Abstract
A major challenge in evolutionary ecology is to explain extensive natural variation in transmission rates and virulence across pathogens. Host and pathogen ecology is a potentially important source of that variation. Theory of its effects has been developed through the study of non-spatial models, but host population spatial structure has been shown to influence evolutionary outcomes. To date, the effects of basic host and pathogen demography on pathogen evolution have not been thoroughly explored in a spatial context. Here we use simulations to show that space produces novel predictions of the influence of the shape of the pathogen’s transmission–virulence tradeoff, as well as host reproduction and mortality, on the pathogen’s evolutionary stable transmission rate. Importantly, non-spatial models predict that neither the slope of linear transmission–virulence relationships, nor the host reproduction rate will influence pathogen evolution, and that host mortality will only influence it when there is a transmission–virulence tradeoff. We show that this is not the case in a spatial context, and identify the ecological conditions under which spatial effects are most influential. Thus, these results may help explain observed natural variation among pathogens unexplainable by non-spatial models, and provide guidance about when space should be considered. We additionally evaluate the ability of existing analytical approaches to predict the influence of ecology, namely spatial moment equations closed with an improved pair approximation (IPA). The IPA is known to have limited accuracy, but here we show that in the context of pathogens the limitations are substantial: in many cases, IPA incorrectly predicts evolution to pathogen-driven extinction. Despite these limitations, we suggest that the impact of ecology can still be understood within the conceptual framework arising from spatial moment equations, that of “self-shading’’, whereby the spread of highly transmissible pathogens is impeded by local depletion of susceptible hosts.






Similar content being viewed by others
References
Alizon S, Lion S (2011) Within-host parasite cooperation and the evolution of virulence. Proc R Soc Lond Ser B Biol Sci 278(1725):3738–3747
Anderson RM, May RM (1982) Coevolution of hosts and parasites. Parasitology 85:411–426
Bauch CT (2005) The spread of infectious diseases in spatially structured populations: an invasory pair approximation. Math Biosci 198(2):217–237
Best A, Webb S, White A et al (2011) Host resistance and coevolution in spatially structured populations. Proc R Soc Lond Ser B Biol Sci 278(1715):2216–2222
Boerlijst MC, van Ballegooijen WM (2010) Spatial pattern switching enables cyclic evolution in spatial epidemics. PLoS Comput Biol 6(12):e1001030
Bonhoeffer S, Lenski RE, Ebert D (1996) The curse of the pharaoh: the evolution of virulence in pathogens with long living propagules. Proc R Soc Lond Ser B Biol Sci 263(1371):715–721
Boots M, Sasaki A (1999) ‘Small worlds’ and the evolution of virulence: infection occurs locally and at a distance. Proc R Soc Lond Ser B Biol Sci 266(1432):1933–1938
Boots M, Sasaki A (2000) The evolutionary dynamics of local infection and global reproduction in host-parasite interactions. Ecol Lett 3(3):181–185
Boots M, Hudson PJ, Sasaki A (2004) Large shifts in pathogen virulence relate to host population structure. Science 303(5659):842–844
Boots M, Kamo M, Sasaki A (2006) The implications of spatial structure within populations to the evolution of parasites. In: Feng Z, Dieckmann U, Levin SA (eds) Disease evolution: models, concepts, and data analyses. American Mathematical Society, Providence, p 297
Bull JJ (1994) Virulence. Evolution 48(5):1423–1437
Caraco T, Glavanakov S, Li SG et al (2006) Spatially structured superinfection and the evolution of disease virulence. Theor Popul Biol 69(4):367–384
Claessen D, de Roos AM (1995) Evolution of virulence in a host-pathogen system with local pathogen transmission. Oikos 74(3):401–413
Cooper VS, Reiskind MH, Miller JA et al (2002) Timing of transmission and the evolution of virulence of an insect virus. Proc R Soc Lond Ser B Biol Sci 269(1496):1161–1165
Day T (2002) Virulence evolution via host exploitation and toxin production in spore-producing pathogens. Ecol Lett 5(4):471–476
Day T (2003) Virulence evolution and the timing of disease life-history events. Trends Ecol Evol 18(3):113–118
de Aguiar MAM, Rauch EM, Bar-Yam Y (2004) Invasion and extinction in the mean field approximation for a spatial host-pathogen model. J Stat Phys 114(5/6):1417–1451
de Roode JC, Yates AJ, Altizer S (2008) VIrulence–transmission trade-offs and population divergence in virulence in a naturally occuring butterfly parasite. PNAS 105(21):7489–7494
Dieckmann U, Law R (2000) Moment approximations of individual-based models. In: Dieckmann U, Law R, Metz JAJ (eds) The geometry of ecological interactions: simplifying spatial complexity. Cambridge University Press, Cambridge, pp 252–270
Eshelman CM, Vouk R, Stewart JL, Halsne E, Lindsey HA, Schneider S, Gualu M, Dean AM, Kerr B (2010) Unrestricted migration favours virulent pathogens in experimental metapopulations: evolutionary genetics of a rapacious life history. Phil Trans R Soc B Biol Sci 365(1552):2503–2513
Ewald PW (1987) Transmission modes and evolution of the parasitism-mutualism continuum. Ann NY Acad Sci 503:295–306
Ewald PW (1993) Evolution of virulence. Sci Am 268:56–62
Ewald PW (1998) The evolution of virulence and emerging diseases. J Urban Health Bull NY Acad Med 75(3):480–491
Filipe JAN, Maule MM (2003) Analytical methods for predicting the behaviour of population models with general spatial interactions. Math Biosci 183(1):15–35
Frank SA (1996) Models of parasite virulence. Q Rev Biol 71(1):37–78
Gandon S (1998) The curse of the pharaoh hypothesis. Proc R Soc Lond Ser B Biol Sci 265(1405):1545–1552
Gillespie DT (1977) Exact stochastic simulation of coupled chemical reactions. J Phys Chem 81(25):2340–2361
Goodnight C, Rauch E, Sayama H et al (2008) Evolution in spatial predator-prey models and the “prudent predator”: the inadequacy of steady-state organism fitness and the concept of individual and group selection. Complexity 13(5):23–44
Hansen SK, Rainey PB, Haagensen JAJ et al (2007) Evolution of species interactions in a biofilm community. Nature 445(7127):533–536
Harada Y, Iwasa Y (1994) Lattice population-dynamics for plants with dispersing seeds and vegetative propagation. Res Popul Ecol 36(2):237–249
Haraguchi Y, Sasaki A (2000) The evolution of parasite virulence and transmission rate in a spatially structured population. J Theor Biol 203(2):85–96
Johnson CR, Boerlijst MC (2002) Selection at the level of the community: the importance of spatial structure. Trends Ecol Evol 17(2):83–90
Johnson CR, Seinen I (2002) Selection for restraint in competitive ability in spatial competition systems. Proc R Soc Lond Ser B Biol Sci 269:655–663
Kamo M, Boots M (2004) The curse of the pharaoh in space: free-living infectious stages and the evolution of virulence in spatially explicit populations. J Theor Biol 231(3):435–441
Kamo M, Boots M (2006) The evolution of parasite dispersal, transmission, and virulence in spatial host populations. Evol Ecol 8(7):1333–1347
Kamo M, Sasaki A, Boots M (2007) The role of trade-off shapes in the evolution of parasites in spatial host populations: an approximate analytical approach. J Theor Biol 244(4):588–596
Kerr B, Godfrey-Smith P (2002) Individualist and multi-level perspectives on selection in structured populations. Biol Philos 17(4):477–517
Kerr B, Neuhauser C, Bohannan BJM et al (2006) Local migration promotes competitive restraint in a host-pathogen ‘tragedy of the commons’. Nature 442(7098):75–78
Le Galliard JF, Ferriere R, Dieckmann U (2003) The adaptive dynamics of altruism in spatially heterogeneous populations. Evolution 57(1):1–17
Levin BR, Bull JJ (1994) Short-sighted evolution and the virulence of pathogenic microorganisms. Trends Microbiol 2(3):76–81
Levin S, Pimentel D (1981) Selection of intermediate rates of increase in parasite-host systems. Am Nat 117(3):308–315
Lion S, Boots M (2010) Are parasites ‘‘prudent’’ in space? Ecol Lett 13(10):1245–1255
Lion S, van Baalen M (2008) Self-structuring in spatial evolutionary ecology. Ecol Lett 11(3):277–295
Matsuda H, Ogita N, Sasaki A et al (1992) Statistical mechanics of population: the lattice Lotka-Volterra model. Prog Theor Phys 88(6):1035–1049
May RM, Anderson RM (1982) Parasite-host coevolution. In: Futuyma DJ, Slatkin M (eds) Coevolution. Sinauer Associates, Inc., Sunderland, pp 186–206
Messinger SM, Ostling A (2009) The consequences of spatial structure for the evolution of pathogen transmission rate and virulence. Am Nat 174(4):441–454
Morris AJ (1997) Representing spatial interactions in simple ecological models. Dissertation, University of Warwick
Mosquera J, Adler FR (1998) Evolution of virulence: a unified framework for coinfection and superinfection. J Theor Biol 195(3):293–313
Nowak MA, Sigmund K (2002) Super- and coinfection: the two extremes. In: Dieckmann U, Metz JAJ, Sabelis MW, Sigmund K (eds) Adaptive dynamics of infectious diseases: in pursuit of virulence management. Cambridge University Press, Cambridge, pp 124–149
Ohtsuka K, Konno N, Masuda N et al (2006) Phase diagrams and correlation inequalities of a three-state stochastic epidemic model on the square lattice. Int J Bifurc Chaos 16(12):3687–3693
Osnas EE, Dobson AP (2010) Evolution of virulence when transmission occurs before disease. Biol Lett 6(4):505–508
Peyrard N, Dieckmann U, Franc A (2008) Long-range correlations improve understanding of the influence of network structure on contact dynamics. Theor Popul Biol 73(3):383–394
Prado F, Kerr B (2008) The evolution of restraint in bacterial biofilms under nontransitive competition. Evolution 62(3):538–548
Rauch EM, Sayama H, Bar-Yam Y (2002) Relationship between measures of fitness and time scale in evolution. Phys Rev Lett 88(22):1–4
Roche B, Drake JM, Rohani P (2011) The curse of the Pharaoh revisited: evolutionary bi-stability in environmentally transmitted pathogens. Ecol Lett 14(4):569–575
Sato K, Matsuda H, Sasaki A (1994) Pathogen invasion and host extinction in lattice structured populations. J Math Biol 32(3):251–268
Slobodkin LB (1974) Prudent predation does not require group selection. Am Nat 108(963):665–678
van Baalen M, Rand DA (1998) The unit of selection in viscous populations and the evolution of altruism. J Theor Biol 193(4):631–648
van Baalen M, Saebelis MW (1995) The milker-killer dilemma in spatially structured predator–prey interactions. Oikos 74:391–400
Walther BA, Ewald PW (2004) Pathogen survival in the external environment and the evolution of virulence. Biol Rev 79(4):849–869
Webb SD, Keeling MJ, Boots M (2007a) Spatially extended host-parasite interactions: the role of recovery and immunity. Theor Popul Biol 71(2):251–266
Webb SD, Keeling MJ, Boots M (2007b) Host–parasite interactions between the local and the non-spatial: how and when does spatial population structure matter? J Theor Biol 249(1):140–152
White PJ, Norman RA, Hudson PJ (2002) Epidemiological consequences of a pathogen having both virulent and avirulent modes of transmission: the case of rabbit haemorrhagic disease virus. Epidemiol Infect 129(3):665–677
Acknowledgments
We would like to thank the Associate Editor and the reviewers for their comments and the Rackham Graduate School of the University of Michigan for funding.
Author information
Authors and Affiliations
Corresponding author
Appendices
Appendix 1
Inferring ES transmission rate from simulations
By definition, the ES transmission rate is the transmission rate that cannot be invaded by any other transmission rate. Using simulations, one can infer the ES transmission rate from the results of a multitude of discrete pair-wise invasions (see for example Kamo et al. 2007). The ability of one pathogen strain to invade another is taken as the average number of successful invasions out of some number of trial invasions. One can also infer the ES transmission rate from the results of continuous invasion by mutation (see for example Webb et al. 2007a, b). Once the pathogen strain with the ES transmission rate invades, no other mutant strain will be able to invade and the average transmission rate in the population will stabilize at the ES value. The notable, and potentially important difference between the two methods is that the under pair-wise invasion there are never more than two pathogen strains in the population at one time while under continuous invasion many strains are present. To be sure our results were not dependent on our method of inference, we tested three different methods of inferring the ES transmission rate from simulations: pair-wise invasion, continuous invasion by small mutations, and continuous invasion by random mutation.
For the pair-wise invasion method we started the host population with a single resident strain at equilibrium. We then allowed a second pathogen strain with a random transmission rate to invade. Once either the invading or resident strain had been excluded from the host population, the process was repeated. Although this method is not exactly the same as a complete pair-wise invasion analysis, it does capture the essential feature that only two pathogen strains reside in the population at any given time. For the two continuous invasion methods, the ES transmission rate is taken as the average transmission rate of the pathogen population over 2,000 time steps when it is no longer changing over time (a slope between −0.001 and 0.001). For the continuous invasion by random mutation method, we allowed mutant pathogens to differ randomly (rather than only by a small amount) from the parent strain. In all cases, for at least one set of parameters we started the simulation from a range of initial transmission rates. Regardless of the initial transmission rate, the predicted ES transmission rate was the same, suggesting that evolutionary bi-stability is not a fundamental feature of the model.
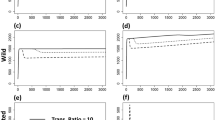
To determine whether the methods yielded different results, we compared the ES transmission rate across multiple realizations of each method for the case of no transmission–virulence tradeoff. The three methods did not give significantly different predictions (Fig. 7). Thus, using the ES transmission rate predicted by the less computationally intensive continuous evolution method is an accurate comparison point for the ES transmission rate predicted by an IPA of the spatial moment equations.
Comparing methods of inferring evolutionary stable transmission rate (ES β) for no transmission–virulence relationship. The left panel shows the average pathogen transmission rate (β) over time for six different stochastic simulations where evolution is modeled as continuous evolution by small mutations. The middle panel shows the same for continuous evolution with randomly sized mutations. The right panel shows the average β over time for simulations consisting of a sequence of pair-wise invasions—there are only ever two pathogen strains in the population at one time. The dashed line shows the ES β inferred by averaging across the last 500 time steps of the continuous evolution by small mutation data. In the analyses here we found no substantial difference in the ES β obtained across these methods. Other parameters are r = 20, α = 1, and d = 1
Appendix 2
Spatial moment equations
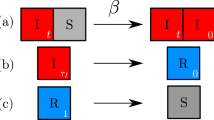
On a regular two-dimensional lattice where sites can be occupied by a susceptible host (S), an infected host (I), or be empty (O), the change in the number of paired states on the lattice (e.g., SO pairs) can be described by the following set of equations.
In these equations, r is the host reproduction rate, d is the natural host death rate, α is the pathogen virulence, β is the pathogen transmission rate, and n is the neighborhood size. Q(X|YZ) is the number of lattice sites in state X in the neighborhood of a site in state Y with a neighboring lattice site in state Z. This set of equations does not incorporate any approximations but is not closed. Further, these equations can be scaled to the size of the lattice and interaction neighborhood to represent the proportion of state pairs on the lattice. Summing Eqs. 11, 12, and 15 yields an equation for the time derivative of the number of susceptible hosts on the lattice, summing Eqs. 14–16 yields the time derivative of the number of infected hosts, and summing Eqs. 12, 13, and 16 yields the time derivative of the number of empty sites.
Appendix 3
Improved pair approximation (IPA)
Equations 11–16 are not closed. Quantities involving state triples (Q(X|YZ)) depend on higher order state combinations. To solve these equations, a moment closure method is required. An ordinary pair approximation (OPA) simply assumes that the presence of a site in state X next to a site in state Y does not depend on the state of other sites in the neighborhood of Y. Thus,
where θ is 1/n and scales the relationship to account for the fact that the maximum Q(X|Y) is 4 but the maximum Q(X|YZ) is 3. However, when host reproduction is local, Q(S|OO) should be smaller than Q(S|O) and Q(S|OS) should be larger than Q(S|O) because susceptible hosts will tend to cluster in space. The improved pair approximation (IPA) takes this clustering into account. When the size of the neighborhood on the lattice is 4, the ratio of Q(S|OO) to Q(S|O) has been shown to be approximately 0.8093. Thus, the IPA assumes that
Using the relationship in Eq. 19 and the following identity relationship
the IPA also assumes that
Appendix 4
Solving for quasi-equilibrium Q(S|I)
The quasi-equilibrium Q(S|I) for a pathogen strain invading a resident strain in equilibrium with the host population must be obtained numerically. Using IPA, for two pathogen strains J and K, where J is the invading strain and K is the resident strain, the time derivatives of paired states involving strain J are:
Parameters are as defined above and θ is 1/n. Numerically solving these equations assuming equilibrium values for all pairs and Q(X|Y) that do not involve strain J (e.g., in Eq. 22, [SS], Q(S|O), and Q(I K |S)), yields the quasi-equilibrium number of state pairs for the invading pathogen. The quasi-equilibrium Q S/I for strain J can then be calculated as:
Appendix 5
Supplementary data and figures
Shape of the transmission–virulence relationship
Figure 8 shows the predicted effect of the shape of the transmission–virulence relationship on the evolutionary stable (ES) transmission rate. For linear relationships (Fig. 8a), simulations and the IPA predict a decrease in the ES transmission rate with increasing relationship steepness (higher values of C in the relationship α = C·β). The non-spatial model predicts no trend with relationship steepness. The IPA predicts higher ES transmission rates than simulations and the predictions diverge for less steep relationships. For concave-down transmission–virulence tradeoffs (Fig. 8b), simulations, the IPA, and the non-spatial model all predict a decrease in the ES transmission rate with increasing tradeoff steepness (higher values of z in the relationship α = C·β z). As with linear relationships, the IPA and non-spatial model predict higher ES transmission rates than simulations and the predictions diverge for less steep tradeoffs (Fig. 8b, lower panel).
ES transmission rates across transmission–virulence relationships. The ES transmission rates (ES β) obtained from simulation (data points) and predictions from improved pair approximation (IPA; thin lines) and a non-spatial model (thick lines) are shown for different transmission–virulence relationships and for two host reproduction rates (r = 30, squares and dashed line; r = 100, triangles and solid line). a Linear transmission–virulence relationships, shape controlled by constant of proportionality C in α = C·β z when z = 1, b concave-down transmission–virulence tradeoffs, shape controlled by tradeoff exponent z in α = C·β z when C = 0.1. The top panel shows ES β and bottom panel shows the percent difference between the spatial and non-spatial predictions
Host reproduction rate
Figure 9 shows the predicted effect of the host reproduction rate on the ES transmission rate for no transmission–virulence relationship across a range of natural host death rates and pathogen virulence. In particular, simulations predict that the ES transmission rate initially decreases and then increases with host reproduction rate for low host death rates (d = 1, d = 2) and pathogen virulence (α = 0, α = 1). For higher host death rates (d = 3, d = 4) and pathogen virulence (α = 2, α = 3), simulations predict that the ES transmission rate increases with host reproduction. In all cases, the non-spatial model and the IPA predict an infinite ES transmission rate (and pathogen-driven extinction) and thus no trend with host reproduction.
Effect of host reproduction rate for no transmission–virulence relationship. The ES transmission rates (ES β) obtained from simulation (circles with dashed spline fit) are shown for a range of host reproduction rates, pathogen virulence, and host natural death rates along with predictions from the non-spatial model and improved pair approximation (IPA; thick lines). The shaded areas are regions where the IPA predicts extinction of the host, pathogen, or both. In the region to the left, the host will go extinct due to an insufficient reproduction rate. In the upper region the pathogen drives the host population and itself to extinction. In the lower region the pathogen cannot invade the host population due to an insufficient transmission rate. In some cases (high d or α) data could not be obtained for lower values of r, likely because the predicted invasion threshold is underestimated. For all cases, the non-spatial model and IPA predict evolution to infinite ES transmission rates and pathogen-driven extinction
Figure 10 shows the predicted effect of the host reproduction rate on the ES transmission rate for linear transmission–virulence relationships. Simulations predict an increase in the ES transmission rate with the host reproduction rate. The non-spatial model predicts an infinite ES transmission rate (and pathogen-driven extinction) and thus no trend with host reproduction. The IPA correctly predicts an intermediate ES transmission rate for all host reproduction rates. However, the trends with host reproduction are qualitatively wrong, especially when the relationship is less steep and the host reproduction rate is low: the ES transmission rate is predicted to be greater than the extinction threshold transmission rate, effectively predicting pathogen-driven extinction.
Effect of host reproduction rate for linear transmission–virulence relationships. The ES transmission rates (ES β) obtained from simulation (circles with dashed spline fit) are shown across a range of host reproduction rates and tradeoff steepness, along with predictions from the non-spatial model (thick solid line) and improved pair approximation (IPA; thin solid line). The shaded areas are as in Fig. 9. Data could not be obtained for lower values of r, likely because the IPA predicted invasion threshold is underestimated. For all graphs, d = 1
Figure 11 shows the predicted effect of the host reproduction rate on the ES transmission rate for concave-down transmission–virulence tradeoffs. Simulations predict an increase in the ES transmission rate with the host reproduction rate. Though the non-spatial model correctly predicts an intermediate transmission rate across host reproduction rates, it predicts no trend with the host reproduction rate. The IPA correctly predicts that the ES transmission rate varies with the host reproduction rate. Further, the predicted trends are qualitatively similar to the trends predicted by simulation. Quantitatively, however, the relative difference between the ES transmission rate predicted by IPA and simulations varies depending on the steepness of transmission–virulence tradeoff as well as the host reproduction rate and natural death rate. As the transmission–virulence tradeoff becomes steeper (higher z in α = C·β z where C = 0.1) the relative difference decreases. In fact, for the steepest tradeoff (α = 0.1·β 1.7), the difference between predictions is nearly 0 across all host reproduction rates. For less steep tradeoffs, the relative difference is highest for low host reproduction rates, but decreases as host reproduction increases.
Effect of host reproduction rate for concave-down transmission–virulence tradeoffs. The ES transmission rates (ES β) obtained from simulation (circles with dashed spline fit, error bars shown when larger than data points) are shown across a range of host reproduction rates and tradeoff steepness, along with predictions from the non-spatial model (thick solid line) and improved pair approximation (IPA; thin solid line). The shaded areas are as in Fig. 9
Host mortality
Figure 12 shows the effect of host mortality (either natural host death rate or virulence) on the ES transmission rate for no transmission–virulence relationship. Simulations predict that the ES transmission rate will increase with increased host mortality. However, the factor of increase declines as the host reproduction rate increases, and for high host reproduction rates the ES transmission rate may actually decrease. The non-spatial model predicts an infinite ES transmission rate for all mortality rates and thus predicts no trend with either the host natural death rate or virulence. Similarly, the IPA incorrectly predicts an infinite ES transmission rate and pathogen-driven extinction regardless of the host natural death rate or virulence. Thus, the ES transmission rate predicted by IPA is both qualitatively and quantitatively inaccurate.
Effect of host mortality on ES transmission rate for no transmission–virulence relationship. The factor of increase in the ES transmission rate (ES β) with a unit increase in the death rate (d; top left panel α = 0 and top right panel α = 1) and virulence (α; bottom left panel d = 1 and bottom right panel d = 2) from simulations (bars) along with predictions from the non-spatial model and improved pair approximation (IPA; thick lines) is shown for a range of host reproduction rates. A value of 1 represents no change in the ES β with death rate or virulence, a value greater than 1 is an increase, and a value less than 1 is a decrease. For example, the first bar in the top left graph (the d = 1–2 bar) indicates that when α = 0 the ES transmission rate is 3.5 times higher when d = 2 than when d = 1. The non-spatial model and IPA predict no change across host reproduction rates
Figure 13 (top two panels and lower left panel) shows the effect of host mortality (natural host death rate) on the ES transmission rate for linear transmission–virulence relationships. Simulations predict that the ES transmission rate will increase with increased host mortality. In some cases the factor of increase is greater for low host reproduction rates, but the trend is not consistent across relationship shapes. For some parameter sets the IPA correctly predicts that the ES transmission rate should increase with host death rate; however, overall the predicted trends are qualitatively and quantitatively inaccurate.
Effect of host mortality on ES transmission rate for transmission–virulence relationships, linear and concave-down tradeoffs. The factor of increase in the ES transmission rate (ES β) with an increase in the death rate from d = 1 to d = 2 is shown for a range of host reproduction rates and tradeoff types: three linear and one concave down (simulation data = bars, predictions from non-spatial model = thick solid line, predictions from IPA = thin solid line). The factor of increase predicted by simulations is always greater than or equal to 1 (dashed line). For the least steep linear tradeoff, the dotted portion of the curve indicates that the IPA predicts an ES β above the extinction threshold for d = 1 or d = 2 at that host reproduction rate
Figure 13 (lower right panel) also shows the effect of host mortality (natural host death rate) on the ES transmission rate for a concave-down transmission–virulence tradeoff. Simulations, the non-spatial model, and the IPA predict that the ES transmission rate will increase with increased host mortality. The non-spatial model predicts the same factor of increase in the ES transmission rate with host mortality across host reproduction rates. In contrast, though the trends are not strong, the IPA predicts a slight decrease in the influence of host death with host reproduction while simulations predict a slight increase with host reproduction.
Rights and permissions
About this article
Cite this article
Messinger, S.M., Ostling, A. The influence of host demography, pathogen virulence, and relationships with pathogen virulence on the evolution of pathogen transmission in a spatial context. Evol Ecol 27, 353–380 (2013). https://doi.org/10.1007/s10682-012-9594-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10682-012-9594-y