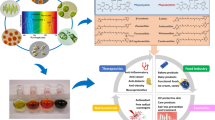
Abstract
Thermus filiformis is an aerobic thermophilic bacterium isolated from a hot spring in New Zealand. The experimental study of the mechanisms of thermal adaptation is important to unveil response strategies of the microorganism to stress. In this study, the main pathways involved on T. filiformis thermoadaptation, as well as, thermozymes with potential biotechnological applications were revealed based on omics approaches. The strategy adopted in this study disclosed that pathways related to the carbohydrate metabolism were affected in response to thermoadaptation. High temperatures triggered oxidative stress, leading to repression of genes involved in glycolysis and the tricarboxylic acid cycle. During heat stress, the glucose metabolism occurred predominantly via the pentose phosphate pathway instead of the glycolysis pathway. Other processes, such as protein degradation, stringent response, and duplication of aminoacyl-tRNA synthetases, were also related to T. filiformis thermoadaptation. The heat-shock response influenced the carotenoid profile of T. filiformis, favoring the synthesis of thermozeaxanthins and thermobiszeaxanthins, which are related to membrane stabilization at high temperatures. Furthermore, antioxidant enzymes correlated with free radical scavenging, including superoxide dismutase, catalase and peroxidase, and metabolites, such as oxaloacetate and α-ketoglutarate, were accumulated at 77 °C.





Similar content being viewed by others
References
Adamberg K, Kask S, Laht T, Paalme T (2003) The effect of temperature and pH on the growth of lactic acid bacteria: a pH-auxostat study. Int J Food Microbiol 85:171–183. doi:10.1016/S0168-1605(02)00537-8
Atkinson GC, Tenson T, Hauryliuk V (2011) The RelA/SpoT Homolog (RSH) superfamily: distribution and functional evolution of ppgpp synthetases and hydrolases across the tree of life. PLoS One. doi:10.1371/journal.pone.0023479
Becker HD, Kern D (1998) Thermus thermophilus: a link in evolution of the tRNA-dependent amino acid amidation pathways. Proc Natl Acad Sci USA 95:12832–12837. doi:10.1073/pnas.95.22.12832
Becker HD, Roy H, Moulinier L et al (2000) Thermus thermophilus contains an eubacterial and an archaebacterial aspartyl-tRNA synthetase. Biochemistry 39:3216–3230. doi:10.1021/bi992573y
Benitez A, Sánchez O (2009) Evaluation of agro-industrial by-products for α-galactosidase production by aspergillus oryzae in submerged culture. In: 9th international conference on chemical and process engineering, pp 1155–1160
Blasco F, Kauffmann I, Schmid RD (2004) CYP175A1 from Thermus thermophilus HB27, the first beta-carotene hydroxylase of the P450 superfamily. Appl Microbiol Biotechnol 64:671–674. doi:10.1007/s00253-003-1529-7
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. doi:10.1093/bioinformatics/btu170
Brüggemann H, Chen C (2006) Comparative genomics of Thermus thermophilus: plasticity of the megaplasmid and its contribution to a thermophilic lifestyle. J Biotechnol 124:654–661. doi:10.1016/j.jbiotec.2006.03.043
Caldas TD, El Yaagoubi A, Richarme G (1998) Chaperone properties of bacterial elongation factor EF-Tu. J Biol Chem 273:11478–11482. doi:10.1074/jbc.273.19.11478
Chen J, Shen J, Solem C, Jensen PR (2013) Oxidative stress at high temperatures in lactococcus lactis due to an insufficient supply of riboflavin. Appl Environ Microbiol 79:6140–6147. doi:10.1128/AEM.01953-13
Cuadros-Inostroza A, Caldana C, Redestig H et al (2009) TargetSearch—a bioconductor package for the efficient preprocessing of GC–MS metabolite profiling data. BMC Bioinform 10:428. doi:10.1186/1471-2105-10-428
Dalebroux ZD, Swanson MS (2012) ppGpp: magic beyond RNA polymerase. Nat Rev Microbiol 10:203–212. doi:10.1038/nrmicro2720
de Souza TA, Soprano AS, de Lira NPV et al (2012) The TAL effector PthA4 interacts with nuclear factors involved in RNA-dependent processes including a HMG protein that selectively binds poly(U) RNA. PLoS One 7:e32305. doi:10.1371/journal.pone.0032305
Developement Core Team R (2015) R: a language and environment for statistical computing. R Found Stat Comput 1:409. doi:10.1007/978-3-540-74686-7
Egas M, Da Costa M, Cowan D, Pires E (1998) Extracellular α-amylase from Thermus filiformis Ork A2: purification and biochemical characterization. Extremophiles 2:23–32. doi:10.1007/s007920050039
Farr SB, Kogoma T (1991) Oxidative stress responses in Escherichia coli and Salmonella typhimurium. Microbiol Rev 55:561–585
Fiorentino A, D’Abrosca B, Pacifico S et al (2006) Spectroscopic identification and antioxidant activity of glucosylated carotenoid metabolites from Cydonia vulgaris fruits. J Agric Food Chem 54:9592–9597. doi:10.1021/jf062125e
Fridovich I (1998) Oxygen toxicity: a radical explanation. J Exp Biol 201:1203–1209
Fu H, Xie B, Fan G et al (2010) Effect of esterification with fatty acid of B-cryptoxanthin on its thermal stability and antioxidant activity by chemiluminescence method. Food Chem 122:602–609. doi:10.1016/j.foodchem.2010.03.019
Fuciños P, Abadín CM, Sanromán A et al (2005) Identification of extracellular lipases/esterases produced by Thermus thermophilus HB27: partial purification and preliminary biochemical characterisation. J Biotechnol 117:233–241. doi:10.1016/j.jbiotec.2005.01.019
Gandhi A, Shah NP (2016) Integrating omics to unravel the stress response mechanisms in probiotic bacteria: approaches, challenges, and prospects. Crit Rev Food Sci Nutr. doi:10.1080/10408398.2015.1136805
Gao H, Wang Y, Liu X et al (2004) Global transcriptome analysis of the heat shock response of Shewanella oneidensis. J Bacteriol 186:7796–7803
Giavalisco P, Li Y, Matthes A et al (2011) Elemental formula annotation of polar and lipophilic metabolites using (13) C, (15) N and (34) S isotope labelling, in combination with high-resolution mass spectrometry. Plant J 68:364–376. doi:10.1111/j.1365-313X.2011.04682.x
Grüning N-M, Du D, Keller MA et al (2014) Inhibition of triosephosphate isomerase by phosphoenolpyruvate in the feedback-regulation of glycolysis. Open Biol 4:130232. doi:10.1098/rsob.130232
Hamid AA, Aiyelaagbe OO, Usman LA et al (2010) Antioxidants: its medicinal and pharmacological applications. Afr J Pure Appl Chemostry 4:142–151
Hanna SL, Sherman NE, Kinter MT, Goldberg JB (2000) Comparison of proteins expressed by Pseudomonas aeruginosa strains representing initial and chronic isolates from a cystic fibrosis patient: an analysis by 2-D gel electrophoresis and capillary column liquid chromatography-tandem mass spectrometry. Microbiology 146:2495–2508
Hara M, Yuan H, Yang Q et al (1999) Stabilization of liposomal membranes by thermozeaxanthins: carotenoid-glucoside esters. Biochim Biophys Acta 1461:147–154
Hauryliuk V, Atkinson GC, Murakami KS et al (2015) Recent functional insights into the role of (p)ppGpp in bacterial physiology. Nat Rev Microbiol 13:298–309. doi:10.1038/nrmicro3448
Hickey DA, Singer GAC (2004) Genomic and proteomic adaptations to growth at high temperature. Genome Biol 5:1–7. doi:10.1186/gb-2004-5-10-117
Hirshfield IN, Bloch PL, Van Bogelen RA, Neidhardt FC (1981) Multiple forms of lysyl-transfer ribonucleic acid synthetase in Escherichia coli. J Bacteriol 146:345–351
Jia B, Liu J, Van Duyet L et al (2015) Proteome profiling of heat, oxidative, and salt stress responses in Thermococcus kodakarensis KOD1. Front Microbiol 6:1–12. doi:10.3389/fmicb.2015.00605
Jozefczuk S, Klie S, Catchpole G et al (2010) Metabolomic and transcriptomic stress response of Escherichia coli. Mol Syst Biol 6:364. doi:10.1038/msb.2010.18
Kanehisa M, Goto S (2000) Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30. doi:10.1093/nar/28.1.27
Kanehisa M, Sato Y, Kawashima M et al (2016) KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44:D457–D462. doi:10.1093/nar/gkv1070
Keller A, Nesvizhskii AI, Kolker E, Aebersold R (2002) Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal Chem 74:5383–5392. doi:10.1021/ac025747h
Kopylova E, Noe L, Touzet H (2012) SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28(24):3211–3217
Kriel A, Bittner AN, Kim SH et al (2012) Direct regulation of GTP homeostasis by (p)ppGpp: a critical component of viability and stress resistance. Mol Cell 48:231–241. doi:10.1016/j.molcel.2012.08.009
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359. doi:10.1038/nmeth.1923
Leveque F, Gazeau M, Fromant M et al (1991) Control of Escherichia coli lysyl-tRNA synthetase expression by anaerobiosis. J Bacteriol 173:7903–7910
Li K, Jiang T, Yu B et al (2012) Transcription elongation factor GreA has functional chaperone activity. PLoS One 7:e47521. doi:10.1371/journal.pone.0047521
Liao Y, Smyth GK, Shi W (2013) The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. doi:10.1093/nar/gkt214
Liu C, Niu Y, Zhou X et al (2015) Streptococcus mutans copes with heat stress by multiple transcriptional regulons modulating virulence and energy metabolism. Sci Rep 5:12929. doi:10.1038/srep12929
Long LH, Halliwell B (2012) The effects of oxaloacetate on hydrogen peroxide generation from ascorbate and epigallocatechin gallate in cell culture media: potential for altering cell metabolism. Biochem Biophys Res Commun 417:446–450. doi:10.1016/j.bbrc.2011.11.136
Mailloux RJ, Bériault R, Lemire J et al (2007) The tricarboxylic acid cycle, an ancient metabolic network with a novel twist. PLoS One 2:e690. doi:10.1371/journal.pone.0000690
Mandelli F, Miranda VS, Rodrigues E, Mercadante AZ (2012a) Identification of carotenoids with high antioxidant capacity produced by extremophile microorganisms. World J Microbiol Biotechnol 28:1781–1790. doi:10.1007/s11274-011-0993-y
Mandelli F, Yamashita F, Pereira J, Mercadante A (2012b) Evaluation of biomass production, carotenoid level and antioxidant capacity produced by Thermus filiformis using fractional factorial design. Braz J Microbiol 43:126–134. doi:10.1590/S1517-83822012000100014
Mandelli F, Franco Cairo JPL, Citadini APS et al (2013) The characterization of a thermostable and cambialistic superoxide dismutase from Thermus filiformis. Lett Appl Microbiol 57:40–46. doi:10.1111/lam.12071
Mandelli F, Ramires BO, Couger MB et al (2015) Draft genome sequence of the thermophile Thermus filiformis ATCC 43280, producer of carotenoid—(di) glucoside-branched fatty acid (di) esters and source of hyperthermostable enzymes of biotechnological interest. Genome Announc 3(3):e00475–15. doi:10.1128/genomeA.00475-15
Mandelstam I (1963) Protein turnover and its function in the economy of the cell. Ann N Y Acad Sci 102:621–636
Marçais G, Kingsford C (2011) A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27:764–770. doi:10.1093/bioinformatics/btr011
Mariutti LRB, Rodrigues E, Mercadante AZ (2013) Carotenoids from Byrsonima crassifolia: identification, quantification and in vitro scavenging capacity against peroxyl radicals. J Food Compos Anal 31:155–160. doi:10.1016/j.jfca.2013.05.005
Martens JH, Barg H, Warren MJ, Jahn D (2002) Microbial production of vitamin B12. Appl Microbiol Biotechnol 58:275–285. doi:10.1007/s00253-001-0902-7
Miyazaki K (2005) A hyperthermophilic laccase from Thermus thermophilus HB27. Extremophiles 9:415–425. doi:10.1007/s00792-005-0458-z
Miyoshi A, Rochat T, Gratadoux J-J et al (2003) Oxidative stress in Lactococcus lactis. Genet Mol Res 2:348–359
Nesvizhskii AI, Keller A, Kolker E, Aebersold R (2003) A statistical model for identifying proteins by tandem mass spectrometry abilities that proteins are present in a sample on the basis. Anal Chem 75:4646–4658. doi:10.1021/ac0341261
Panda T, Gowrishankar BS (2005) Production and applications of esterases. Appl Microbiol Biotechnol 67:160–169. doi:10.1007/s00253-004-1840-y
Pintea A, Bunea A, Andrei S (2013) Impact of esterification on the antioxidant capacity of β-cryptoxanthin. Bull USAMV Anim Sci Biotechnol 70:79–85
Piper PW (1995) The heat shock and ethanol stress responses of yeast exhibit extensive similarity and functional overlap. FEMS Microbiol Lett 134:121–127
Pruneau L, Moumène A, Meyer DF et al (2014) Understanding Anaplasmataceae pathogenesis using “Omics” approaches. Front Cell Infect Microbiol 4:86. doi:10.3389/fcimb.2014.00086
Ralser M, Wamelink MM, Kowald A et al (2007) Dynamic rerouting of the carbohydrate flux is key to counteracting oxidative stress. J Biol 6:10. doi:10.1186/jbiol61
Ramaley RF, Hixson J (1970) Isolation of a nonpigmented, thermophilic bacterium similar to Thermus aquaticus. J Bacteriol 103:526–528
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. doi:10.1093/bioinformatics/btp616
Rodrigues E, Mariutti LRB, Chisté RC, Mercadante AZ (2012a) Development of a novel micro-assay for evaluation of peroxyl radical scavenger capacity: application to carotenoids and structure–activity relationship. Food Chem 135:2103–2111. doi:10.1016/j.foodchem.2012.06.074
Rodrigues E, Mariutti LRB, Mercadante AZ (2012b) Scavenging capacity of marine carotenoids against reactive oxygen and nitrogen species in a membrane-mimicking system. Mar Drugs 10:1784–1798. doi:10.3390/md10081784
Roessner U, Luedemann A, Brust D et al (2001) Metabolic profiling allows comprehensive phenotyping of genetically or environmentally modified plant systems. Plant Cell 13:11–29. doi:10.1105/tpc.13.1.11
Shimizu K (2013) Regulation systems of bacteria such as Escherichia coli in response to nutrient limitation and environmental stresses. Metabolites 4:1–35. doi:10.3390/metabo4010001
Sinnott ML (1990) Catalytic mechanism of enzymic glycosyl transfer. Chem Rev 90:1171–1202. doi:10.1021/cr00105a006
Sun J, Zhou J, Wang Z et al (2015) Multi-omics based changes in response to cadmium toxicity in Bacillus licheniformis A. R Soc Chem 5:7330–7339
Takano H, Kondo M, Usui N et al (2011) Involvement of CarA/LitR and CRP/FNR family transcriptional regulators in light-induced carotenoid production in Thermus thermophilus. J Bacteriol 193:2451–2459. doi:10.1128/JB.01125-10
Topanurak S, Sinchaikul S, Phutrakul S et al (2005) Proteomics viewed on stress response of thermophilic bacterium Bacillus stearothermophilus TLS33. Proteomics 5:3722–3730. doi:10.1002/pmic.200401254
Tsouko E, Khan A, White M et al (2014) Regulation of the pentose phosphate pathway by an androgen receptor-mTOR-mediated mechanism and its role in prostate cancer cell growth. Oncogenesis. doi:10.1038/oncsis.2014.18
Weckwerth W, Wenzel K, Fiehn O (2004) Process for the integrated extraction, identification and quantification of metabolites, proteins and RNA to reveal their co-regulation in biochemical networks. Proteomics 4:78–83. doi:10.1002/pmic.200200500
Wickham H (2009) Elegant graphics for data analysis. Media 35:211. doi:10.1007/978-0-387-98141-3
Willetts NS (1967) Intracellular protein breakdown in non-growing cells of Escherichia coli. Biochem J 103:453–461
Yokoyama A (1995) Thermozeaxanthins, new carotenoid-glycoside-esters from thermophilic eubacterium Thermus thermophilus. Tetrahedron Lett 36:4901–4904. doi:10.1016/00404-0399(50)0881C-
Zhou A, Chen YI, Zane GM et al (2012) Functional characterization of Crp/Fnr-type global transcriptional regulators in Desulfovibrio vulgaris hildenborough. Appl Environ Microbiol 78:1168–1177. doi:10.1128/AEM.05666-11
Acknowledgements
We gratefully acknowledge the provision of time at the NGS and MAS facilities (CTBE and LNBio, respectively) of the National Center for Research in Energy and Materials. This work was financially supported by grants from CNPq (442333/2014-5 and 310186/2014-5) and FAPESP (10/18198-3). FM received a fellowship from CNPq (142685/2010-0).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by A. Driessen.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mandelli, F., Couger, M.B., Paixão, D.A.A. et al. Thermal adaptation strategies of the extremophile bacterium Thermus filiformis based on multi-omics analysis. Extremophiles 21, 775–788 (2017). https://doi.org/10.1007/s00792-017-0942-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-017-0942-2