Abstract
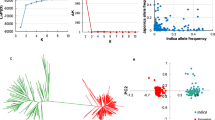
Improvement of rice eating quality is an important objective in current breeding programs. In this study, 130 rice accessions of diverse origin were genotyped using 170 SSR markers to identify marker–trait associations with physicochemical traits on eating quality. Analysis of population structure revealed four subgroups in the population. Linkage disequilibrium (LD) patterns and distributions are of fundamental importance for genome-wide mapping associations. The mean r 2 value for all intrachromosomal loci pairs was 0.0940. LD between linked markers decreased with distance. Marker–trait associations were investigated using the unified mixed-model approach, considering both population structure (Q) and kinship (K). In total, 101 marker–trait associations (p < 0.05) were identified using 52 different SSR markers covering 12 chromosomes. The results suggest that association mapping in rice is a viable alternative to quantitative trait loci mapping, and detection of new marker–trait associations associated with rice eating quality will also provide important information for marker-assisted breeding and functional analysis of rice grain quality.


Similar content being viewed by others
References
Abdurakhmonov IY, Abdukarimov A (2008) Application of association mapping to understanding the genetic diversity of plant germplasm resources. Int J Plant Genomics 2008:574927. doi:10.1155/2008/574927
Abdurakhmonov IY, Kohel RJ, Yu JZ, Pepper AE, Abdullaev AA, Kushanov FN, Salakhutdinov IB, Buriev ZT, Saha S, Scheffler BE, Jenkins JN, Abdukarimov A (2008) Molecular diversity and association mapping of fiber quality traits in exotic G. hirsutum L. germplasm. Genomics 92(6):478–487. doi:10.1016/j.ygeno.2008.07.013
Agrama HA, Eizenga GC (2008) Molecular diversity and genome-wide linkage disequilibrium patterns in a worldwide collection of Oryza sativa and its wild relatives. Euphytica 160(3):339–355. doi:10.1007/s10681-007-9535-y
Agrama HA, Eizenga GC, Yan W (2007) Association mapping of yield and its components in rice cultivars. Mol Breeding 19(4):341–356. doi:10.1007/s11032-006-9066-6
Amarawathi Y, Singh R, Singh A, Singh V, Mohapatra T, Sharma T, Singh N (2008) Mapping of quantitative trait loci for basmati quality traits in rice (Oryza sativa L.). Mol Breeding 21(1):49–65. doi:10.1007/s11032-007-9108-8
Aranzana MJ, Kim S, Zhao K, Bakker E, Horton M, Jakob K, Lister C, Molitor J, Shindo C, Tang C, Toomajian C, Traw B, Zheng H, Bergelson J, Dean C, Marjoram P, Nordborg M (2005) Genome-wide association mapping in Arabidopsis identifies previously known flowering time and pathogen resistance genes. PLoS Genet 1(5):e60. doi:10.1371/journal.pgen.0010060
Bao JS, Xia YW (1999) Genetic control of paste viscosity characteristics in indica rice (Oryza sativa L.). Theor Appl Genet 98(6):1120–1124. doi:10.1007/s001220051175
Bao JS, Sun M, Corke H (2002) Analysis of the genetic behavior of some starch properties in indica rice (Oryza sativa L.): thermal properties, gel texture, swelling volume. Theor Appl Genet 104(2):408–413. doi:10.1007/s001220100688
Bao JS, Corke H, Sun M (2006) Microsatellites, single nucleotide polymorphisms and a sequence tagged site in starch-synthesizing genes in relation to starch physicochemical properties in nonwaxy rice (Oryza sativa L.). Theor Appl Genet 113(7):1185–1196. doi:10.1007/s00122-006-0394-z
Bao JS, Jin L, Xiao P, Shen SQ, Sun M, Corke H (2008) Starch physicochemical properties and their associations with microsatellite alleles of starch-synthesizing genes in a rice RIL population. J Agric Food Chem 56(5):1589–1594. doi:10.1021/jf073128+
Breseghello F, Sorrells ME (2006) Association mapping of kernel size and milling quality in wheat (Triticum aestivum L.) cultivars. Genetics 172(2):1165–1177. doi:10.1534/genetics.105.044586
Cagampang GB, Perez CM, Juliano BO (1973) A gel consistency test for eating quality of rice. J Sci Food Agr 24(12):1589–1594. doi:10.1002/jsfa.2740241214
Cardon LR, Bell JI (2001) Association study designs for complex diseases. Nat Rev Genet 2(2):91–99. doi:10.1038/35052543
Cho YH, Lee SY, Kim SM, Yu JW, Lee JR, Hong HC, Kim JB, Ma KH, Kwon TR, Kang HK, Lee GA, Gwag JG, Kim TS, Park YJ (2008) Amylose, tocopherol, free sugar and fatty acid content in selected mutant lines of Oryza sativa cv. Shindongjin. J Crop Sci Biotechnol 11(3):83–90
Churchill GA, Doerge RW (1994) Empirical threshold values for quantitative trait mapping. Genetics 138(3):963–971
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14(8):2611–2620. doi:10.1111/j.1365-294X.2005.02553.x
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164(4):1567–1587
Fan CC, Yu XQ, Xing YZ, Xu CG, Luo LJ, Zhang QF (2005) The main effects, epistatic effects and environmental interactions of QTLs on the cooking and eating quality of rice in a doubled-haploid line population. Theor Appl Genet 110(8):1445–1452. doi:10.1007/s00122-005-1975-y
Farnir F, Coppieters W, Arranz JJ, Berzi P, Cambisano N, Grisart B, Karim L, Marcq F, Moreau L, Mni M, Nezer C, Simon P, Vanmanshoven P, Wagenaar D, Georges M (2000) Extensive genome-wide linkage disequilibrium in cattle. Genome Res 10(2):220–227. doi:10.1101/gr.10.2.220
Flint-Garcia SA, Thornsberry JM, Buckler ES IV (2003) Structure of linkage disequilibrium in plants. Annu Rev Plant Biol 54(1):357–374. doi:10.1146/annurev.arplant.54.031902.134907
Garris AJ, McCouch SR, Kresovich S (2003) Population structure and its effect on haplotype diversity and linkage disequilibrium surrounding the xa5 locus of rice (Oryza sativa L.). Genetics 165(2):759–769
Gaut BS, Long AD (2003) The lowdown on linkage disequilibrium. Plant Cell 15(7):1502–1506. doi:10.1105/tpc.150730
Gupta PK, Rustgi S, Kulwal PL (2005) Linkage disequilibrium and association studies in higher plants: present status and future prospects. Plant Mol Biol 57(4):461–485. doi:10.1007/s11103-005-0257-z
Hanashiro I, Abe J, Hizukuri S (1996) A periodic distribution of the chain length of amylopectin as revealed by high performance anion-exchange chromatography. Carbohydr Res 283:151–159
Hansen M, Kraft T, Ganestam S, Säll T, Nilsson NO (2001) Linkage disequilibrium mapping of the bolting gene in sea beet using AFLP markers. Genet Res 77(1):61–66. doi:10.1017/S0016672300004857
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2(4):618–620. doi:10.1046/j.1471-8286.2002.00305.x
Hasan M, Friedt W, Pons-Kühnemann J, Freitag NM, Link K, Snowdon RJ (2008) Association of gene-linked SSR markers to seed glucosinolate content in oilseed rape (Brassica napus ssp. napus). Theor Appl Genet 116(8):1035–1049. doi:10.1007/s00122-008-0733-3
He P, Li SG, Qian Q, Ma YQ, Li JZ, Wang WM, Chen Y, Zhu LH (1999) Genetic analysis of rice grain quality. Theor Appl Genet 98(3):502–508. doi:10.1007/s001220051098
Huang FS, Sun ZX, Hu PS, Tang SQ (1998) Present situations and prospects for the research on rice grain quality forming. Chin J Rice Sci 12(3):172–176
Jantaboona J, Siangliwa M, Im-markb S, Jamboonsria W, Vanavichitc A, Toojindaa T (2011) Ideotype breeding for submergence tolerance and cooking quality by marker-assisted selection in rice. Field Crop Res 123:206–213. doi:10.1016/j.fcr.2011.05.001
Jin L, Lu Y, Shao YF, Zhang G, Xiao P, Shen SQ, Corke H, Bao JS (2010) Molecular marker assisted selection for improvement of the eating, cooking and sensory quality of rice (Oryza sativa L.). J Cereal Sci 51:159–164. doi:10.1016/j.jcs.2009.11.007
Juliano BO (1971) A simplified assay for milled-rice amylose. Cereal Sci Today 16:334–338
Juliano BO (1985) Criteria and test for rice grain quality. In: Juliano BO (ed) Rice chemistry and technology. American Association of Cereal Chemists, Saint Paul, pp 443–513
Jun TH, Van K, Kim MY, Lee SH, Walker DR (2008) Association analysis using SSR markers to find QTL for seed protein content in soybean. Euphytica 162(2):179–191. doi:10.1007/s10681-007-9491-6
Kang HJ, Hwang IK, Kim KS, Choi HC (2003) Comparative structure and physicochemical properties of Ilpumbyeo, a high-quality japonica rice, and its mutant, Suweon 464. J Agric Food Chem 51(22):6598–6603. doi:10.1021/jf0344946
Kim KW, Chung HK, Cho GT, Ma KH, Chandrabalan D, Gwag JG, Kim TS, Cho EG, Park YJ (2007) PowerCore: a program applying the advanced M strategy with a heuristic search for establishing core sets. Bioinformatics 23(16):2155–2162. doi:10.1093/bioinformatics/btm313
Koyama ML, Levesley A, Koebner RMD, Flowers TJ, Yeo AR (2001) Quantitative trait loci for component physiological traits determining salt tolerance in rice. Plant Physiol 125(1):406–422. doi:10.1104/pp.125.1.406
Kraakman ATW, Niks RE, Van den Berg PMMM, Stam P, Van Eeuwijk FA (2004) Linkage disequilibrium mapping of yield and yield stability in modern spring barley cultivars. Genetics 168(1):435–446. doi:10.1534/genetics.104.026831
Kraakman ATW, Martínez F, Mussiraliev B, van Eeuwijk FA, Niks RE (2006) Linkage disequilibrium mapping of morphological, resistance, and other agronomically relevant traits in modern spring barley cultivars. Mol Breeding 17(1):41–58. doi:10.1007/s11032-005-1119-8
Lanceras JC, Huang ZL, Naivikul O, Vanavichit A, Ruanjaichon V, Tragoonrung S (2000) Mapping of genes for cooking and eating qualities in Thai jasmine rice (KDML105). DNA Res 7(2):93–101. doi:10.1093/dnares/7.2.93
Larkin PD, McClung AM, Ayres NM, Park WD (2003) The effect of the Waxy locus (granule bound starch synthase) on pasting curve characteristics in specialty rices (Oryza sativa L.). Euphytica 131(2):243–253. doi:10.1023/a:1023962406605
Lestari PJ, Ham TH, Lee HH, Woo MO, Jiang WZ, Chu SH, Kwon SW, Ma KH, Lee JH, Cho YC, Koh HJ (2009) PCR marker-based evaluation of the eating quality of japonica rice (Oryza sativa L.). J Agric Food Chem 57(7):2754–2762. doi:10.1021/jf803804k
Li J, Xiao JH, Grandillo S, Jiang LY, Wan YZ, Deng QY, Yuan LP, McCouch SR (2004) QTL detection for rice grain quality traits using an interspecific backcross population derived from cultivated Asian (O. sativa L.) and African (O. glaberrima S.) rice. Genome 47(4):697–704. doi:10.1139/g04-029
Little RR, Hilder GB, Dawson EH (1958) Differential effect of dilute alkali on 25 varieties of milled white rice. Cereal Chem 35:111–126
Liu KJ, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21(9):2128–2129. doi:10.1093/bioinformatics/bti282
Malysheva-Otto LV, Ganal MW, Roder MS (2006) Analysis of molecular diversity, population structure and linkage disequilibrium in a worldwide survey of cultivated barley germplasm (Hordeum vulgare L.). BMC Genet 7(1):6. doi:10.1186/1471-2156-7-6
Mather DE, Hyes PM, Chalmers KJ, Eglinton J, Matus I, Richardson K, Von Zitzewitz J, Marquez-Cedillo L, Hearnden P, Pal N (2004) Use of SSR marker data to study linkage disequilibrium and population structure in Hordeum vulgare: prospects for association mapping in barley. In: International barley genetics symposium. Brno, Czech Republic, pp 302–307
Mather KA, Caicedo AL, Polato NR, Olsen KM, McCouch S, Purugganan MD (2007) The extent of linkage disequilibrium in rice (Oryza sativa L.). Genetics 177(4):2223–2232. doi:10.1534/genetics.107.079616
Nagamine T, Komae K (1996) Improvement of a method for chain-length distribution analysis of wheat amylopectin. J Chromatogr A 732:255–259
Nakamura Y, Kubo A, Shimamune T, Matsuda T, Harada K, Satoh H (1997) Correlation between activities of starch debranching enzyme and α-polyglucan structure in endosperms of sugary-1 mutants of rice. Plant J 12(1):143–153. doi:10.1046/j.1365-313X.1997.12010143.x
Nordborg M, Borevitz JO, Bergelson J, Berry CC, Chory J, Hagenblad J, Kreitman M, Maloof JN, Noyes T, Oefner PJ, Stahl EA, Weigel D (2002) The extent of linkage disequilibrium in Arabidopsis thaliana. Nat Genet 30(2):190–193. doi:10.1038/ng813
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Rakshit S, Rakshit A, Matsumura H, Takahashi Y, Hasegawa Y, Ito A, Ishii T, Miyashita N, Terauchi R (2007) Large-scale DNA polymorphism study of Oryza sativa and O. rufipogon reveals the origin and divergence of Asian rice. Theor Appl Genet 114(4):731–743. doi:10.1007/s00122-006-0473-1
Remington DL, Thornsberry JM, Matsuoka Y, Wilson LM, Whitt SR, Doebley J, Kresovich S, Goodman MM, Buckler ES (2001) Structure of linkage disequilibrium and phenotypic associations in the maize genome. PNAS USA 98(20):11479–11484. doi:10.1073/pnas.201394398
Risch NJ (2000) Searching for genetic determinants in the new millennium. Nature 405(6788):847–856. doi:10.1038/35015718
Ross-Ibarra J, Morrell PL, Gaut BS (2007) Plant domestication, a unique opportunity to identify the genetic basis of adaptation. PNAS USA 104(Suppl 1):8641–8648. doi:10.1073/pnas.0700643104
Roy JK, Bandopadhyay R, Rustgi S, Balyan HS, Gupta PK (2006) Association analysis of agronomically important traits using SSR, SAMPL and AFLP markers in bread wheat. Curr Sci 90(5):683–689
Schuelke M (2000) An economic method for the fluorescent labeling of PCR fragments. Nat Biotechnol 18(2):233–234. doi:10.1038/72708
Stella A, Boettcher PJ (2004) Optimal designs for linkage disequilibrium mapping and candidate gene association tests in livestock populations. Genetics 166(1):341–350. doi:10.1534/genetics.166.1.341
Stich B, Melchinger AE, Frisch M, Maurer HP, Heckenberger M, Reif JC (2005) Linkage disequilibrium in European elite maize germplasm investigated with SSRs. Theor Appl Genet 111(4):723–730. doi:10.1007/s00122-005-2057-x
Stich B, Maurer HP, Melchinger AE, Frisch M, Heckenberger M, van der Voort JR, Peleman J, Sørensen AP, Reif JC (2006) Comparison of linkage disequilibrium in elite European maize inbred lines using AFLP and SSR markers. Mol Breeding 17(3):217–226. doi:10.1007/s11032-005-5296-2
Sun SY, Hao W, Lin HX (2006) Identification of QTLs for cooking and eating quality of rice grain. Rice Sci 13(3):161–169
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24(8):1596–1599. doi:10.1093/molbev/msm092
Tan YF, Li JX, Yu SB, Xing YZ, Xu CG, Zhang QF (1999) The three important traits for cooking and eating quality of rice grains are controlled by a single locus in an elite rice hybrid, Shanyou 63. Theor Appl Genet 99(3):642–648. doi:10.1007/s001220051279
Thornsberry JM, Goodman MM, Doebley J, Kresovich S, Nielsen D, Buckler ES (2001) Dwarf8 polymorphisms associate with variation in flowering time. Nat Genet 28(3):286–289. doi:10.1038/90135
Tommasini L, Schnurbusch T, Fossati D, Mascher F, Keller B (2007) Association mapping of Stagonospora nodorum blotch resistance in modern European winter wheat varieties. Theor Appl Genet 115(5):697–708. doi:10.1007/s00122-007-0601-6
Umemoto T, Yano M, Satoh H, Shomura A, Nakamura Y (2002) Mapping of a gene responsible for the difference in amylopectin structure between japonica-type and indica-type rice varieties. Theor Appl Genet 104(1):1–8. doi:10.1007/s001220200000
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93(1):77–78. doi:10.1093/jhered/93.1.77
Wada T, Uchimura Y, Ogata T, Tsubone M, Matsue Y (2006) Mapping of QTLs for physicochemical properties in japonica rice. Breeding Sci 56(3):253–260. doi:10.1270/jsbbs.56.253
Wan XY, Wan JM, Su CC, Wang CM, Shen WB, Li JM, Wang HL, Jiang L, Liu SJ, Chen LM, Yasui H, Yoshimura A (2004) QTL detection for eating quality of cooked rice in a population of chromosome segment substitution lines. Theor Appl Genet 110(1):71–79. doi:10.1007/s00122-004-1744-3
Whitt SR, Buckler ES (2003) Using natural allelic diversity to evaluate gene function. In: Grotewald E (ed) Plant functional genomics: methods and protocols. Humana Press, New York, pp 123–140
Yu JM, Pressoir G, Briggs WH, Vroh Bi I, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38(2):203–208. doi:10.1038/ng1702
Yuan PR, Kim HJ, Chen QH, Ju HG, Ji SD, Ahn SN (2010) Mapping QTLs for grain quality using an introgression line population from a cross between Oryza sativa and O. rufipogon. J Crop Sci Biotechnol 13:205–212. doi:10.1007/s12892-010-0094-8
Zhao JJ, Wang XW, Deng B, Lou P, Wu J, Sun RF, Xu ZY, Vromans J, Koornneef M, Bonnema G (2005) Genetic relationships within Brassica rapa as inferred from AFLP fingerprints. Theor Appl Genet 110(7):1301–1314. doi:10.1007/s00122-005-1967-y
Zondervan KT, Cardon LR (2004) The complex interplay among factors that influence allelic association. Nat Rev Genet 5(2):89–100. doi:10.1038/nrg1270
Acknowledgments
This study was supported by a grant from the BioGreen 21 Program (No. PJ009099), Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10681_2012_820_MOESM4_ESM.tif
Supplementary material 4 Model-based ancestry for each of the 130 rice accessions examined based on the 170 SSR markers used to build the Q matrix. The accession numbers correspond to those listed on supplemental Table 1 (TIFF 1883 kb)
Rights and permissions
About this article
Cite this article
Zhao, WG., Chung, JW., Kwon, SW. et al. Association analysis of physicochemical traits on eating quality in rice (Oryza sativa L.). Euphytica 191, 9–21 (2013). https://doi.org/10.1007/s10681-012-0820-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-012-0820-z