II molecule at high levels, whose properties can determine the immune response (Kaufman 2018b, Kaufman et al. 1999a).
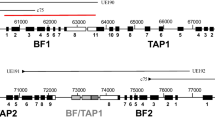
The chicken MHC is known for strong associations with resistance and susceptibility to economically important pathogens (Kaufman 2013, Miller and Taylor 2016), and so it perhaps is not too surprising that the focus of most chicken MHC research has been on the genes important for adaptive immunity. However, there are some genes in the chicken MHC which are likely to be important for innate immunity and/or for recognition by cells of the adaptive immune system with semi-invariant receptors (called by some “adaptate responses”, Hayday 2019; Hayday and Vantourout 2020). Moreover, the chicken MHC is embedded in a larger region and on the same chromosome with other genomic regions, and some of the genes in these regions may also be important for innate or adaptate immunity. This brief review will describe some aspects of these genes which have been neglected up to now, contrasting the complexity of the various innate systems with the simplicity of the adaptive immune system genes on this chromosome (Fig. 1).
A schematic diagram to indicate the genes in the chicken MHC and adjacent chromosomal regions that are likely to be involved in innate or adaptate immune responses (downward-pointing arrows underneath the line depicting the genomic sequence), in comparison to the classical class I gene BF2 and the classical class II B gene BLB2 involved in the adaptive response (upward-pointing arrows above the line). To be clear, some multigene families have copy number variation, so the exact number of genes is not implied in this diagram. There are genes that are involved in supplying peptides to the class I and class II molecules that are not depicted
The chicken MHC, the B locus and the Y locus
Altogether, the genomic regions on the long arm of chromosome 16 are understood in different levels of detail. These levels vary from mapping and cytogenetics to detailed molecular sequence (Fig. 2).
Organisation of regions on chicken chromosome 16, as currently understood. A Depiction of chromosome 16, based on analysis by FISH, radiation hybrids, genetics, southern blotting and sequencing. B, B locus; GC, G + C-rich region of PO1 repeats; Y, Rfp-Y region; NOR, nucleolar organiser region; BLA, class II A gene; fB, factor B gene; ORs, olfactory receptor genes; SRCRs, scavenger receptor with cysteine repeat genes. Double-headed arrows indicate recombination frequencies between B and BLA, fB and Rfp-Y, and B and Rfp-Y. B Region of the B locus currently sequenced, including the BF-BL region, the TRIM region and the BG region. Genes represented by boxes. Rising and falling stripes indicate genes of the classical class I and class II presentation system, respectively; stippled indicate class II region genes; black indicates lectin-like genes and pseudogenes; horizontal stripes indicate TRIM family genes; vertical stripes indicate BG genes. Names of genes above indicate transcription from left to right, below indicate transcription from right to left. (Figure modified from Kaufman 2013)
The chicken MHC is embedded in a larger historical genetic region called the B locus (or sometimes, the B complex), which was originally described based on the reactivity of alloantisera raised against blood cells and assessed by reactivity to erythrocytes (reviewed in Afrache et al. 2020). By a variety of methods, it was determined that two genetic loci separated by some level of recombination were involved, the BG region and the BF/BL region (Pink et al. 1977; Simonsen et al. 1982; Ziegler and Pink 1976). The BG antigens, whose closest relatives are the butyrophilins, were originally thought to be erythrocyte antigens, until it was shown that they are found on other cells of the hemopoietic lineage (including lymphoid and myeloid cells) as well as epithelial cells (Miller et al. 1990; Salomonsen et al. 1991, 2014). The BF antigens, originally described on erythrocytes and lymphocytes, are the chicken classical class I molecules. The BL antigens, originally described on B cells, are the chicken classical class II molecules. The B locus was found to determine resistance to the tumours induced by Marek’s disease virus (an oncogenic herpesvirus), with recombinants mapping the response to the BF/BL region (Briles et al. 1977, 1983; Plachy et al. 1992). The B locus was shown to be located on a microchromosome (now numbered as chromosome 16) along with the nucleolar organiser region (NOR) that contains many ribosomal RNA (rRNA) genes (Bloom and Bacon 1985).
The first step in understanding the B locus at the molecular level was through cosmids isolated by Francois Guillemot in the lab of Charles Auffray, who defined four cosmid clusters from the B12 haplotype, one of which is now known to cover much of the BF-BL region (Guillemot et al. 1988). When cosmids from this cluster were sequenced, it became clear that they constituted the BF/BL region which was the chicken MHC, defined as the genomic region responsible for rapid graft rejection. Compared to the MHC of typical mammals, this region was small and simple, containing a few polymorphic classical class I and class II B genes, polymorphic genes involved as in peptide loading and a single C4 gene, so that the BF-BL region was dubbed a “minimal essential MHC” (Kaufman et al. 1995, 1999a, b).
The other three cosmids, originally thought to be part of the MHC, were eventually understood to be derived from the Rfp-Y region that was discovered by Marcia Miller and colleagues based on restriction fragment polymorphism (Rfp) with BF gene probes (Briles et al. 1993; Miller et al. 1994, 1996). These Rfp-Y cosmids contained class I sequences related to the classical BF genes as well as class II B sequences related to the classical BLB genes (Miller et al. 1994; Zoorob et al. 1993); later, lectin-like genes related to Blec genes were described as well (Rogers et al. 2003). The Y region was found to have moderate effects on transplantation (at the level of minor histocompatibility antigens in mammals) and on responses to Rous sarcomas (LePage et al. 2000; Thoraval et al. 2003). Despite being on the same chromosome, the BF/BL region and the Y region were separated by sufficient recombination as to segregate entirely independently. Originally, this separation was attributed to the NOR, but eventually cytogenetics showed that the two loci are separated by a region of repeats, perhaps originally described as sub-telomeric repeats (Delany et al. 2009).
Additional cloning and sequencing filled out some of the regions on either side of the BF/BL region. On one side, additional genes joined the C4 gene as the class III region of the MHC, followed by a pair of CD1 genes, with CD1 genes in mammals found in an MHC paralogous region on a chromosome other than the MHC (Maruoka et al. 2005; Miller et al. 2005; Salomonsen et al. 2005). On the other side, cosmids and BACs were used to define a region with tripartite motif (TRIM) and several other kinds of genes, followed by the BG region which contained primarily BG genes along with a few lectin-like genes related to Blec genes (Ruby et al 2005; Salomonsen et al. 2014; Shiina et al. 2007). Some TRIM genes are found in and around the MHC of mammals, with other TRIM genes located elsewhere. Two other genes known to be in the MHC of mammals had been cloned as chicken cDNAs and used to map their location: the class II A gene known as BLA was found roughly 5 cM from the BF/BL region, and the complement component factor B was located roughly 12 cM from the Y region (Kaufman et al. 1999; Koch et al. 1986; Salomonsen et al. 2003). Neither of these genes has yet been identified in any genomic sequence. Finally, BAC cloning, cytogenetics and sequencing were used to locate two other groups of genes, olfactory receptors and scavenger receptors (along with other genes), the first found in the class I regions and the second not on MHC chromosomes in mammals (Miller et al 2014; Warren et al 2017).
It is perhaps helpful to describe a schism in the names given to these various regions. The original paper reporting the sequence of the BF/BL region (Kaufman et al. 1999a) described it as the chicken MHC, due to the presence of the polymorphic classical class I and class II genes responsible for graft rejection as well as a variety of in vivo and in vitro assays emblematic of the MHC of mammals. However, the organisation of this MHC was arranged differently than in mammals, with the class III region on the outside and the TAP genes in between the two class I genes, an organisation suggested to be ancestral. Moreover, some additional genes were present, including a pair of lectin-like genes (BNK and Blec) whose orthologs are located in the natural killer complex (NKC), sometimes considered to be another MHC paralogous region. CD1 genes, which in mammals are also in an MHC paralogous region, were later found outside of the chicken class III region (Maruoka et al. 2005; Miller et al. 2005; Salomonsen et al. 2005). However, many genes expected from the MHC of mammals were missing. Following up some examples, the C2 gene of the complement cascade, which is found next to factor B and C4 genes in class III region of mammals, was found on chicken chromosome 20 (Kaufman 2013). A more recent example is the TNF (or TNFα) gene, which in mammals is found in the class III region, located on an unplaced scaffold, which in a crow genome is surrounded by many of the same genes as mammals (Rohde et al. 2018). Early on, this led to the notion that the genes missing from the BF/BL region were either absent or moved elsewhere in the chicken genome (Kaufman et al. 1999a, b).
In contrast to this view based on function, some colleagues more influenced by genomics consider that all regions on chicken chromosome 16 with genes associated with any MHC of mammals should rightly be called MHC regions (Miller and Taylor 2016; Miller et al. 2004). Thus, the region around the BF-BL region has been called MHC-B, while regions around the Rfp-Y region have been named MHC-Y. Included in MHC-B and MHC-Y would be those genes which in mammals are located on different chromosomes, such as the lectin-like genes BNK and Blec. Thus far, no comment has been made for those chicken regions with orthologs that in mammals are in the MHC, but in chickens are in different chromosomes; the region on chromosome 20 with C2 gene(s) could be called MHC-C2 in this schema. The differences in nomenclature based on genomics and on function are not of any ultimate importance, although the functional view is that the MHC is defined by the iconic gene systems present, with all other genes being part of the MHC syntenic region in which different genes come and go over long periods of evolutionary history (Kaufman 2018a, b).
Innate immunity genes of the BF/BL region
Most of the genes in the chicken MHC are involved in adaptive immunity. The genes of the classical class I system include the classical class I genes BF1 and BF2, the genes for the transporter associated with antigen presentation TAP1 and TAP2, and the bespoke class I chaperone and peptide editor gene tapasin (also called, TAP binding protein or TAPBP) (Kaufman 2015a, b, 2018b, 1999b). The genes of the class II system include the classical class II B genes BLB1 and BLB2, and the bespoke class II chaperone and peptide editor genes DMA, DMB1 and DMB2 (Parker and Kaufman 2017; Kaufman et al. 1999b). Despite not knowing the exact location for the classical class II A gene BLA, whose nearly monomorphic gene product BLα forms a heterodimer with polymorphic BLβ chains, it has sometimes been considered part of the chicken MHC. Another way to think of it is the same as β2-microglobulin (β2m) gene, found outside of the MHC in all vertebrates except cartilaginous fish, encoding the monomorphic partner of polymorphic class I heavy (α) chains; in this sense, β2m and BLA genes encode average best fits for whatever partner chains are expressed (Kaufman 1999). CD1 heavy chains also bind β2m to form CD1 molecules in both mammals and chickens (Pickel et al. 1990; Salomonsen et al. 2005), which present lipids to T cells as part of the adaptive immune system.
There are other genes in the chicken MHC that are not obviously involved in immunity, such as those that encode the somewhat mysterious serine/threonine kinase called BRD2 (or sometimes RING3) involved in epigenetics (Denis 2010; Thorpe et al. 1996), the enzyme steroid hydroxylase which is a cytochrome P450 monooxygenase with a major role in adrenal steroidogenesis (Haider et al. 2013), the extracellular matrix glycoprotein tenascin B (TNXB) involved in collagen binding and the centromere protein A (CenpA) which is a Histone H3–like nucleosomal protein involved in the kinetochore (which does seem to be involved in autoimmunity, Vakdivia et al. 2009). However, such genes can be involved in resistance and susceptibility to pathogens, such as the interaction of chicken BRD2 with Newcastle disease virus replication (Duan et al. 2020), and some tenascin family members are involved in binding chemokines (Orend and Tucker 2021).
A third group of chicken MHC genes are (potentially) involved with innate immunity. One gene next to the CD1 genes encodes a leukotriene B4 receptor for potent chemoattractants involved in inflammation (Saeki and Yokomizo 2017), some genes are involved with NK cells, one gene with the complement cascade and a couple of genes potentially with γδ T cells.
Studies of NK cells in mammals have shown that many viruses evade cytotoxic T lymphocytes (CTLs) by downregulating class I molecules in one fashion or another, so a host response is to detect the lack of class I molecules by inhibitory NK cell receptors on NK cells. Some viruses encode class I–like molecules as decoys to fool the NK cells with inhibitory receptors; one host counter-strategy is to evolve activating NK cell receptors that bind directly to the decoy molecules. In humans, killer immunoglobulin–like receptors (KIRs) recognize classical class I molecules, including some HLA-A alleles (A3 and A11), some HLA-B alleles (those with Bw4 epitopes) and all HLA-C alleles (divided into C1 and C2 allelic lineages). On most human cell types, HLA-A and HLA-B molecules are much better expressed than HLA-C molecules (Djaoud and Parham 2020). In chickens, the classical class I molecules encoded by the BF2 gene are far better expressed than those expressed by the BF1 gene (Shaw et al. 2007; Wallny et al. 2006). Chicken immunoglobulin-like receptor (ChIR) genes, some of which are very much like inhibitory and activating KIRs (along with others whose gene products bind the immunoglobulin IgY), are found on chromosome 31 in a region considered to be the equivalent of the leukocyte receptor complex (LRC) (Jansen et al. 2016; Laun et al. 2006; Lochner et al. 2010; Straub et al. 2013; Viertlboeck et al. 2005, 2009). There are not many studies examining either CTLs or NK cells in chickens; the limited data suggests that BF2 molecules are generally CTL ligands while BF1 molecules are NK ligands (Kim et al. 2018).
Another set of NK receptor genes are based on extracellular lectin-like domains, including the NKPR1 genes (also known as KLRB1 or CD161), encoded in the NKC. In humans, mice and rats, the NKRP1 genes are located next to (or near) the genes that encode lectin-like ligands: A single NKRP1 gene is next to a single LLT1 gene in humans, while Nkrp1/clr gene pairs are found in mice and rats, with variation driven by immune evasion with at least one virus (Bialoszewska and Malejczyk 2018; Kirkham and Carlyle 2014; Voigt et al. 2007). In chickens, the ortholog of NKRP1 is BNK, which sits next to the LLT1/clr homolog Blec (Kaufman et al. 1999a; Rogers et al. 2005) (with BNK and Blec sometimes referred to as Blec2 and Blec1, respectively, by some bioinformaticians, Shiina et al. 2007). In addition, other genes similar to Blec are found in the BG region and in the Y region (Rogers et al. 2003; Salomonsen et al. 2014) (later confusingly referred to by some bioinformaticians as Blec3 and so on, such that Blec2 in this nomenclature is structurally quite different from Blec1, Blec3 and so on, Shiina et al. 2007). BNK is highly polymorphic, while Blec is monomorphic (Rogers and Kaufman 2008). Contrary to expectations, the BNK molecule from a particular MHC haplotype was not stimulated by Blec (or by BF1 or BF2) molecules; instead, a protease-sensitive ligand was detected on spleen cells from young chickens (Viertlboeck et al. 2008). Whether different members of the Blec family are recognised by different BNK alleles, and whether Blec molecules are protease-sensitive has not yet been determined. As mentioned above, NKRP1-ligand gene pairs are found in the mammalian NKC; the two lectin-like genes found in the syntenic region in the chicken NKC were found to likely have different functions (Chiang et al. 2007; Neulen and Gȍbel 2012). The presence of the BNK-Blec gene pair was suggested as evidence that NKRP1-ligand gene pairs were originally in the MHC and translocated elsewhere during evolution (Rogers et al. 2005). Interestingly, some passerine birds have the BNK-Blec gene pair on the Z chromosome (Ekblom et al. 2011; Rogers and Kaufman 2016), so these genes may translocate with some facility, for reasons as yet unknown.
A single C4 gene marks the start of the class III region of the chicken MHC (Kaufman et al. 1999a, b), but isolation of the C4 protein in chicken blood revealed a completely different N-terminal sequence (Y. Palarasah, J. Kaufman and K Skjødt, unpublished). The cDNAs of both genes were cloned and sequenced, and used to map the second blood C4 to chicken chromosome 1. The two sequences were very different, but the single amino acid position that determines the specificity of thioester reactivity showed that the non-MHC C4 is the equivalent of C4A in mammals and the MHC C4 is the equivalent of C4B; these preferences were confirmed by biochemical assays. Specific monoclonal antibodies were derived and used to show that both genes are expressed in the liver, but that the non-MHC C4 was much better expressed in blood than the MHC C4, for reasons that are still being examined. Upon the first sequence of the chicken genome, the genes around the non-MHC C4 were found to define a syntenic region which was conserved at least to bony fish, with a similar gene in various vertebrates but lacking in mammals (Y. Palarasah, J. Kaufman and K Skjødt, unpublished). Strikingly, the C4A and C4B genes of humans have long been known to be extremely similar and likely the product of a recent duplication, leading to the evolutionary scenario that the genomes in the lineage leading to placental mammals lost the non-MHC C4 gene, duplicated the MHC C4 gene and then selected for the one copy to reproduce the enzymatic specificity of the lost non-MHC C4 gene. As this work was ongoing, another group of bioinformaticians came to similar conclusions about the evolutionary history (Nonaka et al. 2017).
Mammalian C4 molecules have long been known to have central roles in innate immunity, through cell lysis, through opsonisation of bacteria and fungi by macrophages and other myeloloid cells, and through attraction of certain cell types through an anaphylotoxin. However, the C4 genes have been found to exhibit variation that affects not only innate immunity, but also other physiological processes such as autoimmunity and synaptic pruning in the central nervous system (Johnson and Stevens 2018; Wang and Liu 2021). Indeed, single nucleotide polymorphisms (SNPs) in and around the C4 gene are among the top hits in genome-wide association studies (GWAS) with schizophrenia and other mental conditions, as well as sex-linked autoimmunity (Kamitaki et al. 2020; Sekar et al. 2016). The polymorphism of the chicken C4 gene and possible importance in various physiological functions has yet to be investigated.
Although the boundary of the opposite end of the BF/BL region has not been agreed upon, there is a single BG1 gene present along with an authentic butyrophilin gene (Kaufman et al. 1999a, b). Butyrophilin (and butyrophilin-like) genes in the MHC of humans and mice encode chains of heterodimers that play central roles in the maturation, tissue distribution and stimulation of γδ T cells (Di Marco Barros et al. 2016; Jandke et al. 2020; Ventourout et al. 2018). Of course, γδ T cells are created as part of the adaptive immune system, but like NKT cells with semi-invariant T cell receptors, some γδ T cells act in an innate manner, with the name “adaptate” (rather than “inaptive”) being coined to name this usage (Hayday 2019; Hayday and Vantourout 2020). In any case, there are only two butyrophilin genes known to be present in the chicken genome, one on chromosome 28 where it is known as Tvc, a receptor for avian leucosis virus (ALV) subtype C, and the other at the edge of the BF/BL region, with unknown function (Elleder et al. 2005; Kaufman et al. 1999). It has long been speculated that BG genes, which have some features in common with butyrophilins, might encode disulphide-linked dimers on the cell surface that play roles for γδ T cells similar to butyrophilins in mammals. Both the extracellular V domain and the long cytoplasmic tail formed of heptad repeats in BG molecules are polymorphic, but only the variation in the cytoplasmic tail seems to have been selected (Chattaway et al. 2016; Chen et al. 2018). A coiled-coil protein that affects actin-myosin interactions has been shown to originate in the cytoplasmic tail of a BG gene (Bikle et al. 1996). Also, a recombination event in the cytoplasmic tail of the BG1 gene has been identified and proposed to have a significant effect on resistance to tumours induced by Marek’s disease virus (Goto et al. 2009). Whether the BG1 chain might form a heterodimer with the chains encoded by the other BG genes in the BG region is not yet clear.
Innate immunity genes in other regions on chicken chromosome 16
Outside of the BF/BL region is a region with many TRIM genes and then a region of many BG genes, and on the same chromosome is the Y region with at least one expressed non-classical class I gene, some non-polymorphic class II B genes and lectin-like receptor genes, followed by regions with olfactory receptors and scavenger receptors. Some of these genes are likely to be involved in innate immunity.
In mammals, there are at least 60 TRIM genes that function as E3 ligases involved in conjugation of substrates with ubiquitin, small ubiquitin-like modifier (SUMO) or interferon-stimulated protein of 15 kDa (ISG15). TRIM proteins are important for cell cycle, autophagy, development, cancer and autoimmunity (Di Rienzo et al. 2020; Hatakeyama 2017; Ozato et al. 2008). TRIM genes are well-known for acting as viral restriction factors at many levels including uncoating (TRIM5α), transcription (TRIM11, 32) and assembly (TRIM15, 22), and in regulation of interferon signalling (TRIM8), regulation of pattern recognition receptor signalling (TRIM25, 26, 27, 30α, 31, 38, 39 and 40) (Ozato et al. 2008) and mediating intracellular immunity directed by cytoplasmic IgG (TRIM21) (Bottermann and James 2018). All TRIM proteins have the tripartite motif: an N-terminal RING box for interaction with E2 liganses, one or two B-box domains and a coiled-coil region for oligomerisation, followed by various other domains, with most having a PRY-SPRY (also called B30.2) domain for ligand binding (Ozato et al. 2008).
The human MHC has 11 TRIM genes in the class I region, nine in a cluster (TRIM10, 15, 26, 31, 39, 40 and RNF39) and two singletons (TRIM27 and 38). The cluster of TRIM genes is found in the mouse MHC, but Trim27 and Trim38 are located on a different mouse chromosome. The human and mouse genes have some polymorphism, leading to association with various immune responses and autoimmune conditions (Jia et al. 2021). The chicken TRIM region has seven TRIM genes with PRY-SPRY domains, two each identified as having the most similarity to TRIM7, 27 and 39, and one orthologous to TRIM41, as well as three other Zn finger–containing genes (Ruby et al 2005; Shiina et al. 2007). TRIM7 and TRIM41 in the chicken MHC are located next to a guanine nucleotide binding protein homologous to GNB2L1; these three genes are located on human chromosome 5 (Guillemot et al. 1989; Ruby et al 2005). Nothing is yet known about the polymorphism or function of any of these chicken TRIM genes. The comparison of the human, mouse and chicken genomes underscores the fact that the TRIM genes come and go from the MHC syntenic region during evolution.
Next to the TRIM region is the BG region, which has many BG genes as well as a few pairs of a kinesin-like motor protein gene and a lectin-like gene similar to Blec (Salomonsen et al. 2014). As mentioned above, BG genes were discovered as erythrocyte cell surface antigens, but were then found to be expressed on lymphocytes and epithelial cells (Miller et al 1990; Salomonsen et al. 1991, 2014). As members of the B7 gene superfamily, it has long been speculated (Henry et al. 1999) that BG molecules act as their closest relatives in mammals, the butyrophilins and butyrophilin-like molecules, acting as co-stimulators and co-inhibitors for αβ and γδ T cells, and more recently as major regulators of location and activation of γδ T cells in an adaptate manner (Hayday 2019; Arnett and Viney 2014; Hayday and Ventourout 2020; Rhodes et al. 2016).
However, there are major differences in the domain organisation of butyrophilin, butyrophilin-like and BG molecules. The butyrophilin molecules have an extracellular immunoglobulin-like (Ig-like) V domain followed by an Ig-like C2 domain (with V-C2-V-C2 for butyrophilin-like molecules), followed by a transmembrane region, a short region of heptad repeats and a PRY-SPRY (B30.2) domain (Arnett and Viney 2014; Rhodes et al. 2016). In contrast, BG molecules are disulphide-linked dimers of chains with a single Ig-like V domain, a transmembrane region and a very long region of heptad repeats which presumably form a coiled-coil (Kaufman et al. 1990; Miller et al. 1991; Salomonsen et al. 2014). Although most butyrophilins are heterodimers, it is not yet clear whether BG molecules expressed on cells are homodimers or heterodimers, and if the latter, heterodimers of which gene products. Whether additional proteins (for example, orphan B30.2 domains) bind to the intracellular tail to provide function and whether BG molecules are recognised by T cells are both unclear.
An additional challenge is to understand the diversity of the BG genes, which exhibit much copy number variation (CNV) as well as sequence polymorphism selected in the intracellular cytoplasmic tails. While there are two stable singleton BG genes, BG1 in the MHC and BG0 on another chromosome, the BG genes in the BG region are arranged tandemly in the same transcriptional orientation, an arrangement ideal for recombination and deletion leading to expansion and contraction of the BG multigene family, manifested as CNV (Salomonsen et al. 2014). Although there is sequence variation throughout each BG gene, comparison of apparent alleles (that is, BG genes expressed in a particular cell type) suggests that selection for diversity is found only in the intracellular coiled-coils (Chattaway et al. 2016; Chen et al. 2018). This is even more surprising given that the promoter at the 5′ end of the gene determines in which cells a particular BG chain is expressed, but recombination leads to hybrids of the extracellular and intracellular parts of the protein (Salomonsen et al. 2014).
The Y locus is located on chicken chromosome 16, although a region of repeats leads to sufficient recombination that it is genetically unlinked to the B locus, as though they were on different chromosomes. The Y region has one or more class I genes encoding polymorphic YF chains, which bind β2m and have hydrophobic binding sites (Afanassieff et al. 2001; Hee et al. 2010). In mammals, the monomorphic CD1 and MR1 molecules also bind β2m and have binding sites for hydrophobic lipid tails and for bacterial metabolites, respectively (D’Souza et al. 2019; Ogg et al. 2019). The sequences of YF genes are not particularly like either CD1 or MR1. Recently, evidence has been presented that the Y locus is highly repetitive with many YF genes (Zhang et al. 2020). What kind of T cell recognises YF molecules and for what purpose is also a mystery. Also in the Y region are a variable number of class II B genes, apparently monomorphic and related to the polymorphic BLB genes in the chicken MHC, and some lectin-like genes related to Blec, which might be ligands for BNK.
Beyond the Y locus are regions of olfactory receptors and cysteine-rich domain scavenger receptors, as defined by cytogenetics and BAC sequences (Delany et al 2009; Miller et al 2014; Warren et al 2017). There is a huge superfamily of scavenger receptors in mammals that have many functions, including as pattern recognition receptors involved in innate immunity (Canton et al. 2013; Yu et al 2015). Many of the cysteine-rich domain scavenger receptors on chicken chromosome 16 are reported to have immunotyrosine-based inhibitory motifs (ITIMs) that are important for signalling (Miller et al 2014). Moreover, some ChIR genes are reported also to be in this region (Miller et al. 2014), although most are located in the LRC on chromosome 31 (Straub et al. 2013).
Conclusions
Although the main focus of research on the BF/BL region of the B locus, the chicken MHC, has been those genes involved in adaptive immunity, there is a long list of genes actually or potentially important in innate immunity, or in those immune responses that combine features of innate immunity within the adaptive system, called by some adaptate. It appears that the simplicity of the adaptive immune system genes on this chicken microchromosome contrasts with the complexity of the genes potentially involved in innate and adaptate immunity. This contrast may be typical for non-mammalian vertebrates, where a single or a dominantly expressed classical class I molecule compares to many non-classical class I and innate immune genes (Kaufman 2018; Ohta et al. 2006; Flajnik et al. 1993; Grimholt et al. 2015; Langavin et al. 2019; Ohta et al. 2006).
A few seminal studies, such as the importance of the BG1 locus for resistance to Marek’s disease (Goto et al. 2009) and the identification of BF1 as an NK ligand (Kim et al. 2018), have provided enlightenment about function beyond just describing the molecular structures at the genomic and protein levels. Otherwise, there are mysteries everywhere to be understood, and the next years may be exciting, as genetic techniques are used to relate infection outcomes to genomic features. Among the more general approaches, the development of SNP typing methods (particularly those that cover the TRIM region, Fulton et al. 2016) may help link structure to function for such innate and adaptate genes. It will be more difficult in the BF/BL region due to the low level of recombination and in the BG region due to the rampant CNV, so there is much work but also potentially exciting results ahead.
References
Afanassieff M, Goto RM, Ha J, Sherman MA, Zhong L, Auffray C, Coudert F, Zoorob R, Miller MM (2001) At least one class I gene in restriction fragment pattern-Y (Rfp-Y), the second MHC gene cluster in the chicken, is transcribed, polymorphic, and shows divergent specialization in antigen binding region. J Immunol 166(5):3324–3333. https://doi.org/10.4049/jimmunol.166.5.3324
Afrache H, Tregaskes CA, Kaufman J (2020) A potential nomenclature for the ImmunoPolymorphism Database (IPD) of chicken MHC genes: progress and problems. Immunogenetics 72(1–2):9–24. https://doi.org/10.1007/s00251-019-01145-6
Arnett HA, Viney JL (2014) Immune modulation by butyrophilins. Nat Rev Immunol 14(8):559–569. https://doi.org/10.1038/nri3715
Bialoszewska A, Malejczyk J (2018) Biological and clinical significance of human NKRP1A/LLT1 receptor/ligand interactions. Crit Rev Immunol 38(6):479–489. https://doi.org/10.1615/CritRevImmunol.2019029559
Bikle DD, Munson S, Komuves L (1996) Zipper protein, a B-G protein with the ability to regulate actin/myosin 1 interactions in the intestinal brush border. J Biol Chem 271(15):9075–9083. https://doi.org/10.1074/jbc.271.15.9075
Bloom SE, Bacon LD (1985) Linkage of the major histocompatibility (B) complex and the nucleolar organizer in the chicken. Assignment to a microchromosome. J Hered 76(3):146–154
Bottermann M, James LC (2018) Intracellular antiviral immunity. Adv Virus Res 100:309–354. https://doi.org/10.1016/bs.aivir.2018.01.002
Briles WE, Briles RW, Taffs RE, Stone HA (1983) Resistance to a malignant lymphoma in chickens is mapped to subregion of major histocompatibility (B) complex. Science 219(4587):977–979. https://doi.org/10.1126/science.6823560
Briles WE, Goto RM, Auffray C, Miller MM (1993) A polymorphic system related to but genetically independent of the chicken major histocompatibility complex. Immunogenetics 37(6):408–414. https://doi.org/10.1007/BF00222464
Briles WE, Stone HA, Cole RK (1977) Marek’s disease: effects of B histocompatibility alloalleles in resistant and susceptible chicken lines. Science 195(4274):193–195. https://doi.org/10.1126/science.831269
Canton J, Neculai D, Grinstein S (2013) Scavenger receptors in homeostasis and immunity. Nat Rev Immunol 13(9):621–634. https://doi.org/10.1038/nri3515
Chattaway J, Ramirez-Valdez RA, Chappell PE, Caesar JJ, Lea SM, Kaufman J (2016) Different modes of variation for each BG lineage suggest different functions. Open Biol 6(9):160188. https://doi.org/10.1098/rsob.160188
Chen L, Fakiola M, Staines K, Butter C, Kaufman J (2018) Functional alleles of chicken BG genes, members of the butyrophilin gene family, in peripheral T cells. Front Immunol 1(9):930. https://doi.org/10.3389/fimmu.2018.00930
Chiang HI, Zhou H, Raudsepp T, Jesudhasan PR, Zhu JJ (2007) Chicken CD69 and CD94/NKG2-like genes in a chromosomal region syntenic to mammalian natural killer gene complex. Immunogenetics 59(7):603–611. https://doi.org/10.1007/s00251-007-0220-z
Delany ME, Robinson CM, Goto RM, Miller MM (2009) Architecture and organization of chicken microchromosome 16: order of the NOR, MHC-Y, and MHC-B subregions. J Hered 100(5):507–514. https://doi.org/10.1093/jhered/esp044
Denis GV (2010) Bromodomain coactivators in cancer, obesity, type 2 diabetes, and inflammation. Discov Med 10(55):489–499
Di Marco BR, Roberts NA, Dart RJ, Vantourout P, Jandke A, Nussbaumer O, Deban L, Cipolat S, Hart R, Iannitto ML, Laing A, Spencer-Dene B, East P, Gibbons D, Irving PM, Pereira P, Steinhoff U, Hayday A (2016) Epithelia use butyrophilin-like molecules to shape organ-specific γδ T cell compartments. Cell 167(1):203-218.e17. https://doi.org/10.1016/j.cell.2016.08.030
Di Rienzo M, Romagnoli A, Antonioli M, Piacentini M, Fimia GM (2020) TRIM proteins in autophagy: selective sensors in cell damage and innate immune responses. Cell Death Differ 27(3):887–902. https://doi.org/10.1038/s41418-020-0495-2
Djaoud Z, Parham P (2020) HLAs, TCRs, and KIRs, a triumvirate of human cell- mediated immunity. Annu Rev Biochem 20(89):717–739. https://doi.org/10.1146/annurev-biochem-011520-102754
D’Souza MP, Adams E, Altman JD, Birnbaum ME, Boggiano C, Casorati G, Chien YH, Conley A, Eckle SBG, Früh K, Gondré-Lewis T, Hassan N, Huang H, Jayashankar L, Kasmar AG, Kunwar N, Lavelle J, Lewinsohn DM, Moody B, Picker L, Ramachandra L, Shastri N, Parham P, McMichael AJ, Yewdell JW (2019) Casting a wider net: immunosurveillance by nonclassical MHC molecules. PLoS Pathog 15(2):e1007567. https://doi.org/10.1371/journal.ppat.1007567
Duan Z, Han Y, Zhou L, Yuan C, Wang Y, Zhao C, Tang H, Chen J (2020) Chicken bromodomain-containing protein 2 interacts with the Newcastle disease virus matrix protein and promotes viral replication. Vet Res 51(1):120. https://doi.org/10.1186/s13567-020-00846-1
Ekblom R, Stapley J, Ball AD, Birkhead T, Burke T, Slate J (2011) Genetic mapping of the major histocompatibility complex in the zebra finch (Taeniopygia guttata). Immunogenetics 63(8):523–530. https://doi.org/10.1007/s00251-011-0525-9
Elleder D, Stepanets V, Melder DC, Senigl F, Geryk J, Pajer P, Plachý J, Hejnar J, Svoboda J, Federspiel MJ (2005) The receptor for the subgroup C avian sarcoma and leukosis viruses, Tvc, is related to mammalian butyrophilins, members of the immunoglobulin superfamily. J Virol 79(16):10408–10419. https://doi.org/10.1128/JVI.79.16.10408-10419.2005
Flajnik MF, Kasahara M, Shum BP, Salter-Cid L, Taylor E, Du Pasquier L (1993) A novel type of class I gene organization in vertebrates: a large family of non- MHC-linked class I genes is expressed at the RNA level in the amphibian Xenopus. EMBO J 12(11):4385–4396
Fulton JE, McCarron AM, Lund AR, Pinegar KN, Wolc A, Chazara O, Bed’Hom B, Berres M, Miller MM (2016) A high-density SNP panel reveals extensive diversity, frequent recombination and multiple recombination hotspots within the chicken major histocompatibility complex B region between BG2 and CD1A1. Genet Sel Evol 48:1. https://doi.org/10.1186/s12711-015-0181-x.
Goto RM, Wang Y, Taylor RL Jr, Wakenell PS, Hosomichi K, Shiina T, Blackmore CS, Briles WE, Miller MM (2009) BG1 has a major role in MHC-linked resistance to malignant lymphoma in the chicken. Proc Natl Acad Sci U S A 106(39):16740–16745. https://doi.org/10.1073/pnas.0906776106
Grimholt U, Tsukamoto K, Azuma T, Leong J, Koop BF, Dijkstra JM (2015) A comprehensive analysis of teleost MHC class I sequences. BMC Evol Biol 6(15):32. https://doi.org/10.1186/s12862-015-0309-1
Guillemot F, Billault A, Auffray C (1989) Physical linkage of a guanine nucleotide- binding protein-related gene to the chicken major histocompatibility complex. Proc Natl Acad Sci U S A 86(12):4594–4598. https://doi.org/10.1073/pnas.86.12.4594
Guillemot F, Billault A, Pourquié O, Béhar G, Chaussé AM, Zoorob R, Kreibich G, Auffray C (1988) A molecular map of the chicken major histocompatibility complex: the class II beta genes are closely linked to the class I genes and the nucleolar organizer. EMBO J 7(9):2775–2785
Haider S, Islam B, D’Atri V, Sgobba M, Poojari C, Sun L, Yuen T, Zaidi M, New MI (2013) Structure-phenotype correlations of human CYP21A2 mutations in congenital adrenal hyperplasia. Proc Natl Acad Sci U S A 110(7):2605–2610. https://doi.org/10.1073/pnas.1221133110
Hatakeyama S (2017) TRIM family proteins: roles in autophagy, immunity, and carcinogenesis. Trends Biochem Sci 42(4):297–311. https://doi.org/10.1016/j.tibs.2017.01.002
Hayday AC (2019) γδ T cell update: adaptate orchestrators of immune surveillance. J Immunol 203(2):311–320. https://doi.org/10.4049/jimmunol.1800934
Hayday AC, Vantourout P (2020) The innate biologies of adaptive antigen receptors. Annu Rev Immunol 26(38):487–510. https://doi.org/10.1146/annurev-immunol-102819-023144
Hee CS, Gao S, Loll B, Miller MM, Uchanska-Ziegler B, Daumke O, Ziegler A (2010) Structure of a classical MHC class I molecule that binds “non-classical” ligands. PLoS Biol 8(12):e1000557. https://doi.org/10.1371/journal.pbio.1000557
Henry J, Miller MM, Pontarotti P (1999) Structure and evolution of the extended B7 family. Immunol Today 20(6):285–288. https://doi.org/10.1016/s0167-5699(98)01418-2
Jandke A, Melandri D, Monin L, Ushakov DS, Laing AG, Vantourout P, East P, Nitta T, Narita T, Takayanagi H, Feederle R, Hayday A (2020) Butyrophilin-like proteins display combinatorial diversity in selecting and maintaining signature intraepithelial γδ T cell compartments. Nat Commun 11(1):3769. https://doi.org/10.1038/s41467-020-17557-y
Jansen CA, van Haarlem DA, Sperling B, van Kooten PJ, de Vries E, Viertlboeck BC, Vervelde L, Göbel TW (2016) Identification of an activating chicken Ig-like receptor recognizing avian influenza viruses. J Immunol 197(12):4696–4703. https://doi.org/10.4049/jimmunol.1600401
Jia X, Zhao C, Zhao W (2021) Emerging roles of MHC class I region-encoded E3 ubiquitin ligases in innate immunity. Front Immunol 10(12):687102. https://doi.org/10.3389/fimmu.2021.687102
Johnson MB, Stevens B (2018) Pruning hypothesis comes of age. Nature 554(7693):438–439. https://doi.org/10.1038/d41586-018-02053-7
Kamitaki N, Sekar A, Handsaker RE, de Rivera H, Tooley K, Morris DL, Taylor KE, Whelan CW, Tombleson P, Loohuis LMO, Schizophrenia Working Group of the Psychiatric Genomics Consortium, Boehnke M, Kimberly RP, Kaufman KM, Harley JB, Langefeld CD, Seidman CE, Pato MT, Pato CN, Ophoff RA, Graham RR, Criswell LA, Vyse TJ, McCarroll SA (2020) Complement genes contribute sex-biased vulnerability in diverse disorders. Nature 582(7813):577–581. https://doi.org/10.1038/s41586-020-2277-x
Kaufman J (1999) Co-evolving genes in MHC haplotypes: the “rule” for nonmammalian vertebrates? Immunogenetics 50(3–4):228–236. https://doi.org/10.1007/s002510050597
Kaufman J (2013) The Avian MHC (Chapter 8). In: Schat KA, Kaiser P, and Kaspers B (eds) Avian immunology, 2nd end, Elsevier, Ltd, p 149–167
Kaufman J (2015) Co-evolution with chicken class I genes. Immunol Rev 267(1):56–71. https://doi.org/10.1111/imr.12321
Kaufman J (2015) What chickens would tell you about the evolution of antigen processing and presentation. Curr Opin Immunol 34:35–42. https://doi.org/10.1016/j.coi.2015.01.001
Kaufman J (2018) Generalists and specialists: a new view of how MHC class I molecules fight infectious pathogens. Trends Immunol 39(5):367–379. https://doi.org/10.1016/j.it.2018.01.001
Kaufman J (2018) Unfinished business: evolution of the MHC and the adaptive immune system of jawed vertebrates. Annu Rev Immunol 26(36):383–409. https://doi.org/10.1146/annurev-immunol-051116-052450
Kaufman J, Jacob J, Shaw I, Walker B, Milne S, Beck S, Salomonsen J (1999) Gene organisation determines evolution of function in the chicken MHC. Immunol Rev 167:101–117. https://doi.org/10.1111/j.1600-065x.1999.tb01385.x
Kaufman J, Milne S, Göbel TW, Walker BA, Jacob JP, Auffray C, Zoorob R, Beck S (1999) The chicken B locus is a minimal essential major histocompatibility complex. Nature 401(6756):923–925. https://doi.org/10.1038/44856
Kaufman J, Salomonsen J, Skjødt K, Thorpe D (1990) Size polymorphism of chicken major histocompatibility complex-encoded B-G molecules is due to length variation in the cytoplasmic heptad repeat region. Proc Natl Acad Sci U S A 87(21):8277–8281. https://doi.org/10.1073/pnas.87.21.8277
Kaufman J, Völk H, Wallny HJ (1995) A “minimal essential Mhc” and an “unrecognized Mhc”: two extremes in selection for polymorphism. Immunol Rev 143:63–88. https://doi.org/10.1111/j.1600-065x.1995.tb00670.x
Kim T, Hunt HD, Parcells MS, van Santen V, Ewald SJ (2018) Two class I genes of the chicken MHC have different functions: BF1 is recognized by NK cells while BF2 is recognized by CTLs. Immunogenetics 70(9):599–611. https://doi.org/10.1007/s00251-018-1066-2
Kirkham CL, Carlyle JR (2014) Complexity and diversity of the NKR-P1: Clr (Klrb1:Clec2) recognition systems. Front Immunol 2(5):214. https://doi.org/10.3389/fimmu.2014.00214
Koch C (1986) A genetic polymorphism of the complement component factor B in chickens not linked to the major histocompatibility complex (MHC). Immunogenetics 23(6):364–367. https://doi.org/10.1007/BF00372668
Langevin C, Levraud JP, Boudinot P (2019) Fish antiviral tripartite motif (TRIM) proteins. Fish Shellfish Immunol 86:724–733. https://doi.org/10.1016/j.fsi.2018.12.008
Laun K, Coggill P, Palmer S, Sims S, Ning Z, Ragoussis J, Volpi E, Wilson N, Beck S, Ziegler A, Volz A (2006) The leukocyte receptor complex in chicken is characterized by massive expansion and diversification of immunoglobulin-like loci. PLoS Genet 2(5):e73. https://doi.org/10.1371/journal.pgen.0020073
LePage KT, Miller MM, Briles WE, Taylor RL Jr (2000) Rfp-Y genotype affects the fate of Rous sarcomas in B2B5 chickens. Immunogenetics 51(8–9):751–754. https://doi.org/10.1007/s002510000180
Lochner KM, Viertlboeck BC, Göbel TW (2010) The red jungle fowl leukocyte receptor complex contains a large, highly diverse number of chicken immunoglobulin-like receptor (CHIR) genes. Mol Immunol 47(11–12):1956–1962. https://doi.org/10.1016/j.molimm.2010.05.001
Maruoka T, Tanabe H, Chiba M, Kasahara M (2005) Chicken CD1 genes are located in the MHC: CD1 and endothelial protein C receptor genes constitute a distinct subfamily of class-I-like genes that predates the emergence of mammals. Immunogenetics 57(8):590–600. https://doi.org/10.1007/s00251-005-0016-y
Miller MM, Bacon LD, Hala K, Hunt HD, Ewald SJ, Kaufman J, Zoorob R, Briles WE (2004) 2004 Nomenclature for the chicken major histocompatibility (B and Y) complex. Immunogenetics 56(4):261–279. https://doi.org/10.1007/s00251-004-0682-1
Miller MM, Goto R, Bernot A, Zoorob R, Auffray C, Bumstead N, Briles WE (1994) Two Mhc class I and two Mhc class II genes map to the chicken Rfp-Y system outside the B complex. Proc Natl Acad Sci U S A 91(10):4397–4401. https://doi.org/10.1073/pnas.91.10.4397
Miller MM, Goto R, Young S, Chirivella J, Hawke D, Miyada CG (1991) Immunoglobulin variable-region-like domains of diverse sequence within the major histocompatibility complex of the chicken. Proc Natl Acad Sci U S A 88(10):4377–4381. https://doi.org/10.1073/pnas.88.10.4377
Miller MM, Goto R, Young S, Liu J, Hardy J (1990) Antigens similar to major histocompatibility complex B-G are expressed in the intestinal epithelium in the chicken. Immunogenetics 32(1):45–50. https://doi.org/10.1007/BF01787328
Miller MM, Goto RM, Taylor RL Jr, Zoorob R, Auffray C, Briles RW, Briles WE, Bloom SE (1996) Assignment of Rfp-Y to the chicken major histocompatibility complex/NOR microchromosome and evidence for high-frequency recombination associated with the nucleolar organizer region. Proc Natl Acad Sci U S A 93(9):3958–3962. https://doi.org/10.1073/pnas.93.9.3958
Miller MM, Robinson CM, Abernathy J, Goto RM, Hamilton MK, Zhou H, Delany ME (2014) Mapping genes to chicken microchromosome 16 and discovery of olfactory and scavenger receptor genes near the major histocompatibility complex. J Hered 105(2):203–215. https://doi.org/10.1093/jhered/est091
Miller MM, Taylor RL Jr (2016) Brief review of the chicken major histocompatibility complex: the genes, their distribution on chromosome 16, and their contributions to disease resistance. Poult Sci 95(2):375–392. https://doi.org/10.3382/ps/pev379
Miller MM, Wang C, Parisini E, Coletta RD, Goto RM, Lee SY, Barral DC, Townes M, Roura-Mir C, Ford HL, Brenner MB, Dascher CC (2005) Characterization of two avian MHC-like genes reveals an ancient origin of the CD1 family. Proc Natl Acad Sci U S A 102(24):8674–8679. https://doi.org/10.1073/pnas.0500105102
Neulen ML, Göbel TW (2012) Identification of a chicken CLEC-2 homologue, an activating C-type lectin expressed by thrombocytes. Immunogenetics 64(5):389–397. https://doi.org/10.1007/s00251-011-0591-z
Nonaka MI, Terado T, Kimura H, Nonaka M (2017) Evolutionary analysis of two complement C4 genes: ancient duplication and conservation during jawed vertebrate evolution. Dev Comp Immunol 68:1–11. https://doi.org/10.1016/j.dci.2016.11.009
Ogg G, Cerundolo V, McMichael AJ (2019) Capturing the antigen landscape: HLA-E, CD1 and MR1. Curr Opin Immunol 59:121–129. https://doi.org/10.1016/j.coi.2019.07.006
Ohta Y, Goetz W, Hossain MZ, Nonaka M, Flajnik MF (2006) Ancestral organization of the MHC revealed in the amphibian Xenopus. J Immunol 176(6):3674–3685. https://doi.org/10.4049/jimmunol.176.6.3674
Orend G, Tucker RP (2021) Did tenascin-C co-evolve with the general immune system of vertebrates? Front Immunol 12(12):663902. https://doi.org/10.3389/fimmu.2021.663902
Ozato K, Shin DM, Chang TH, Morse HC 3rd (2008) TRIM family proteins and their emerging roles in innate immunity. Nat Rev Immunol 8(11):849–860. https://doi.org/10.1038/nri2413
Parker A, Kaufman J (2017) What chickens might tell us about the MHC class II system. Curr Opin Immunol 46:23–29. https://doi.org/10.1016/j.coi.2017.03.013
Pickel JM, Chen CL, Cooper MD (1990) An avian B-lymphocyte protein associated with beta 2-microglobulin. Immunogenetics 32(1):1–7. https://doi.org/10.1007/BF01787321
Pink JRL, Droege W, Hala K, Miggiano VC, Ziegler A (1977) A three-locus model for the chicken major histocompatibility complex. Immunogenetics 5:203–216
Plachy J, Pink JR, Hála K (1992) Biology of the chicken MHC (B complex). Crit Rev Immunol 12(1–2):47–79
Rhodes DA, Reith W, Trowsdale J (2016) Regulation of immunity by butyrophilins. Annu Rev Immunol 20(34):151–172. https://doi.org/10.1146/annurev-immunol-041015-055435
Rogers SL, Göbel TW, Viertlboeck BC, Milne S, Beck S, Kaufman J (2005) Characterization of the chicken C-type lectin-like receptors B-NK and B-lec suggests that the NK complex and the MHC share a common ancestral region. J Immunol 174(6):3475–3483. https://doi.org/10.4049/jimmunol.174.6.3475
Rogers SL, Kaufman J (2008) High allelic polymorphism, moderate sequence diversity and diversifying selection for B-NK but not B-lec, the pair of lectin-like receptor genes in the chicken MHC. Immunogenetics 60(8):461–475. https://doi.org/10.1007/s00251-008-0307-1
Rogers SL, Kaufman J (2016) Location, location, location: the evolutionary history of CD1 genes and the NKR-P1/ligand systems. Immunogenetics 68(8):499–513. https://doi.org/10.1007/s00251-016-0938-6
Rogers S, Shaw I, Ross N, Nair V, Rothwell L, Kaufman J, Kaiser P (2003) Analysis of part of the chicken Rfp-Y region reveals two novel lectin genes, the first complete genomic sequence of a class I alpha-chain gene, a truncated class II beta-chain gene, and a large CR1 repeat. Immunogenetics 55(2):100–108. https://doi.org/10.1007/s00251-003-0553-1
Rohde F, Schusser B, Hron T, Farkašová H, Plachý J, Härtle S, Hejnar J, Elleder D, Kaspers B (2018) Characterization of chicken tumor necrosis factor-α, a long missed cytokine in birds. Front Immunol 17(9):605. https://doi.org/10.3389/fimmu.2018.00605
Ruby T, Bed’Hom B, Wittzell H, Morin V, Oudin A, Zoorob R (2005) Characterisation of a cluster of TRIM-B30.2 genes in the chicken MHC B locus. Immunogenetics 57(1–2):116–128. https://doi.org/10.1007/s00251-005-0770-x
Saeki K, Yokomizo T (2017) Identification, signaling, and functions of LTB4 receptors. Semin Immunol 33:30–36. https://doi.org/10.1016/j.smim.2017.07.010
Salomonsen J, Chattaway JA, Chan AC, Parker A, Huguet S, Marston DA, Rogers SL, Wu Z, Smith AL, Staines K, Butter C, Riegert P, Vainio O, Nielsen L, Kaspers B, Griffin DK, Yang F, Zoorob R, Guillemot F, Auffray C, Beck S, Skjødt K, Kaufman J (2014) Sequence of a complete chicken BG haplotype shows dynamic expansion and contraction of two gene lineages with particular expression patterns. PLoS Genet 10(6):e1004417. https://doi.org/10.1371/journal.pgen.1004417
Salomonsen J, Dunon D, Skjødt K, Thorpe D, Vainio O, Kaufman J (1991) Chicken major histocompatibility complex-encoded B-G antigens are found on many cell types that are important for the immune system. Proc Natl Acad Sci U S A 88(4):1359–1363. https://doi.org/10.1073/pnas.88.4.1359
Salomonsen J, Marston D, Avila D, Bumstead N, Johansson B, Juul-Madsen H, Olesen GD, Riegert P, Skjødt K, Vainio O, Wiles MV, Kaufman J (2003) The properties of the single chicken MHC classical class II alpha chain (B-LA) gene indicate an ancient origin for the DR/E-like isotype of class II molecules. Immunogenetics 55(9):605–614. https://doi.org/10.1007/s00251-003-0620-7
Salomonsen J, Sørensen MR, Marston DA, Rogers SL, Collen T, van Hateren A, Smith AL, Beal RK, Skjødt K, Kaufman J (2005) Two CD1 genes map to the chicken MHC, indicating that CD1 genes are ancient and likely to have been present in the primordial MHC. Proc Natl Acad Sci U S A 102(24):8668–8673. https://doi.org/10.1073/pnas.0409213102
Sekar A, Bialas AR, de Rivera H, Davis A, Hammond TR, Kamitaki N, Tooley K, Presumey J, Baum M, Van Doren V, Genovese G, Rose SA, Handsaker RE, Schizophrenia Working Group of the Psychiatric Genomics Consortium, Daly MJ, Carroll MC, Stevens B, McCarroll SA (2016) Schizophrenia risk from complex variation of complement component 4. Nature 530(7589):177–183. https://doi.org/10.1038/nature16549
Shaw I, Powell TJ, Marston DA, Baker K, van Hateren A, Riegert P, Wiles MV, Milne S, Beck S, Kaufman J (2007) Different evolutionary histories of the two classical class I genes BF1 and BF2 illustrate drift and selection within the stable MHC haplotypes of chickens. J Immunol 178(9):5744–5752. https://doi.org/10.4049/jimmunol.178.9.5744
Shiina T, Briles WE, Goto RM, Hosomichi K, Yanagiya K, Shimizu S, Inoko H, Miller MM (2007) Extended gene map reveals tripartite motif, C-type lectin, and Ig superfamily type genes within a subregion of the chicken MHC-B affecting infectious disease. J Immunol 178(11):7162–7172. https://doi.org/10.4049/jimmunol.178.11.7162
Simonsen M, Crone M, Koch C, Hála K (1982) The MHC haplotypes of the chicken. Immunogenetics 16(6):513–532. https://doi.org/10.1007/BF00372021 (PMID: 6763913)
Straub C, Neulen ML, Sperling B, Windau K, Zechmann M, Jansen CA, Viertlboeck BC, Göbel TW (2013) Chicken NK cell receptors. Dev Comp Immunol 41(3):324–333. https://doi.org/10.1016/j.dci.2013.03.013
Thoraval P, Afanassieff M, Bouret D, Luneau G, Esnault E, Goto RM, Chaussé AM, Zoorob R, Soubieux D, Miller MM, Dambrine G (2003) Role of nonclassical class I genes of the chicken major histocompatibility complex Rfp-Y locus in transplantation immunity. Immunogenetics 55(9):647–651. https://doi.org/10.1007/s00251-003-0618-1
Thorpe KL, Abdulla S, Kaufman J, Trowsdale J, Beck S (1996) Phylogeny and structure of the RING3 gene. Immunogenetics 44(5):391–396. https://doi.org/10.1007/BF02602785
Trowsdale J, Knight JC (2013) Major histocompatibility complex genomics and human disease. Annu Rev Genomics Hum Genet 14:301–323. https://doi.org/10.1146/annurev-genom-091212-153455
Valdivia MM, Hamdouch K, Ortiz M, Astola A (2009) CENPA a genomic marker for centromere activity and human diseases. Curr Genomics 10(5):326–335. https://doi.org/10.2174/138920209788920985
Vantourout P, Laing A, Woodward MJ, Zlatareva I, Apolonia L, Jones AW, Snijders AP, Malim MH, Hayday AC (2018) Heteromeric interactions regulate butyrophilin (BTN) and BTN-like molecules governing γδ T cell biology. Proc Natl Acad Sci U S A 115(5):1039–1044. https://doi.org/10.1073/pnas.1701237115
Viertlboeck BC, Habermann FA, Schmitt R, Groenen MA, Du Pasquier L, Göbel TW (2005) The chicken leukocyte receptor complex: a highly diverse multigene family encoding at least six structurally distinct receptor types. J Immunol 175(1):385–393. https://doi.org/10.4049/jimmunol.175.1.385
Viertlboeck BC, Schweinsberg S, Schmitt R, Herberg FW, Göbel TW (2009) The chicken leukocyte receptor complex encodes a family of different affinity FcY receptors. J Immunol 182(11):6985–6992. https://doi.org/10.4049/jimmunol.0803060
Viertlboeck BC, Wortmann A, Schmitt R, Plachý J, Göbel TW (2008) Chicken C-type lectin-like receptor B-NK, expressed on NK and T cell subsets, binds to a ligand on activated splenocytes. Mol Immunol 45(5):1398–1404. https://doi.org/10.1016/j.molimm.2007.08.024
Voigt S, Mesci A, Ettinger J, Fine JH, Chen P, Chou W, Carlyle JR (2007) Cytomegalovirus evasion of innate immunity by subversion of the NKR-P1B:Clr-b missing-self axis. Immunity 26:617–627. https://doi.org/10.1016/j.immuni.2007.03.013
Wallny HJ, Avila D, Hunt LG, Powell TJ, Riegert P, Salomonsen J, Skjødt K, Vainio O, Vilbois F, Wiles MV, Kaufman J (2006) Peptide motifs of the single dominantly expressed class I molecule explain the striking MHC-determined response to Rous sarcoma virus in chickens. Proc Natl Acad Sci U S A 103(5):1434–1439. https://doi.org/10.1073/pnas.0507386103
Wang H, Liu M (2021) Complement C4, infections, and autoimmune diseases. Front Immunol 14(12):694928. https://doi.org/10.3389/fimmu.2021.694928
Warren WC, Hillier LW, Tomlinson C, Minx P, Kremitzki M, Graves T, Markovic C, Bouk N, Pruitt KD, Thibaud-Nissen F, Schneider V, Mansour TA, Brown CT, Zimin A, Hawken R, Abrahamsen M, Pyrkosz AB, Morisson M, Fillon V, Vignal A, Chow W, Howe K, Fulton JE, Miller MM, Lovell P, Mello CV, Wirthlin M, Mason AS, Kuo R, Burt DW, Dodgson JB, Cheng HH (2017) A new chicken genome assembly provides insight into avian genome structure. G3 (Bethesda) 7(1):109–117. https://doi.org/10.1534/g3.116.035923
Yu X, Guo C, Fisher PB, Subjeck JR, Wang XY (2015) Scavenger receptors: emerging roles in cancer biology and immunology. Adv Cancer Res 128:309–364. https://doi.org/10.1016/bs.acr.2015.04.004
Zhang J, Goto RM, Miller MM (2020) A simple means for MHC-Y genotyping in chickens using short tandem repeat sequences. Immunogenetics 72(5):325–332. https://doi.org/10.1007/s00251-020-01166-6
Ziegler A, Pink R (1976) Chemical properties of two antigens controlled by the major histocompatibility complex of the chicken. J Biol Chem 251(17):5391–5396
Zoorob R, Bernot A, Renoir DM, Choukri F, Auffray C (1993) Chicken major histocompatibility complex class II B genes: analysis of interallelic and interlocus sequence variance. Eur J Immunol 23(5):1139–1145. https://doi.org/10.1002/eji.1830230524
Acknowledgements
Thanks to the Wellcome Trust for an Investigator Award (110106/Z/15/Z) that supported this work, and to Nicolas Rocos and Maria Danysz for comments on the manuscript before submission.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.