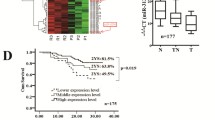
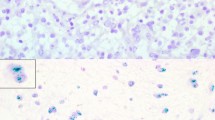
Abstract
RAS oncogenes are master regulator genes in many cancers. In general, RAS-driven cancers have an oncogenic RAS mutation that promotes disease progression (colon, lung, pancreas). In contrast, brain tumors are not necessarily RAS-driven cancers because RAS mutations are rarely observed. In particular, glioblastomas (the most lethal brain tumor) do not appear to have dominant genetic mutations that are suitable for targeted therapy. Standard treatment for most brain tumors continues to focus on maximal surgical resection, radiotherapy and chemotherapy. Yet the convergence of genomic aberrations such as EGFR, PDGFR and NF1 (some of which are clinically effective) with activation of the RAS/MAPK cascade is still considered a key point in gliomagenesis, and KRAS is undoubtedly a driving gene in gliomagenesis in mice. In cancer, microRNAs (miRNA) are small, non-coding RNAs that regulate carcinogenesis. However, the functional consequences of aberrant miRNA expression in cancer are still poorly understood. let-7 encodes an intergenic miRNA that is classified as a tumour suppressor, at least in lung cancer. Let-7 suppresses a plethora of oncogenes such as RAS, HMGA, c-Myc, cyclin-D and thus suppresses cancer development, differentiation and progression. let-7 family members are direct regulators of certain RAS family genes by binding to the sequences in their 3′untranslated region (3′UTR). let-7 miRNA is involved in the malignant behaviour in vitro—proliferation, migration and invasion—of gliomas and stem-like glioma cells as well as in vivo models of glioblastoma multiforme (GBM) via KRAS inhibition. It also increases resistance to certain chemotherapeutic agents and radiotherapy in GBM. Although let-7 therapy is not yet established, this review updates the current state of knowledge on the contribution of miRNA let-7 in interaction with KRAS to the oncogenesis of brain tumours.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MicroRNAs (miRNAs) are a class of non-coding RNAs that function as endogenous triggers of the RNA interference pathway. In general, aberrant expression of microRNAs has been linked to all human cancers. miRNAs are dysregulated in a plethora of diseases, including cancers; poorly differentiated tumors displayed lower miRNA expression compared to tumors with a higher differentiation. The Slack laboratory showed one of the first pieces of evidence that lethal-7 (let-7) plays a key role in cancer. By showing that RAS is regulated by the let-7 miRNA family in Caenorhabditis elegans and by highlighting let-7 complementary sites in human RAS 3′UTRs, they first validated that the let-7 miRNA family negatively regulates RAS in two different C. elegans tissues and human cell lines and lung tissue. This was the first report of the mechanism of action of a miRNA acting as a tumor suppressor [1, 2].
The heterochronic miRNA let-7 display tumor suppressive properties at least in lung, colon, breast, and leukemia cancers. It is categorized as a tumor suppressor because it reduces cancer aggressiveness, chemoresistance, and radio-resistance. Particularly, studies inferred from human cancer databases report let-7 family member expression associated with poor overall survival in some cancers; but regretfully with unsignificant results in glioblastoma multiforme (GBM) [3].
By contrast, many in vitro studies report that let-7 inhibits the malignant behavior of glioma cells and stem-like cells [4,5,6,7,8,9,10,11,12,13] (Table 1). Given the promiscuous and context-specific nature of miRNA targeting, many mechanisms of interactions remain to be elucidated. Moreover, regulation of RAS protein level and RAS/MAPK cascade are regulated by a myriad of miRNAs family without a clear mechanistic link [11, 12, 14,15,16]. Three recent reports focused on miRNAs targeting RAS in GBM and showed that miR-143-3p directly targets NRAS [17], let-7a-5p directly targets KRAS [7], and both R-Ras and N-Ras (related RAS viral oncogene homolog, HRAS homolog) are direct targets of miR-124-3p [18].
let-7 is a bona fide tumor suppressor gene, but this categorization is rarely straightforward, since some miRNA can have both oncogenic and tumor-suppressive mechanisms, depending on the context. Let-7 targets multiple oncogenes (RAS, c-MYC, HMGA2 to date) and the prominent mechanisms by which let-7 exerts a tumor suppressive role is by repressing the translation of the three RAS proteins (HRAS, NRAS, and KRAS) and c-MYC, a downstream effector of RAS-ERK signaling [1, 23,24,25,26,27]. Moreover, oncogenic functions for let-7 have been also reported in some cases [28, 29]. Indeed, the functions of let-7 have been reported not only in cancer but also in other diseases, including viral infection, immune diseases, neurological diseases, diabetes, and cardiovascular diseases. In vivo studies with Cre-inducible let-7-transgenic mice have reported a strong phenotypic read-out both in diabetic retinopathy and in diabetes [30, 31].
As a proto-oncogene that regulates several oncogenic pathways, KRAS has always been considered a central signaling modulator. Thus, there is a flourishing literature on studying miRNAs that regulate K-RAS expression, with let-7 being of prime importance. Our knowledge of how miRNAs can modulate activation of the RAS-ERK signalling pathway continues to grow as potential miRNA-mRNA regulatory networks are identified using different strategies [32]. From these studies, three main paradigms of miRNA-mediated RAS-ERK regulation have emerged. miRNAs can affect the translation of (i) core components of the RAS-ERK pathway (e.g., let-7 targets HRAS, NRAS and KRAS) [1, 33], (ii) critical proteins that regulate the pathway and are required for proper spatial and temporal control of RAS-ERK signalling [29, 34, 35], and (iii) upstream drivers and downstream effector/regulator molecules [36, 37]. A myriad of miRNAs regulates RAS-ERK pathway activity in a variety of cancer contexts [38, 39].
In general, there is a very clear correlation between the loss of let-7 expression and the development of poorly differentiated and aggressive cancers, this is the case for let-7a in lung carcinomas. In prototypic studies in non-small cell lung cancer (NSCLC) models, let-7 expression was analysed in vitro and in vivo and its role in KRAS-mediated NSCLC tumorigenesis was demonstrated [40]. let-7 inhibits tumour development and the RAS-ERK signalling pathway in an autochthonous model of NSCLC driven by activated KRAS (KRASG12D) [41, 42]. In xenograft models of NSCLC, let-7 exerts a tumour suppressive role [24] and increased expression of let-7a significantly reduces tumour burden in a K-Ras lung cancer model in mice [43]. Furthermore, let-7 counteracts the maintenance, survival and self-renewal of cancer stem-like cells (CSCs) in ovarian and breast cancer, and this suppressive activity correlated with reduced expression of RAS and HMGA2 [43,44,45,, 45, 46, 47]. Thus, by suppressing RAS expression, let-7 can attenuate RAF-MEK-ERK signalling and its dependent oncogenic phenotypes independent of RAS mutation status. Functional studies have confirmed that miRNA let-7 dysregulation is causative in many cancer cases, highlighting its potential impact on RAS-ERK signalling from a mechanism perspective.
Main text
Let-7 expression in brain tumors
Brain tumours comprise a broad spectrum of over 120 histologically, demographically, clinically and molecularly different diseases (overview in [48]). Glioblastoma is the most common malignant primary brain tumour in adults and remains incurable [49]. Standard treatment for most brain tumours remains focused on maximal surgical resection, radiotherapy and chemotherapy with temozolomide (TMZ) as first-line therapy. Indirect targeting of the tumour by anti-angiogenics (e.g. bevacizumab) and immunotherapies (vaccines, adoptive therapies, immune checkpoint inhibitors and oncolytic viruses) have shown mixed efficacy (or inactivity) in preclinical studies [5048,49,50,51,, 51, 52, 53, 54]. Alternative studies focusing on molecular profiling of GBM identified neurofibromin-1 (NF1) as targeting mutations that contribute to activated KRAS signalling [55, 56].
The members of the let-7 family are direct and strong regulators of K-RAS, N-RAS and H-RAS mRNAs through their 3′UTR sequences [15, 57, 58]. A single miRNA can control entire cellular signalling pathways by interacting with a broad spectrum of target genes. For example, let-7 inhibits GBM tumour growth by interacting with a broad spectrum of target genes such as Ras, c-Myc, Stat3, Cyclin D1 [4, 6, 7, 10, 22] (see Table 1). Mature let-7 can be blocked by the LIN28 protein, increased levels of which are associated with poorer survival in gliomas, and let-7b can serve as a marker for chemoresistance [20, 21]. In irradiated human glioblastoma cells, let-7 mediated resistance to radiotherapy by regulating its relative expression [19]. Remarkably, the resulting loss of let-7 enhances the expression of oncogenic targets such as RAS in loss-of-function and gain-of-function experiments [36].
The detection of miRNA has rapidly emerged as a potential biomarker in patients with glioblastoma [59, 60]. In particular, decreased let-7b has recently been associated with poor prognosis in gliomas [61]. Based on several high-throughput genomic technologies, The cancer genome atlas (TCGA) has defined RAS/MAPK as one of the central pathways involved in GBM [62] and let-7 miRNA family expression levels are not reduced in GBM (summarised by [11, 12]). Regardless of its expression level, let-7 miRNA can impair glioblastoma growth and cell migration through RAS inhibition [7, 10]. Specifically, forced expression of let-7 miRNA reduced the expression of pan-RAS, N-RAS and K-RAS, thereby reducing proliferation and migration as well as tumour size in xenograft-transplanted GBM in nude mice [10]. In another study, overexpressed let-7a inhibited glioma cell malignancy by directly targeting KRAS independently of PTEN [7], and let-7b in turn inhibits malignant behaviour (proliferation, migration and invasion) of glioma cells and stem-like glioma cells [6]. Indeed, focal deletion of members of the let-7 family (let-7a-2 and let-7e) has been found in medulloblastoma (MB) [63] and the let-7 family has been validated in spontaneous and radiation-induced MB [64]. Conversely, miR-let7g was found to be upregulated in anaplastic and differentially expressed in desmoplastic MB [65, 66], although the functional consequences of its dysregulation have not yet been investigated. Recently, let-7 miRNA activity has been identified as a prognostic biomarker of SHH medulloblastoma [67]. Furthermore, in a paediatric brain tumour (Diffuse intrinsic pontine gliomas, DIPG), RAS signalling has recently been identified as a novel therapeutic vulnerability and the RNA-binding protein LIN28B is overexpressed in DMG and suppresses the let-7 family of microRNAs [68, 69]. Numerous oncogenes and signalling pathways besides RAS (e.g. MYC) have been shown to be targets of let‐7 miRNAs, and KRAS is the target of many other miRNAs in gliomogenesis [5, 16]. Which of these targets genes determine the overall phenotype in glioblastoma remains to be investigated.
RAS oncogenes in brain tumors
Comprehensive molecular profiling has dramatically changed the diagnostic neuropathology of brain tumours. Several of the key molecular alterations that are critical for glioma classification involve epigenetic dysregulation at a fundamental level and involve areas of biology not previously thought to play an important role in glioma pathogenesis [48]. The biological functions of the RAS family [the viral oncogene homologue of Harvey rat sarcoma (HRAS), the viral oncogene homologue of Kirsten rat sarcoma (KRAS) and the viral oncogene homologue of neuroblastoma (NRAS)] have been studied in detail for decades. Although only 1% of GBM tumours exhibit RAS mutation or amplification, 10% of GBM tumours contain inactivating genetic alterations of neurofibromin 1 (NF1) that lead to hyperactive RAS activity by enhancing intrinsic GTPase activity [62, 70]. Remarkably, dysfunctional signalling in tumours arises not only from gene mutations but also from epigenetic changes or pathway rewiring, which probably explains why there appear to be no dominant driver mutations in certain tumour types, most notably glioblastoma. Furthermore, no evidence of oncogenic mutations affecting NRAS, KRAS, HRAS, BRAF or PDGFR was found in medulloblastoma [71].
Deregulated RAS signalling is an important step in carcinogenesis, with activating RAS mutations playing a role in 30% of all cancers [72]. In contrast to many human tumours, RAS mutation is not common in human gliomas, with some exceptions such as cerebellar GBM [73]. Hyperactive RAS signalling alone is sufficient to generate gliomas that closely resemble human tumours in glioma mouse models. The RAS signalling pathway is therefore of central importance for human gliomagenesis. Primary GBMs are associated with impaired RAS signalling and expression of the oncogenic HRAS leads to a malignant phenotype in glioma cell lines [74, 75, 76, 77]. In particular, the KRAS oncogene is strongly involved in tumourigenesis in glioblastomas [75, 78, 79, 80], although KRAS mutations are almost absent in malignant gliomas [81]. Therefore, the observed deregulation of the Ras-RAF-ERK signalling pathway in gliomas is generally attributed to its upstream positive regulators, including EGFR and PDGFR, which are known to be highly active in many malignant gliomas [70]. It is likely that mechanisms other than mutations contribute to the activation of the RAS-MAPK signalling pathway in wild-type cancer. Indeed, epigenetic modifications have been described to enhance this activation in human tumours [82], and dysregulation of physiological miRNA activity has been shown to play an important role in gliomagenesis, but the functional significance of this regulatory level is currently unknown [11, 12, 75,76,77,78,, 84, 85, 86, 87].
Therapeutic and diagnostic potential of mirnas
Clinical studies on the benefits of miRNAs in diagnosis and therapy are already underway [88, 89]. MiRNAs are abundant non-coding RNAs, and their short length increases their stability compared to longer RNA molecules. In addition, miRNAs are secreted alone into the extracellular fluid or encapsulated by vesicles such as microvesicles and exosomes. Subsequently, the secreted miRNAs are found in the circulation. In summary, cancer-specific and circulating miRNAs are attractive diagnostic markers. Besides diagnostic miRNAs, miRNAs that can be used to predict drug efficacy and patient prognosis will greatly assist the advancement of precision cancer medicine. Despite the increasing interest in miRNA therapies as potential approaches for cancer treatment, the actual deployment of these molecules into solid tumors remains a formidable obstacle [89]. Stabilty of let-7 mimics for cancer therapy have been improved [90], but delivery issues remain [91].
Therapeutic anti-miRs are currently being developed for cancer therapy, such as miR-155 for treating leukemia and lymphoma [92]. Several clinical trials of improved miRNA drug strategies, such as synthetic RNA molecules and advanced delivery technologies, are ongoing despite the failure of the first-in-human clinical trial of miRNA cancer therapy [93]. Thus far, one clinical trial in gliobastoma to assess miR-10b expression in patients with several subtypes of brain cancer is still ongoing (recruiting, NCT01849952). A miR-10b inhibitor, RGLS5579, was developed and preclinically tested for glioblastoma multiforme (GBM) [94, 86,87,, 96, 97].
Since there is currently only one miRNA drug in clinical trials, there is little direct evidence of that miRNAs can be applied with minimal side-effects. In fact, their mechanism of action is tuning expression rather than blunting their targets which reasonably should be less detrimental to healthy tissues. In contrast to the selectivity of enzymatic protein inhibitors, miRNA drugs are developed with the idea of controlling multiple gene-components in the same or overlapping signaling-pathways. Such gene-products are not limited to proteins with enzymatic activity but could include any deregulated genes or proteins in a given disease.
Conclusion
The role of KRAS, when activated through canonical mutations, has been well established in cancer. Research on RAS-driven cancers has focused almost exclusively on RAS coding mutations. However, KRAS without canonical mutations is still largely unexplored. KRAS regulation by miRNA is now emerging as a new layer of regulation [98]. Since the discovery of let-7 miRNA, work is currently in its pre-clinical era. The control of KRAS expression by the let-7 family of miRNAs is well documented. Low expression of let-7 in human cancer correlates with high KRAS expression (at least in lung, breast and colorectal). The control over KRAS expression and activity in glioblastomas suggests that let-7 can regulate RAS activity even without oncogenic mutation and that this may be a general phenomenon related to the interactions between tumour suppressor genes (let-7) and proto-oncogenes (KRAS). Open questions remain to be answered: (i) is the functional output of KRAS signaling essential for promoting tumorigenesis?; (ii) is the suppression of KRAS the dominant signalling effect in glioblastoma?; (iii) what are the general physiological effects of let-7 miRNA dysregulation in brain tumours (if any)?; (iv) are oncogenic inhibitors currently in clinical trials for KRAS-driven cancers suitable for therapeutics in KRAS wild-type tumours? Moreover, a large fraction of miRNAs bind their targets independent of seed match complementarity at nucleotides 2–7 [99], this observation greatly affects our ability to accurately predict miRNA targets using existing algorithms. Besides the 3′UTR, miRNAs have been demonstrated to target the 5′UTR and coding sequence of mRNAs as well as other RNA species such as lncRNAs, pseudogenes, rRNAs and tRNAs [100]. We still have little work on the roles of other miRNAs and ncRNAs in brain tumors. While let-7 shows promise as a therapeutic in brain tumors, we are left with the question of how this therapy will be implemented to help patients.
Data availability
No data associated in the manuscript.
Abbreviations
- miRNA:
-
MicroRNA
- let-7 :
-
Lethal-7
- GBM:
-
Glioblastoma
- MB:
-
Medulloblastoma
- SHH:
-
Sonic Hedgehog-activated
- DIPG:
-
Diffuse intrinsic pontine gliomas
- NF1:
-
Neurofibromin 1
- PDGFR:
-
Platelet-derived growth factor receptor
- EGFR:
-
Epidermal growth factor receptor
- DMG:
-
Diffuse midline glioma
References
Johnson SM, Grosshans H, Shingara J, Byrom M, Jarvis R, Cheng A, Labourier E, Reinert KL, Brown D, Slack FJ (2005) RAS is regulated by the let-7 microRNA family. Cell 120(5):635–647. https://doi.org/10.1016/j.cell.2005.01.014
Johnson CD, Esquela-Kerscher A, Stefani G, Byrom M, Kelnar K, Ovcharenko D, Wilson M, Wang X, Shelton J, Shingara J, Chin L, Brown D, Slack FJ (2007) The let-7 microRNA represses cell proliferation pathways in human cells. Cancer Res 67(16):7713–7722. https://doi.org/10.1158/0008-5472.CAN-07-1083
Gilles ME, Slack FJ (2018) Let-7 microRNA as a potential therapeutic target with implications for immunotherapy. Expert Opin Ther Targets 22(11):929–939. https://doi.org/10.1080/14728222.2018.1535594
Degrauwe N, Schlumpf TB, Janiszewska M, Martin P, Cauderay A, Provero P, Riggi N, Suvà ML, Paro R, Stamenkovic I (2016) The RNA binding protein IMP2 preserves glioblastoma stem cells by preventing let-7 target gene silencing. Cell Rep 15(8):1634–1647. https://doi.org/10.1016/j.celrep.2016.04.086
Li Y, Zhang X, Chen D, Ma C (2016) Let-7a suppresses glioma cell proliferation and invasion through TGF-β/Smad3 signaling pathway by targeting HMGA2. Tumour Biol 37(6):8107–8119. https://doi.org/10.1007/s13277-015-4674-6
Song H, Zhang Y, Liu N, Zhang D, Wan C, Zhao S, Kong Y, Yuan L (2016) Let-7b inhibits the malignant behavior of glioma cells and glioma stem-like cells via downregulation of E2F2. J Physiol Biochem. https://doi.org/10.1007/s13105-016-0512-6
Wang XR, Luo H, Li HL, Cao L, Wang XF, Yan W, Wang YY, Zhang JX, Jiang T, Kang CS, Liu N, You YP, Chinese Glioma Cooperative Group (CGCG) (2013) Overexpressed let-7a inhibits glioma cell malignancy by directly targeting K-ras, independently of PTEN. Neuro Oncol 15(11):1491–1501. https://doi.org/10.1093/neuonc/not107
Buonfiglioli A, Efe IE, Guneykaya D, Ivanov A, Huang Y, Orlowski E, Krüger C, Deisz RA, Markovic D, Flüh C, Newman AG, Schneider UC, Beule D, Wolf SA, Dzaye O, Gutmann DH, Semtner M, Kettenmann H, Lehnardt S (2019) let-7 microRNAs regulate microglial function and suppress glioma growth through toll-like receptor 7. Cell Rep 29(11):3460–3471. https://doi.org/10.1016/j.celrep.2019.11.029
Huang Y, Liu P, Luo J, Zhu C, Lu C, Zhao N, Zhao W, Cui W, Yang X (2023) Par6 enhances glioma invasion by activating MEK/ERK pathway through a LIN28/let-7d positive feedback loop. Mol Neurobiol 60(3):1626–1644. https://doi.org/10.1007/s12035-022-03171-0
Lee ST, Chu K, Oh HJ, Im WS, Lim JY, Kim SK, Park CK, Jung KH, Lee SK, Kim M, Roh JK (2011) Let-7 microRNA inhibits the proliferation of human glioblastoma cells. J Neurooncol 102(1):19–24. https://doi.org/10.1007/s11060-010-0286-6
Wang X, Xin Z, Xu Y, Ma J (2016) Upregulated miRNA-622 inhibited cell proliferation, motility, and invasion via repressing Kirsten rat sarcoma in glioblastoma. Tumour Biol 37(5):5963–5970. https://doi.org/10.1007/s13277-015-4455-2
Wang Z, Lin S, Zhang J, Xu Z, Xiang Y, Yao H, Ge L, Xie D, Kung HF, Lu G et al (2016) Loss of MYC and E-box3 binding contributes to defective MYC-mediated transcriptional suppression of human MC-let-7a- 1$let-7d in glioblastoma. Oncotarget 7:56266–56278. https://doi.org/10.18632/oncotarget.10517
Xie C, Chen W, Zhang M, Cai Q, Xu W, Li X, Jiang S (2015) MDM4 regulation by the let-7 miRNA family in the DNA damage response of glioma cells. FEBS Lett 589(15):1958–1965. https://doi.org/10.1016/j.febslet.2015.05.030
Li Y, Li Y, Ge P, Ma C (2017) Mir-126 regulates the ERK pathway via targeting KRAS to inhibit the glioma cell proliferation and invasion. Mol Neurobiol 54(1):137–145. https://doi.org/10.1007/s12035-015-9654-8
Yu ML, Wang JF, Wang GK, You XH, Zhao XX, Jing Q, Qin YW (2011) Vascular smooth muscel cell proliferation is influenced by let-7d microRNA and its interaction with KRAS. Circ J 75:703–709. https://doi.org/10.1253/circj.cj-10-0393
Zhao Y, Pang D, Wang C, Zhong S, Wang S (2016) MicroRNA-134 modulates glioma cell U251 proliferation and invasion by targeting KRAS and suppressing the ERK pathway. Tumour Biol 37(8):11485–11493. https://doi.org/10.1007/s13277-016-5027-9
Wang L, Shi ZM, Jiang CF, Liu X, Chen QD, Qian X, Li DM, Ge X, Wang XF, Liu LZ, You YP, Liu N, Jiang BH (2014) MiR-143 acts as a tumor suppressor by targeting N-RAS and enhances temozolomide-induced apoptosis in glioma. Oncotarget 5(14):5416–5427. https://doi.org/10.18632/oncotarget.2116
Shi Z, Chen Q, Li C, Wang L, Qian X, Jiang C, Liu X, Wang X, Li H, Kang C, Jiang T, Liu LZ, You Y et al (2014) MiR-124 governs glioma growth and angiogenesis and enhances chemosensitivity by targeting R-Ras and N-Ras. Neuro Oncol 16:1341–1353. https://doi.org/10.1093/neuonc/nou084
Chaudhry MA, Sachdeva H, Omaruddin RA (2010) Radiation-induced micro-RNA modulation in glioblastoma cells differing in DNA-repair pathways. DNA Cell Biol 29(9):553–561. https://doi.org/10.1089/dna.2009.0978
Evers L, Schäfer A, Pini R, Zhao K, Stei S, Nimsky C, Bartsch JW (2023) Identification of dysregulated microRNAs in glioblastoma stem-like cells. Brain Sci 13(2):350. https://doi.org/10.3390/brainsci13020350
Guo Y, Yan K, Fang J, Qu Q, Zhou M, Chen F (2013) Let-7b expression determines response to chemotherapy through the regulation of cyclin D1 in glioblastoma. J Exp Clin Cancer Res 32(1):41. https://doi.org/10.1186/1756-9966-32-41
Mao XG, Hütt-Cabezas M, Orr BA, Weingart M, Taylor I, Rajan AK, Odia Y, Kahlert U, Maciaczyk J, Nikkhah G et al (2013) LIN28A facilitates the transformation of human neural stem cells and promotes glioblastoma tumorigenesis through a pro-invasive genetic program. Oncotarget 4:1050–1064. https://doi.org/10.18632/oncotarget.1131
Gunzburg MJ, Sivakumaran A, Pendini NR, Yoon JH, Gorospe M, Wilce MC, Wilce JA (2015) Cooperative interplay of let-7 mimic and HuR with MYC RNA. Cell Cycle 14(17):2729–2733. https://doi.org/10.1080/15384101.2015.1069930
He XY, Chen JX, Zhang Z, Li CL, Peng QL, Peng HM (2010) The let-7a microRNA protects from growth of lung carcinoma by suppression of k-Ras and c-Myc in nude mice. J Cancer Res Clin Oncol 136(7):1023–1028
Maldotti M, Incarnato D, Neri F, Krepelova A, Rapelli S, Anselmi F, Parlato C, Basile G, Dettori D, Calogero R, Oliviero S (2016) The long intergenic non-coding RNA CCR492 functions as a let-7 competitive endogenous RNA to regulate c-Myc expression. Biochim Biophys Acta 1859(10):1322–1332. https://doi.org/10.1016/j.bbagrm.2016.06.010
Sampson VB, Rong NH, Han J, Yang Q, Aris V, Soteropoulos P, Petrelli NJ, Dunn SP, Krueger LJ (2007) MicroRNA let-7a down-regulates MYC and reverts MYC-induced growth in Burkitt lymphoma cells. Cancer Res 67(20):9762–9770. https://doi.org/10.1158/0008-5472.CAN-07-2462
Wong TS, Man OY, Tsang CM, Tsao SW, Tsang RK, Chan JY, Ho WK, Wei WI, To VS (2011) MicroRNA let-7 suppresses nasopharyngeal carcinoma cells proliferation through downregulating c-Myc expression. J Cancer Res Clin Oncol 137(3):415–422. https://doi.org/10.1007/s00432-010-0898-4
Brueckner B, Stresemann C, Kuner R, Mund C, Musch T, Meister M, Sültmann H, Lyko F (2007) The human let-7a-3 locus contains an epigenetically regulated microRNA gene with oncogenic function. Cancer Res 67(4):1419–1423. https://doi.org/10.1158/0008-5472.CAN-06-4074
Meng F, Henson R, Wehbe-Janek H, Smith H, Ueno Y, Patel T (2007) The MicroRNA let-7a modulates interleukin-6-dependent STAT-3 survival signaling in malignant human cholangiocytes. J Biol Chem 282(11):8256–8264. https://doi.org/10.1074/jbc.M607712200
Zhou Q, Frost RJA, Anderson C, Zhao F, Ma J, Yu B, Wang S (2017) let-7 contributes to diabetic retinopathy but represses pathological ocular angiogenesis. Mol Cell Biol 37(16):00001–00017. https://doi.org/10.1128/MCB.00001-17
Frost RJ, Olson EN (2011) Control of glucose homeostasis and insulin sensitivity by the Let-7 family of microRNAs. Proc Natl Acad Sci USA 108(52):21075–21080. https://doi.org/10.1073/pnas.1118922109
Shui B, La Rocca G, Ventura A, Haigis KM (2022) Interplay between K-RAS and miRNAs. Trends Cancer 8(5):384–396. https://doi.org/10.1016/j.trecan.2022.01.002
Danac JMC, Garcia RL (2021) CircPVT1 attenuates negative regulation of NRAS by let-7 and drives cancer cells towards oncogenicity. Sci Rep 11(1):9021. https://doi.org/10.1038/s41598-021-88539-3
Hatley ME, Patrick DM, Garcia MR, Richardson JA, Bassel-Duby R, van Rooij E, Olson EN (2010) Modulation of K-Ras-dependent lung tumorigenesis by microRNA-21. Cancer Cell 18(3):282–293. https://doi.org/10.1016/j.ccr.2010.08.013
Sharma V, Dixit D, Koul N, Mehta VS, Sen E (2011) Ras regulates interleukin-1β-induced HIF-1α transcriptional activity in glioblastoma. J Mol Med 89(2):123–136. https://doi.org/10.1007/s00109-010-0683-5
Stainthorp AK, Lin CC, Wang D, Medhi R, Ahmed Z, Suen KM, Miska EA, Whitehouse A, Ladbury JE (2023) Regulation of microRNA expression by the adaptor protein GRB2. Sci Rep 13(1):9784. https://doi.org/10.1038/s41598-023-36996-3
Zawistowski JS, Nakamura K, Parker JS, Granger DA, Golitz BT, Johnson GL (2013) MicroRNA 9-3p targets β1 integrin to sensitize claudin-low breast cancer cells to MEK inhibition. Mol Cell Biol 33(11):2260–2274. https://doi.org/10.1128/MCB.00269-13
Asl ER, Amini M, Najafi S, Mansoori B, Mokhtarzadeh A, Mohammadi A, Lotfinejad P, Bagheri M, Shirjang S, Lotfi Z, Rasmi Y, Baradaran B (2021) Interplay between MAPK/ERK signaling pathway and MicroRNAs: a crucial mechanism regulating cancer cell metabolism and tumor progression. Life Sci 278:119499. https://doi.org/10.1016/j.lfs.2021.119499
Masliah-Planchon J, Garinet S, Pasmant E (2016) RAS-MAPK pathway epigenetic activation in cancer: miRNAs in action. Oncotarget 7(25):38892–38907. https://doi.org/10.18632/oncotarget.6476.Review
Jinesh G, Sambandam V, Vijayaraghavan S et al (2018) Molecular genetics and cellular events of K-Ras-driven tumorigenesis. Oncogene 37:839–846. https://doi.org/10.1038/onc.2017.377
Esquela-Kerscher A, Trang P, Wiggins JF, Patrawala L, Cheng A, Ford L, Weidhaas JB, Brown D, Bader AG, Slack FJ (2008) The let-7 microRNA reduces tumor growth in mouse models of lung cancer. Cell Cycle 7(6):759–764. https://doi.org/10.4161/cc.7.6.5834
Kumar MS, Erkeland SJ, Pester RE, Chen CY, Ebert MS, Sharp PA, Jacks T (2008) Suppression of non-small cell lung tumor development by the let-7 microRNA family. Proc Natl Acad Sci USA 105(10):3903–3908. https://doi.org/10.1073/pnas.0712321105
Trang P, Medina PP, Wiggins JF, Ruffino L, Kelnar K, Omotola M, Homer R, Brown D, Bader AG, Weidhaas JB, Slack FJ (2010) Regression of murine lung tumors by the let-7 microRNA. Oncogene. 29(11):1580–7. https://doi.org/10.1038/onc.2009.445
Chirshev E, Oberg KC, Ioffe YJ, Unternaehrer JJ (2019) Let-7 as biomarker, prognostic indicator, and therapy for precision medicine in cancer. Clin Transl Med 8(1):24. https://doi.org/10.1186/s40169-019-0240-y
Ma L, Li GZ, Wu ZS, Meng G (2014) Prognostic significance of let-7b expression in breast cancer and correlation to its target gene of BSG expression. Med Oncol 31(1):773. https://doi.org/10.1007/s12032-013-0773-7
Petrillo M, Zannoni GF, Beltrame L, Martinelli E, DiFeo A, Paracchini L, Craparotta I, Mannarino L, Vizzielli G, Scambia G, D’Incalci M, Romualdi C, Marchini S (2016) Identification of high-grade serous ovarian cancer miRNA species associated with survival and drug response in patients receiving neoadjuvant chemotherapy: a retrospective longitudinal analysis using matched tumor biopsies. Ann Oncol 27(4):625–634. https://doi.org/10.1093/annonc/mdw007
Yu F et al (2007) let-7 regulates self-renewal and tumorigenicity of breast cancer cells. Cell 131:1109–1123. https://doi.org/10.1016/j.cell.2007.10.054
Louis DN, Perry A, Wesseling P, Brat DJ, Cree IA, Figarella-Branger D, Hawkins C, Ng HK, Pfister SM, Reifenberger G, Soffietti R, von Deimling A, Ellison DW (2021) The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol 23(8):1231–1251. https://doi.org/10.1093/neuonc/noab106
Bagley SJ, Kothari S, Rahman R, Lee EQ, Dunn GP, Galanis E, Chang SM, Nabors LB, Ahluwalia MS, Stupp R, Mehta MP, Reardon DA, Grossman SA, Sulman EP, Sampson JH, Khagi S, Weller M, Cloughesy TF, Wen PY, Khasraw M (2022) Glioblastoma clinical trials: current landscape and opportunities for improvement. Clin Cancer Res 28(4):594–602. https://doi.org/10.1158/1078-0432.CCR-21-2750
Omuro A, DeAngelis LM (2013) Glioblastoma and other malignant gliomas: a clinical review. JAMA. 310(17):1842–50. https://doi.org/10.1001/jama.2013.280319
Huse JT, Holland EC (2010) Targeting brain cancer: advances in the molecular pathology of malignant glioma and medulloblastoma. Nat Rev Cancer 10(5):319–331. https://doi.org/10.1038/nrc2818
Thomas AA, Brennan CW, DeAngelis LM, Omuro AM (2014) Emerging therapies for glioblastoma. JAMA Neurol 71(11):1437–1444. https://doi.org/10.1001/jamaneurol.2014.1701
Yang K, Wu Z, Zhang H et al (2022) Glioma targeted therapy: insight into future of molecular approaches. Mol Cancer 21:39. https://doi.org/10.1186/s12943-022-01513-z
Khosla D (2016) Concurrent therapy to enhance radiotherapeutic outcomes in glioblastoma. Ann Transl Med 4(3):54. https://doi.org/10.3978/j.issn.2305-5839.2016.01.25
Behnan J, Finocchiaro G, Hanna G (2019) The landscape of the mesenchymal signature in brain tumours. Brain 142(4):847–866. https://doi.org/10.1093/brain/awz044
Blomquist MR, Ensign SF, D’Angelo F, Phillips JJ, Ceccarelli M, Peng S et al (2020) Temporospatial genomic profiling in glioblastoma identifies commonly altered core pathways underlying tumor progression. Neurooncol Adv 2(1):vdaa078. https://doi.org/10.1093/noajnl/vdaa078
Büssing I, Slack FJ, Grosshans H (2008) Let-7 microRNAs in development stem cells and cancer. Trends Mol Med 14:400–409. https://doi.org/10.1016/j.molmed.2008.07.001
Khodayari N, Mohammed KA, Goldberg EP, Nasreen N (2011) EphrinA1 inhibits malignant mesothelioma tumor growth via let-7 microRNA-mediated repression of the RAS oncogene. Cancer Gene Ther 18(11):806–816. https://doi.org/10.1038/cgt.2011.50
Ahir BK, Ozer H, Engelhard HH, Lakka SS (2017) MicroRNAs in glioblastoma pathogenesis and therapy: a comprehensive review. Crit Rev Oncol Hematol 120:22–33. https://doi.org/10.1016/j.critrevonc.2017.10.003
Chen M, Medarova Z, Moore A (2021) Role of microRNAs in glioblastoma. Oncotarget 12(17):1707–1723. https://doi.org/10.18632/oncotarget.28039
Zhang W, Zhao W, Ge C, Li X, Yang X, Xiang Y, Sun Z (2019) Decreased let-7b is associated with poor prognosis in glioma. Medicine 98(22):e15784. https://doi.org/10.1097/MD.0000000000015784
Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR, Zheng S, Chakravarty D, Sanborn JZ, Berman SH, Beroukhim R, Bernard B, Wu CJ, Genovese G, Shmulevich I, Barnholtz-Sloan J, Zou L, Vegesna R, Shukla SA, Ciriello G, Yung WK, Zhang W, Sougnez C, Mikkelsen T, Aldape K, Bigner DD, Van Meir EG, Prados M, Sloan A, Black KL, Eschbacher J, Finocchiaro G, Friedman W, Andrews DW, Guha A, Iacocca M, O'Neill BP, Foltz G, Myers J, Weisenberger DJ, Penny R, Kucherlapati R, Perou CM, Hayes DN, Gibbs R, Marra M, Mills GB, Lander E, Spellman P, Wilson R, Sander C, Weinstein J, Meyerson M, Gabriel S, Laird PW, Haussler D, Getz G, Chin L (2013) TCGA Research Network. The somatic genomic landscape of glioblastoma. Cell. 155(2):462–77. https://doi.org/10.1016/j.cell.2013.09.034
Wang Y, Hu X, Greshock J, Shen L, Yang X, Shao Z, Liang S, Tanyi JL, Sood AK, Zhang L (2012) Genomic DNA copy-number alterations of the let-7 family in human cancers. PLoS ONE 7(9):e44399. https://doi.org/10.1371/journal.pone.0044399
Tanno B, Babini G, Leonardi S, Giardullo P, De Stefano I, Pasquali E, Ottolenghi A, Atkinson MJ, Saran A, Mancuso M (2016) Ex vivo miRNome analysis in Ptch1+/ cerebellum granule cells reveals a subset of miRNAs involved in radiation-induced medulloblastoma. Oncotarget. https://doi.org/10.18632/oncotarget.11938
Turner JD, Williamson R, Almefty KK, Nakaji P, Porter R, Tse V, Kalani MY (2010) The many roles of microRNAs in brain tumor biology. Neurosurg Focus. 28(1):E3. https://doi.org/10.3171/2009.10.FOCUS09207
Shahab SW, Roggeveen CM, Sun J, Kunhiraman H, McSwain LF, Juraschka K, Kumar SA, Saulnier O, Taylor MD, Schniederjan M, Schnepp RW, MacDonald TJ, Kenney AM (2023) The LIN28B-let-7-PBK pathway is essential for group 3 medulloblastoma tumor growth and survival. Mol Oncol. https://doi.org/10.1002/1878-0261.13477
Westphal MS, Lee E, Schadt EE, Sholler GS, Zhu J (2022) Identification of Let-7 miRNA activity as a prognostic biomarker of SHH medulloblastoma. Cancers 14:139. https://doi.org/10.3390/cancers14010139
Knowles T, Huang T, Qi J, An S, Burket N, Cooper S, Nazarian J, Saratsis AM (2023) LIN28B and Let-7 in diffuse midline glioma: a review. Cancers 15(12):3241. https://doi.org/10.3390/cancers15123241
Koncar RF, Dey BR, Stanton AJ, Agrawal N, Wassell ML, McCarl LH, Locke AL, Sanders L, Morozova-Vaske O, Myers MI, Hamilton RL, Carcaboso AM, Kohanbash G, Hu B, Amankulor NM, Felker J, Kambhampati M, Nazarian J, Becher OJ, James CD, Hashizume R, Broniscer A, Pollack IF, Agnihotri S (2019) Identification of novel RAS signaling therapeutic vulnerabilities in diffuse intrinsic pontine gliomas. Cancer Res 79(16):4026–4041. https://doi.org/10.1158/0008-5472.CAN-18-3521
Lo HW (2010) Targeting Ras-RAF-ERK and its interactive pathways as a novel therapy for malignant gliomas. Curr Cancer Drug Targets. 10(8):840–8
Gilbertson RJ, Langdon JA, Hollander A, Hernan R, Hogg TL, Gajjar A, Fuller C, Clifford SC (2006) Mutational analysis of PDGFR-RAS/MAPK pathway activation in childhood medulloblastoma. Eur J Cancer 42(5):646–649. https://doi.org/10.1016/j.ejca.2005.11.023
Mukhopadhyay S, Vander Heiden MG, McCormick F (2021) The metabolic landscape of RAS-driven cancers from biology to therapy. Nat Cancer 2(3):271–283. https://doi.org/10.1038/s43018-021-00184-x
Milinkovic VP, Skender Gazibara MK, Manojlovic Gacic EM, Gazibara TM, Tanic NT (2014) The impact of TP53 and RAS mutations on cerebellar glioblastomas. Exp Mol Pathol. 97(2):202–7. https://doi.org/10.1016/j.yexmp.2014.07.009
Eleveld TF et al (2015) Relapsed neuroblastomas show frequent RAS-MAPK pathway mutations. Nat Genet. 47(8):864–71
Ding H, Roncari L, Shannon P, Wu X, Lau N, Karaskova J, Gutmann DH, Squire JA, Nagy A, Guha A (2001) Astrocyte-specific expression of activated p21-ras results in malignant astrocytoma formation in a transgenic mouse model of human gliomas. Cancer Res 61(9):3826–3836
Knobbe CB, Reifenberger J, Reifenberger G (2004) Mutation analysis of the Ras pathway genes NRAS, HRAS, KRAS and BRAF in glioblastomas. Acta Neuropathol 108(6):467–470
Vitucci M, Karpinich NO, Bash RE, Werneke AM, Schmid RS, White KK, McNeill RS, Huff B, Wang S, Van Dyke T, Miller CR (2013) Cooperativity between MAPK and PI3K signaling activation is required for glioblastoma pathogenesis. Neuro Oncol 15(10):1317–1329. https://doi.org/10.1093/neuonc/not084
Dasgupta B, Li W, Perry A, Gutmann DH (2005) Glioma formation in neurofibromatosis 1 reflects preferential activation of K-RAS in astrocytes. Cancer Res 65(1):236–245.
Holmen SL, Williams BO (2005) Essential role for Ras signaling in glioblastoma maintenance. Cancer Res 65(18):8250–8255. https://doi.org/10.1158/0008-5472.CAN-05-1173
Bunda S, Burrell K, Heir P, Zeng L, Alamsahebpour A, Kano Y, Raught B, Zhang ZY, Zadeh G, Ohh M (2015) Inhibition of SHP2-mediated dephosphorylation of Ras suppresses oncogenesis. Nat Commun 6:8859. https://doi.org/10.1038/ncomms9859
Gömöri E, Dóczi T, Pajor L, Matolcsy A (1999) Sporadic p53 mutations and absence of ras mutations in glioblastomas. Acta Neurochir 141(6):593–599. https://doi.org/10.1007/s007010050348
Masliah-Planchon J, Garinet S, Pasmant E (2016) RAS-MAPK pathway epigenetic activation in cancer: miRNAs in action. Oncotarget. 7(25):38892–38907. https://doi.org/10.18632/oncotarget.6476(Review)
Gabriely G, Yi M, Narayan RS, Niers JM, Wurdinger T, Imitola J, Ligon KL, Kesari S, Esau C, Stephens RM et al (2011) Human glioma growth is controlled by microRNA-10b. Cancer Res 71(10):3563–3572. https://doi.org/10.1158/0008-5472.CAN-10-3568
Godlewski J, Nowicki MO, Bronisz A, Williams S, Otsuki A, Nuovo G, Raychaudhury A, Newton HB, Chiocca EA, Lawler S (2008) Targeting of the Bmi-1 oncogene/stem cell renewal factor by microRNA-128 inhibits glioma proliferation and self-renewal. Cancer Res 68(22):9125–9130. https://doi.org/10.1158/0008-5472.CAN-08-2629
Kim H, Huang W, Jiang X, Pennicooke B, Park PJ, Johnson MD (2010) Integrative genome analysis reveals an oncomir/oncogene cluster regulating glioblastoma survivorship. Proc Natl Acad Sci USA 107:2183–2188. https://doi.org/10.1158/0008-5472.CAN-08-2629
Kim TM, Huang W, Park R, Park PJ, Johnson MD (2011) A developmental taxonomy of glioblastoma defined and maintained by microRNAs. Cancer Res 71:3387–3399. https://doi.org/10.1073/pnas.0909896107
Kwak HJ, Kim YJ, Chun KR, Woo YM, Park SJ, Jeong JA, Jo SH, Kim TH, Min HS, Chae JS, Choi EJ, Kim G, Shin SH, Gwak HS, Kim SK, Hong EK, Lee GK, Choi KH, Kim JH, Yoo H, Park JB, Lee SH (2011) Downregulation of Spry2 by miR-21 triggers malignancy in human gliomas. Oncogene 30(21):2433–2442. https://doi.org/10.1038/onc.2010.620
Hydbring P, Badalian-Very G (2013) Clinical applications of microRNAs. F1000Res 2:136. https://doi.org/10.12688/f1000research.2-136.v3
Diener C, Keller A, Meese E (2022) Emerging concepts of miRNA therapeutics: from cells to clinic. Trends Genet 38(6):613–626. https://doi.org/10.1016/j.tig.2022.02.006
Segal M, Biscans A, Gilles ME, Anastasiadou E, De Luca R, Lim J, Khvorova A, Slack FJ (2020) Hydrophobically modified let-7b miRNA enhances biodistribution to NSCLC and downregulates HMGA2 in vivo. Mol Ther Nucleic Acids 19:267–277. https://doi.org/10.1016/j.omtn.2019.11.008
Segal M, Slack FJ (2020) Challenges identifying efficacious miRNA therapeutics for cancer. Expert Opin Drug Discov 15(9):987–992. https://doi.org/10.1080/17460441.2020.1765770
Anastasiadou E, Seto AG, Beatty X, Hermreck M, Gilles ME, Stroopinsky D, Pinter-Brown LC, Pestano L, Marchese C, Avigan D, Trivedi P, Escolar DM, Jackson AL, Slack FJ (2021) Cobomarsen, an oligonucleotide inhibitor of miR-155, slows DLBCL tumor cell growth in vitro and in vivo. Clin Cancer Res 27(4):1139–1149. https://doi.org/10.1158/1078-0432.CCR-20-3139
Chakraborty C, Sharma AR, Sharma G, Lee S-S (2021) Therapeutic advances of miRNAs: a preclinical and clinical update. J Adv Res 28:127–138. https://doi.org/10.1016/j.jare.2020.08.012
Romano G, Acunzo M, Nana-Sinkam P (2021) microRNAs as Novel Therapeutics in Cancer. Cancers (Basel). 13(7):1526. https://doi.org/10.3390/cancers13071526
Liang L, He X (2021) A narrative review of microRNA therapeutics: understanding the future of microRNA research. Precis Cancer Med 4:33. https://doi.org/10.21037/pcm-21-28
Kim T, Croce CM (2023) MicroRNA: trends in clinical trials of cancer diagnosis and therapy strategies. Exp Mol Med. https://doi.org/10.1038/s12276-023-01050-9
Teplyuk NM, Uhlmann EJ, Gabriely G, Volfovsky N, Wang Y, Teng J, Karmali P, Marcusson E, Peter M, Mohan A, Kraytsberg Y, Cialic R, Chiocca EA, Godlewski J, Tannous B, Krichevsky AM (2016) Therapeutic potential of targeting microRNA-10b in established intracranial glioblastoma: first steps toward the clinic. EMBO Mol Med 8(3):268–287. https://doi.org/10.15252/emmm.201505495
Stalnecker CA, Der CJ (2023) KRAS regulation of miRNA: stepping on the brake to go faster. Mol Cell 83(14):2390–2392. https://doi.org/10.1016/j.molcel.2023.06.029
Helwak A, Kudla G, Dudnakova T, Tollervey D (2013) Mapping the human miRNA interactome by CLASH revealsfrequent noncanonical binding. Cell. 153(3):654–65. https://doi.org/10.1016/j.cell.2013.03.043
Diener C, Keller A, Meese E (2023) The miRNA-target interactions: an underestimated intricacy. Nucleic Acids Res. https://doi.org/10.1093/nar/gkad1142
Acknowledgements
The author is grateful to Frank Slack for critical review of the manuscript.
Funding
Open access funding provided by Università degli Studi Roma Tre within the CRUI-CARE Agreement. This work was supported by the Grant of Excellence Departments, MUR (ARTICOLO 1, COMMI 314337 LEGGE 232/2016) to Department of Science.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Messina, S. The RAS oncogene in brain tumors and the involvement of let-7 microRNA. Mol Biol Rep 51, 531 (2024). https://doi.org/10.1007/s11033-024-09439-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-024-09439-z