Abstract
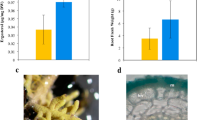
An increased knowledge on the real impacts of ectomycorrhizal symbiosis in forest species is needed to optimize forest sustainable productivity and thus to improve forest services and their capacity to act as carbon sinks. In this study, we investigated the response of an oak species to ectomycorrhizae formation using a proteomics approach complemented by biochemical analysis of carbohydrate levels. Comparative proteome analysis between mycorrhizal and nonmycorrhizal cork oak plants revealed no differences at the foliar level. However, the protein profile of 34 unique oak proteins was altered in the roots. Consistent with the results of the biochemical analysis, the proteome analysis of the mycorrhizal roots suggests a decreasing utilization of sucrose for the metabolic activity of mycorrhizal roots which is consistent with an increased allocation of carbohydrates from the plant to the fungus in order to sustain the symbiosis. In addition, a promotion of protein unfolding mechanisms, attenuation of defense reactions, increased nutrient mobilization from the plant-fungus interface (N and P), as well as cytoskeleton rearrangements and induction of plant cell wall loosening for fungal root accommodation in colonized roots are also suggested by the results. The suggested improvement in root capacity to take up nutrients accompanied by an increase of root biomass without apparent changes in aboveground biomass strongly re-enforces the potential of mycorrhizal inoculation to improve cork oak forest resistance capacity to cope with coming climate change.




Similar content being viewed by others
References
Abdallah C, Valot B, Guillier C, Mounier A, Balliau T, Zivy M, van Tuinen D, Renaut J, Wipf D, Dumas-Gaudot E, Recorbet G (2014) The membrane proteome of Medicago truncatula roots displays qualitative and quantitative changes in response to arbuscular mycorrhizal symbiosis. J Proteomics 108:354–368. doi:10.1016/j.jprot.2014.05.028
Alvarez M, Huygens D, Díaz LM, Villanueva CA, Heyser W, Boeckx P (2012) The spatial distribution of acid phosphatase activity in ectomycorrhizal tissues depends on soil fertility and morphotype, and relates to host plant phosphorus uptake. Plant Cell Environ 35:126–135. doi:10.1111/j.1365-3040.2011.02422.x
Alvarez M, Huygens D, Fernandez C, Gacitua Y, Olivares E, Saavedra I, Alberdi M, Valenzuela E (2009) Effect of ectomycorrhizal colonization and drought on reactive oxygen species metabolism of Nothofagus dombeyi roots. Tree Physiol 29:1047–1057. doi:10.1093/treephys/tpp038
Baptista P, Martins A, Pais MS, Tavares RM, Lino-Neto T (2007) Involvement of reactive oxygen species during early stages of ectomycorrhiza establishment between Castanea sativa and Pisolithus tinctorius. Mycorrhiza 17:185–193. doi:10.1007/s00572-006-0091-4
Bestel-Corre G, Dumas-Gaudot E, Gianinazzi S (2004) Proteomics as a tool to monitor plant-microbe endosymbioses in the rhizosphere. Mycorrhiza 14:1–10. doi:10.1007/s00572-003-0280-3
Boca S, Koestler F, Ksas B, Chevalier A, Leymarie J, Fekete A, Mueller MJ, Havaux M (2014) Arabidopsis lipocalins AtCHL and AtTIL have distinct but overlapping functions essential for lipid protection and seed longevity. Plant Cell Environ 37:368–381. doi:10.1111/pce.12159
Book AJ, Gladman NP, Lee S, Scalf M, Smith LM, Vierstra RD (2010) Affinity purification of the Arabidopsis 26 S proteasome reveals a diverse array of plant proteolytic complexes. J Biol Chem 285:25554–25569. doi:10.1074/jbc.M110.136622
Burgess T, Dell B (1996) Changes in protein biosynthesis during the differentiation of Pisolithus–Eucaliptus grandis ectomycorrhiza. Can J Bot 74:553–560
Cairney JWG (2011) Ectomycorrhizal fungi: the symbiotic route to the root for phosphorus in forest soils. Plant Soil 344:51–71. doi:10.1007/s11104-011-0731-0
Cairney JWG, Chambers SM (1997) Interactions between Pisolithus tinctorius and its hosts: a review of current knowledge. Mycorrhiza 7:117–131. doi:10.1007/s005720050172
Cangahuala-Inocente GC, Da Silva MF, Johnson JM, Manga A, van Tuinen D, Henry C, Lovato PE, Dumas-Gaudot E (2011) Arbuscular mycorrhizal symbiosis elicits proteome responses opposite of P-starvation in SO4 grapevine rootstock upon root colonisation with two Glomus species. Mycorrhiza 21:473–493. doi:10.1007/s00572-010-0352-0
Caplan JL, Zhu X, Mamillapalli P, Marathe R, Anandalakshmi R, Dinesh-Kumar SP (2009) Induced ER-chaperones regulate a novel receptor-like kinase to mediate a viral innate immune response. Cell Host Microbe 19:457–469. doi:10.1016/j.chom.2009.10.005
Charron JB, Ouellet F, Houde M, Sarhan F (2008) The plant apolipoprotein D ortholog protects Arabidopsis against oxidative stress. BMC Plant Biol 8:86. doi:10.1186/1471-2229-8-86
Chen R, Zhao X, Shao Z, Zhu L, He G (2007) Multiple isoforms of UDP-glucose pyrophosphorylase in rice. Physiol Plant 12:725–736. doi:10.1111/j.1399-3054.2007.00865.x
Dickie IA, Montgomery RA, Reich PB, Schnitzer SA (2007) Physiological and phenological responses of oak seedlings to oak forest soil in the absence of trees. Tree Physiol 27:133–140. doi:10.1093/treephys/27.1.133
Dixon RA, Harrison MJ, Paiva NL (1995) The isoflavonoid phytoalexin pathway: from enzymes to genes to transcription factors. Physiol Plant 93:385–392. doi:10.1111/j.1399-3054.1995.tb02243.x
Duddridge JA (1986) The development and ultrastructure of ectomycorrhizas. III. Compatible and incompatible interactions between Suillus grevillei (Klotzsch) Sing. and 11 species of ectomycorrhizal hosts in vitro in the absence of exogenous carbohydrate. New Phytol 103:457–464
Espinosa F, Garrido I, Ortega A, Casimiro I, Alvarez-Tinaut MC (2014) Redox activities and ROS, NO and phenylpropanoids production by axenically cultured intact olive seedling roots after interaction with a mycorrhizal or a pathogenic fungus. PLoS One 9(6):e100132. doi:10.1371/journal.pone.0100132
Felten J, Kohler A, Morin E, Bhalerao RP, Palme K, Martin F, Ditengou FA, Legue V (2009) The ectomycorrhizal fungus Laccaria bicolor stimulates lateral root formation in poplar and Arabidopsis through auxin transport and signalling. Plant Physiol 151:1991–2005. doi:10.1104/pp.109.147231
Fini A, Frangi P, Amoroso G, Piatti R, Faoro M, Bellasio C, Ferrini F (2011) Effect of controlled inoculation with specific mycorrhizal fungi from the urban environment on growth and physiology of containerized shade tree species growing under different water regimes. Mycorrhiza 21:703–719. doi:10.1007/s00572-011-0370-6
Fiorilli V, Catoni M, Miozzi L, Novero M, Accotto GP, Lanfranco L (2009) Global and cell-type gene expression profiles in tomato plants colonized by an arbuscular mycorrhizal fungus. New Phytol 184:975–987. doi:10.1111/j.1469-8137.2009.03031.x
Flores-Monterroso A, Canales J, de la Torre F, Ávila C, Cánovas FM (2013) Identification of genes differentially expressed in ectomycorrhizal roots during the Pinus pinaster–Laccaria bicolor interaction. Planta 237:1637–1650. doi:10.1007/s00425-013-1874-4
Frettinger P, Derory J, Herrmann S, Plomion C, Lapeyrie F, Oelmüller R, Martin F, Buscot F (2007) Transcriptional changes in two types of pre-mycorrhizal roots and in ectomycorrhizas of oak microcuttings inoculated with Piloderma croceum. Planta 225:331–340. doi:10.1007/s00425-006-0355-4
Garcia-Garrido JM, Ocampo J (2002) Regulation of the plant defence response in arbuscular mycorrhizal symbiosis. J Exp Bot 53:1377–1386. doi:10.1093/jexbot/53.373.1377
Gobom J, Nordhoff E, Mirgorodskaya E, Ekman R, Roepstorff P (1999) Sample purification and preparation technique based on nano-scale reversed-phase columns for the sensitive analysis of complex peptide mixtures by matrix-assisted laser desorption/ionization mass spectrometry. J Mass Spectrom 34:105–116. doi:10.1002/(SICI)1096-9888(199902)34:2<105::AID-JMS768>3.0.CO;2-4
Grant WD, West AW (1986) Measurement of ergosterol, diaminopimelic acid and glucosamine in soil: evaluation as indicators of microbial biomass. J Microbiol Methods 6:47–54. doi:10.1016/0167-7012(86)90031-X
Guy CL, Huber JLA, Huber SC (1992) Sucrose phosphate synthase and sucrose accumulation at low temperature. Plant Physiol 100:502–508, 10.1104/pp.100.1.502
Harrison MD, Jones CE, Dameron CT (1999) Copper chaperones: function, structure and copper-binding properties. J Biol Inorg Chem 4:145–153
Hartung W, Ratcliffe RG (2002) Utilization of glycine and serine as nitrogen sources in the roots of Zea mays and Chamaegigas intrepidus. J Exp Bot 53:2305–2314. doi:10.1093/jxb/erf092
Heller G, Adomas A, Li G, Osborne J, van Zyl L, Sederoff R, Finlay RD, Stenlid J, Asiegbu FO (2008) Transcriptional analysis of Pinus sylvestris roots challenged with the ectomycorrhizal fungus Laccaria bicolor. BMC Plant Biol 8:19. doi:10.1186/1471-2229-8-19
Hilbert JL, Costa G, Martin F (1991) Ectomycorrhizin synthesis and polypeptide changes during the early stage of eucalypt mycorrhiza development. Plant Physiol 97:977–984, 10.1104/pp.97.3.977
Hemmingsen SM, Woolford C, van der Vies SM, Tilly K, Dennis DT, Georgopoulos CP, Hendrix RW, Ellis RJ (1988) Homologous plant and bacterial proteins chaperone oligomeric protein assembly. Nature 333:330–334. doi:10.1038/333330a0
Hinsinger P (2001) Bioavailability of soil inorganic P in the rhizosphere as affected by root-induced chemical changes: a review. Plant Soil 237:173–195. doi:10.1023/A:1013351617532
Hughes AL, Powell DW, Bard M, Eckstein J, Barbuch R, Link AJ, Espenshade PJ (2007) Dap1/PGRMC1 binds and regulates cytochrome P450 enzymes. Cell Metab 5:143–149. doi:10.1016/j.cmet.2006.12.009
Inohara N, Nuñez G (2002) ML—a conserved domain involved in innate immunity and lipid metabolism. Trends Biochem Sci 27:219–221, 10.1016/S0968-0004(02)02084-4
Jelenska J, Kang Y, Greenberg JT (2014) Plant pathogenic bacteria target the actin microfilament network involved in the trafficking of disease defense components. BioArchitecture 4:149–153. doi:10.4161/19490992.2014.980662
Johansson T, Le Quere A, Ahren D, Soderstrom B, Erlandsson R, Lundeberg J, Uhlen M, Tunlid A (2004) Transcriptional responses of Paxillus involutus and Betula pendula during formation of ectomycorrhizal root tissue. Mol Plant Microbe Interact 17:202–215. doi:10.1094/MPMI.2004.17.2.202
Kang Y, Jelenska J, Cecchini NM, Li Y, Lee MW, Kovar DR, Greenberg JT (2014) HopW1 from Pseudomonas syringae disrupts the actin cytoskeleton to promote virulence in Arabidopsis. PLoS Pathog 10:e1004232, 10.1371/journal.ppat.1004232
Kleczkowski LA, Geisler M, Ciereszko I, Johansson H (2004) UDP-glucose pyrophosphorylase. An old protein with new tricks. Plant Physiol 134:912–918. doi:10.1104/pp.103.036053
Kuhn H, Küster H, Requena N (2010) Membrane steroid-binding protein 1 induced by a diffusible fungal signal is critical for mycorrhization in Medicago truncatula. New Phytol 185:716–733. doi:10.1111/j.1469-8137.2009.03116.x
Lambertson D, Chen L, Madura K (2003) Investigating the importance of proteasome-interaction for Rad23 function. Curr Genet 42:199–208. doi:10.1007/s00294-002-0350-7
Laurent P, Voiblet C, Tagu D, de Carvalho D, Nehls U, De Bellis R, Balestrini R, Bauw G, Bonfante P, Martin F (1999) A novel class of ectomycorrhiza-regulated cell wall polypeptides in Pisolithus tinctorius. Mol Plant-Microbe Interact 12:862–871. doi:10.1094/MPMI.1999.12.10.862
Le Quéré A, Wright D, Soderstrom B, Tunlid A, Johansson T (2005) Global patterns of gene regulation associated with the development of ectomycorrhiza between birch (Betula pendula Roth.) and Paxillus involutus (Batsch) Fr. Mol Plant Microbe Interact 18:659–673. doi:10.1094/MPMI-18-0659
Li L, Cheng XF, Leshkevich J, Umezawa T, Harding SA, Chiang VL (2001) The last step of syringyl monolignol biosynthesis in angiosperms is regulated by a novel gene encoding sinapyl alcohol dehydrogenase. Plant Cell 13:1567–1586, 10.1105/TPC.010111
Liu JJ, Ekramoddoullah AKM (2006) The family 10 of plant pathogenesis-related proteins: their structure, regulation, and function in response to biotic and abiotic stresses. Physiol Mol Plant Pathol 68:3–13, 10.1016/j.pmpp.2006.06.004
Luo ZB, Janz D, Jiang X, Gobel C, Wildhagen H, Tan Y, Rennenberg H, Feussner I, Polle A (2009) Upgrading root physiology for stress tolerance by ectomycorrhizas: insights from metabolite and transcriptional profiling into reprogramming for stress anticipation. Plant Physiol 151:1902–1917. doi:10.1104/pp.109.143735
Luo Z-B, Lic K, Gaic Y, Göbeld C, Wildhagene H, Jiangc X, Feußnerd I, Rennenberge H, Polle A (2011) The ectomycorrhizal fungus Paxillus involutus modulates leaf physiology of poplar towards improved salt tolerance. Environ Exp Bot 72:304–311, doi:10.1016/j.envexpbot.2011.04.008
Martin F, Aerts A, Ahren D et al (2008) Symbiosis insights from the genome of the mycorrhizal basidiomycete Laccaria bicolor. Nature 452:88–92. doi:10.1038/nature06556
Martin F, Deraruelle C, Hilbert JL (1990) An improved ergosterol assay to estimate fungal biomass in ectomycorrhizas. Mycol Res 94:1059–1064. doi:10.1016/S0953-7562(09)81333-6
Martin F, Kohler A, Murat C et al (2010) Périgord black truffle genome uncovers evolutionary origins and mechanisms of symbiosis. Nature 464:1033–1038. doi:10.1038/nature08867
Martin F, Nehls U (2009) Harnessing ectomycorrhizal genomics for ecological insights. Curr Opin Plant Biol 12:508–515, doi:10.1016/j.pbi.2009.05.007
Martins A, Casimiro A, Pais MS (1997) Influence of mycorrhization on physiological parameters of micropropagated Castanea sativa Mill. plants. Mycorrhiza 7:161–165. doi:10.1007/s005720050176
Meyer Y, Verdoucq L, Vignols F (1999) Plant thioredoxins and glutaredoxins: identity and putative roles. Trends Plant Sci 4:10, 10.1016/S1360-1385(99)01475-2
Miedes E, Vanholme R, Boerjan W, Molina A (2014) The role of the secondary cell wall in plant resistance to pathogens. Front Plant Sci 5:358. doi:10.3389/fpls.2014.00358
Miura K, Jin JB, Hasegawa PM (2007) Sumoylation, a posttranslational regulatory process in plants. Curr Opin Plant Biol 10:495–502, 10.1016/j.pbi.2007.07.002
Monteiro F, Sebastiana M, Pais MS, Figueiredo A (2013) Reference gene selection and validation for the early responses to downy mildew infection in susceptible and resistant Vitis vinifera cultivars. PLoS One 8:e72998, 10.1371/journal.pone.0072998
Müller T, Avolio M, Olivi M, Benjdia M, Rikirsch E, Kasaras A, Fitz M, Chalot M, Wipf D (2007) Nitrogen transport in the ectomycorrhiza association: the Hebeloma cylindrosporum-Pinus pinaster model. Phytochemistry 68:41–51. doi:10.1016/j.phytochem.2006.09.021
Nehls U, Gohringer F, Wittulsky S, Dietz S (2010) Fungal carbohydrate support in the ectomycorrhizal symbiosis: a review. Plant Biol 12:292–301. doi:10.1111/j.1438-8677.2009.00312.x
Novatchkova M, Budhiraja R, Coupland G, Eisenhaber F, Bachmair A (2004) SUMO conjugation in plants. Planta 220:1–8. doi:10.1007/s00425-004-1370-y
Penheiter AR, Klucas RV, Sarath G (1998) Purification and characterization of a soybean root nodule phosphatase expressed in Pichia pastoris. Protein Expr Purif 14:125–130. doi:10.1006/prep.1998.0935
Prasad TK, Stewart CR (1992) cDNA clones encoding Arabidopsis thaliana and Zea mays mitochondrial chaperonin HSP60 and gene expression during seed germination and heat shock. Plant Mol Biol 18:873–885. doi:10.1007/BF00019202
Qiang X, Zechmann B, Reitz MU, Kogel KH, Schäfer P (2012) The mutualistic fungus Piriformospora indica colonizes Arabidopsis roots by inducing an endoplasmic reticulum stress-triggered caspase-dependent cell death. Plant Cell 24:794–809. doi:10.1105/tpc.111.093260
Raasi S, Pickart CM (2003) Rad23 ubiquitin-associated domains (UBA) inhibit 26 S proteasome-catalyzed proteolysis by sequestering lysine 48-linked polyubiquitin chains. J Biol Chem 278:8951–8959. doi:10.1074/jbc.M212841200
Radauer C, Lackner P, Breiteneder H (2008) The Bet v 1 fold: an ancient, versatile scaffold for binding of large, hydrophobic ligands. BMC Evol Biol 8:286. doi:10.1186/1471-2148-8-286
Ray S, Anderson JM, Urmeev FI, Goodwin SB (2003) Rapid induction of a protein disulphide isomerase and defense-related genes in wheat in response to the hemibiotrophic fungal pathogen Mycosphaerella graminicola. Plant Mol Biol 53:701–714. doi:10.1023/B:PLAN.0000019120.74610.52
Rodrigues CI, Maia R, Maguas C (2010) Comparing total nitrogen and crude protein content of green coffee beans (Coffea spp.) from different geographical origins. Coffee Sci 5:197–205
Rose CM, Venkateshwaran M, Volkening JD et al (2012) Rapid phosphoproteomic and transcriptomic changes in the rhizobia legume symbiosis. Mol Cell Proteomics 11:724–744. doi:10.1074/mcp.M112.019208
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C (T) method. Nat Protoc 3:1101–1108. doi:10.1038/nprot.2008.73
Sebastiana M, Figueiredo A, Monteiro F et al (2013a) A possible approach for gel-based proteomic studies in recalcitrant woody plants. SpringerPlus 2:210. doi:10.1186/2193-1801-2-210
Sebastiana M, Pereira V, Alcantara A, Pais M, Silva A (2013b) Ectomycorrhizal inoculation with Pisolithus tinctorius increases the performance of Quercus suber L. (cork oak) nursery and field seedlings. New For 44:937–949. doi:10.1007/s11056-013-9386-4
Sebastiana M, Vieira B, Lino-Neto T, Monteiro F, Figueiredo A, Sousa L, Pais MS, Tavares R, Paulo OS (2014) Oak root response to ectomycorrhizal symbiosis establishment: RNA-Seq derived transcript identification and expression profiling. PLoS One 9:e98376. doi:10.1371/journal.pone.0098376
Selles B, Jacquot J, Rouhier N (2011) Comparative genomic study of protein disulfide isomerases from photosynthetic organisms. Genomics 97:37–50. doi:10.1016/j.ygeno.2010.10.001
Serrato AJ, Guilleminot J, Meyer Y, Vignols F (2008) AtCXXS: atypical members of the Arabidopsis thaliana thioredoxin h family with a remarkably high disulfide isomerase activity. Physiol Plant 133:611–622. doi:10.1111/j.1399-3054.2008.01093.x
Shin L, Lo J, Yeh K (2012) Copper chaperone antioxidant protein1 is essential for copper homeostasis. Plant Physiol 159:1099–1110. doi:10.1104/pp.112.195974
Simoneau P, Viemont JD, Moreau JC, Strullu DG (1993) Symbiosis-related polypeptides associated with the early stages of ectomycorrhiza organogenesis in birch (Betula pendula Roth). New Phytol 124:495–504
Slajcherova K, Fiserova J, Fischer L, Schwarzerova K (2012) Multiple actin isotypes in plants: diverse genes for diverse roles? Front Plant Sci 3:226. doi:10.3389/fpls.2012.00226
Smith S, Read D (1997) Mycorrhizal symbiosis. Academic, London
Tada Y, Spoel SH, Pajerowska-Mukhtar K et al (2008) Plant immunity requires conformational charges of NPR1 via S-nitrosylation and thioredoxins. Science 321:952–956. doi:10.1126/science.1156970
Timonen S, Peterson RL (1993) Cytoskeleton in mycorrhizal symbiosis. Plant Soil 244:199–210. doi:10.1023/A:1020209213524
Tran HT, Hurley BA, Plaxton WC (2010) Feeding hungry plants: the role of purple acid phosphatases in phosphate nutrition. Plant Sci 179:14–27, 10.1016/j.plantsci.2010.04.005
Unlu M, Morgan ME, Minden JS (1997) Difference gel electrophoresis: a single gel method for detecting changes in protein extracts. Electrophoresis 18:2071–2077. doi:10.1002/elps.1150181133
Valot B, Dieu M, Recorbet G, Raes M, Gianinazzi S, Dumas-Gaudot E (2005) Identification of membrane-associated proteins regulated by the arbuscular mycorrhizal symbiosis. Plant Mol Biol 59:565–580. doi:10.1007/s11103-005-8269-2
Vincent D, Kohler A, Claverol S, Solier E, Joets J, Gibon J, Lebrun M-H, Plomion C, Martin F (2012) Secretome of the free-living mycelium from the ectomycorrhizal basidiomycete Laccaria bicolor. J Proteome Res 11:157–171. doi:10.1021/pr200895f
Vizcaino JA, Deutsch EW, Wang R et al (2014) ProteomeXchange provides globally coordinated proteomics data submission and dissemination. Nat Biotechnol 32:223–226. doi:10.1038/nbt.2839
Wan CY, Wilkins TA (1994) A modified hot borate method significantly enhances the yield of high-quality RNA from cotton (Gossypium hirsutum L.). Anal Biochem 223:7–12. doi:10.1006/abio.1994.1538
Wang D, Weaver ND, Kesarwani M, Dong X (2005) Induction of protein secretory pathway is required for systemic acquired resistance. Science 308:1036–1040. doi:10.1126/science.1108791
Wang L, Li Z, Qian W, Guo W, Gao X, Huang L, Wang H, Zhu H, Wu J, Wang D, Liu D (2011) The Arabidopsis purple acid phosphatase AtPAP10 is predominantly associated with the root surface and plays an important role in plant tolerance to phosphate limitation. Plant Physiol 157:1283–1299, doi:10.1104/pp.111.183723
Wilkinson B, Gilbert HF (2004) Protein disulfide isomerase. Biochim Biophys Acta 1699:35–44. doi:10.1016/j.bbapap.2004.02.017
Xu X-H, Wang C, Li S-X et al (2015) Friend or foe: differential responses of rice to invasion by mutualistic or pathogenic fungi revealed by RNAseq and metabolite profiling. Sci Rep 5:13624. doi:10.1038/srep13624
Zouari I, Salvioli A, Chialva M, Novero M, Miozzi L, Tenore GC, Bagnaresi P, Bonfante P (2014) From root to fruit: RNA-Seq analysis shows that arbuscular mycorrhizal symbiosis may affect tomato fruit metabolism. BMC Genomics 15:221. doi:10.1186/1471-2164-15-221
Acknowledgments
The authors would like to acknowledge Dr. Andrea Lorentzen from the Department of Biochemistry and Molecular Biology of the University of Southern Denmark for the help provided with protein identification. This study was supported by Fundação para a Ciência e Tecnologia (FCT/MCTES/PIDDAC, Portugal) with the project PTDC/AGR-AAM/105531/2008. JS and JP were funded by the European Research Council Synergy grant ERC-2013-SyG 610028-IMBALANCE-P.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online resource 1 (Supplementary Table 1)
Quantitative analysis of the protein spots. Column A: spot number; Column B: normalised spot volume in mycorrhizal roots according to Progenesis SameSpot; Column C: spot volume upon correction for the 0.93:0.07 plant-fungal proportion in mycorrhizal roots; Column D: normalised spot volume in non-mycorrhizal roots according to Progenesis SameSpot; Column E: Fold Change (FC) between mycorrhizal and non-mycorrhizal roots calculated as C/D; Column F: for representation proposes, a −1/FC transformation was applied to FC values between 0 and 1 (down-accumulated spots). (XLSX 12 kb)
Online resource 2 (Supplementary Table 2)
Target genes for real-time PCR analysis: accession in cork oak transcriptome database (www.corkoakdb.org), primers sequences, annealing (Ta) and melting (Tm) temperature. (DOCX 12 kb)
Online resource 3 (Supplementary Fig. 1)
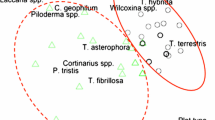
The interaction between P. tinctorius and the roots of cork oak. (a) Example of a colonized (right) and a non-colonized (left) plant. (b) Non-inoculated roots. (c) Colonized root, 2 months after inoculation with P. tinctorius. (d) Transverse section of a colonized root (2 months after inoculation) showing the fungal mantle (m) surrounding the root and the Hartig net (hn) on root epidermal cells; scale 50 μm (Sebastiana et al. 2014). (GIF 232 kb)
Online resource 4 (Supplementary Fig. 2)
Representative images of Cy labelled 2-DE gels for mycorrhizal leaves (A) and non-mycorrhizal leaves (B) (GIF 140 kb)
Online resource 5 (Supplementary Table 3)
The identity of differentially expressed protein spots as determined by tandem mass spectrometry. (XLSX 31 kb)
Rights and permissions
About this article
Cite this article
Sebastiana, M., Martins, J., Figueiredo, A. et al. Oak protein profile alterations upon root colonization by an ectomycorrhizal fungus. Mycorrhiza 27, 109–128 (2017). https://doi.org/10.1007/s00572-016-0734-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00572-016-0734-z