Abstract
The explosive increase in the worldwide prevalence of diabetes over recent years has transformed the disease into a major public health concern. While diabetes can be screened for and diagnosed by reliable biological tests based on blood glucose levels, by and large there are no means of detecting at-risk patients or of following diabetic complications. The recent discovery that microRNAs are not only chief intracellular players in many biological processes, including insulin secretion and action, but are also circulating, has put them in the limelight as possible biological markers. Here we discuss the potential role of circulating microRNAs as biomarkers in the context of diabetes and its associated complications.
Similar content being viewed by others
Introduction
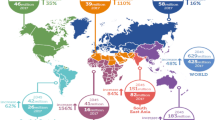
The worldwide prevalence of diabetes has drastically increased over the past 50 years and the disease is expected to affect 592 million people by 2035 making diabetes a major public health concern [1]. Diabetes is a heterogeneous disease resulting from a variety of mechanisms leading to chronic hyperglycaemia [2, 3]. The disease can be classified in general categories including type 1, type 2, gestational diabetes mellitus (GDM) and other specific types associated with disorders affecting the liver, the pancreas or endocrine glands such as the adrenal glands. While the molecular and cellular mechanisms underlying the disease are not totally understood, the role of microRNAs (miRNAs) in diabetes pathophysiology has recently emerged. miRNAs form a class of small endogenous non-coding RNAs that act as post-transcriptional regulators of gene expression [4]. Hence, miRNAs impact on physiological processes, including cell proliferation, differentiation, apoptosis and metabolism [4–6]. Consequently, their dysregulation appears to be involved in various disorders, including metabolic diseases such as diabetes (for reviews, see [5, 7]).
Interestingly, miRNAs have been detected in several biological fluids, including blood, urine, saliva, breast milk, tears, seminal, amniotic and cerebrospinal fluids [8]. While the origins of extracellular miRNAs are not totally clear, with some investigators reporting that they are the result of cellular degradation and others reporting them to be selectively secreted, it is understood that their relative compositions and concentrations vary between these fluids. Interestingly, extracellular miRNAs are very stable even under harsh conditions [9, 10]. The stability of circulating miRNAs may be explained, at least in part, by their association with proteins such as Argonaute-2 [11], HDL [12] and Nucleophosmin-1 [13] or their packaging in vesicles (e.g. apoptotic bodies, microvesicles, exosomes) [14, 15]. Owing to their stability, circulating miRNAs could be used as biomarkers, and this possibility is currently being explored in a wide range of pathologies, including diabetes [5, 16, 17]. Indeed, they can be easily collected using non-invasive procedures. Moreover, their levels can be measured by quantitative RT-PCR (qRT-PCR), which allows easy, rapid, sensitive and specific detection and quantification [18].
The aim of our review is to summarise the knowledge on circulating miRNAs in diabetes, and to analyse their potential relevance as biomarkers in medical practice. In view of the different studies, current limitations and perspectives are discussed.
Circulating miRNAs as biomarkers of the diabetic condition
In general, the diagnosis of diabetes is defined as fasting plasma glucose (FPG) >7.0 mmol/l (>126 mg/dl) or as a 2 h plasma glucose value after a 75 g oral glucose tolerance test (OGTT) >11.1 mmol/l (>200 mg/dl) [19]. The use of HbA1c reflecting the average glycaemia over a 3 month period, has also been recommended to diagnose diabetes, with a threshold of 6.5% [19]. While the above-mentioned methods are the gold standards for the biological diagnosis of diabetes, they have several shortcomings, two of which are major ones. First, the direct or indirect measurement of increased glycaemia reflects the settled disease situation. Moreover, these methods do not reflect the presence or absence of diabetic complications. Therefore, the identification of novel biomarkers that could predict the development of diabetes and identify diabetic complications, would represent a leap forward in terms of diabetes care.
In this context, the question arises of whether circulating miRNAs could be such biomarkers. The observation of the deregulation of miRNA profiles in different biological fluids of diabetic patients compared with healthy controls was the first evidence supporting the idea that miRNAs could be biomarkers of the diabetic disease (Table 1). Indeed, a differential miRNA profile between type 2 diabetic patients and healthy controls was found in whole blood [20, 21], serum [22–26], plasma [27–33] and plasma exosomes [34]. Studies revealed a different miRNA pattern in serum of type 1 diabetic patients compared with healthy individuals [35], as well as in plasma [36, 37] and urine [36]. Furthermore, the miRNA content of different types of blood cells were also deregulated in type 1 and type 2 diabetic patients compared with controls [38–42]. Finally, analysis of circulating miRNAs in women with gestational diabetes mellitus has revealed that in both serum and plasma some miRNAs have an altered profile compared with those in control women [43, 44].
Circulating miRNAs as early predictive biomarkers of diabetes
In addition to its use for the diagnosis of diabetes, blood glucose measurement also allows the detection of impaired fasting glucose (IFG; FPG 5.6–6.9 mmol/l [100–125 mg/dl]) and impaired glucose tolerance (IGT; 2 h plasma glucose value after OGTT 7.8–11.0 mmol/l [140–199 mg/dl]) [19]. IFG and IGT correspond to a prediabetic state and as such represent harbingers of future diabetes and its complications. An HbA1c range of 5.7–6.4% categorises individuals as being prediabetic [19]. However, FPG, OGTT and HbA1c measurements have some limitations in regards to predicting diabetes. Indeed, glycaemia monitoring does not allow the identification of individuals who are susceptible to developing diabetes when glucose homeostasis is still normal. Discovering new biomarkers to detect prediabetic individuals and individuals at high risk of becoming diabetic is relevant for clinical practice, as several reports indicate that intervention strategies can delay diabetes onset in individuals at high risk of the disease [45, 46].
Interestingly, some reports show that the miRNA profile in plasma or serum is deregulated in the prediabetic state before the development of overt type 2 diabetes. Indeed, type 2 diabetic and prediabetic individuals have a differential profile of circulating miRNAs compared with controls [23, 25, 30, 42]. Interestingly, for some miRNAs, such as miR-126 and miR-23a, the difference was significantly greater in type 2 diabetic than in prediabetic individuals, suggesting that their levels could be correlated with the worsening of the glycaemic status [23, 25]. Furthermore, in one study miR-192 and miR-193b were reported to be deregulated in the serum of prediabetic individuals but not in that of diabetic individuals, suggesting that prediabetic, diabetic and healthy individuals have distinct profiles [26]. Finally, altered miRNA expression (miR-15a, -29b, -126, -223, -28-3p) has been observed in serum from normoglycaemic individuals who developed type 2 diabetes over a 10 year period [27]. While additional investigations are clearly needed to strengthen these findings, the results are promising, as circulating miRNAs could be early predictive biomarkers of type 2 diabetes and could help to distinguish healthy individuals from prediabetic and diabetic patients.
To the best of our knowledge, there has been no report to date identifying circulating miRNAs as early predictive biomarkers of type 1 prediabetic individuals. In contrast, an analysis of circulating miRNAs in GDM before diagnosis revealed the dysregulation of some in the serum and plasma of women who went on to develop GDM compared with the controls [43, 44]. These results suggest that circulating miRNAs could be used as biomarkers for the early diagnosis or as a predictive factor for GDM. Hence, the measurement of circulating miRNAs could facilitate GDM detection, allowing early interventions to reduce maternal and fetal complications.
Circulating miRNAs as biomarkers of the disease process
Insulin resistance associated with type 2 diabetes is not evaluated by biomarkers as only indirect tools are available. For example, HOMA estimates steady-state beta cell function and insulin sensitivity based on algorithms that use fasting glycaemia and fasting insulinaemia measurements (www.dtu.ox.ac.uk/homacalculator/). Even if several studies have revealed a metabolic signature of insulin resistance in blood [47], no ‘simple’ nor bona fide biomarker of insulin resistance is currently available and systematically used in routine medical practice. Identifying and estimating the degree of insulin resistance is relevant as this phenomenon occurs very early in the pathogenesis of type 2 diabetes. Moreover, it could help in deciding the most appropriate approach to combat the development of overt diabetes.
Several studies have investigated circulating miRNAs of potential interest as biomarkers of insulin resistance. One study revealed that the deregulation of plasma miR-140-5p and miR-222 in type 2 diabetic patients was partially reversed after 3 months of metformin treatment, which improved insulin sensitivity as evidenced by a decrease in fasting glycaemia and HbA1c [31]. Interestingly, in healthy male volunteers, the plasma level of these two miRNAs appears to be altered after insulin infusion followed by intralipid and heparin mixture infusion, which is known to induce insulin resistance [31]. A second study reported a correlation between levels of circulating miR-484 and metabolites related to insulin resistance, which points to the potential benefit of circulating miRNAs as biomarkers of insulin resistance [48].
The possibility that circulating miRNAs could represent biomarkers of the disease process itself has also been investigated for type 1 diabetes. Levels of miR-326 were measured in peripheral blood lymphocytes from type 1 diabetic patients [49]. Note that these miRNAs were extracted from whole blood samples and were in fact associated with blood cells. Interestingly, the miR-326 level was increased in type 1 diabetic patients who tested positive for islet antibodies vs those who were antibody negative. Thus, increased miR-326 levels appear to be linked to ongoing islet autoimmunity and could be a candidate biomarker for the autoimmune process in type 1 diabetes.
Pancreatic failure is the common feature of all forms of diabetes, although the mechanisms involved are different depending on the diabetes type. Interestingly, an association between miR-25 levels and residual beta cell function was identified 3 months after diabetes onset in type 1 diabetic patients, suggesting that circulating miRNAs could reflect pancreatic endocrine function [35]. Among the miRNAs expressed in the endocrine pancreas, miR-375 has one of the highest levels of expression both in humans and rodents. In animal studies, miR-375 seems to play a complex role in beta cell function and physiology [5, 7]. Under physiological conditions its overexpression leads to reduced beta cell growth [50], while in stressed beta cells its decreased expression appears to counteract the beta cell hyperplasia [51]. Its altered expression in islets and/or pancreas has been observed in animal models with a predisposition to type 2 diabetes [52] and overt diabetes [7]. While miR-375 expression is not significantly augmented in human islets from glucose intolerant donors [53], increased levels have been observed in the pancreas [54] and blood of type 2 diabetic patients [22, 33]. Given that miRNAs have been described as being responsive to cellular and extracellular stress and to be used by cells to adjust changes in their environment, it is conceivable that, under conditions of harsh stress, such as type 2 diabetes, miR-375 is increased, while under circumstances of mild stress, such as glucose intolerance, its expression remains in the normal range. Its altered expression thus corresponds to beta cell failure. However, since miR-375 is highly expressed in both alpha and beta cells, it cannot be used to accurately estimate beta cell mass. Therefore, a beta cell-specific biomarker is needed to develop appropriate prevention strategies.
Circulating miRNAs as biomarkers of therapeutic follow-up
The biological evaluation of the efficacy of the treatment of diabetes is largely based on glycaemia monitoring and measurement of HbA1c [55]. Reflecting average glycaemia over several months, HbA1c is a relevant marker of long-term treatment efficacy. However, it is technically difficult to measure and has several limitations. First, the normal range may vary depending on the patient’s ethnic origin [56]. Second, when interpreting HbA1c levels confounders that may modify the results, such as the presence of haemoglobinopathies, need to be taken into account. Third, in situations of abnormal erythrocyte turnover, such as anaemia owing to haemolysis, transfusion or recent blood loss, the HbA1c value is not reliable and should therefore not be assessed [55]. Finally, given the worldwide pressures on healthcare systems, its cost precludes it from becoming a routine screening test in the near future. Therefore, novel biomarkers to evaluate the effectiveness of treatment on glucose homeostasis would be a great asset.
Several studies have reported a link between the level of circulating miRNAs and glycaemic control. In one study a correlation was found between miR-25 measured at 1 month after type 1 diabetes diagnosis and the HbA1c levels 3 months after onset [35]. In another cohort, consisting of 68 type 1 diabetic patients treated with insulin for at least 6 months, urinary miR-126 was lower in type 1 diabetic patients compared with controls and was inversely correlated with HbA1c levels [36]. In a third study, blood levels of miR-144-5p and miR-1207-5p reflected glucose and HbA1c levels, respectively [48].
In addition, according to other observations, the profile of circulating miRNAs changes upon treatment. Treatment, including lifestyle adjustments and medications, can partially normalise the deregulated miRNA profile observed in type 2 diabetic patients, as demonstrated by the modification of exosomal levels of miR-let-7a, miR-let-7f [34], miR-126 [23], miR-140-5p, miR-222 and miR-192 [31]. Surgical interventions such as kidney transplantation or simultaneous pancreas–kidney transplantation appear to partially normalise the deregulated miRNA profile observed in type 1 diabetic patients [37]. Furthermore, a partial normalisation of deregulated miRNAs (miR-126, -192, -193b) was also observed after lifestyle interventions in prediabetic type 2 patients [23, 26], and this was concomitant with an improvement of glycaemic control [26]. These results suggest that miRNAs could be potential biomarkers of treatment efficacy. In healthy volunteers, changes in miR-222 levels have been reported after an i.v. infusion of insulin [31]. Hence, diabetes treatment can modify the miRNA pattern, not only in type 2 diabetic and prediabetic individuals, but also in healthy people.
Circulating miRNAs as biomarkers to assess diabetic complications
Diabetes is associated with microvascular complications including nephropathy, retinopathy, and neuropathy, and with macrovascular complications such as atherosclerotic cardiovascular diseases. Both lead to increased risks of morbidity and mortality.
In medical practice, specific reliable biomarkers to predict and detect diabetic cardiovascular complications are lacking. Indeed, global cardiovascular risk can be estimated by a score index that takes into account age, sex, family history of cardiovascular disease, smoking and the presence of hypertension or dyslipidaemia. Moreover, the detection of cardiovascular problems is mainly based on clinical symptoms and medical tests, which in general provide late markers of the disease process. Thus, the identification of early and more specific biomarkers is eagerly awaited to allow optimal treatment and to combat progression of the complications.
Several studies have recommended plasma miRNAs as potential biomarkers of peripheral artery complications in type 2 diabetic patients [27, 32]. Among them, miR-126 is expressed in endothelial cells and hence is enriched in highly vascularised tissues such as the heart. Interestingly, miR-126 plays a role in the processes of angiogenesis and inflammation, and hence has been proposed as an intercellular mediator between endothelial cells and vascular smooth muscle cells [57]. Studies focusing on limb ischaemia have reported increased expression of miR-503 in the limb muscles and in the plasma of diabetic patients with critical ischaemia, suggesting that miR-503 could be a biomarker of ongoing ischaemia in diabetic patients [58]. In addition, miR-15a and miR-16 have been found to be increased in the serum and in proangiogenic cells of patients with critical limb ischaemia [59]. Levels of these two markers were found to be correlated with the risk of amputation after restenosis at 12 months post-revascularisation in type 2 diabetic patients with critical limb ischaemia. Levels of miR-15a were also correlated with the risk of post-revascularisation restenosis in those patients. Besides their potential use as biomarkers of peripheral artery disease and critical limb ischaemia, circulating miRNAs have also been investigated in atherosclerosis-related diseases. For example, miR-24 has been identified as a potential predictor of neointimal hyperplasia after coronary stenting in type 2 diabetics [60]. Furthermore, analysing urinary miRNAs in type 2 diabetic patients, miR-29b levels were found to correlate significantly with carotid intima–media thickness, pointing to the potential interest of this miRNA as a biomarker for atherosclerosis [61].
In addition to cardiovascular complications, nephropathy is another major diabetes-related public health concern, as diabetes is the leading cause of end-stage renal disease in the Western world. In general, detection of renal complications in diabetic patients is based on urinary albumin excretion measurements. However, these measurements vary under certain circumstances such as infection, fever, pronounced hyperglycaemia, severe hypertension or after physical exercise [55]. Moreover, a substantial percentage of diabetic patients have decreased glomerular filtration, reflecting kidney impairment in the absence of increased urine albumin excretion [62]. As yet, no biomarker in the urine is available to predict the decline in kidney function in those patients.
Assessment of miRNA expression in the urinary sediment of patients with chronic kidney diseases revealed a significant difference in miR-15 level between patients with diabetic glomerulosclerosis and those with other renal diseases [63]. The levels of 27 miRNAs vary significantly at different stages of untreated nephropathy in the urine of patients with type 1 diabetes [64]. In addition, a differential miRNA level in urine exosomes was found between type 1 diabetic patients with microalbuminuria and type 1 diabetic patients with normoalbuminuria [65]. While most of the investigations related to markers for renal complications have been based on urinary samples, studies on blood samples have identified an alteration of miRNA levels in type 1 and type 2 diabetic patients with diabetic nephropathy [21, 37]. Thus, the findings to date suggest that circulating miRNAs could help to diagnose and/or to monitor diabetic renal complications.
Among the diabetic complications, retinopathy carries a heavy social and economic burden as it is the leading cause of adult blindness in the Western world. HbA1c is a biomarker currently used by clinicians as it has a strong predictive value for diabetic retinopathy [19]. However, some patients develop eye complications despite satisfactory glycaemic control. Thus, new biomarkers would be useful to improve the detection of these complications. Interestingly, a study revealed three serum miRNAs (miR-21, -181c, -1179) that are deregulated in patients with proliferative diabetic retinopathy vs those with non-proliferative retinopathy suggesting that circulating miRs could be non-invasive biomarkers for the detection and/or the follow-up of retinopathy [66]. Furthermore, in a small cohort, a higher level of several miRNAs related to angiogenesis (miR-15a, -320a, -320b, -93, -29a) was found in vitreous humour of patients with proliferative diabetic retinopathy compared with patients with macular hole, with some of these miRNAs also deregulated in the serum [67].
To the best of our knowledge, no reports concerning the relevance of circulating miRNAs in diabetic neuropathy have been published to date. As intracellular miRNAs appear to be deregulated in this diabetic complication, it would be worthwhile to investigate whether circulating miRNAs could be potential biomarkers of this disease [68, 69].
Current limits to the use of circulating miRNAs in clinical practice
Diversity of study design
While promising results have been gathered concerning the potential use of circulating miRNAs as biomarkers, the studies conducted to date need to be critically evaluated at the level of the patient populations analysed and the methodology used.
First, the investigations concern different diabetic populations, with some analysing type 1 patients and others analysing type 2 or GDM patients (Table 1). Only a few miRNAs appear to show a comparable change in levels in type 2 and type 1 diabetic patients compared with controls. For example, miR-29a is increased in the serum of both type 2 [22] and type 1 [35] diabetic patients. In contrast, several groups have reported a decrease in the plasma level of miR-126 in type 2 diabetes [27, 30, 31], but no difference or an increase in type 1 diabetes [36, 37]. Comparison of several studies reveals that the majority of the deregulated miRNAs in type 1 diabetes are different from those deregulated in type 2 diabetes. This suggests that there is a distinct pattern of circulating miRNAs in type 1 diabetes vs type 2 diabetes. Indeed, a study comparing miRNA profiles between type 1 diabetes, type 2 diabetes and GDM in peripheral blood mononuclear cells confirmed divergent profiles in the different forms of diabetes despite the fact that a few miRNAs are shared among the three groups [70].
A second limitation concerns the nature of the samples. Two types of biological fluid were generally investigated (blood and urine) in most studies, as these fluids are easily collected by non-invasive methods and provide a sufficient volume for analysis. Depending on the study, when blood samples were used, analyses were usually performed on plasma or serum, but rarely on whole blood or on blood cells (Table 1). Even though, according to the manufacturers, miRNA profiles obtained from serum and plasma should be comparable, one group of investigators reported higher concentrations of circulating miRNAs in serum than in plasma from the same individual, suggesting that some miRNAs are lost during plasma preparation [71]. In addition, blood cells had more detectable miRNAs species than either serum or plasma. Therefore, haemolysis should be carefully prevented when preparing plasma or serum.
An additional issue with confounding effects relates to the nature of the circulating miRNAs. Indeed, circulating miRNAs travel in different forms of packaging, including vesicle-like exosomes, microparticles or apoptotic bodies, or they are associated with proteins. It is possible to extract all the miRNAs in the sample or to specifically extract miRNAs associated with particular vesicles or proteins. The large majority of the studies extract all the miRNAs, whereas some investigate only those associated with exosomes [20, 34, 65]. However, the profile of extracellular miRNAs associated with vesicles is different from that of vesicle-free miRNAs [13]. Hence, it is important to take into account the packaging of circulating miRNAs as this will modify the observed pattern.
Another issue to consider is that RNA isolation and miRNA profiling with qRT-PCR are performed using different commercial kits having distinct specifications. The method of RNA isolation can affect the miRNA profile depending on the protocol chosen for either total RNA or specific miRNA purification [72]. The most commonly used miRNA profiling assays are from Applied Biosystems (Taqman technology) and Exiqon (LNA technology). Both approaches are designed to enhance qRT-PCR specificity and sensitivity. Taqman technology combines miRNA-specific RT and PCR primers containing a fluorophore and a quencher for the qPCR. Exiqon combines a universal RT with locked nucleotide acid (LNA)-enhanced PCR primers, a class of high-affinity RNA analogues in which the ribose ring is ‘locked’ in the ideal conformation for Watson–Crick binding. Examining the miRNA level from plasma and serum from four individuals, one study compared the miRNA profiles obtained with these technologies [71]. For 67 commonly detectable miRNAs in plasma and serum, a low correlation between the two qRT-PCR methods was found. A second study revealed that the two methods yielded significantly different copy number estimations of some miRNAs, even though the efficiency appears to be similar [73]. To the best of our knowledge, at present, no consensus has been reached regarding which assay should be preferentially used. Thus, depending on the technique applied, the results may vary and this should be taken into account when analysing the results.
Another issue linked to the miRNA profiling technology is that at present there is no well-established housekeeping RNA for data normalisation of circulating miRNAs, therefore variable references are utilised. Urine measurements apply exogenous controls such as miR-39, Uni SP3 or endogenous controls such as RNA U6, while studies based on blood utilise exogenous controls such as miR-39 or endogenous controls such as RNU 6b, RNU 48, RNA U6, miR-16, miR-19b, miR-30c, miR-103, miR-106a, miR-191, miR-146a, miR-221, miR-223, miR-238, miR-423-3p, miR-425, miR-454, or miRNAs belonging to the let-7 family. Although these miRNAs have been selected for their relative stability, the levels of some may vary in diabetic populations such as miR-let-7a [21, 34] and miR-let-7f [34], miR-let-7i [25], miR-191 [27, 38], miR-223 [27] or miR-146a [22, 29, 41]. Given these shortcomings, a consensus for data normalisation is urgently needed based, for example, on the geometric mean of a common set of miRNAs.
Because of the heterogeneity in study design, the comparison of the results of the different reports should take into account that divergent results may be explained by differences in the techniques applied and the populations analysed. Unfortunately, only a few studies can be compared on the basis of having similar diabetic populations, samples, RNA extraction methods and miRNAs investigated. Among the reports on the same circulating miRNAs, some show similar results. For example, miR-126 was found by several authors to be decreased in plasma from type 2 diabetic patients [27, 30, 31]. At the same time, observations made in some studies were not totally reproducible in others. Although it was reported that plasma levels of miR-15a and miR-223 were significantly different between type 2 diabetic patients and controls [27], this finding was not replicated by other investigators [30]. Remarkably, a meta-analysis including 38 studies in human and animal models comparing miRNA profiles between type 2 diabetics and non-diabetics identified 40 dysregulated miRNAs among different tissues, including blood. Of these, eight miRNAs emerged as potential circulating biomarkers of type 2 diabetes [74]. In summary, the investigated populations and the methodologies used in different reports suffer from important dissimilarities. These need to be standardised to allow the study of circulating miRNAs. Even though there are recent reports that seem to address this issue, to the best of our knowledge no global consensus exists at this time [72, 75, 76].
Potential confounding factors
Besides the impact of the methodology on the observations made, there are some recently revealed confounding factors that should be taken into account when interpreting miRNA profiling results (ESM Table 1). Several studies have identified age-related modifications in circulating miRNA levels [77–82]. Notably, among these, miR-29b [27, 28, 70], miR-130b [31], miR-222 [31], miR-375 [22], miR-21 [26, 27, 36] and miR-126 [23, 27, 30, 31] were found to be deregulated in diabetic patients. These results underline that age could induce a bias, which has to be dealt with.
In the same way, sex should be taken in consideration, as indicated by a study of a cohort of 102 individuals with and without the metabolic syndrome where the profile of circulating miRNAs was found to be different only in women with the metabolic syndrome [83]. Another potential confounding factor is linked to ethnicity. Indeed, several reports revealed ethnicity-related plasma miRNA levels in diabetic populations [28, 84].
Lifestyle can also influence the circulating miRNA pattern. In small cohorts of patients, some studies illustrate the impact of nutrition such as zinc or 25-hydroxyvitamin D levels on circulating miRNAs [85, 86]. These observations suggest that nutrition can be a confounding factor, which is difficult to evaluate. Indeed, the long-term impact of diet on the circulating miRNA profile is cumbersome to consider in studies investigating miRNAs in disease states. Another lifestyle factor that can potentially affect the results is physical exercise. Several studies have reported that acute physical activity and prolonged exercise training programmes can modify the circulating miRNA profile [87]. Furthermore, smoking appears to alter the pattern of circulating miRNAs [88–91], and of the miRNAs affected, miR-29b was identified to be deregulated in diabetic patients [27, 28, 70]. Environmental exposure to air pollutants and chemicals seems to modify circulating miRNAs [92], inducing a bias that is difficult to estimate.
Finally, the patterns of circulating miRNAs in diabetes can be changed by a wide range of associated conditions such as cancer [93], and cardiovascular, neurodegenerative, metabolic [72] and autoimmune diseases [94].
The potential confounding factors discussed here should urge clinical investigators to exercise caution when designing the controls, and they should be obliged to take these factors into account in their analyses. By using the appropriate questionnaires before the clinical study, most confounding factors, such as age, sex, ethnic origin, comorbidity, lifestyle (including smoking or physical exercise) can be revealed. Therefore, these factors should be ruled out using exclusion criteria or controlled for by stratifying the population or using multivariate analysis. However, the impact of other factors, such as nutrition or environmental exposure, appear to be much more challenging to evaluate or to detect, and would need further characterisation to reach a consensus for including them.
While the effects of confounding factors can, at least in part, be limited when running a clinical study, the question of the specificity of circulating miRNAs as a biomarker of diabetes remains. As demonstrated by previous studies, an miRNA pattern can be confounded by an associated disease, which is a current limit for the use of circulating miRNAs as biomarker for multi-disease patients [95]. Therefore, it might be useful not to focus on changes in the level of one particular miRNA, but rather to look for a disease-related signature of a series of miRNAs. Interestingly, novel miRNAs have been identified, many of which are human specific and tissue specific, offering a promising opportunity to identify such biomarkers [96]. Finally, while this is a burgeoning area of research, the recent discovery of the high tissue specificity of long non-coding RNAs [97] and their identification in biological fluids such as plasma could offer new perspectives [98].
Conclusions
While the field is still in its infancy, circulating miRNAs appear to offer various potential applications in medical practice as biomarkers of diabetes and associated complications. Different applications can be envisioned, such as prediction of the disease, detection and monitoring of its complications, evaluation of treatment efficacy and the characterisation of pathogenic traits such as insulin resistance for type 2 diabetes and the autoimmune process for type 1 diabetes. However, the observations from the currently available studies are difficult to compare because of the heterogeneity of the technical methods used and the small and heterogeneous cohorts investigated. As the impact of these shortcomings is not precisely evaluated and not considered in most studies, a top priority is to establish a consensus to standardise the methodology used and to improve reproducibility. While improvements in the standardisation of preanalytical and analytical methods are needed and additional studies on larger cohorts are required before circulating miRNAs can be introduced into medical practice, miRNAs appear to be bona fide biomarkers for diabetes in the future. Even if the measurement of circulating miRNAs currently has limitations, miRNAs are likely to become useful in the near future. Indeed, one can anticipate that they will complement existing biomarkers, especially in the context of the early identification of individuals at high risk to develop diabetes, and in the detection and the follow-up of diabetic complications.
More than being simple markers, which vary depending on physiological or pathological processes, recent observations suggest a functional role of circulating miRNAs. Indeed, circulating miRNAs contained in vesicles or associated with HDL can be transferred into recipient cells and thus could be involved in cell-to-cell communication [12, 14]. To date, a series of studies has investigated the role of circulating miRNAs as intercellular mediators in diabetic conditions, pointing to possible new mechanisms in diabetes pathogenesis [99–101]. It is clear that the precise potential of circulating miRNAs in complex and multifactorial diseases such as diabetes requires further confirmation both in terms of a biomarker and its involvement in disease mechanisms. A considerable amount of research efforts need to be invested to determine whether circulating miRNAs profiles are of use in routine hospital practice, but given the potential advantages, the challenge is undoubtedly worth it.
Abbreviations
- FPG:
-
Fasting plasma glucose
- GDM:
-
Gestational diabetes mellitus
- IFG:
-
Impaired fasting glucose
- IGT:
-
Impaired glucose tolerance
- miRNA:
-
microRNA
- PAC:
-
Proangiogenic cells
References
IDF (2014) http://www.idf.org/sites/default/files/Atlas-poster-2014_EN.pdf
Stumvoll M, Goldstein BJ, van Haeften TW (2005) Type 2 diabetes: principles of pathogenesis and therapy. Lancet 365:1333–1346
van Belle TL, Coppieters KT, von Herrath MG (2011) Type 1 diabetes: etiology, immunology, and therapeutic strategies. Physiol Rev 91:79–118
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Dumortier O, Hinault C, Van Obberghen E (2013) MicroRNAs and metabolism crosstalk in energy homeostasis. Cell Metab 18:312–324
Fernandez-Hernando C, Ramirez CM, Goedeke L, Suarez Y (2013) MicroRNAs in metabolic disease. Arterioscler Thromb Vasc Biol 33:178–185
Guay C, Regazzi R (2015) Role of islet microRNAs in diabetes: which model for which question? Diabetologia 58:456–463
Weber JA, Baxter DH, Zhang S et al (2010) The microRNA spectrum in 12 body fluids. Clin Chem 56:1733–1741
Mitchell PS, Parkin RK, Kroh EM et al (2008) Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A 105:10513–10518
Chen X, Ba Y, Ma L et al (2008) Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 18:997–1006
Arroyo JD, Chevillet JR, Kroh EM et al (2011) Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc Natl Acad Sci U S A 108:5003–5008
Vickers KC, Palmisano BT, Shoucri BM, Shamburek RD, Remaley AT (2011) MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat Cell Biol 13:423–433
Wang K, Zhang S, Weber J, Baxter D, Galas DJ (2010) Export of microRNAs and microRNA-protective protein by mammalian cells. Nucleic Acids Res 38:7248–7259
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO (2007) Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9:654–659
Zernecke A, Bidzhekov K, Noels H et al (2009) Delivery of microRNA-126 by apoptotic bodies induces CXCL12-dependent vascular protection. Sci Signal 2:ra81
Guay C, Regazzi R (2013) Circulating microRNAs as novel biomarkers for diabetes mellitus. Nat Rev Endocrinol 9:513–521
Chien HY, Lee TP, Chen CY et al (2015) Circulating microRNA as a diagnostic marker in populations with type 2 diabetes mellitus and diabetic complications. J Chin Med Assoc JCMA 78:204–211
de Planell-Saguer M, Rodicio MC (2013) Detection methods for microRNAs in clinic practice. Clin Biochem 46:869–878
American Diabetes A (2014) Diagnosis and classification of diabetes mellitus. Diabetes Care 37(Suppl 1):S81–S90
Karolina DS, Tavintharan S, Armugam A et al (2012) Circulating miRNA profiles in patients with metabolic syndrome. J Clin Endocrinol Metab 97:E2271–E2276
Zhou J, Peng R, Li T et al (2013) A potentially functional polymorphism in the regulatory region of let-7a-2 is associated with an increased risk for diabetic nephropathy. Gene 527:456–461
Kong L, Zhu J, Han W et al (2011) Significance of serum microRNAs in pre-diabetes and newly diagnosed type 2 diabetes: a clinical study. Acta Diabetol 48:61–69
Liu Y, Gao G, Yang C et al (2014) The role of circulating microRNA-126 (miR-126): a novel biomarker for screening prediabetes and newly diagnosed type 2 diabetes mellitus. Int J Mol Sci 15:10567–10577
Pescador N, Perez-Barba M, Ibarra JM, Corbaton A, Martinez-Larrad MT, Serrano-Rios M (2013) Serum circulating microRNA profiling for identification of potential type 2 diabetes and obesity biomarkers. PLoS One 8, e77251
Yang Z, Chen H, Si H et al (2014) Serum miR-23a, a potential biomarker for diagnosis of pre-diabetes and type 2 diabetes. Acta Diabetol 51:823–831
Parrizas M, Brugnara L, Esteban Y et al (2014) Circulating miR-192 and miR-193b are markers of prediabetes and are modulated by an exercise intervention. J Clin Endocrinol Metab 100:E407–E415
Zampetaki A, Kiechl S, Drozdov I et al (2010) Plasma microRNA profiling reveals loss of endothelial miR-126 and other microRNAs in type 2 diabetes. Circ Res 107:810–817
Wang X, Sundquist J, Zoller B et al (2014) Determination of 14 circulating microRNAs in Swedes and Iraqis with and without diabetes mellitus type 2. PLoS One 9, e86792
Rong Y, Bao W, Shan Z et al (2013) Increased microRNA-146a levels in plasma of patients with newly diagnosed type 2 diabetes mellitus. PLoS One 8, e73272
Zhang T, Lv C, Li L et al (2013) Plasma miR-126 is a potential biomarker for early prediction of type 2 diabetes mellitus in susceptible individuals. Biomed Res Int 2013:761617
Ortega FJ, Mercader JM, Moreno-Navarrete JM et al (2014) Profiling of circulating microRNAs reveals common microRNAs linked to type 2 diabetes that change with insulin sensitization. Diabetes Care 37:1375–1383
Dangwal S, Stratmann B, Bang C et al (2015) Impairment of wound healing in patients with type 2 diabetes mellitus influences circulating microRNA patterns via inflammatory cytokines. Arterioscler Thromb Vasc Biol 35:1480–1488
Sun K, Chang X, Yin L et al (2014) Expression and DNA methylation status of microRNA-375 in patients with type 2 diabetes mellitus. Mol Med Rep 9:967–972
Santovito D, De Nardis V, Marcantonio P et al (2014) Plasma exosome microRNA profiling unravels a new potential modulator of adiponectin pathway in diabetes: effect of glycemic control. J Clin Endocrinol Metab 99:E1681–E1685
Nielsen LB, Wang C, Sorensen K et al (2012) Circulating levels of microRNA from children with newly diagnosed type 1 diabetes and healthy controls: evidence that miR-25 associates to residual beta-cell function and glycaemic control during disease progression. Exp Diabetes Res 2012:896362
Osipova J, Fischer DC, Dangwal S et al (2014) Diabetes-associated microRNAs in paediatric patients with type 1 diabetes mellitus: a cross-sectional cohort study. J Clin Endocrinol Metab 99:E1661–E1665
Bijkerk R, Duijs JM, Khairoun M et al (2015) Circulating microRNAs associate with diabetic nephropathy and systemic microvascular damage and normalize after simultaneous pancreas–kidney transplantation. Am J Transplant Off J Am Soc Transplant Am Soc Transplant Surg 15:1081–1090
Hezova R, Slaby O, Faltejskova P et al (2010) MicroRNA-342, microRNA-191 and microRNA-510 are differentially expressed in T regulatory cells of type 1 diabetic patients. Cell Immunol 260:70–74
Salas-Perez F, Codner E, Valencia E, Pizarro C, Carrasco E, Perez-Bravo F (2013) MicroRNAs miR-21a and miR-93 are down regulated in peripheral blood mononuclear cells (PBMCs) from patients with type 1 diabetes. Immunobiology 218:733–737
Takahashi P, Xavier DJ, Evangelista AF et al (2014) MicroRNA expression profiling and functional annotation analysis of their targets in patients with type 1 diabetes mellitus. Gene 539:213–223
Balasubramanyam M, Aravind S, Gokulakrishnan K et al (2011) Impaired miR-146a expression links subclinical inflammation and insulin resistance in Type 2 diabetes. Mol Cell Biochem 351:197–205
Luo M, Li R, Deng X et al (2015) Platelet-derived miR-103b as a novel biomarker for the early diagnosis of type 2 diabetes. Acta Diabetol. doi:10.1007/s00592-015-0733-0
Zhao C, Dong J, Jiang T et al (2011) Early second-trimester serum miRNA profiling predicts gestational diabetes mellitus. PLoS One 6, e23925
Zhu Y, Tian F, Li H, Zhou Y, Lu J, Ge Q (2015) Profiling maternal plasma microRNA expression in early pregnancy to predict gestational diabetes mellitus. Int J Gynaecol Obstet Off Organ Int Fed Gynaecol Obstet 130:49–53
Li G, Zhang P, Wang J et al (2008) The long-term effect of lifestyle interventions to prevent diabetes in the China Da Qing Diabetes Prevention Study: a 20-year follow-up study. Lancet 371:1783–1789
Knowler WC, Barrett-Connor E, Fowler SE et al (2002) Reduction in the incidence of type 2 diabetes with lifestyle intervention or metformin. N Engl J Med 346:393–403
Wurtz P, Makinen VP, Soininen P et al (2012) Metabolic signatures of insulin resistance in 7,098 young adults. Diabetes 61:1372–1380
Raitoharju E, Seppala I, Oksala N et al (2014) Blood microRNA profile associates with the levels of serum lipids and metabolites associated with glucose metabolism and insulin resistance and pinpoints pathways underlying metabolic syndrome: the cardiovascular risk in Young Finns Study. Mol Cell Endocrinol 391:41–49
Sebastiani G, Grieco FA, Spagnuolo I, Galleri L, Cataldo D, Dotta F (2011) Increased expression of microRNA miR-326 in type 1 diabetic patients with ongoing islet autoimmunity. Diabetes Metab Res Rev 27:862–866
El Ouaamari A, Baroukh N, Martens GA, Lebrun P, Pipeleers D, Van Obberghen E (2008) miR-375 targets 3'-phosphoinositide-dependent protein kinase-1 and regulates glucose-induced biological responses in pancreatic beta-cells. Diabetes 57:2708–2717
Poy MN, Eliasson L, Krutzfeldt J et al (2004) A pancreatic islet-specific microRNA regulates insulin secretion. Nature 432:226–230
Dumortier O, Hinault C, Gautier N, Patouraux S, Casamento V, Van Obberghen E (2014) Maternal protein restriction leads to pancreatic failure in offspring: role of misexpressed microRNA-375. Diabetes 63:3416–3427
Bolmeson C, Esguerra JL, Salehi A, Speidel D, Eliasson L, Cilio CM (2011) Differences in islet-enriched miRNAs in healthy and glucose intolerant human subjects. Biochem Biophys Res Commun 404:16–22
Zhao H, Guan J, Lee HM et al (2010) Up-regulated pancreatic tissue microRNA-375 associates with human type 2 diabetes through beta-cell deficit and islet amyloid deposition. Pancreas 39:843–846
American Diabetes Association (2014) Standards of medical care in diabetes—2014. Diabetes Care 37(Suppl 1):S14–S80
Ziemer DC, Kolm P, Weintraub WS et al (2010) Glucose-independent, black–white differences in hemoglobin A1c levels: a cross-sectional analysis of 2 studies. Ann Intern Med 152:770–777
Zhou J, Li YS, Nguyen P et al (2013) Regulation of vascular smooth muscle cell turnover by endothelial cell-secreted microRNA-126: role of shear stress. Circ Res 113:40–51
Caporali A, Meloni M, Vollenkle C et al (2011) Deregulation of microRNA-503 contributes to diabetes mellitus-induced impairment of endothelial function and reparative angiogenesis after limb ischemia. Circulation 123:282–291
Spinetti G, Fortunato O, Caporali A et al (2013) MicroRNA-15a and microRNA-16 impair human circulating proangiogenic cell functions and are increased in the proangiogenic cells and serum of patients with critical limb ischemia. Circ Res 112:335–346
Hong SJ, Choi SC, Cho JY et al (2015) Pioglitazone increases circulating microRNA-24 with decrease in coronary neointimal hyperplasia in type 2 diabetic patients – optical coherence tomography analysis. Circ J Off J Jpn Circ Soc 79:880–888
Peng H, Zhong M, Zhao W et al (2013) Urinary miR-29 correlates with albuminuria and carotid intima-media thickness in type 2 diabetes patients. PLoS One 8, e82607
Kramer HJ, Nguyen QD, Curhan G, Hsu CY (2003) Renal insufficiency in the absence of albuminuria and retinopathy among adults with type 2 diabetes mellitus. JAMA 289:3273–3277
Szeto CC, Ching-Ha KB, Ka-Bik L et al (2012) Micro-RNA expression in the urinary sediment of patients with chronic kidney diseases. Dis Markers 33:137–144
Argyropoulos C, Wang K, McClarty S et al (2013) Urinary microRNA profiling in the nephropathy of type 1 diabetes. PLoS One 8, e54662
Barutta F, Tricarico M, Corbelli A et al (2013) Urinary exosomal microRNAs in incipient diabetic nephropathy. PLoS One 8, e73798
Qing S, Yuan S, Yun C et al (2014) Serum miRNA biomarkers serve as a fingerprint for proliferative diabetic retinopathy. Cell Physiol Biochem 34:1733–1740
Hirota K, Keino H, Inoue M, Ishida H, Hirakata A (2015) Comparisons of microRNA expression profiles in vitreous humor between eyes with macular hole and eyes with proliferative diabetic retinopathy. Graefes Arch Clin Exp Ophthalmol 253:335–342
Wang L, Chopp M, Szalad A et al (2014) The role of miR-146a in dorsal root ganglia neurons of experimental diabetic peripheral neuropathy. Neuroscience 259:155–163
Zhang X, Gong X, Han S, Zhang Y (2014) miR-29b protects dorsal root ganglia neurons from diabetic rat. Cell Biochem Biophys 70:1105–1111
Collares CV, Evangelista AF, Xavier DJ et al (2013) Identifying common and specific microRNAs expressed in peripheral blood mononuclear cell of type 1, type 2, and gestational diabetes mellitus patients. BMC Res Notes 6:491
Wang K, Yuan Y, Cho JH, McClarty S, Baxter D, Galas DJ (2012) Comparing the microRNA spectrum between serum and plasma. PLoS One 7, e41561
Chevillet JR, Lee I, Briggs HA, He Y, Wang K (2014) Issues and prospects of microRNA-based biomarkers in blood and other body fluids. Molecules 19:6080–6105
Redshaw N, Wilkes T, Whale A, Cowen S, Huggett J, Foy CA (2013) A comparison of miRNA isolation and RT-qPCR technologies and their effects on quantification accuracy and repeatability. BioTechniques 54:155–164
Zhu H, Leung SW (2015) Identification of microRNA biomarkers in type 2 diabetes: a meta-analysis of controlled profiling studies. Diabetologia 58:900–911
Farina NH, Wood ME, Perrapato SD et al (2014) Standardizing analysis of circulating microRNA: clinical and biological relevance. J Cell Biochem 115:805–811
Witwer KW (2015) Circulating MicroRNA biomarker studies: pitfalls and potential solutions. Clin Chem 61:56–63
ElSharawy A, Keller A, Flachsbart F et al (2012) Genome-wide miRNA signatures of human longevity. Aging Cell 11:607–616
Gombar S, Jung HJ, Dong F et al (2012) Comprehensive microRNA profiling in B-cells of human centenarians by massively parallel sequencing. BMC Genomics 13:353
Noren Hooten N, Fitzpatrick M, Wood WH 3rd et al (2013) Age-related changes in microRNA levels in serum. Aging 5:725–740
Olivieri F, Spazzafumo L, Santini G et al (2012) Age-related differences in the expression of circulating microRNAs: miR-21 as a new circulating marker of inflammaging. Mech Ageing Dev 133:675–685
Serna E, Gambini J, Borras C et al (2012) Centenarians, but not octogenarians, up-regulate the expression of microRNAs. Sci Rep 2:961
Zhang H, Yang H, Zhang C et al (2015) Investigation of microRNA expression in human serum during the aging process. J Gerontol A: Biol Med Sci 70:102–109
Wang YT, Tsai PC, Liao YC, Hsu CY, Juo SH (2013) Circulating microRNAs have a sex-specific association with metabolic syndrome. J Biomed Sci 20:72
Chang X, Li S, Li J et al (2014) Ethnic differences in microRNA-375 expression level and DNA methylation status in type 2 diabetes of Han and Kazak populations. J Diabetes Res 2014:761938
Ryu MS, Langkamp-Henken B, Chang SM, Shankar MN, Cousins RJ (2011) Genomic analysis, cytokine expression, and microRNA profiling reveal biomarkers of human dietary zinc depletion and homeostasis. Proc Natl Acad Sci U S A 108:20970–20975
Enquobahrie DA, Williams MA, Qiu C, Siscovick DS, Sorensen TK (2011) Global maternal early pregnancy peripheral blood mRNA and miRNA expression profiles according to plasma 25-hydroxyvitamin D concentrations. J Matern Fetal Neonatal Med 24:1002–1012
Flowers E, Won GY, Fukuoka Y (2015) MicroRNAs associated with exercise and diet: a systematic review. Physiol Genomics 47:1–11
Badrnya S, Baumgartner R, Assinger A (2014) Smoking alters circulating plasma microvesicle pattern and microRNA signatures. Thromb Haemost 112:128–136
Banerjee A, Waters D, Camacho OM, Minet E (2015) Quantification of plasma microRNAs in a group of healthy smokers, ex-smokers and non-smokers and correlation to biomarkers of tobacco exposure. Biomark Biochem Indic Expo Response Susceptibility Chem 1–9
Huang J, Wu J, Li Y et al (2014) Deregulation of serum microRNA expression is associated with cigarette smoking and lung cancer. Biomed Res Int 2014:364316
Takahashi K, Yokota S, Tatsumi N, Fukami T, Yokoi T, Nakajima M (2013) Cigarette smoking substantially alters plasma microRNA profiles in healthy subjects. Toxicol Appl Pharmacol 272:154–160
Vrijens K, Bollati V, Nawrot TS (2015) MicroRNAs as potential signatures of environmental exposure or effect: a systematic review. Environ Health Perspect 123:399–411
Wang J, Zhang KY, Liu SM, Sen S (2014) Tumor-associated circulating microRNAs as biomarkers of cancer. Molecules 19:1912–1938
Zeng L, Cui J, Wu H, Lu Q (2014) The emerging role of circulating microRNAs as biomarkers in autoimmune diseases. Autoimmunity 1–11
Keller A, Leidinger P, Bauer A et al (2011) Toward the blood-borne miRNome of human diseases. Nat Methods 8:841–843
Londin E, Loher P, Telonis AG et al (2015) Analysis of 13 cell types reveals evidence for the expression of numerous novel primate- and tissue-specific microRNAs. Proc Natl Acad Sci U S A 112:E1106–E1115
Cabili MN, Trapnell C, Goff L et al (2011) Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev 25:1915–1927
Lorenzen JM, Schauerte C, Kielstein JT et al (2015) Circulating long noncoding RNATapSaki is a predictor of mortality in critically ill patients with acute kidney injury. Clin Chem 61:191–201
Wang X, Huang W, Liu G et al (2014) Cardiomyocytes mediate anti-angiogenesis in type 2 diabetic rats through the exosomal transfer of miR-320 into endothelial cells. J Mol Cell Cardiol 74:139–150
Chaturvedi P, Kalani A, Medina I, Familtseva A, Tyagi SC (2015) Cardiosome mediated regulation of MMP9 in diabetic heart: role of mir29b and mir455 in exercise. J Cell Mol Med. doi:10.1111/jcmm.12589
Guay C, Menoud V, Rome S, Regazzi R (2015) Horizontal transfer of exosomal microRNAs transduce apoptotic signals between pancreatic beta-cells. Cell Commun Sig CCS 13:17
Acknowledgements
The authors would like to thank E. Corsetti (University Nice Sophia Antipolis, INSERM, U1081, CNRS, UMR7284, IRCAN, Nice, France) and F. Lareyre (University Nice Sophia Antipolis, Nice, France) for their constructive discussions of the manuscript.
Funding
The authors are supported by INSERM, Université Nice Sophia Antipolis, Conseil Régional PACA, Conseil Général des Alpes-Maritimes, Aviesan/AstraZeneca, ‘Diabetes and the vessel wall injury’ programme by the Agence Nationale de la Recherche (ANR) through ANR-RPV12004AAA, ‘Investments for the Future’ LABEX SIGNALIFE #ANR-11-LABX-0028-01 and European Foundation for the Study of Diabetes (EFSD/Lilly; European Diabetes Research Programme). We apologise for being unable to discuss all the relevant papers due to space limitations.
Duality of interest
The authors declare that there is no duality of interest associated with the manuscript.
Contribution statement
All authors confirm they contributed to the intellectual content of this article and approved the final version.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM Table 1
(PDF 37 kb)
Rights and permissions
About this article
Cite this article
Raffort, J., Hinault, C., Dumortier, O. et al. Circulating microRNAs and diabetes: potential applications in medical practice. Diabetologia 58, 1978–1992 (2015). https://doi.org/10.1007/s00125-015-3680-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-015-3680-y