Abstract
Background
Gliomas represent about 80% of primary brain tumours and about 30% of malignant ones, which today don’t have a resolution therapy because of their variability. A valid model for the study of new personalized therapies can be represented by primary cultures from patient’s tumour biopsies.
Methods
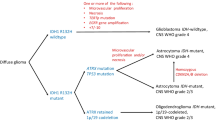
In this study we consider 12 novel cell lines established from patients’ gliomas and immunohistochemically and molecularly characterized according to the newly updated 2016 CNS Tumour WHO classification.
Results
Eight of these lines were glioblastoma cells, two grade III glioma cells (anaplastic astrocytoma and oligo astrocytoma) and two low grade glioma cells (grade II astrocytoma and oligodendroglioma). All cell lines were analysed by immunohistochemistry for specific glioma markers, respectively VIMENTIN, GFAP, IDH1R132, and ATRX. The methylation status of the MGMT gene promoter was also determined in all lines. The comparison of the immunohistochemical characteristics and of the MGMT methylation status of the lines with the tissues of origin shows that the cells in culture maintain the same characteristics.
Conclusions
Human cancer cell lines represent a support in the knowledge of tumour biology and in drug discovery through its facile experimental manipulation.
Trial registration
NCT 04180046.



Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the authors upon reasonable request.
Abbreviations
- GBM:
-
Glioblastoma
- TMZ:
-
Temozolomide
- BBB:
-
Blood–brain barrier
- IDH:
-
Isocitrate dehydrogenase
- MGMT:
-
O6-methylguanine–DNA methyltransferase
References
Ciceroni C, Bonelli M, Mastrantoni E, Niccolini C, Laurenza M, Larocca LM, Pallini R, Traficante A, Spinsanti P, Ricci-Vitiani L, Arcella A, De Maria R, Nicoletti F, Battaglia G, Melchiorri D (2013) Type-3 metabotropic glutamate receptors regulate chemoresistance in glioma stem cells, and their levels are inversely related to survival in patients with malignant gliomas. Cell Death Differ 20(3):396–407
De Almeida SF, Lunardi Brunetto A, Schwartsmann G, Roesler R, Abujamra AL (2012) Glioma revisited: from neurogenesis and cancer stem cells to the epigenetic regulation of theniche. J Oncol 2012:537861
Ohgaki H, Kleihues P (2005) Epidemiology and etiology of gliomas. Acta Neuropathol 109(1):93–108
Surawicz TS, Davis F, Freels S, Laws ER Jr., Menck HR (1998) Brain tumor survival: results from the National Cancer Data Base. J Neurooncol 40(2):151–160
Stupp R, Weber DC (2005) The role of radio- and chemotherapy in glioblastoma. Onkologie 28(6–7):315–317
Dai C, Holland EC (2003) Astrocyte differentiation states and glioma formation. Cancer J. 9(2):72–81
Louis DN, Perry A, Burger P, Ellison DW, Reifenberger G, von Deimling A, Aldape K, Brat D, Collins VP, Eberhart C, Figarella-Branger D, Fuller GN, Giangaspero F, Giannini C, Hawkins C, Kleihues P, Korshunov A, Kros JM, Lopes MB, Ng HK, Ohgaki H, Paulus W, Pietsch T, Rosenblum M, Rushing E, Soylemezoglu F, Wiestler O, Wesseling P (2014) International Society of Neuropathology-Haarlem. International Society of Neuropathology-Haarlem Consensus Guidelines for Nervous System Tumor Classification and Grading. Brain Pathol 24(5):429–435
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131(6):803–820
van den Bent MJ, Weller M, Wen PY, Kros JM, Aldape K, Chang S (2017) A clinical perspective on the 2016 WHO brain tumor classification and routine molecular diagnostics. Neuro Oncol 19(5):614–624
Ohgaki H, Kleihues P (2013) The definition of primary and secondary glioblastoma. Clin Cancer Res 19(4):764–772
Deng L, Xiong P, Luo Y, Bu X, Qian S, Zhong W, Lv S (2018) Association between IDH1/2 mutations and brain glioma grade. Oncol Lett 16(4):5405–5409
Cancer Genome Atlas Research (2008) Network comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455:1061–1068
Yan H, Parsons DW, Jin G, Mc LR, Rasheed BA, Yuan W, Kos I, Batinic-Haberle I, Jones S, Riggins GJ et al (2009) IDH1 and IDH2 mutations in gliomas. N Engl J Med 360:765–773
Balss J, Meyer J, Mueller W, Korshunov A, Hartmann C, von Deimling A (2008) Analysis of the IDH1 codon 132 mutation in brain tumors. Acta Neuropathol 116:597–602
Ichimura K, Pearson DM, Kocialkowski S, Bäcklund LM, Chan R, Jones DT, Collins VP (2009) IDH1 mutations are present in the majority of common adult gliomas but rare in primary glioblastomas. Neuro Oncol 11:341–347
Horbinski C (2013) What do we know about IDH1/2 mutations so far, and how do we use it? Acta Neuropathol 125:621–636
Dang L, White DW, Gross S, Bennett BD, Bittinger MA, Driggers EM, Fantin VR, Jang HG, Jin S, Keenan MC, Marks KM, Prins RM, Ward PS, Yen KE, Liau LM, Rabinowitz JD, Cantley LC, Thompson CB, Vander Heiden MG, Su SM (2009) Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462(7274):739
Turcan S, Rohle D, Goenka A, Walsh LA, Fang F, Yilmaz E, Campos C, Fabius AWM, Lu C, Ward PS, Thompson CB, Kaufman A, Guryanova O, Levine R, Heguy A, Viale A, Morris LGT, Huse JT, Mellinghoff IK, Chan TA (2012) IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature 483:479–483
Lu C, Ward PS, Kapoor GS, Rohle D, Turcan S, Abdel-Wahab O, Edwards CR, Khanin R, Figueroa ME, Melnick A, Wellen KE, O’Rourke DM, Berger SL, Chan TA, Levine RL, Mellinghoff IK, Thompson CB (2012) IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483(7390):474–478
Reiter-Brennan C, Semmler L, Klein A (2018) The effects of 2-hydroxyglutarate on the tumorigenesis of gliomas. ContempOncol (Pozn) 22(4):215–222
Sanson M, Marie Y, Paris S, Idbaih A, Laffaire J, Ducray F, El Hallani S, Boisselier B, Mokhtari K, Hoang-Xuan K, Delattre JY (2009) Isocitrate Dehydrogenase 1 Codon 132 mutation is an important prognostic biomarker in gliomas. J Clin Oncol 27(25):4150–4154
Hartmann C, Hentschel B, Simon M, Westphal M, Schackert G, Tonn JC, Loeffler M, Reifenberger G, Pietsch T, von Deimling A, Weller M, German Glioma Network (2013) Long-term survival in primary glioblastoma with versus without isocitrate dehydrogenase mutations. Clin Cancer Res. 19(18):5146–5157
Cohen A, Holmen S, Colman H (2013) IDH1 and IDH2 mutations in gliomas. Curr Neurol Neurosci Rep. 13(5):345
Minniti G, Salvati M, Arcella A, Buttarelli F, D’Elia A, Lanzetta G, Esposito V, Scarpino S, MauriziEnrici R, Giangaspero F (2011) Correlation between O6-methylguanine-DNA methyltransferase and survival in elderly patients with glioblastoma treated with radiotherapy plus concomitant and adjuvant temozolomide. J Neurooncol 102:311–316
Hegi ME, Diserens AC, Gorlia T, Hamou MF, de Tribolet N, Weller M, Kros JM, Hainfellner JA, Mason W, Mariani L, Bromberg JEC, Hau P, Mirimanoff RO, Cairncross JG, Janzer RC, Stupp R (2005) MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med. 352:997–1003
Wong LH, McGhie JD, Sim M, Anderson MA, Ahn S, Hannan RD, George AJ, Morgan KA, Mann JR, Andy Choo KH (2010) ATRX interacts with H3.3 in maintaining telomere structural integrity in pluripotent embryonic stem cells. Genome Res. 20(3):351–360
Voon HP, Hughes JR, Rode C, De La Rosa-Velázquez IA, Jenuwein T, Feil R et al (2015) ATRX plays a key role in maintaining silencing at interstitial heterochromatic loci and imprinted genes. Cell Rep 11(3):405–418
Elsässer SJ, Noh KM, Diaz N, Allis CD, Banaszynski LA (2015) Histone H3.3 is required for endogenous retroviral element silencing in embryonic stem cells. Nature 522(7555):240–244
Udugama M, Chang FT, Chan FL, Tang MC, Pickett HA, McGhie JD et al (2015) Histone variant H3.3 provides the heterochromatic H3 lysine 9 tri-methylation mark at telomeres. Nucleic Acids Res 43(21):10227–10237
Bérubé NG, Smeenk CA, Picketts DJ (2000) Cell cycle-dependent phosphorylation of the ATRX protein correlates with changes in nuclear matrix and chromatin association. Hum Mol Genet 9(4):539–547
Gibbons R (2006) Alpha thalassaemia-mental retardation, X linked. Orphanet J Rare Dis 1:15
Liu XY, Gerges N, Korshunov A, Sabha N, Khuong-Quang DA, Fontebasso AM, Fleming A, Hadjadj D, Schwartzentruber J, Majewski J, Dong Z, Siegel P, Albrecht S, Croul S, Jones DTW, Kool M, Tonjes M, Reifenberger G, Faury D, Zadeh G, Pfister S, Frequent NJ (2012) ATRX mutations and loss of expression in adult diffuse astrocytic tumors carrying IDH1/IDH2 and TP53 mutations. Acta Neuropathol 124:615–625
Ramirez C, Bowman C, Maurage CA, Dubois F, Blond S, Porchet N, Escande F (2010) Loss of 1p, 19q, and 10q heterozygosity prospectively predicts prognosis of oligodendroglial tumors—towards individualized tumor treatment? Neuro Oncol 12(5):490–499
Bax DA, Little SE, Gaspar N, Perryman L, Marshall L, Viana-Pereira M, Jones TA, Williams RD, Grigoriadis A, Vassal G, Workman P, Sheer D, Reis RM, Pearson ADJ, Hargrave D, Jones C (2009) Molecular and phenotypic characterisation of paediatric glioma cell lines as models for preclinical drug development. PLoS ONE 4(4):e5209
Akuta N, Suzuki F, Kobayashi M, Fujiyama S, Kawamura Y, Sezaki H, Hosaka T, Saitoh S, Arase Y, Ikeda K, Suzuki Y, Kumada H (2020) Detection of TERT promoter mutation in serum cell-free DNA using wild-type blocking PCR combined with Sanger sequencing in hepatocellular carcinoma. J Med Virol. https://doi.org/10.1002/jmv.25724
Loussouarn D, Le Loupp AG, Frenel JS, Leclair F, Von Deimling A, Aumont M, Martin S, Campone M, Denis MG (2012) Comparison of immunohistochemistry, DNA sequencing and allele-specific PCR for the detection of IDH1 mutations in gliomas. Int J Oncol 40:2058–2062
Agarwal S, Sharma MC, Jha P, Pathak P, Suri V, Sarkar C, Chosdol K, Suri A, Kale SS, Mahapatra AK, Jha P (2013) Comparative study of IDH1 mutations in gliomas by immunohistochemistry and DNA sequencing. Neuro Oncol 15(6):718–726
Waitkus MS, Yan H (2020) Targeting isocitrate dehydrogenase mutations in cancer: emerging evidence and diverging strategies. Clin Cancer Res. https://doi.org/10.1158/1078-0432.CCR-20-1827
Han S, Liu Y, Cai SJ, Qian M, Ding J, Larion M, Gilbert MR, Yang C (2020) IDH mutation in glioma: molecular mechanisms and potential therapeutic targets. Br J Cancer 122:1580–1589
Arcella A, Palchetti S, Digiacomo L, Pozzi D, Capriotti A, Frati L, Oliva MA, Tsaouli G, Rota R, Screpanti I, MahmoudiMandCaracciolo G (2018) Brain targeting by liposome-biomolecular corona boosts anticancer efficacy of temozolomide in glioblastoma cells. ACS Chem Neurosci 9(12):3166–3174
Funding
This work has been supported by Italian Ministry of Health with Ricerca Corrente.
Author information
Authors and Affiliations
Contributions
MAO performed immunofluorescence, immunohistochemistry and proliferation assay; SS performed MGMT methylation specific PCR and DNA sequencing; SC contributed to the expansion of cell cultures; VE collected and provided glioma biopsies; FG performed the histological diagnosis of gliomas and contributed to the discussion of the data; AA performed the molecular analysis of gliomas, coordinated this study, prepared primary cell cultures from biopsies and wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Consent for submission
The work described here has not been published before and is not under consideration for publication elsewhere. The authors declare that they have no conflict of interest. All authors have approved the manuscript.
Ethical approval
This study was approved by Neuromed Ethics Committee on 20 February 2020 and registered on Clinical Trial.gov with identification number NCT 04180046.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11060_2020_3673_MOESM1_ESM.tif
Figure S1 Primary cell culture and corresponding tumour tissue FL : low grade human glioma (grade II oligodendroglioma) IDH1R132 mutant. A) Cells and tissues express glioma markers VIMENTIN, GFAP, wild type ATRX and IDH1R132H mutation. B) Electropherogram shows cDNA Sequencing of IDH1 in primary cell line (lower panel) and DNA Sequencing of IDH1 in corresponding tumor tissue (upper panel). C) Representative electropherogram of Loss of heterozygosity analysis on 1p chromosome; it shows a deletion at locus D1S508 both in cells and corresponding patient's tissue respect to the patient's blood (used as control wild type). Supplementary file1 (TIF 55990 KB)
11060_2020_3673_MOESM2_ESM.tif
Figure S2 Doubling time of primary glioblastoma cell lines (A) and low grade glioma cell lines (B). Low grade glioma cells (MI, FL) proliferate slowly than glioblastoma cells (PAP, CL, NULU, COGI). Growth curves of low grade glioma cells IDH1 mutant treated with TMZ (C) Both glioma cell lines (MI, FL) were responsive to TMZ treatment, expecially at the higher concentration used (50 μM). Western blot analysis of IDH1R132H mutation in low grade glioma cells (D). Western blot of MI and FL protein’s lysates showed a 46kDa band corresponding to the mutated protein in both cell lines analysed. Supplementary file2 (TIF 20839 KB)
Rights and permissions
About this article
Cite this article
Oliva, M.A., Staffieri, S., Castaldo, S. et al. Characterization of primary glioma cell lines derived from the patients according to 2016 CNS tumour WHO classification and comparison with their parental tumours. J Neurooncol 151, 123–133 (2021). https://doi.org/10.1007/s11060-020-03673-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-020-03673-8