Abstract
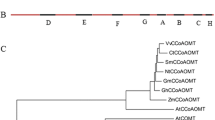
The biosynthesis of salvianolic acid B shares the phenylpropanoid pathway with lignin, and cinnamoyl CoA reductase (CCR; EC 1.2.1.44) is a specific enzyme in the lignin pathway. In this study, a CCR gene (SmCCR1) from Salvia miltiorrhiza Bunge was cloned using DNA walking technology (GenBank ID: JF798634). The full-length SmCCR1 is 2,489 bp long and consists of four introns and five exons encoding a polypeptide of 324 amino acid residues. Sequence alignment revealed that SmCCR1 shares 83 % identity with CCR sequences reported in Camellia oleifera and other plant species. Expression pattern analysis indicated that expression of SmCCR1 can be induced by exposure to Xanthomonas campestris pv. Campestris or methyl jasmonate. To demonstrate its functioning, we selected a 296-bp fragment and established an RNA interference construct that was introduced into S. miltiorrhiza by Agrobacterium tumefaciens-mediated gene transfer. Transgenic plants exhibited dwarfing phenotypes, and both syringyl and guaiacyl lignin monomers were decreased more than 60 %. In contrast, biosynthesis of phenolic acids—danshensu, rosmarinic acid, and salvianolic acid B—was strongly induced by 2.03-, 1.41-, and 1.45-fold, respectively, in the roots of transgenic plants from line CCR-10. Consistent with these phytochemical changes, downregulation of SmCCR1 also affected the expression of related genes in the phenolics and lignin biosynthetic pathways. Our results also provide potential opportunities for engineering danshensu and salvianolic acid B production in S. miltiorrhiza.







Similar content being viewed by others
References
Bi CL, Fang C, Jackson L, Gill BS, Li WL (2011) Expression of lignin biosynthetic genes in wheat during development and upon infection by fungal pathogens. Plant Mol Biol Report 29:149–161
Capeleti I, Bonini EA, Ferrarese MDL, Teixeira ACN, Krzyzanowski FC, Ferrarese-Filho O (2005) Lignin content and peroxidase activity in soybean seed coat susceptible and resistant to mechanical damage. Acta Physiol Plant 27:103–108
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure from small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Escamilla-Treviño L, Shen H, Uppalapati SR, Ray T, Tang YH, Hernandez T, Yin YB, Xu Y, Dixon RA (2010) Switchgrass (Panicum virgatum) possesses a divergent family of cinnamoyl CoA reductases with distinct biochemical properties. New Phytol 185:143–155
Fan L, Linker R, Gepstein S, Tanimoto E, Yamamoto R, Neumann PM (2006) Progressive inhibition by water deficit of cell wall extensibility and growth along the elongation zone of maize roots is related to increased lignin metabolism and progressive stelar accumulation of wall phenolics. Plant Physiol 140:603–612
Gleave AP (1992) A versatile binary vector system with a T-DNA organisational structure conducive to efficient integration of cloned DNA into the plant genome. Plant Mol Biol 20:1203–1207
Goujon T, Ferret V, Mila I, Pollet B, Ruel K, Burlat V, Joseleau JP, Barrière Y, Lapièrre C, Jouanin L (2003) Down-regulation of the AtCCR1 gene in Arabidopsis thaliana: effects on phenotype, lignins and cell wall degradability. Planta 217:218–228
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Hoffmann L, Besseau S, Geoffroy P, Ritzenthaler C, Meyer D, Lapierre C, Pollet B, Legranda M (2004) Silencing of hydroxycinnamoyl-coenzyme A shikimate/quinate hydroxycinnamoyltransferase affects phenylpropanoid biosynthesis. Plant Cell 16:1446–1465
Holsters M, Waele D, Depicker A, Messens E, Montagu M, Schell J (1978) Transfection and transformation of Agrobacterium tumefaciens. Mol Gen Genet 163:181–187
Hu P, Luo G, Zhao Z, Jiang Z (2005) Quality assessment of Radix alviae Miltiorrhizae. Chem Pharm Bull (Tokyo) 53:481–486
Hu YS, Di P, Chen JF, Xiao Y, Zhang L, Chen WS (2010) Isolation and characterization of a gene encoding cinnamoyl-CoA reductase from Isatis indigotica Fort. Mol Biol Rep. doi:10.1007/s11033-010-0333-6
Kim KH, Janiak V, Petersen M (2004) Purification, cloning and functional expression of hydroxyphenylpyruvate reductase involved in rosmarinic acid biosynthesis in cell cultures of Coleus blumei. Plant Mol Biol 54:311–323
Kirsi M, Oksman C, Dirk L (2004) Plant cell factories in the post-genomic era: new ways to produce designer secondary metabolites. Trends Plant Sci 9:432–440
Lacombe E, Hawkins S, Doorsselaere J, Piquemal J, Goffner D, Poeydomenge O, Boudet AM, Grima-Pettenati J (1997) Cinnamoyl CoA reductase, the first committed enzyme of the lignin branch biosynthetic pathway: cloning, expression and phylogenetic relationships. Plant J 11:429–441
Larsen K (2004) Molecular cloning and characterization of cDNAs encoding cinnamoyl CoA reductase (CCR) from barley (Hordeum vulgare) and potato (Solanum tuberosum). J Plant Physiol 161:105–121
Lauvergeat V, Lacomme C, Lacombe E, Lasserre E, Roby D, Grima-Pettenati J (2001) Two cinnamoyl-CoA reductase (CCR) genes from Arabidopsis thaliana are differentially expressed during development and in response to infection with pathogenic bacteria. Phytochemistry 57:1187–1195
Leple JC, Dauwe R, Morreel K, Storme V, Lapierre C, Pollet B, Naumann A, Kang KY, Kim H, Ruel K et al (2007) Downregulation of cinnamoyl-coenzyme A reductase in poplar: multiple-level phenotyping reveals effects on cell wall polymer metabolism and structure. Plant Cell 19:3669–3691
Li YG, Song L, Liu M, Hu ZB, Wang ZT (2009) Advancement in analysis of Salviae miltiorrhizae Radix et Rhizoma (Danshen). J Chromatogr 1216:1941–1953
Liu AH, Li L, Xu M, Lin YH, Guo HZ, Guo DA (2006) Simultaneous quantification of six major phenolic acids in the roots of Salvia miltiorrhiza and four related traditional Chinese medicinal preparations by HPLC-DAD method. J Pharm Biomed Anal 41:48–56
Lummerzheim M, De Oliveira D, Castresana C, Miguens FC, Louzada E, Roby D, Van Montagu M, Timmerman B (1993) Identification of compatible and incompatible interactions between Arabidopsis thaliana and Xanthomonas campestris pv. campestris and characterization of the hypersensitive response. Mol Plant Microbe Interact 6:532–544
Ma L, Zhang X, Guo H, Gan Y (2006) Determination of fourwater-soluble compounds in Salvia miltiorrhiza Bunge by high performance liquid chromatography with a coulometric electrode array system. J Chromatogr B Analyt Technol Biomed Life Sci 833:260–263
Minami E, Ozeki Y, Matsuoka M, Koiruka N, Tanaka Y (1989) Structure and some characterization of the gene for phenylalanineammonia-lyase from rice plants. Eur J Biochem 185:19–25
Mir Derikvand M, Sierra JB, Ruel K, Pollet B, Do CT, Thevenin J, Buffard D, Jouanin L, Lapierre C (2008) Redirection of the phenylpropanoid pathway to feruloyl malate in Arabidopsis mutants deficient for cinnamoyl-CoA reductase 1. Planta 227:943–956
Petersen M, Haüsler E, Karwatzki B, Meinhard J (1993) Proposed biosynthetic pathway for rosmarinic acid in cell cultures of Coleus blumei Benth. Planta 189:10–14
Pichon M, Courbou I, Beckert M, Boudet AM, Grima-Pettenati J (1998) Cloning and characterization of two maize cDNAs encoding cinnamoyl-CoA reductase (CCR) and differential expression of the corresponding genes. Plant Mol Biol 38:671–676
Pincon G, Chabannes M, Lapierre C, Pollet B, Ruel K, Joseleau JP, Boudet AM, Legrand M (2001) Simultaneous down-regulation of caffeic/5-hydroxy ferulic acid-O-methyltransferase I and cinnamoyl-coenzyme A reductase in the progeny from a cross between tobacco lines homozygous for each transgene. Consequences for plant development and lignin synthesis. Plant Physiol 126:145–155
Rest B, Danoun S, Boudet AM, Rochange SF (2006) Down-regulation of cinnamoyl-CoA reductase in tomato (Solanum lycopersicum L.) induces dramatic changes in soluble phenolic pools. J Exp Bot 57:1399–1411
Shadle G, Chen F, Srinivasa Reddy MS, Jackson L, Nakashima J, Dixon RA (2007) Down-regulation of hydroxycinnamoyl CoA: shikimate hydroxycinnamoyl transferase in transgenic alfalfa affects lignification, development and forage quality. Phytochemistry 68:1521–1529
Tu Y, Rochfort S, Liu ZQ, Ran YD, Griffith M, Badenhorst P, Louie GV, Bowman EB, Smith KF, Noel JP, Mouradov A, Spangenberg G (2010) Functional analyses of caffeic acid O-methyltransferase and cinnamoyl-CoA-reductase genes from perennial ryegrass (Lolium perenne). Plant Cell 22:3357–3373
Vance CP, Kirk TK, Sherwood RT (1980) Lignification as a defence mechanism of disease resistance. Annu Rev Phytopathol 18:259–288
Vanholme R, Morreel K, Ralph J, Boerjan W (2008) Lignin engineering. Curr Opin Plant Biol 11:278–285
Wadenbäck J, Arnold S, Egertsdotter U, Walter MH, Grima-Pettenati J, Goffner D, Gellerstedt G, Gullion T, Clapham D (2008) Lignin biosynthesis in transgenic Norway spruce plants harboring an antisense construct for cinnamoyl CoA reductase (CCR). Transgen Res 17:379–392
Wang XJ, Wang ZB, Xu JX (2005) Effect of salvianic acid A on lipid peroxidation and membrane permeability in mitochondria. J Ethnopharmacol 97:441–445
Wang G, Sun B, Zhu H, Gao Y, Li XR, Xue DB, Jiang HC (2010) Protective effects of emodin combined with danshensu on experimental severe acute pancreatitis. Inflamm Res 59:479–488
Wu L, Qiao H, Li Y, Li L (2007) Protective roles of puerarin and Danshensu on acute ischemic myocardial injury in rats. Phytomedicine 14:652–658
Yan YP, Wang ZZ (2007) Genetic transformation of the medicinal plant Salvia miltiorrhiza by Agrobacterium tumefaciens-mediated method. Plant Cell Tiss Org Cult 88:175–184
Zhang Y, Yan YP, Wang ZZ (2010) The Arabidopsis PAP1 transcription factor plays an important role in the enrichment of phenolic acids in Salvia miltiorrhiza. J Agric Food Chem 58:12168–12175
Acknowledgments
This work was supported by the “Fundamental Research Funds for the Central Universities” (Program No. GK200901014) and “Innovation Funds of Graduate Programs, Shaanxi Normal University” (2011CXS034).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, Z., Cui, L., Chen, C. et al. Downregulation of Cinnamoyl CoA Reductase Affects Lignin and Phenolic Acids Biosynthesis in Salvia miltiorrhiza Bunge. Plant Mol Biol Rep 30, 1229–1236 (2012). https://doi.org/10.1007/s11105-012-0444-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-012-0444-4