Abstract
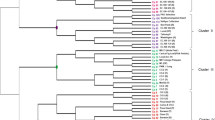
A set of 81 new microsatellite markers for Carica papaya L. previously identified by data mining using freely available sequence information from Genbank were tested for polymorphism using 30 germplasm accessions from the Papaya Germplasm Bank (PGM) at Embrapa Mandioca e Fruticultura Tropical (CNPMF) and 18 landraces. The data were used to estimate pairwise genetic distances between the genotypes. A neighbor-joining based dendrogram was used to define clusters and infer possible genetic structuring of the collection. Most microsatellites were polymorphic (73%), with an observed number of alleles per locus ranging from one to eleven. The levels of observed and expected heterozygosity for 51 polymorphic loci varied from 0.00 to 0.85 and from 0.08 to 0.82, averaging 0.19 and 0.59, respectively. Forty-four percent of microsatellites showed polymorphism information content (PIC) higher than 0.50. The compound microsatellites seem to be more informative than dinucleotide and trinucleotide repeats in average alleles per locus and PIC. Among dinucleotides, AG/TC or GA/CT repeat motifs exhibited more informativeness than TA/AT, GT/CA and TG/AC repeat motifs. The neighbor-joining analysis based on shared allele distance could differentiate all the papaya accessions and landraces as well as differences in their genetic structure. This set of markers will be useful for examining parentage, inbreeding and population structure in papaya.

Similar content being viewed by others
References
Alvizo VHF, Rojkind MC (1987) Resistencia al virus mancha anular del papaya en Carica cauliflora. Rev Mex Fitopat 5:61–62
Aradhya MK, Manshardt RM, Zee F, Morden CW (1999) A phylogenetic analysis of the genus Carica L (Caricaceae) based on restriction fragment length variation in a cpDNA intergenic spacer region. Genet Resour Crop Evol 46:579–586
Arumuganathan K, Earle ED (1991) Nuclear DNA content of some important plant species. Plant Mol Biol Rep 9:208–218
Badillo VM (2000) Carica L vs Vasconcella St Hil (Caricaceae): con la rehabilitación de este úlltimo. Ernstia 10:74–79
Baumung R, Simianer H, Hoffmann I (2004) Genetic diversity studies in farm animals - a survey. J Anim Breed Genet 121:361–373
Brondani RPV, Brondani C, Tarchini R, Grattapaglia D (1998) Development characterization and mapping of microsattelite markers in Eucalyptus grandis and E. urophylla. Theor Appl Genet 97:816–827
Carrasco B, Avila P, Perez-Diaz J, Muñoz P, Garcia R, Lavandero B, Zurita-Silva A, Retamales JB, Caligari PDS (2009) Genetic structure of highland papayas (Vasconcellea pubescens (Lenné et C Koch) Badillo) cultivated along a geographic gradient in Chile as revealed by Inter Simple Sequence Repeats (ISSR). Genet Resour Crop Evol 56:331–337
Collevatti RG, Brondani RV, Grattapaglia D (1999) Development and characterisation of microsatellite markers for genetic analysis of a Brazilian endangered tree species Caryocar brasiliense. Heredity 83:748–756
Condit R, Hubbel SP (1991) Abundance and DNA sequence of two-base repeat regions in tropical tree genomes. Genome 34:66–71
Creste S, Tulmann Neto A, Figueira A (2001) Detection of single sequence repeat polymorfisms in denaturing polyacrylamide sequencing gels by silver staining. Plant Mol Biol Rep 19:299–306
Deputy JC, Ming R, Ma H, Liu Z, Fitch MMM, Wang M, Manshardt R, Stiles JI (2002) Molecular markers for sex determination in papaya (Carica papaya L.). Theor Appl Genet 106:107–111
Dillon S, Ramage C, Drew R, Ashmore S (2005) Genetic mapping of a PRSV-P resistance gene in “highland papaya” based on inheritance of RAF markers. Euphytica 145:11–23
Dow BD, Ashley MV (1996) Microsatellite analysis of seed dispersal and sapling parentage in bur oak Quercus macrocarpa. Mol Ecol 5:615–627
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Drew RA, O’Brien CM, Magdalita PM (1998) Development of interspecific Carica hybrids. Acta Hortic 461:285–292
Eustice M, Yu Q, Lai CW, Hou S, Thimmapuram J, Liu L, Alam M, Moore PH, Presting GG, Ming R (2008) Development and application of microsatellite markers for genomic analysis of papaya. Tree Genet Genomes 4:333–341
Faostat: The United Nations: Food and Agriculture Organization of The United Nations 2007 Available via: http://faostatfaoorg/site/567/defaultaspx#ancor Accessed 04 June 2009
Ferguson ME, Burow MD, Schultz SR, Bramel PJ, Paterson AH, Kresovich S, Mitchell S (2004) Microsatellite identification and characterization in peanut (Arachis hypogaea L). Theor Appl Genet 108:1064–1070
Gianfranceschi L, Seglias N, Tarchini R, Komjanc M, Gessler C (1998) Simple sequence repeats for the genetic analysis of apple. Theor Appl Genet 96:1069–1076
Guilford P, Prakash S, Zhu JM, Rikkerink E, Gardiner S, Bassett H, Forster R (1997) Microsatellites in Malus × domestica (apple): abundance polymorphism and cultivar identification. Theor Appl Genet 94:249–254
Hamrick JL, Godt MJ (1996) Conservation genetics of endemic plant species. In: Avise JC, Hamrick JL (eds) Conservation genetics case histories from nature. Chapman and Hall, New York USA, pp 281–304
Horovitz S, Jiménez H (1967) Cruzamientos interespecificos e intergenericos en Caricaceas y sus implicaciones fitotecnicas. Agron Trop 17:323–344
Jansen J (2005) Construction of linkage maps in full-sib families of diploid outbreeding species by minimizing the number of recombinations in hidden inheritance vectors. Genetics 170:2013–2025
Jobin-Decor MP, Graham GC, Henry RJ, Drew RA (1997) RAPD and isozyme analysis of genetic relationships between Carica papaya and wild relatives. Genet Resour Crop Evol 44:471–477
Karp A (2002) The new genetic Era: will it help us in managing genetic diversity? In: Engels JMM, Ramanatha Rao V, Brown AHD, Jackson MT (eds) Managing plant genetic diversity. IPGRI, Rome, pp 43–56
Kim M, Moore P, Zee F, Fitch MMM, Steiger D, Manshardt R, Paull R, Drew R, Sekioka T, Ming R (2002) Genetic diversity of Carica papaya as revealed by AFLP markers. Genome 45:503–512
Kyndt T, Haegeman A, Van Glabeke S, Maertens I, Van Droogenbroeck B, Roldán-Ruiz I, Gheysen G (2005) Isolation and characterization of microsatellite loci in the highland papaya Vasconcelle × heilbornii V. Badillo (Caricaceae). Mol Ecol Notes 5:590–592
Kyndt T, Van Droogenbroeck B, Haegeman A, Roldán-Ruiz I, Gheysen G (2006) Cross-species microsatellite amplification in Vasconcellea and related genera and their use in germplasm classification. Genome 49:786–798
Liang F, Deng Q, Wang Y, Xiong Y, Jin D, Li J, Wang B (2004) Molecular marker-assisted selection for yield-enhancing genes in the progeny of “9311 × O rufipogon” using SSR. Euphytica 139:159–165
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Liu Z, Moore PH, Ma H, Ackerman CM, Makandar R, Yu Q, Pear HM, Kim MS, Charlton JW, Stiles JI, Zee FT, Paterson AH, Ming R (2004) A primitive Y chromosome in papaya marks incipient sex chromosome evolution. Nature 427:348–352
Lowe A, Harris S, Ashton P (2004) Ecological genetics: design analysis and application. Blackwell Publishing, Oxford
Luo ZW, Hackett CA, Bradshaw JE, Mcnicol JW, Milbourne D (2001) Construction of a genetic linkage map in tetraploid species using molecular markers. Genetics 157:1369–1385
Lynch M, Milligan BG (1994) Analysis of population genetic structure with RAPD markers. Mol Ecol 3:91–99
Manifesto M, Schlatter AR, Hopp HE, Suárez EY, Dubcovsky J (2001) Quantitative evaluation of genetic diversity in wheat germplasm using molecular markers. Crop Sci 41:682–690
Martins-Lopes P, Lima-Brito J, Gomes S, Meirinhos J, Santos L, Guedes-Pinto H (2007) RAPD and ISSR molecular markers in Olea europaea L: genetic variability and molecular cultivar identification. Genet Resour Crop Evol 54:117–128
Milczarski P, Banek-Tabor A, Lebiecka K, Stojaowski S, Mysków B, Masojc P (2007) New genetic map of rye composed of PCR-based molecular markers and its alignment with the reference map of the DS2 × RXL10 intercross. J Appl Genet 48:11–24
Moretzsohn MC, Leoi L, Proite K, Guimaraes PM, Leal-Bertioli SCM, Gimanes MA, Martin WS, Valls JFM, Grattapaglia D, Bertioli D (2005) A microsatellite-based gene-rich linkage map for the AA genome of Arachis (Fabaceae). Theor Appl Genet 111:1060–1071
Morshidi M (1998) Genetic control of isozymes in Carica papaya L. Theor Appl Genet 103:89–94
Nishijima W (1994) Papaya. In: Ploetz RC, Zentmyer GA, Nishijima WT, Rohrbach KG, Ohr HD (eds) Compendium of tropical fruit disease. American Phytopath Soc Press, St. Paul, Minnesota, pp 54–70
Ocampo Pérez J, Dambier D, Ollitrault P, d’Eeckenbrugge GC, Brottier P, Froelicher Y, Risterucci AM (2006) Microsatellite markers in Carica papaya L: isolation characterization and transferability to Vasconcellea species. Mol Ecol Notes 6:212–217
Ocampo Pérez J, d'Eeckenbrugge GC, Risterucci AM, Dambier D, Ollitrault P (2007) Papaya genetic diversity assessed with microsatellite markers in germplasm from the Caribbean Region. Acta Hortic 740:93–101
Oliveira EJ, Marin ALA, Borém A, Fagundes SA, Barros EG, Moreira MA (2005) Molecular-marker assisted selection for development of common bean lines resistant to angular leaf spot. Plant Breed 124:572–575
Oliveira EJ, Dantas JLL, Castellen MS, Machado MD (2008a) Identificação de microssatélites para o mamoeiro por meio da exploração do banco de dados de DNA. Rev Bras Frutic 30:841–845
Oliveira EJ, Vieira MLC, Garcia AAF, Munhoz CF, Margarido GRA, Consoli L, Matta FP, Moraes MC (2008b) An integrated molecular map of yellow passion fruit based on simultaneous maximum-likelihood estimation of linkage and linkage phases. J Am Soc Hortic Sci 133:35–41
Paniego N, Echaide M, Marianne M, Ferrandez L, Tolales S, Faccio P, Fuxan I, Carrera M, Zandomeni R, Suarez E, Hopp HE (2002) Microsatellite isolation and characterization in sunflower (Helianthus annuus L). Genome 45:34–43
Parasnis AS, Rama Krishna W, Chowdari KY, Gupta VS, Ranjekar PK (1999) Microsatellite (GATA)4 reveals sex-specic differences in papaya. Theor Appl Genet 99:1047–1052
Powell W, Machray GC, Provan J (1996) Polymorphism revealed by simple sequence repeats. Trends Plant Sci 1:215–222
Rafalski JA, Tingey SV (1993) Genetic diagnostics in plant breeding: RAPDs microsatellites and machines. Trends Genet 9:275–279
Rallo P, Dorado G, Martín A (2000) Development of simple sequence repeats (SSRs) in olive tree (Olea europaea L). Theor Appl Genet 101:984–989
Ren J, Lu L, Liu X, Tao Z, Zhang C, Wang D, Shen J, Liu W, Tian Y, Zhu Z (2008) Paternity assessment: application on estimation of breeding value in body-weight at first egg trait of egg-laying duck (Anas platyrhynchos). Mol Biol Rep. doi:10.1007/s11033-008-9432-z
Santos SC, Ruggiero C, Silva CLSP, Lemos EGM (2003) A microsatellite library for Carica papaya L cv Sunrise Solo. Rev Bras Frutic 25:263–267
Scheldeman X, Van Damme P, Urena Alvarez JV, Romero M (2003) Horticultural potential of Andean fruit crops exploring their centre of origin. Acta Hortic 598:97–102
Sondur SN, Manshardt RM, Stiles JI (1996) A genetic linkage map of papaya based on randomly amplified polymorphic DNA markers. Theor Appl Genet 93:547–553
Stajner N, Jakse J, Kozjak P, Javornik B (2005) The isolation and characterisation of microsatellites in hop (Humulus lupulus L). Plant Sci 168:213–221
Stiles JI, Lemme C, Sondur S, Morshidi MB, Manshardt R (1993) Using randomly amplified polymorphic DNA for evaluating genetic relationships among papaya cultivars. Theor Appl Genet 85:697–701
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4. Mol Biol Evol 24:1596–1599
Tan SC, Weinheimer EA (1976) The isoenzyme patterns of developing fruit and leaf of papaya (Carica papaya L). Sains Malays 5:7–14
Van Droogenbroeck B, Breyne P, Gotghebeur P, Romeijn-Peeters E, Kyndt T, Gheysen G (2002) AFLP analysis of genetic relationships among papaya and its wild relatives (Caricaceae) from Ecuador. Theor Appl Genet 105:289–297
Van Droogenbroeck B, Kyndt T, Maertens I, Romeijn-Peeters E, Scheldeman X, Romero-Motochi J, Van Damme P, Goetghebeur P, Gheysen G (2004) Phylogenetic analysis of the highland papayas (Vasconcellea) and allied genera (Caricaceae) using PCR-RFLP. Theor Appl Genet 108:1473–1486
Wang X, Wadl PA, Rinehart TA, ScheZer BE, Windham MT, Spiers JM, Johnson DH, Trigiano RN (2009) A linkage map for Xowering dogwood (Cornus Xorida L) based on microsatellite markers. Euphytica 165:165–175
Yonemaru J-I, Ando T, Mizubayashi T, Kasuga S, Matsumoto T, Yano M (2009) Development of genome-wide simple sequence repeat markers using whole-genome shotgun sequences of sorghum (Sorghum bicolor (L) Moench). DNA Res 16:187–193
Zhivotovsky LA (1999) Estimating population structure in diploids with multilocus dominant DNA markers. Mol Ecol 8:907–913
Zhou H-F, Xie Z-W, Ge S (2003) Microsatellite analysis of genetic diversity and population genetic structure of a wild rice (Oryza rufipogon Griff) in China. Theor Appl Genet 107:332–339
Acknowledgements
This work was supported by Embrapa Macroprograma 2, CNPq-Conselho Nacional de Desenvolvimento Científico e Tecnológico (National Counsel of Technological and Scientific Development) and FAPESB–Fundação de Amparo à Pesquisa do Estado da Bahia (Bahia State Research Funding Institution).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
de Oliveira, E.J., Amorim, V.B.O., Matos, E.L.S. et al. Polymorphism of Microsatellite Markers in Papaya (Carica papaya L.). Plant Mol Biol Rep 28, 519–530 (2010). https://doi.org/10.1007/s11105-010-0180-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-010-0180-6