Abstract
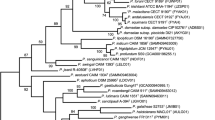
Six strains were isolated from the hemolymph of the spider crab Maja brachydactyla, captured in Spain, and one from a diseased blue mussel, Mytilus edulis. The 16S rRNA gene sequences showed close similarity to the recently described Photobacterium swingsii (98.1 %) and to a lesser degree to Photobacterium aquimaris (97.8 %). MLSA analyses showed a monophyletic group including P. swingsii that form a new subclade. All genomic analyses (Average Nucleotide Identity, Average Amino Acid Identity, and in silico DNA–DNA) clearly separate the strains analysed from P. swingsii with values below the thresholds to delimit a new species. The phenotypic, genotypic and genomic data presented here clearly place these strains as a coherent group within the genus Photobacterium, for which we propose the name Photobacterium sanguinicancri sp. nov. Strain CAIM 1827T (=CECT 7579T, =DSM 24670T) is proposed as the type strain of the species.


Similar content being viewed by others
References
Auch AF, von Jan M, Klenk H-P, Göker M (2010) Digital DNA–DNA hybridization for microbial species delineation by means of genome-to-genome sequence comparison. Stand Genomic Sci 2:117–134. doi:10.4056/sigs.531120
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parrello B, Pusch GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genom 9:75
Cano-Gómez A, Goulden EF, Owens L, Høj L (2010) Vibrio owensii sp. nov., isolated from cultured crustaceans in Australia. FEMS Microbiol Lett 302:175–181. doi:10.1111/j.1574-6968.2009.01850.x
Chimetto L, Cleenwerck I, Thompson CC, Brocchi M, Willems A, De Vos P, Thompson FL (2010) Photobacterium jeanii sp. nov., isolated from corals and zoanthids. Int J Syst Evol Microbiol 60:2843–2848. doi:10.1099/ijs.0.019968-0
Cohn F (1878) Verzameling van stukken betreffende het Geneeskundig Staatstoezicht in Nederland. 126–130
Deep K, Poddar A, Das SK (2014) Photobacterium panuliri sp. nov., an Alkalitolerant Marine Bacterium Isolated from Eggs of Spiny Lobster, Panulirus penicillatus from Andaman Sea. Curr Microbiol 69:660–668. doi:10.1007/s00284-014-0638-0
Eggermont M, Tamanji A, Nevejan N, Bossier P, Sorgeloos P, Defoirdt T (2014) Stimulation of heterotrophic bacteria associated with wild-caught blue mussel (Mytilus edulis) adults results in mass mortality. Aquaculture 431:136–138. doi:10.1016/j.aquaculture.2014.01.014
Gomez-Gil B, Roque A, Lacuesta B, Rotllant G (2010) Diversity of vibrios in the haemolymph of the spider crab Maja brachydactyla. J Appl Microbiol 109(3):918–926. doi:10.1111/j.1365-2672.2010.04718.x
Gomez-Gil B, Thompson CC, Matsumura Y, Sawabe T, Iida T, Christen R, Thompson FL, Sawabe T (2014) The family Vibrionaceae. In: Rosenberg E (ed) The prokaryotes. Springer, Berlin, pp 659–747
González-Castillo A, Enciso-Ibarrra J, Bolán-Mejia MC, Balboa S, Lasa A, Romalde JL, Cabanillas-Beltrán H, Gomez-Gil B (2015) Vibrio mexicanus sp. nov., isolated from a cultured oyster Crassostrea corteziensis. Antonie Van Leeuwenhoek 108:355–364
Hasegawa M, Kishino H, Yano T (1985) Dating the human-ape split by a molecular clock of mitochondrial DNA. Evolution 22:160–174
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267. doi:10.1093/molbev/msj030
Kämpfer P, Kroppenstedt MR (1996) Numerical analysis of fatty acid patterns of coryneform bacteria and related taxa. Can J Microbiol 42:989–1005. doi:10.1139/m96-128
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16s rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Krebs JE, Gale AN, Sontag TC, Keyser VK, Peluso EM, Newman JD (2013) A web-based method to calculate average amino acid identity (AAI) between prokaryotic genomes. Biotechniques
Lee I, Kim YO, Park S-C, Chun J (2015) OrthoANI: An improved algorithm and software for calculating average nucleotide identity. Int J Syst Bacteriol. In press
Lucena T, Ruvira MA, Pascual J, Garay E, Macián MC, Arahal DR, Pujalte MJ (2011) Photobacterium aphoticum sp. nov., isolated from coastal water. Int J Syst Evol Microbiol 61:1579–1584
Macián MC, Ludwig W, Aznar R, Grimont PAD, Schleifer KH, Garay E, Pujalte MJ (2001) Vibrio lentus sp. nov., isolated from Mediterranean oysters. Int J Syst Evol Microbiol 51:1449–1456. doi:10.1099/00207713-51-4-1449
Meier-Kolthoff JP, Auch AF, Klenk H-P, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60. doi:10.1186/1471-2105-14-60
Pascual J, Macián MC, Arahal DR, Garay E, Pujalte MJ (2010) Multilocus sequence analysis of the central clade of the genus Vibrio by using the 16S rRNA, recA, pyrH, rpoD, gyrB, rctB and toxR genes. Int J Syst Evol Microbiol 60:154–165
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106:19126–19131. doi:10.1073/pnas.0906412106
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. USFCC News 20:16
Sawabe T, Kita-Tsukamoto K, Thompson FL (2007) Inferring the evolutionary history of vibrios by means of multilocus sequence analysis. J Bacteriol 189:7932–7936. doi:10.1128/JB.00693-07
Sawabe T, Ogura Y, Matsumura Y, Feng G, Amin AR, Mino S, Nakagawa S, Sawabe T, Kumar R, Fukui Y, Satomi M, Matsushima R, Thompson FL, Gomez-Gil B, Christen R, Maruyama F, Kurokawa K, Hayashi T (2013) Updating the Vibrio clades defined by multilocus sequence phylogeny: proposal of eight new clades, and the description of Vibrio tritonius sp. nov. Front Microbiol 4:1–14
Stackebrandt E, Frederiksen W, Garrity GM, Grimont PAD, Kämpfer P, Maiden MCJ, Nesme X, Rosselló-Mora R, Swings J, Trüper HG, Vauterin L, Ward AC, Whitman WB (2002) Report of the ad hoc committee for the re-evaluation of the species definition in bacteriology. Int J Syst Evol Microbiol 52:1043–1047
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Thompson FL, Iida T, Swings J (2004) Biodiversity of vibrios. Microbiol Mol Biol Rev 68:403–431. doi:10.1128/MMBR.68.3.403-431.2004
Thompson CC, Vicente ACP, Souza RC, Vasconcelos ATR, Vesth T, Alves N, Ussery DW, Iida T, Thompson FL (2009) Genomic taxonomy of Vibrios. BMC Evol Biol 9:258
Varghese NJ, Mukherjee S, Ivanova N, Konstantinidis KT, Mavrommatis K, Kyrpides NC, Pati A (2015) Microbial species delineation using whole genome sequences. Nucleic Acids Res 43:6761–6771
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE, Stackebrandt E, Starr MP, Truper HG (1987) Report of the Ad Hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464. doi:10.1099/00207713-37-4-463
Yoshizawa S, Wada M, Kita-Tsukamoto K, Yokota A, Kogure K (2009) Photobacterium aquimaris sp. nov., a luminous marine bacterium isolated from seawater. Int J Syst Evol Microbiol 59:1438–1442. doi:10.1099/ijs.0.004309-0
Yoshizawa S, Wada M, Yokota A, Kogure K (2010) Vibrio sagamiensis sp. nov., luminous marine bacteria isolated from sea water. J Gen Appl Microbiol 56:499–507
Acknowledgments
This work was partially funded by CONACYT Project CB-2009-01 132328. Thanks to Carmen Bolán Mejía and Julissa Enciso Barrón for technical help.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gomez-Gil, B., Roque, A., Rotllant, G. et al. Photobacterium sanguinicancri sp. nov. isolated from marine animals. Antonie van Leeuwenhoek 109, 817–825 (2016). https://doi.org/10.1007/s10482-016-0681-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-016-0681-x