Abstract
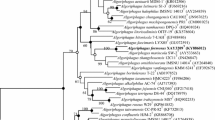
A novel Gram-staining positive, aerobic, rod-shaped, non-motile and yellow-pigmented actinobacterium, designated strain WY83T, was isolated from a marine sediment of Indian Ocean. Strain WY83T grew optimally at 30–35 °C, pH 7–8 and with 0–3% (w/v) NaCl. The predominant menaquinones were MK-10, MK-11 and MK-12, and the major fatty acids were C19:1 ω9c/C19:1 ω11c, anteiso-C15:0, C17:0 3OH, and iso-C16:0. The polar lipids consisted of diphosphatidylglycerol, phosphatidylglycerol and one unidentified glycolipid. The cell-wall peptidoglycan contained lysine as a diamino acid. The DNA G + C content was 72.3 mol%. Phylogenetic analysis based on 16S rRNA gene sequences and ninety-two bacterial core genes indicated that strain WY83T formed an evolutionary lineage with Chryseoglobus frigidaquae JCM 14730T, Chryseoglobus indicus CTD02-10-2T, Yonghaparkia alkaliphila JCM 15138T, Microcella alkaliphila DSM 18851T and Microcella putealis DSM 19627T within the radiation enclosing members of the family Microbacteriaceae. All pairwise percentage of conserved proteins between strain WY83T and the closely related phylogenetic neighbors were greater than 65%. The average nucleotide identity and in silico DNA–DNA hybridization values were both below the thresholds used for the delineation of a new species. On the basis of the evidence presented, strains WY83T, Y. alkaliphila JCM 15138T, C. frigidaquae JCM 14730T, M. alkaliphila DSM 18851T and M. putealis DSM 19627T should belong to different species of the same genus. Strain WY83T represents a novel species of the genus Microcella, for which the name Microcella flavibacter sp. nov. is proposed. The type strain is WY83T (= KCTC 39637T = MCCC 1A07099T). Furthermore, Chryseoglobus frigidaquae, Chryseoglobus indicus, and Yonghaparkia alkaliphila were reclassified as Microcella frigidaquae comb. nov., Microcella indica nom. nov., and Microcella alkalica nom. nov., respectively.


Similar content being viewed by others
Availability of data and material
All data generated or analysed during this study are included in this published article and its supplementary information files.
Code availability
Not applicable.
Change history
05 May 2022
A Correction to this paper has been published: https://doi.org/10.1007/s10482-022-01739-3
23 February 2022
A Correction to this paper has been published: https://doi.org/10.1007/s10482-022-01717-9
Abbreviations
- ANI:
-
Average nucleotide identity
- DAB:
-
2,4-Diaminobutyric acid
- DDH:
-
DNA–DNA hybridization
- MZ2:
-
Modified Zobell 2216E
- ML:
-
Maximum-likelihood
- MP:
-
Maximum-parsimony
- NJ:
-
Neighbour-joining
- POCP:
-
Percentage of conserved proteins
- UBCG:
-
Up to date bacterial core gene
References
Baik KS, Park SC, Kim HJ, Lee KH, Seong CN (2010) Chryseoglobus frigidaquae gen. nov., sp. nov., a novel member of the family Microbacteriaceae. Int J Syst Evol Microbiol 60:1311–1316. https://doi.org/10.1099/ijs.0.016212-0
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY, Medema MH, Weber T (2019) antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res 47:W81–W87. https://doi.org/10.1093/nar/gkz310
Cerny G (1978) Studies on the aminopeptidase test for the distinction of gram-negative from gram-positive bacteria. Appl Microbiol Biotechnol 5:113–122. https://doi.org/10.1007/BF00498805
Chin CS, Alexander DH, Marks P, Klammer AA, Drake J, Heiner C, Clum A, Copeland A, Huddleston J, Eichler EE, Turner SW, Korlach J (2013) Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Methods 10:563–569. https://doi.org/10.1038/nmeth.2474
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376. https://doi.org/10.1007/BF01734359
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416. https://doi.org/10.1093/sysbio/20.4.406
Funabashi M, Funa N, Horinouchi S (2008) Phenolic lipids synthesized by type III polyketide synthase confer penicillin resistance on Streptomyces griseus. J Biol Chem 283:13983–13991. https://doi.org/10.1074/jbc.M710461200
Gogichaeva NV, Alterman MA (2019) Amino Acid Analysis by Means of MALDI TOF Mass Spectrometry or MALDI TOF/TOF Tandem Mass Spectrometry. In: Alterman MA (ed) Amino Acid Analysis Methods and protocols. Humana, New York, NY, pp 17–32 doi: https://doi.org/10.1007/978-1-61779-445-2_12
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM (2007a) DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91
Han SK, Nedashkovskaya OI, Mikhailov VV, Kim SB, Bae KS (2003) Salinibacterium amurskyense gen. nov., sp. nov., a novel genus of the family Microbacteriaceae from the marine environment. Int J Syst Evol Microbiol 53:2061–2066. https://doi.org/10.1099/ijs.0.02627-0
Kaiser P, Geyer R, Surmann P, Fuhrmann H (2012) LC-MS method for screening unknown microbial carotenoids and isoprenoid quinones. J Microbiol Methods 88:28–34. https://doi.org/10.1016/j.mimet.2011.10.001
Krubasik P, Sandmann G (2000) A carotenogenic gene cluster from Brevibacterium linens with novel lycopene cyclase genes involved in the synthesis of aromatic carotenoids. Mol Gen Genet 263:423–432. https://doi.org/10.1007/s004380051186
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, Chichester, pp 115–175
Leifson E (1960) Atlas of Bacterial Flagellation. Academic Press, London
Li WJ, Xu P, Schumann P, Zhang YQ, Pukall R, Xu LH, Stackebrandt E, Jiang CL (2007) Georgenia ruanii sp. nov., a novel actinobacterium isolated from forest soil in Yunnan (China), and emended description of the genus Georgenia. Int J Syst Evol Microbiol 57:1424–1428. https://doi.org/10.1099/ijs.0.64749-0
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60–60. https://doi.org/10.1186/1471-2105-14-60
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microb Meth 2:233–241. https://doi.org/10.1016/0167-7012(84)90018-6
Minnikin DE, Patel PV, Alshamaony L, Goodfellow M (1977) Polar lipid composition in the classification of Nocardia and related bacteria. Int J Syst Bacteriol 27:104–117
Na S-I, Kim YO, Yoon S-H, Ha S-m, Baek I, Chun J (2018) UBCG: Up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J Microbiol 56:280–285. https://doi.org/10.1007/s12275-018-8014-6
Park YH, Suzuki K, Yim DG, Lee KC, Kim E, Yoon J, Kim S, Kho YH, Goodfellow M, Komagata K (1993) Suprageneric classification of peptidoglycan group B actinomycetes by nucleotide sequencing of 5S ribosomal RNA. Antonie Van Leeuwenhoek 64:307–313. https://doi.org/10.1007/bf00873089
Pei S, Xie F, Wang W, Zhang S, Zhang G (2021) Chryseoglobus indicus sp. nov., isolated from deep sea water. International Journal of Systematic and Evolutionary Microbiology 71 doi: https://doi.org/10.1099/ijsem.0.004564
Qin QL, Xie BB, Zhang XY, Chen XL, Zhou BC, Zhou J, Oren A, Zhang YZ (2014) A proposed genus boundary for the prokaryotes based on genomic insights. J Bacteriol 196:2210–2215. https://doi.org/10.1128/JB.01688-14
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sasser M (1990) Identification of Bacteria by Gas Chromatography of Cellular Fatty Acids (technical note no. 101). Newark, DE: MIDI Inc.
Schleifer KH, Kandler O (1972) Peptidoglycan types of bacterial cell walls and their taxonomic implications. Bacteriol Rev 36:407–477
Stackebrandt E, Rainey FA, Ward-Rainey NL (1997) Proposal for a new hierarchic classification system, Actinobacteria classis nov. Int J Syst Bacteriol 47:479–491
Tiago I, Morais PV, da Costa MS, Verissimo A (2006) Microcella alkaliphila sp. nov., a novel member of the family Microbacteriaceae isolated from a non-saline alkaline groundwater, and emended description of the genus Microcella. Int J Syst Evol Microbiol 56:2313–2316. https://doi.org/10.1099/ijs.0.64320-0
Tiago I, Pires C, Mendes V, Morais PV, da Costa M, Verissimo A (2005) Microcella putealis gen. nov., sp. nov., a gram-positive alkaliphilic bacterium isolated from a nonsaline alkaline groundwater. Syst Appl Microbiol 28:479–487. https://doi.org/10.1016/j.syapm.2005.03.004
Tizro P, Choi C, Khanlou N (2019) Sample Preparation for Transmission Electron Microscopy. In: Yong WH (ed) Biobanking: Methods and Protocols. Springer New York, New York, NY, pp 417–424 doi: https://doi.org/10.1007/978-1-4939-8935-5_33
Trujillo ME, Goodfellow M (2012) Genus I. Brevibacterium. In: Goodfellow M et al (eds) Bergey’s Manual of Systematic Bacteriology, 2nd edn. Springer, New York, pp 807–813
Wang J, Li W, Wang H, Lu C (2018) Pentaketide Ansamycin Microansamycins A-I from Micromonospora sp. Reveal Diverse Post-PKS Modifications Org Lett 20:1058–1061. https://doi.org/10.1021/acs.orglett.7b04018
Xu LH, Li WJ, Liu ZH, Jiang CL (2007) Physiological and biochemical characteristics. In: Actinomycete Systematic—Principle, Methods and Practice. Science Press, Beijing, p 45 http://www.irgrid.ac.cn/handle/1471x/260835
Yoon JH, Kang SJ, Schumann P, Oh TK (2006) Yonghaparkia alkaliphila gen. nov., sp. nov., a novel member of the family Microbacteriaceae isolated from an alkaline soil. Int J Syst Evol Microbiol 56:2415–2420. https://doi.org/10.1099/ijs.0.64258-0
Yoon S-H, Ha S-M, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017a) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Yoon S-H, Ha S-m, Lim J, Kwon S, Chun J (2017b) A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek 110:1281–1286. https://doi.org/10.1007/s10482-017-0844-4
Zhi XY, Li WJ, Stackebrandt E (2009) An update of the structure and 16S rRNA gene sequence-based definition of higher ranks of the class Actinobacteria, with the proposal of two new suborders and four new families and emended descriptions of the existing higher taxa. Int J Syst Evol Microbiol 59:589–608. https://doi.org/10.1099/ijs.0.65780-0
Zhou Y, Murphy AC, Samborskyy M, Prediger P, Dias LC, Leadlay PF (2015) Iterative Mechanism of Macrodiolide Formation in the Anticancer Compound Conglobatin. Chem Biol 22:745–754. https://doi.org/10.1016/j.chembiol.2015.05.010
Acknowledgements
We are thankful that Lingqi Ma and Siwen Niu contributed to the isolation of strain wy83 of this manuscript.
Funding
This work was supported by grants from China Ocean Mineral Resources R&D Association (COMRA) Program, No. DY135-B2-01 and the Scientific Research Foundation of Third Institute of Oceanography, MNR, No. 2019011.
Author information
Authors and Affiliations
Contributions
FX, SP carried out most of the experiments; LW conducted physiological experiments; XH and JK conducted genomic analysis. GZ, FX and SP designed the experiments and mainly wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The protologue of Microcella indica was corrected from “nom. nov. (nomen novum)” to “comb. nov. (combinatio nova)”. This error was reversed to comply with Rule 25a of the ICNP. Thus, the original version of the article is restored and a new correction article is published.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xie, F., Pei, S., Huang, X. et al. Microcella flavibacter sp. nov., isolated from marine sediment, and reclassification of Chryseoglobus frigidaquae, Chryseoglobus indicus, and Yonghaparkia alkaliphila as Microcella frigidaquae comb. nov., Microcella indica nom. nov., and Microcella alkalica nom. nov.. Antonie van Leeuwenhoek 114, 2133–2145 (2021). https://doi.org/10.1007/s10482-021-01668-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01668-7