Abstract
Fruit firmness and weight are among the most important fruit quality traits in fruit species. Understanding the control of fruit firmness and weight is essential for the development of domestication research approaches and for the implementation of new breeding strategies. A forward genetic study for these traits was performed using two F1 sweet cherry (Prunus avium) progenies derived from modern cultivars. Quantitative trait locus (QTL) analysis allowed the identification of genomic regions accounting for most of the phenotypic variation in both families. In addition, screening the Prunus persica genome v1.0 permitted the identification of putative candidate genes underlying the QTL with the major effect for fruit weight (LG5) and the one for firmness (LG6). A colocalization of QTLs and candidate genes was found in peach, apple, and tomato. These results give new insights of the interaction between fruit firmness and fruit weight and provide new cues for the identification of genes implicated in the control of these traits. The colocalization of genomic regions between progenies issued from modern cultivars and from modern cultivars × wild individuals suggests the absence of allele fixation within genes controlling fruit firmness and size, two traits potentially involved in domestication/diversification in sweet cherry.


Similar content being viewed by others
References
Anastasiou E, Kenz S, Gerstung M, MacLean D, Timmer J, Fleck C, Lenhard M (2007) Control of plant organ size by KLUH/CYP78A5-dependent intercellular signaling. Dev Cell 13:843–856
Arunyawat U, Capdeville G, Decroocq V, Mariette S (2012) Linkage disequilibrium in French wild cherry germplasm and worldwide sweet cherry germplasm. Tree Genet Genome 8:737–755. doi:10.1007/s11295-011-0460-9
Atkinson RG et al (2012) Down-regulation of POLYGALACTURONASE1 alters firmness, tensile strength and water loss in apple (Malus x domestica) fruit. BMC Plant Biol 12
Badenes ML, Martinez-Calvo J, Llacer G (1998) Analysis of apricot germplasm from the European ecogeographical group. Euphytica 102:93–99. doi:10.1023/a:1018332312570
Cabrera A et al (2012) Rosaceae conserved orthologous sequences marker polymorphism in sweet cherry germplasm and construction of a SNP-based map. Tree Genet Genome 8:237–247. doi:10.1007/s11295-011-0436-9
Cantin CM, Crisosto CH, Ogundiwin EA, Gradziel T, Torrents J, Moreno MA, Gogorcena Y (2010a) Chilling injury susceptibility in an intra-specific peach Prunus persica (L.) Batsch progeny. Postharvest Biol Technol 58:79–87
Cantin CM, Gogorcena Y, Moreno AM (2010b) Phenotypic diversity and relationships of fruit quality traits in peach and nectarine Prunus persica (L.) Batsch breeding progenies. Euphytica 171:211–226. doi:10.1007/s10681-009-0023-4
Cao K, Wang L, Zhu G, Fang W, Chen C, Luo J (2012a) Genetic diversity, linkage disequilibrium, and association mapping analyses of peach (Prunus persica) landraces in China. Tree Genet Genome 8:975–990. doi:10.1007/s11295-012-0477-8
Cao Y, Tang X, Giovannoni J, Xiao F, Liu Y (2012b) Functional characterization of a tomato COBRA-like gene functioning in fruit development and ripening BMC Plant Biol 12 doi:10.1186/1471-2229-12-211
Castède S et al (2014) Genetic determinism of phenological traits highly affected by climate change in Prunus avium: flowering date dissected into chilling and heat requirements. New Phytol. doi:10.1111/nph.12658
Cevik V, Ryder CD, Popovich A, Manning K, King GJ, Seymour GB (2010) A FRUITFULL-like gene is associated with genetic variation for fruit flesh firmness in apple (Malus domestica Borkh.). Tree Genet Genome 6:271–279. doi:10.1007/s11295-009-0247-4
Chagne D et al (2012) Genome-wide SNP detection, validation, and development of an 8K SNP array for apple. PLoS ONE 7:e31745–e31745
Chakrabarti M et al (2013) A cytochrome P450 regulates a domestication trait in cultivated tomato. Proc Natl Acad Sci U S A 110:17125–17130
Chapman NH et al (2012) High-resolution mapping of a fruit firmness-related quantitative trait locus in tomato reveals epistatic interactions associated with a complex combinatorial locus. Plant Physiol 159:1644–1657. doi:10.1104/pp. 112.200634
Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676
Cong B, Liu JP, Tanksley SD (2002) Natural alleles at a tomato fruit size quantitative trait locus differ by heterochronic regulatory mutations. Proc Natl Acad Sci U S A 99:13606–13611. doi:10.1073/pnas.172520999
Cosgrove DJ (2005) Growth of the plant cell wall. Nat Rev Mol Cell Biol 6:850–861. doi:10.1038/nrm1746
Costa F et al (2010) QTL dynamics for fruit firmness and softening around an ethylene-dependent polygalacturonase gene in apple (Malusxdomestica Borkh.). J Exp Bot 61:3029–3039. doi:10.1093/jxb/erq130
Cui L, Li J, Zhang T, Guo Q, Xu J, Lou Q, Chen J (2014) Identification and expression analysis of D-type cyclin genes in early developing fruit of cucumber (Cucumis sativus L.). Plant Mol Biol Report 32:209–218. doi:10.1007/s11105-013-0637-5
De Franceschi P et al (2013) Cell number regulator genes in Prunus provide candidate genes for the control of fruit size in sweet and sour cherry. Mol Breed 32:311–326. doi:10.1007/s11032-013-9872-6
Dirlewanger E et al (1999) Mapping QTLs controlling fruit quality in peach (Prunus persica (L.) Batsch). Theor Appl Genet 98:18–31
Dirlewanger E, Graziano E, Joobeur T, Garriga-Caldere F, Cosson P, Howad W, Arus P (2004) Comparative mapping and marker-assisted selection in Rosaceae fruit crops. Proc Natl Acad Sci U S A 101:9891–9896. doi:10.1073/pnas.0307937101
Dirlewanger E et al (2012) Comparison of the genetic determinism of two key phenological traits, flowering and maturity dates, in three Prunus species: peach, apricot and sweet cherry. Heredity 109:280–292
Doebley JF, Gaut BS, Smith BD (2006) The molecular genetics of crop domestication. Cell 127:1309–1321. doi:10.1016/j.cell.2006.12.006
Doganlar S, Frary A, Daunay MC, Lester RN, Tanksley SD (2002) Conservation of gene function in the Solanaceae as revealed by comparative mapping of domestication traits in eggplant. Genetics 161:1713–1726
Doong RL, Mohnen D (1998) Solubilization and characterization of a galacturonosyltransferase that synthesizes the pectic polysaccharide homogalacturonan. Plant J 13:363–374. doi:10.1046/j.1365-313X.1998.00042.x
Eduardo I, Pacheco I, Chietera G, Bassi D, Pozzi C, Vecchietti A, Rossini L (2011) QTL analysis of fruit quality traits in two peach intraspecific populations and importance of maturity date pleiotropic effect. Tree Genet Genome 7:323–335. doi:10.1007/s11295-010-0334-6
Frankel OH, Brown AHD, Burdo JJ (1995) The conservation of plant biodiversity. Cambridge University Press, Cambridge
Frary A et al (2000) fw2.2: a quantitative trait locus key to the evolution of tomato fruit size. Science 289:85–88. doi:10.1126/science.289.5476.85
Goffinet MC, Robinson TL, Lakso AN (1995) A comparison of Empire apple fruit size and anatomy in unthinned and hand-thinned trees. J Hortic Sci 70:375–387
Gross BL, Olsen KM (2010) Genetic perspectives on crop domestication. Trends Plant Sci 15:529–537. doi:10.1016/j.tplants.2010.05.008
Guyer DE, Sinha NK, Chang TS, Cash JN (1993) Physicochemical and sensory characteristics of selected Michigan sweet cherry (Prunus avium L) cultivars. J Food Qual 16:355–370
Holland JB, Nyquist WE, Cervantes-Martinez CT (2003) Estimating and interpreting heritability for plant breeding: an update. Plant Breed Rev 22:9–112
Illa E et al (2011) Saturating the Prunus (stone fruits) genome with candidate genes for fruit quality. Mol Breed 28:667–682. doi:10.1007/s11032-010-9518-x
Johnson DS (1994) Influence of time of flower and fruit thinning on the firmness of Coxs Orange Pippin apples at harvest and after storage. J Hortic Sci 69:197–203
Jung S et al. (2012) Whole genome comparisons of Fragaria, Prunus and Malus reveal different modes of evolution between Rosaceous subfamilies Bmc Genomics 13 doi:10.1186/1471-2164-13-129
Kenis K, Keulemans J, Davey MW (2008) Identification and stability of QTLs for fruit quality traits in apple. Tree Genet Genome 4:647–661. doi:10.1007/s11295-008-0140-6
Klagges C et al. (2013) Construction and comparative analyses of highly dense linkage maps of two sweet cherry intra-specific progenies of commercial cultivars. PLoS ONE 8 doi:10.1371/journal.pone.0054743
Koinange EMK, Singh SP, Gepts P (1996) Genetic control of the domestication syndrome in common bean. Crop Sci 36:1037–1045
Lamb NC (1953) Notes on the inheritance of some characters in the sweet cherry Prunus avium. Proc Am Soc Hortic Sci 61:293–298
Lerceteau-Kohler E, Moing A, Guerin G, Renaud C, Petit A, Rothan C, Denoyes B (2012) Genetic dissection of fruit quality traits in the octoploid cultivated strawberry highlights the role of homoeo-QTL in their control. Theor Appl Genet 124:1059–1077. doi:10.1007/s00122-011-1769-3
Li HH, Bradbury P, Ersoz E, Buckler ES, Wang JK (2011) Joint QTL linkage mapping for multiple-cross mating design sharing one common parent. PLoS ONE 6 doi:10.1371/journal.pone.0017573
Longhi S, Moretto M, Viola R, Velasco R, Costa F (2012) Comprehensive QTL mapping survey dissects the complex fruit texture physiology in apple (Malus x domestica Borkh.). J Exp Bot 63:1107–1121. doi:10.1093/jxb/err326
Mann HS, Alton JJ, Kim S, Tong CBS (2008) Differential expression of cell-wall-modifying genes and novel cDNAs in apple fruit during storage. J Am Soc Hortic Sci 133:152–157
Mariette S, Tavaud M, Arunyawat U, Capdeville G, Millan M, Salin F (2010) Population structure and genetic bottleneck in sweet cherry estimated with SSRs and the gametophytic self-incompatibility locus Bmc Genetics 11 doi:10.1186/1471-2156-11-77
Martinez-Garcia PJ et al (2013) High density SNP mapping and QTL analysis for fruit quality characteristics in peach (Prunus persica L.). Tree Genet Genome 9:19–36. doi:10.1007/s11295-012-0522-7
Meyer RS, Purugganan MD (2013) Evolution of crop species: genetics of domestication and diversification. Nat Rev Genet 14:840–852. doi:10.1038/nrg3605
Miller AJ, Gross BL (2011) From forest to field: perennial fruit crop domestication. Am J Bot 98:1389–1414. doi:10.3732/ajb.1000522
Minic Z, Jouanin L (2006) Plant glycoside hydrolases involved in cell wall polysaccharide degradation. Plant Physiol Biochem 44:435–449. doi:10.1016/j.plaphy.2006.08.001
Munos S et al (2011) Increase in tomato locule number is controlled by two single-nucleotide polymorphisms located near WUSCHEL. Plant Physiol 156:2244–2254. doi:10.1104/pp. 111.173997
Ogundiwin EA, Peace CP, Gradziel TM, Parfitt DE, Bliss FA, Crisosto CH (2009) A fruit quality gene map of Prunus Bmc Genomics 10 doi:10.1186/1471-2164-10-587
Olmstead JW, Lezzoni AF, Whiting MD (2007) Genotypic differences in sweet cherry fruit size are primarily a function of cell number. J Am Soc Hortic Sci 132:697–703
Panda S, Martin JP, Aguinagalde I, Mohanty A (2003) Chloroplast DNA variation in cultivated and wild Prunus avium L: a comparative study. Plant Breed 122:92–94. doi:10.1046/j.1439-0523.2003.00768.x
Paterson AH (2002) What has QTL mapping taught us about plant domestication? New Phytol 154:591–608. doi:10.1046/j.1469-8137.2002.00420.x
Peace CP, Crisosto CH, Gradziel TM (2005) Endopolygalacturonase: a candidate gene for freestone and melting flesh in peach. Mol Breed 16:21–31. doi:10.1007/s11032-005-0828-3
Peace C et al. (2012) Development and evaluation of a genome-wide 6K SNP array for diploid sweet cherry and tetraploid sour cherry. PLoS ONE 7 doi:10.1371/journal.pone.0048305
Pose S, Paniagua C, Cifuentes M, Blanco-Portales R, Quesada MA, Mercado JA (2013) Insights into the effects of polygalacturonase FaPG1 gene silencing on pectin matrix disassembly, enhanced tissue integrity, and firmness in ripe strawberry fruits. J Exp Bot 64:3803–3815
Purugganan MD, Fuller DQ (2009) The nature of selection during plant domestication. Nature 457:843–848. doi:10.1038/nature07895
Quesada MA et al (2009) Antisense down-regulation of the FaPG1 gene reveals an unexpected central role for polygalacturonase in strawberry fruit softening. Plant Physiol 150:1022–1032
Quilot B, Wu BH, Kervella J, Genard M, Foulongne M, Moreau K (2004) QTL analysis of quality traits in an advanced backcross between Prunus persica cultivars and the wild relative species P-davidiana. Theor Appl Genet 109:884–897. doi:10.1007/s00122-004-1703-z
Romano GS, Cittadini ED, Pugh B, Schouten R (2006) Sweet cherry quality in the horticultural production chain. Stewart Postharvest Rev 2:1–9. doi:10.2212/spr.2006.6.2
Rosyara U et al. (2013) Fruit size QTL identification and the prediction of parental QTL genotypes and breeding values in multiple pedigreed populations of sweet cherry Mol Breed:1–13 doi:10.1007/s11032-013-9916-y
Ruess F, Stosser R (1993) Investigations on the intercellular system of apple fruit by digital image-processing methods. Gartenbauwissenschaft 58:197–205
Ruiz D, Egea J (2008) Phenotypic diversity and relationships of fruit quality traits in apricot (Prunus armeniaca L.) germplasm. Euphytica 163:143–158
Sato S et al (2012) The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485:635–641. doi:10.1038/nature11119
Schreckenberg K, Awono A, Degrande A, Mbosso C, Ndoye O, Tchoundjeu Z (2006) Domesticating indigenous fruit trees as a contribution to poverty reduction. Forest Trees Livelihoods 16:35–51
Silva C, Garcia-Mas J, Sanchez AM, Arus P, Oliveira M (2005) Looking into flowering time in almond (Prunus dulcis (Mill) D. A. Webb): the candidate gene approach. Theor Appl Genet 110:959–968
Tavaud M (2000) Diversité génétique du cerisier doux (Prunus avium L.) sur son aire de répartition : comparaison avec ses espèces apparentées (P. cerasus et P. x gondouinii) et son compartiment sauvage., Thèse de Doctorat Ecole Nationale Supérieur Agronomique de Montpellier
Van Tassel DL, DeHaan LR, Cox TS (2010) Missing domesticated plant forms: can artificial selection fill the gap? Evol Appl 3:434–452. doi:10.1111/j.1752-4571.2010.00132.x
Velasco R et al (2010) The genome of the domesticated apple (Malus x domestica Borkh.). Nat Genet 42:833. doi:10.1038/ng.654
Verde I et al. (2012) Development and Evaluation of a 9K SNP Array for Peach by Internationally Coordinated SNP Detection and Validation in Breeding Germplasm PLoS ONE 7 doi:10.1371/journal.pone.0035668
Verde I et al (2013) The high-quality draft genome of peach (Prunus persica) identifies unique patterns of genetic diversity, domestication and genome evolution. Nat Genet 45:487–U447
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78. doi:10.1093/jhered/93.1.77
Wakasa Y, Kudo H, Ishikawa R, Akada S, Senda M, Niizeki M, Harada T (2006) Low expression of an endopolygalacturonase gene in apple fruit with long-term storage potential. Postharvest Biol Technol 39:193–198. doi:10.1016/j.postharvbio.2005.10.005
Wang D, Karle R, Iezzoni AF (2000) QTL analysis of flower and fruit traits in sour cherry. Theor Appl Genet 100:535–544. doi:10.1007/s001229900121
Whiting MD, Ophardt D, McFerson JR (2006) Chemical blossom thinners vary in their effect on sweet cherry fruit set, yield, fruit quality, and crop value. Horttechnology 16:66–70
Yamamoto T et al (2001) Characterization of morphological traits based on a genetic linkage map in peach. Breed Sci 51:271–278. doi:10.1270/jsbbs.51.271
Zhang GR, Sebolt AM, Sooriyapathirana SS, Wang DC, Bink M, Olmstead JW, Iezzoni AF (2010) Fruit size QTL analysis of an F-1 population derived from a cross between a domesticated sweet cherry cultivar and a wild forest sweet cherry. Tree Genet Genome 6:25–36. doi:10.1007/s11295-009-0225-x
Zhang N, Brewer MT, van der Knaap E (2012) Fine mapping of fw3.2 controlling fruit weight in tomato. Theor Appl Genet 125:273–284
Acknowledgments
The authors thank the technical team of the department of “Biologie du Fruit et Pathologie” of the “Institut National de Recherche Agronomique,” Jacques Joly and Lydie Fouilhaux, for producing the hybrids and collaborating actively in the phenotyping, and the Experimental Unit of INRA-Toulenne (UEA) for growing the trees.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online Resource 1
Markers used in the final parental maps. (PDF 336 kb)
Online Resource 2
Mean, s.d., IQR (interquartile range), c.v., value range, number of individuals (n) for the sweet cherry (Prunus avium) populations derived from the crosses between ‘Regina’ x ‘Lapins’ and ‘Regina’ x ‘Garnet’ observed during the different years of evaluation of the fruit firmness and weight. (PDF 101 kb)
Online Resource 3
Fruit firmness (Ff) and weight (Fw) of haplotype combinations for the QTL region on the extreme end of linkage group 5 for ‘Regina’ × ‘Garnet’ progeny. Significant difference between means (P > 0,05) are indicated by a star. (PDF 164 kb)
Online Resource 4
Genes located on the Quantitative Trait Loci for fruit firmness and fruit size in linkage groups 5 and 6. Linkage group, sequence name, sequence description, sequence length, #Hits, min. eValue, mean Similarity, number of gene ontologies, description of gene ontologies, enzyme codes and InterProScan codes are provided for each gene. (PDF 1054 kb)
Online Resource 5
Candidate genes located on the Quantitative Trait Loci for fruit firmness and fruit size in linkage groups 5 and 6. Linkage group, sequence name, sequence description, sequence length, #Hits, min. eValue, mean Similarity, number of gene ontologies, description of gene ontologies, enzyme codes and InterProScan codes are provided for each candidate gene. (PDF 207 kb)
Online Resource 6
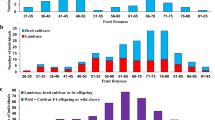
Distribution of fruit weight (left) and firmness (right) measurements for two cross-pollination populations derived from ‘Regina’ and ‘Lapins’ (a) and ‘Regina’ and ‘Garnet’ (b) (PDF 214 kb)
Rights and permissions
About this article
Cite this article
Campoy, J.A., Le Dantec, L., Barreneche, T. et al. New Insights into Fruit Firmness and Weight Control in Sweet Cherry. Plant Mol Biol Rep 33, 783–796 (2015). https://doi.org/10.1007/s11105-014-0773-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-014-0773-6