Abstract
miR-3127-5p is a primate-specific miRNA which is down-regulated in recurrent NSCLC tissue vs. matched primary tumor tissue (N = 15) and in tumor tissue vs. normal lung tissue (N = 177). Reduced miR-3127-5p expression is associated with a higher Ki-67 proliferation index and unfavorable prognosis in NSCLC. Overexpression of miR-3127-5p significantly reduced NSCLC cells proliferation, migration and motility in vitro and in vivo. The oncogene ABL1 was a direct miR-3127-5p target and miR-3127-5p regulated the activation of the Abl/Ras/ERK pathway and transactivated downstream proliferation/metastasis-associated molecules. Overexpression of miR-3127-5p in A549 or H292 cells resulted in enhanced resistance to dasatinib, an Abl/src tyrosine kinase inhibitor. miR-3127-5p expression levels were correlated with dasatinib sensitivity in NSCLC cell lines without K-Ras G12 mutation. In conclusion, miR-3127-5p acts as a tumor suppressor gene and is a potential biomarker for dasatinib sensitivity in the non-mutated Ras subset of NSCLC.
Similar content being viewed by others
Introduction
Lung cancer is a leading cause of cancer deaths worldwide1,2. Non-small-cell lung cancer (NSCLC) accounts for approximately 87% of lung cancer cases3. Dozens of microRNAs (miRNAs) are reportedly involved in NSCLC tumorigenesis and progression and induce post-transcriptional gene silencing to regulate cell proliferation, apoptosis, migration or contribute to drug sensitivity. For example, miR-21, the miR-17-92 cluster and miR-221/222 are up-regulated in NSCLC tumor samples and appear to act as oncogenes4,5, whereas other miRNAs, such as let-7, miR-29, miR-15a, miR-16 and miR-34 function as tumor suppressors6,7,8,9. miR-3127-5p (MELmiRNA-108) was first identified in human melanoma samples by deep sequencing in 201010. This primate-specific miRNA11 is also found in other types of human cancer cells12. Our preliminary studies using miRNA microarrays and Q-PCR revealed that certain species-specific miRNAs, including miR-3127-5p, are down-regulated in relapsed lung cancer tissues compared to levels in paired primary lung cancer samples, however, the exact biological role of miR-3127 in cells was unknown.
In this study, we addressed the relationship between expression levels of miR-3127-5p, clinicopathological characteristics and post-operative overall survivals in NSCLC samples and sought to determine the mechanism of miR-3127-5p action in tumorigenesis through in vivo and in vitro experiments. We also investigated the correlation between miR-3127-5p expression levels and dasatinib sensitivity in NSCLC cell lines. Our results indicate that miR-3127-5p acts as a tumor suppressor gene and suggest that this miRNA has potential for use in diagnosis and therapeutic development for NSCLC.
Results
Patients
The general information of 15 cases of paired primary lung tumor and recurrent tumor is summarized in Table S1. Demographic and clinical information is summarized in Table S2 for 177 NSCLC patients whose tumor samples were evaluated for miRNA expression and IHC analysis; follow-up was completed with 175 patients (98.9%).
miR-3127-5p is down-regulated in recurrent NSCLC
We performed comprehensive miRNA microarray analyses on three pairs of primary and recurrent NSCLC tissue using human miRNA OneArray® microarray (Sanger miRBase release 17.0). A set of miRNAs were found to be expressed at significantly lower levels in relapsed NSCLC samples than in tumor tissue from patients with primary cancer (Figure 1A). We selected four primate-specific miRNAs, i.e. miR-3127-5p, miR-3940-5p, miR-4294 and miR-4447 to validate their expression in 15 pairs of recurrent tumor and matched primary tumor FFPE tissue by Q-PCR. Among them, miR-3127-5p, miR-3940-5p and miR-4294 were notably downregulated in relapsed tumors compared to matched primary tumors (Figure 1B, Figure S1A).
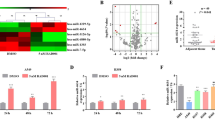
miR-3127-5p expression is down-regulated in NSCLC tumor tissues and low miR-3127-5p levels correlate with unfavorable prognosis in patients with NSCLC.
(A): miRNA expression in primary tumor tissues and tissues from relapse tumors were evaluated by microarray analysis. (N = 3, P: primary tumor; R: recurrent tumor, miRNAs with log2 R/P<-0.8 or log2R/P>0.8 were shown. miR-3127-5p, miR-3940-5p, miR-4294 and miR-4447 for validation were indicated). (B): The expression of miR-3127-5p was significantly down-regulated in relapsed tumor tissue compared to paired primary tumor tissue. Paired Student's t-test (N = 15). (C): The expression of miR-3127-5p in normal, tumor-adjacent and tumor tissues of 177 matched NSCLC samples. N: normal; TN: tumor adjacent normal; T: tumor. Data were analyzed using Wilcoxon Mann Whitney U test; middle line: median; upper and bottom line: min and max; upper and bottom edge of box: 25% and 75% IQR. (D): Reduced miR-3127-5p expression is associated with shorter overall survival in patients with NSCLC (Low: n = 59; Middle: n = 58; High: n = 58; log-rank test; 2YS: 2-year overall survival). (E): Left panel: miR-3127-5p expression was significantly down-regulated in Ki-67-positive NSCLC tumors (N = 177, Wilcoxon Mann Whitney U test). Right panel: Representative photographs of Ki-67 IHC positive and negative NSCLC tissues (ADC: adenocarcinoma; SQC: squamous carcinoma; AD-SQC: adeno-squamous carcinoma; Numbers in parentheses: -ΔΔCT for miR-3127-5p).
The expression of the 4 miRNAs were also detected in the tumor tissue, in tumor-adjacent tissue and in normal tissues taken from 177 patients with NSCLC. We observed markedly lower levels of miR-3127-5p, miR-3940-5p and miR-4294 in tumors than in normal tissue (p < 0.01). The expression of miR-3127-5p and miR-3940-5p were also lower in tumor-adjacent tissue than in normal tissue (p < 0.01) (Figure 1C, Figure S1B).
Reduced miR-3127-5p expression is associated with unfavorable prognosis in NSCLC
To investigate the association of miR-3127-5p, miR-3940-5p and miR-4294 expression with prognosis, 177 NSCLC patients were divided into three groups equally based on miRNA expression levels (high, middle and low). Overall survival was found to be different significantly within groups based on miR-3127-5p expression level (Figure 1D; log-rank test, p = 0.019), while miR-3940-5p and miR-4294 expression were not associated with prognosis in NSCLC (Figure S1C). The 2-year survival rate was significantly worse in the low miR-3127-5p expression group compared to the middle and high expression groups (49.5%, 63.8% and 81.5%, respectively). This finding suggests that reduced miR-3127-5p expression is a predictive factor of poor survival in NSCLC.
We sought correlations between miR-3127-5p expression and other clinicopathological features (Table 1). miR-3127-5p expression levels were significantly lower in clusters of pTNM stage II-IV samples and in tumors with diameters of greater than 3 cm (p = 0.003). The reduced miR-3127-5p expression was also marginally significantly lower in mediastinal lymph metastasis group (p = 0.052).
We also found that miR-3127-5p expressions were specifically down-regulated in tumors that expressed Ki-67, a nuclear marker for proliferation (Figure 1E, Table 2). This finding further confirmed that reduced miR-3127-5p was associated with tumor growth and proliferation in NSCLC.
miR-3127-5p inhibits proliferation, migration and motility in A549 and H292 NSCLC cells in vitro
To examine the biological role of miR-3127-5p in NSCLC cells, we generated stable miR-3127-5p or miR-3127-5p inhibitor transduced NSCLC cell lines A549 and H292 and evaluated effects on proliferation, migration and motility. As shown in Figures 2A and 2B, exogenous overexpression of miR-3127-5p in A549 and H292 significantly reduced cell proliferation rates compared to controls (p< 0.01, repeated measure ANOVA). In contrast, miR-3127-5p inhibitor significantly promoted proliferation. The cell cycle analyses showed that the percentage of cells in S phrase decreased in A549 or H292 that overexpressed miR-3127-5p and was elevated in cells expressed miR-3127-5p inhibitor. (Figure 2C). Taken together, these results indicate that miR-3127-5p acts as a tumor suppressor gene in NSCLC cells.
miR-3127-5p affects proliferation, migration and motility of A549 and H292 cells.
(A) and (B): Left panel: Q-PCR determination of miR-3127-5p expression in A549 cells and H292 cells stably transfected with Lv-GFP, Lv-3127, or Lv-3127-off. Right panel: Growth curves of (A) A549 cells and (B) H292 cells stably transduced with control Lv-GFP; Lv-3127, a vector for overexpression of miR-3127-5p; or Lv-3127-off, a vector for expression of an miR-3127-5p inhibitor. Cells (1000/well) were cultured in a 96-well plate and cell growth was monitored every 24 h for 7 days using a CCK-8 assay, each type of cell was analyzed in quadruplicate (p < 0.01 for A549 cells and H292 cells, repeated measure ANOVA). (C): Cell cycle analysis. The percentage of cells in G0/G1 phase increased and that in S phase decreased in Lv-3127 transduced A549 (left panel) and H292 (right panel) cells; the opposite was observed in Lv-3127-off transduced cells. (D): miR-3127-5p overexpression reduced cell motility and miR-3127-5p knockdown increased cell motility in the wound scratch assay in both A549 (left panel) and H292 (right panel) cells. A uniform scratch was made in each confluent layer culture; the extent of the wound closure was monitored under a phase-contrast microscope and photographs were taken at 0, 12 and 24 h. Triplicate experimentation generated similar results. (E): A549 (upper panels) and H292 (lower panels) cells were loaded onto the top well of the transwell inserts in triplicate for cell migration assay. Photographs are representative of cells that had migrated to the bottom chamber after 48 h. Cells were stained with hematoxylin and observed under a microscope (×200). Invasion was quantified by determining the total number of cells that had migrated through the membrane.
Transwell invasion assay and wound healing assay were performed to investigate whether miR-3127-5p could prevent NSCLC cell migration and invasion. As shown in Figure 2D, motility of cells at 12 and 24 hours after generation of the wound was monitored under a microscope in the scratch wound healing assay. Closure of the wound was complete within 24 hours in inhibitor transduced A549 and H292. In contrast, miR-3127-5p overexpression resulted in cells that migrated toward the wound much more slowly than did the control cells. In the transwell invasion assay, few cells (less than 5%) migrated to the lower chamber in cells in which miR-3127-5p was overexpressed and significantly more cells migrated to the lower chamber in inhibitor transduced A549 and H292 cells than did control cells (Figure 2E; p<0.01). Collectively, these results demonstrate that miR-3127-5p overexpression notably reduces and miR-3127-5p knockdown promotes the invasion and migration ability of NSCLC cells.
miR-3127-5p expression levels influence NSCLC tumorigenicity and metastasis in mouse xenograft model
The effect of miR-3127-5p overexpression or knockdown on NSCLC tumorigenicity and metastasis in vivo were evaluated in mouse A549 or H292 xenograft models. The growth of the xenografted tumors were measured for 5 weeks. As shown in Figures 3A and 3B, the growth of tumors of cells overexpressing miR-3127-5p were significantly slower than that of controls. Growth of xenografts of cells expressing miR-3127-5p inhibitor was faster than that of the Lv-GFP control tumors (repeated measure ANOVA,p < 0.01).
miR-3127-5p knockdown promotes the tumorigenicity and metastasis in mouse xenograft model.
(A): Growth curve of subcutaneous tumors from A549 (upper panels) or H292 (bottom panels) cells transduced with Lv-3127, Lv-3127-off, or Lv-GFP in the first 5 weeks after inoculation. (B): Xenografted tumors were observed using a whole-body fluorescent imaging system at 30 days after inoculation. (C): After 13 weeks, animals were sacrificed and the number of visible neoplastic nodules (metastases) on each lung was counted. Representative photographs of the whole lungs and HE staining for lung metastasis lesion sections (100×) are shown. (D): Representative photographs of c-Abl, Ki-67, cyclin D1, MMP-9 and VEGF IHC of A549 or H292 xenograft tumors (200×).
The number of visible neoplastic nodules (metastases) on each lung was counted and evaluated at 13th weeks after inoculation. Significantly fewer lung metastases were observed in miR-3127-5p overexpressing xenografts and more were observed in miR-3127-5p inhibitor xenografts than in the Lv-GFP control groups (Figure 3C).
Several metastasis-associated molecules, cyclin D1, MMP-9 and VEGF are reportedly trans-activated by Abl kinase signaling13,14,15,16, we found that levels of cyclin D1, VEGF and MMP-9 increased in the miR-3127-5p inhibitor A549 or H292 xenografts and were decreased in miR-3127-5p–overexpressing xenografts compared to those in controls (Figure 3D).
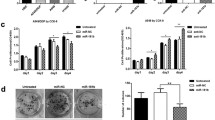
ABL1 is a direct miR-3127-5p target and miR-3127-5p regulates the activation of c-Abl/Ras/ERK pathway in NSCLC cells
To delineate the mechanism by which miR-3127-5p inhibits NSCLC cell proliferation, invasion and migration, we searched for the target genes of miR-3127-5p targets using TargetScan 6.2. We found that the 3′ untranslated region (3′-UTR) of the human oncogene ABL1 possesses two regions of 7 bases that are complementary to miR-3127-5p and conserved in human and other mammalians (Figure 4A). We cloned the wild-type ABL1 3′-UTR or a 3′-UTR with mutations in the putative seed sites into the luciferase reporter vector. The overexpression of miR-3127-5p significantly decreased the luciferase activity from the construct with the wild-type 3′-UTR but had little effect on activity of the construct with the mutant ABL1 3′-UTR in 293T cells; expression of miR-3127-5p inhibitor had the reverse effect (Figure 4B). c-Abl protein levels also decreased in A549 and in H292 cells with overexpression of miR-3127-5p and increased in miR-3127-5p knockdown A549 or H292 cells (Figure 4C). In functional complementation assay, stable H292 overexpressing miR-3127-5p was electroporated with the pGV142-CMV-ABL1 pasmid. The results showed that the cell cycle distribution, inhibition of migration and invasion caused by miR-3127-5p could be reversed by exogenou c-Abl overexpression (Figure 4D, E). To investigate the role of miR-3127-5p-c-Abl axis in NSCLC clinical samples, we examined ABL1 mRNA expression in the 177 cases of paired NSCLC samples by Q-PCR. ABL1 mRNA was highly expressed in NSCLC tumor samples (Figure S2A) and ABL1 mRNA expression levels were much higher in groups with low levels of miR-3127-5p expression than in groups with moderate or high expression (Figure S2B, N = 177; Kruskal-Wallis H test). As shown in Figure 4F and Table 2, miR-3127-5p expression was significantly down-regulated in samples with high levels of IHC staining for c-Abl. There were no significant correlations in levels of p53 or EGFR IHC staining with miR-3127-5p expression (Table 2).
Oncogene c-Abl is a direct target of miR-3127-5p.
(A): Two sites in the ABL1 3′-UTR are complementary to miR-3127-5p as analyzed by Targetscan. (B): 293T cells were co-transfected with pmiR-RB-Report-ABL1_3′UTR or pmiR-ABL1_3′UTR-Mut plasmids and pL-GFP, pL-miR-3127-5p, or pL-miR-3127-5p-off plasmids. A Renilla-TK luciferase reporter (5 ng) was added as an internal control. The luciferase activity of the cells was determined after 48 h. (C): A549 or H292 cells transduced with Lv-3127, Lv-3127-off, or Lv-GFP were subjected to immunoblotting for c-Abl (The full-length blots were presented in the supplementary Figure S4). (D): H292-3127 was transducted with pGV142-CMV-ABL1 and control vectors, the cell cycle analysis and immunoblotting of c-abl were performed at 36hrs after eletroporation. (E): Wound scratch assay (left panel) and transwell invasion assay (right panel) of H292-3127 transduced with pGV142-CMV-ABL1 (Magnification, ×200). Triplicate experiments were done. (F): Left panels: Representative photographs of c-Abl IHC staining grades in NSCLC tumor samples (200×). Right panels: miR-3127-5p expression was low in c-Abl IHC positive staining samples (mean ± SEM, ANOVA).
It is well known that oncogenic c-Abl can activate Ras/ERK signaling by interaction with Grb2 and induce constitutive activation of Ras17; therefore, the c-Abl/Ras/ERK signaling axis is critical for malignant progression in some tumors. We found that there was an decrease/increase in levels of the active Ras-GTP form in the miR-3127-5p/inhibitor transduced A549 or H292 as shown after precipitation with Raf1-RBD and immunoblotting with a Ras antibody (Figure 5A). We found that by silencing ABL1 expression, miR-3127-5p overexpression down-regulated p-ERK in A549 and H292 cells; miR-3127-5p inhibitor had the opposite effect on p-ERK levels. We also examined several proliferation and metastasis-associated molecules (i.e., cyclin D1, MMP-9 and VEGF) that are trans-activated by c-Abl/Ras/ERK signaling and found that these factors were down- or up-regulated by miR-3127-5p overexpression or knockdown, respectively (Figure 5B). Taken together, these results indicate that miR-3127-5p regulates the activation of the c-Abl/Ras/ERK signaling by targeting ABL1 (as illustrated schematically in Figure 5C), thus miR-3127-5p plays a role in tumorigenesis, invasion and metastasis of NSCLC.
miR-3127-5p regulates c-Abl/Ras/ERK pathway activation by targeting oncogene c-Abl.
(A): A549 or H292 cells transduced with Lv-3127, Lv-3127-off, or Lv-GFP were lysed. Lysates (1.0 mg) were incubated with 50 μL of GST-Raf1-RBD beads for 30 min to pull down active Ras. Samples were immunoblotted using an anti-K-Ras antibody, thereby allowing levels of GTP-bound Ras within the cells to be assessed. Whole cell lysates (50 µg) were also subjected to western blot for detection of total K-Ras. The numbers under bands indicate the ratio of integrated optical density to GAPDH normalized by GFP control (The full-length blots were presented in the supplementary Figure S5). (B): A549 or H292 cells transduced with Lv-3127, Lv-3127-off, or Lv-GFP were subjected to immunoblotting with antibodies to ERK, p-ERK and downstream factors MMP-9, VEGF and cyclin D1 (The full-length blots were presented in the supplementary Figure S4, 5). (C): Schematic diagram illustrating the effect of reduced miR-3127-5p levels on activation of c-Abl/Ras/ERK pathway and promotion of tumor proliferation and metastasis.
miR-3127-5p down-regulation is associated with dasatinib sensitivity in NSCLC cell lines without the K-Ras G12 mutation
Dasatinib is a small-molecule, ATP-competitive inhibitor of c-Abl and src tyrosine kinases. We evaluated the dasatinib sensitivity in miR-3127-5p or inhibitor transduced A549 or H292. Interestingly, we found that cells that overexpress miR-3127-5p were much more resistant to dasatinib than control cells; the IC50 value (the half-maximum growth inhibitory concentration) for A549 cells that overexpress miR-3127-5p was 15.0 μg/mL compared to 9.5 μg/mL for control A549 cells and 0.4 μg/mL compared to 0.06 μg/mL for H292 cells. In contrast, miR-3127-5p inhibitor resulted in enhanced sensitivity to dasatinib with IC50 of 0.5 μg/mL compared to 9.5 μg/mL for A549 cells and IC50 of 0.02 μg/mL compared to 0.06 μg/mL for H292 cells (Figure 6A, p < 0.01, repeated measure ANOVA). In H292 cells that harbor wild-type K-Ras, overexpression of miR-3127-5p enhanced resistance by approximately 7 fold, a significantly greater enhancement than observed in A549 cells harboring the K-Ras G12 mutation. The results imply that lower miR-3127-5p expression levels contribute to dasatinib sensitivity, suggesting that miR-3127-5p may be a candidate biomarker for prediction of dasatinib sensitivity in NSCLC.
between expression of miR-3127-5p and response to dasatinib in NSCLC cell lines.
(A): Dasatinib sensitivity of A549 (left) and H292 (right) cells transduced with Lv-3127, Lv-3127-off, or Lv-GFP as determined in a CCK8 cell viability assay. (p < 0.01, repeated measure ANOVA). (B): miR-3127-5p expression in the 14 NSCLC cell lines as determined using an miRNA poly (A) tailing-based Sybr Green Q-PCR assay. (C): Immunoblotting of c-Abl protein for NSCLC cell lines. The number under each band refers to the integrated band density (The full-length blots were presented in the supplementary Figure S6). (D): Negative relationship between c-Abl protein and miR-3127-5p expression in the NSCLC cell lines without K-Ras condon 12 mutation (Pearson's correlation, p < 0.01, r2 = 0.443). (E): IC50 values for dasatinib in NSCLC cell lines determined by CCK8 cell viability assay. (F): miR-3127-5p expression level was linearly associated with dasatinib IC50 for the 11 NSCLC cell lines after exclusion of the three cell lines with a K-Ras codon 12 mutation (K-Ras mutation status: H23(G12C), H358(G12C), A549(G12S), H460(Q61H), SPC-A1(Q61H). Pearson's correlation, p < 0.01, r2 = 0.895).
To confirm the association of miR-3127-5p expression level with dasatinib sensitivity, we examined the miR-3127-5p, c-Abl protein expression and dasatinib IC50 in 14 NSCLC cell lines. We found that miR-3127-5p expression was linearly associated with dasatinib IC50 for the 11 NSCLC cell lines without the Ras G12 mutation (i.e., A549[G12S], H23[G12C] and H358[G12C]) (Figure 6B, E, F, Pearson's correlation, p < 0.01, r2 = 0.895). MiR-3127-5p expression was also negatively related to c-Abl protein level in the NSCLC cell lines without K-Ras condon 12 mutation (Figure 6C, D, Pearson's correlation, p < 0.01, r2 = 0.433). Reduced miR-3127-5p expression means higher sensitivity to dasatinib in NSCLC cell lines without K-Ras codon 12 mutant.
Discussion
We validated the expression of miR-3127-5p, miR-3940-5p, miR-4294 and miR-4447 in NSCLC samples. Our data show that miR-3127-5p is associated with prognosis of NSCLC patients and acts as a tumor suppressor in NSCLC. miR-3127-5p expression was significantly decreased in tumor tissue from patients with recurrent NSCLC as compared to levels in primary tumors. Levels of miR-3127-5p were also lower in tumor tissues than in tumor-adjacent and normal tissues in samples from 177 patients with NSCLC. Reduced miR-3127-5p expression was markedly associated with advanced stage NSCLC. Ki-67, a marker for proliferation of tumor cells, is strictly associated with cell proliferation and present during all active phases of the cell cycle, but absent from resting cells18. We found that miR-3127-5p was down-regulated in Ki-67-positive NSCLC specimens, further suggesting that miR3127-5p is involved tumor cell proliferation. Finally, reduced expression of miR-3127-5p is associated with poor prognosis in NSCLC patients. The 2-year overall survival was only 58% in the patient group with low levels of miR-3127-5p expression but was 82% in the group with high expression levels (p < 0.05). The overexpression of miR-3127-5p in NSCLC cells significantly reduced tumor cell proliferation in both in vitro experiments and a xenograft animal model, whereas inhibition of miR-3127 expression promoted tumor cell proliferation. We also demonstrated that miR-3127-5p acts as a negative regulator of cellular metastatic potential, motility and invasion were reduced in cells engineered to overexpress miR-3127-5p. Based on these observations, we conclude that miR-3127-5p acts as a tumor suppressor gene. To the best of our knowledge, this is the first report that correlates miR-3127-5p with tumorigenic potential.
To study the molecular mechanism responsible for the suppression of NSCLC cell proliferation and invasion caused by miR-3127-5p, we searched for possible gene targets. We focused on a well-known proto-oncogene ABL1; the ABL1 3′-UTR has two conserved regions complementary to the seed region of miR-3127-5p. We found that miR-3127-5p overexpression reduced the luciferase activity of luciferase reporter containing the ABL1 3′-UTR, but did not alter expression of a luciferase reporter with mutations in the putative binding sites. In addition, c-Abl protein levels were up- or down-regulated by miR-3127-5p knockdown or overexpression, respectively, in A549 and H292 cells. The inhibitive effects of migration and invasion caused by miR-3127-5p could be reversed by over-expressing c-Abl. ABL1 mRNA and protein were highly expressed in NSCLC cell lines and tumor samples and levels were negatively correlated with miR-3127-5p expression. These data support the notion that ABL1 is a direct target of miR-3127-5p.
ABL1 is a proto-oncogene that encodes cytoplasmic and nuclear protein tyrosine kinase that has been implicated in processes of cell differentiation, cell division, cell adhesion and stress response19. Although c-Abl non-receptor tyrosine kinases are known to drive leukemia development, recent research suggests that in some solid tumors c-Abl is activated via unique mechanisms that do not involve gene mutation or translocation20. c-Abl activation can promote matrix degradation, invasion, proliferation, tumorigenesis in breast21, lung22 and liver23. In lung cancer, c-Abl expression is correlated with EGFR, IGF-1R, PDGFR, and/or c-Kit expression24. c-Abl can be activated by receptor tyrosine kinases PDGFR25 and EGFR. Activated c-Abl can phosphorylate the EGFR(Tyr1173), impair EGFR internalization and increase EGFR cell-surface expression26. The high c-Abl expression observed in NSCLC tumor tissues may contribute to tumorigenesis and proliferation through by-pass activation. We found that ABL1 mRNA was overexpressed in about 81.4% (144 of 177 cases determined by Q-PCR) and that the protein was overexpressed in 42.0% (75 of 177 cases determined by IHC), in agreement with the finding that c-Abl activation is a frequent event in NSCLC22. Several downstream signaling pathways affected by the c-Abl, v-Abl, or Bcr-Abl have been identified; among them, Ras constitutive activation has been well studied17,27,28. Proto-oncogene c-Abl can interact directly with adaptor Grb2, which leads to activation of Ras/ERK signaling17. We found that inhibition of miR-3127-5p expression activated K-Ras and resulted in phosphorylation of ERK1/2 in A549 and H292 cells, whereas overexpression of miR-3127-5p had the reverse effect. We examined several metastasis-associated molecules, cyclin D1, MMP-9 and VEGF and found that these factors were also down-regulated by miR-3127-5p overexpression. Although there may be other miR-3127-5p target genes, we suggest that miR-3127-5p regulates cell proliferation and invasion by directly controlling expression of ABL1.
Dasatinib (BMS-354825, Bristol-Myers Squibb), an Abl/Src kinase inhibitor that can bind to both the active and inactive conformations of the ABL kinase domain29, is used first-line treatment for chronic myelogenous leukemia and Ph-positive acute lymphoblastic leukemia. Data from a phase II clinical trial of dasatinib in molecularly unselected patients with NSCLC suggested that dasatinib as a single agent had only a modest therapeutic effect that was lower than platinum-based chemotherapy but that a potential subpopulation of patients with NSCLC may be sensitive to dasatinib30. Therefore, molecular markers are needed to predict patient responses to dasatinib. Genetic variants of EGFR, K-Ras, p53 and Smad4/DPC4 do not predict dasatinib response in NSCLC or other solid tumors31,32. Here, we described that miR-3127-5p inhibitor enhanced the dasatinib sensitivity of NSCLC cells with a K-Ras G12 mutant and cells without any Ras mutation. The contribution of overexpression of miR-3127-5p to dasatinib resistance was more apparent in H292 cells harboring wild-type K-Ras. To verify the association of miR-3127-5p expression with dasatinib sensitivity, we examined the dasatinib sensitivity, miR-3127-5p expression level and the mutation status of K-Ras in 14 NSCLC cell lines (Table S3). We found that miR-3127-5p expression level was linearly correlated with dasatinib IC50 in NSCLC cell lines after exclusion of the three cell lines, A549, H358 and H23, which harbor the K-Ras G12 mutation. Because activating K-Ras mutants can produce independent tumor-driving effects, we think it is reasonable to discriminate cell lines with non-mutated Ras from the mutated when evaluating the association of dasatinib sensitivity with c-Abl/Ras/ERK pathway activation by miR-3127-5p. In the other hand, although dasatinib is also an inhibitor for c-src, we did not find the correlation of dasatinib sensitivity to c-src expression in NSCLC cell lines (Figure S3, Pearson's correlation, p > 0.05, r2 = 0.057). Dasatinib may benefit patients with wild-type Ras and low levels of miR-3127-5p expression. Our results suggest that miR-3127-5p is a candidate for a dasatinib sensitivity biomarker for NSCLC; however, this requires verification in larger numbers of clinical samples.
Recent findings suggest a significant function of species-specific miRNAs in varied biological processes, including tumorigenesis33,34,35, inflammation, apoptosis, or induction of pluripotency36. miR-3127-5p is a primate-specific miRNA which is frequently down-regulated in advanced NSCLC. Reduced miR-3127-5p expression can promote tumor proliferation, enhance cell motility and invasion both in vitro and in vivo by targeting proto-oncogene c-Abl and lead the activation of c-Abl/ras/ERK pathway. We demonstrated that reduced miR-3127-5p expression level in NSCLC cell lines is closely associated with dasatinib sensitivity, especially in samples without the K-Ras G12 mutation. miR-3127-5p may contribute to dasatinib sensitivity and its use as a biomarker is worth further investigation.
Methods
Patients and samples
The procedure of human sample collection was approved by the Ethical Committee of Tongji University. All patients signed informed consent for the collection and use of their tissues for this study. Fifteen pairs of primary and recurrent NSCLC formalin fixed, paraffin-embedded (FFPE) samples were obtained from patients who underwent surgery after relapse of NSCLC in Shanghai Pulmonary Hospital. The 177 fresh frozen specimens used in this study were obtained from Shanghai Pulmonary Hospital between December 2009 and April 2012. None of the patients had been treated with adjuvant chemotherapy prior to the operation. The end-point of follow-up was August 2012, or date of death. Normal (N), tumor-adjacent normal (TN) and tumor (T) tissue were obtained from excision specimens for each lung cancer patient.
Total RNA extraction, microarray and real-time PCR quantification of miRNAs
RNA was extracted from frozen tissue samples or cell lines using a miRNA easy Mini kit (QIAGEN, GmbH; Hilden, Germany). For purification of DNA and RNA from FFPE samples, kits from EPICENTRE Biotechnologies (Madison, WI, USA) were used. Microarray analysis was performed with miRNA OneArray (Sanger miRBase, V17.0; Phalanx; Hsinchu, Taiwan).
For quantitative PCR (Q-PCR) analysis of miRNAs, cDNA was reverse transcribed from total RNA samples using the QIAGEN miScript Reverse Transcription Kit. Specific primers for each miRNA were obtained from QIAGEN: hs-miR-3127-5p (Cat. No. MS 00020503), miR-3940-5p (Cat. No. MS 0003019), miR-4294 (Cat. No. MS 00021427), miR-4447 (Cat. No. MS00041230) and hs-RUN6-2-1 (Cat. No. MS 00033740). Q-PCR for each sample was performed in duplicate. The -ΔΔCt method was used to analyze the relative quantitative expression levels of miRNAs with snRNA U6 as an internal control gene. Target gene ABL1 was amplified from 177 cases of paired tumor and normal samples by SYBR Green real-time PCR; forward primer: TCAACAAGATGTTTTGCCAACTG; reverse primer: ATGTGCTGTGACTGCTTGTAGATG.
Immunohistochemistry of Ki-67, c-Abl, p53, EGFR, cyclin D1, MMP-9 and VEGF
Anti-p53 (Clone DO-7, DAKO; Glostrup, Denmark), anti-EGFR (Clone EGFR.25; Gene Tech; Shanghai, China), anti-c-Abl (HPA028409; Sigma-Aldrich; St. Louis, MO, USA), anti-ki-67 (Clone MIB-1, DAKO), anti-cyclin D1 (#AC853, Beyotime Institute of Biotechnology; Jiangsu, China) were used as primary antibodies for immunohistochemistry (IHC) staining. Immunoreactive counting scores were based on four grades that depended on positive nuclear staining for Ki-6737 and c-Abl, or positive cytoplasmic and nuclear staining for p53, or positive membrane staining for EGFR. Immunohistochemical staining in surgical specimens was independently assessed by two pathologists.
Cell lines, lentivirus and infection
The NSCLC cell lines were provided by the ATCC (Manassas, VA, USA). GFP hsa-mir-3127-5p Lentivector (Lv-3127, Cat. No. mh1036), inhibitor hsa-miR-3127-5p Lentivector (Lv-3127-off, Cat. No. mh30446, expressing an miR-3127-5p inhibitor) and pLenti-III-mir-GFP-Blank (Lv-GFP, Cat. No. m001) were purchased from Applied Biological Materials Inc. (Richmond, BC, Canada). pGV142-CMV-ABL1 was provided by Aodeshi Biological Inc. (Shanghai, China). Lentiviruses were packaged in 293T cells and infected per the manufacturer's recommended protocol. A549 or H292 cells were infected with lentiviruses Lv-3127, Lv-3127-off and Lv-GFP. Stable transduced cells were generated by seeding infected A549 or H292 cells in puromycin-supplemented medium (0.8 μg/mL for A549 and 0.2 μg/mL for H292) for 2 weeks. Transduction of pGV142-CMV-ABL1 and vector control into H292-3127 was performed on a Gene Pulser (Bio-rad, CA, USA) and the electroporated cells were cultured in DMEM with 10% serum.
Cell cycle analysis
Cells were trypsinized and washed, then fixed in 70% ethanol. Cells (5 × 105) were resuspended in 0.5 mL PBS and treated with 50 μg/mL propidium iodide and 1 μg/mL RNase for 30 minutes. Samples were analyzed with a Beckman Coulter FC500 (Beckman Coulter; Brea, CA, USA). Data were collected and processed using CXP and MultiCycle AV software.
Proliferation, migration and invasion assay
Stable transduced A549 and H292 (1000/well) were seeded in a 96-well plate in 100 µl complete DMEM supplemented with 10% fetal bovine serum and cell growth was monitored every 24 h for 7 days using a CCK-8 assay.
Migration assays were performed in transwell inserts with 8-μm pore, uncoated membrane filters (Corning Inc.; Corning, NY, USA). Serum-starved adherent cells are trypsynized, 200 µl cell suspension (2 × 105) in serum-free medium was placed into the upper chamber and the low chamber was filled with 500 µl 10% FCS-DMEM. After 24 h, cells that migrated to the lower chamber were fixed and stained with hematoxylin. Experiments were independently repeated three times.
For the scratch wound healing assay, cells were streaked with a 200 µl sterile pipette tip, washed with PBS and replenished with serum-free DMEM. Wounds were imaged using an inverted microscope (Olympus; Tokyo, Japan) at 12 and 24 hours after wounding.
Immunoblot assay
Protein lysates were prepared, subjected to SDS/PAGE, transferred onto nitrocellulose membranes and blotted according to standard methods. The primary antibodies used were anti-GAPDH 1:1,000 (#sc-47724), anti-K-Ras(F234) 1:500 (#sc-30), anti-c-Abl(K-12) 1:500 (#sc-131), anti-cyclin D1 1:500 (#sc-753), anti-VEGF 1:500 (#sc-152) (Santa Cruz Biotechnology; Santa Cruz, CA, USA); anti-MMP-9 1:4000 (#ab76003) (Abcam Ltd., Cambridge, MA, USA); and ERK1/2(#4695), p-ERK1/2(#4370) (Cell Signaling Technology; Danvers, MA, USA). Secondary antibodies were anti-mouse (#sc-2031) and anti-rabbit (#sc-2317) HRP-conjugated (Santa Cruz Biotechnology) used at 1:5,000 dilutions.
Luciferase reporter assay
pmiR-RB-Report-ABL1_3′UTR Vector was obtained from Ribbio Co. Ltd. (Guangzhou, China). The constructs with mutations in both predicted miR-3127-5p seed sites (pmiR-ABL1_3′UTR-Mut) of the ABL1 3′-UTR were generated with mutagenic oligonucleotide primers (Table S4) according to the protocol supplied with the QuikChange II Site-Directed Mutagenesis Kit (Agilent; Santa Clara, CA, USA). The relative luciferase activity of the cells was determined after 48 h in a dual-luciferase reporter assay as per the manufacturer's instructions (Promega; Madison, WI, USA) on a FLX800 fluorescence and luminescence Reader (Bio-tek, Winooski, VT, USA).
Ras-GTP pull-down assay
The active GTP-bound form of Ras was measured with a K-Ras Activation Assay Kit (Cell Biolabs; San Diego, CA, USA) following the manufacturer's instructions. Briefly, total cell lysates (1.0 mg) were incubated with 50 μL Raf1-RBD-agarose beads for 30 min to pull down activated GTP bound Ras. The Raf1 RBD bound protein was then analyzed by western blot and probed with antibody to K-Ras (#sc-30). Whole cell lysates (50 µg) were also subjected to western blot for detection of total K-Ras.
Immunodeficient mouse xenograft tumor model
The murine studies were conducted in accordance with the Institutional Animal Care and Use Committee (IACUC) guidelines and were approved by the Animal Care and Use Committee of Tongji University Animal Center.
A549 and H292 cells (2 × 106 in phosphate-buffered saline containing 50% Matrigel) stably transduced with Lv-3127, Lv-3127-off, or Lv-GFP were injected subcutaneously into the unilateral hind limb of BALB/C nude mice (6 weeks old, N = 6 for each group). The volumes of tumors were recorded four days after transplantation day: tumor volume = length × width2/2. In vivo imaging (IVIS Lumina II In vivo imaging system, USA) was used to detect GFP fluorescence in transplanted mice at 30 days post-injection. At 13 weeks after injection, mice were anesthetized and sacrificed; the visible neoplastic nodules (metastases) on the lungs were counted and then were paraffin-embedded to perform hematoxylin and eosin (HE) staining.
Cell viability and cell sensitivity assay to dasatinib
Stable transduced A549 or H292 cells and 14 NSCLC cell lines (H460, PC9, A549, H1975, H1703, H1650, H23, H1299, H292, H358, H1395, SK-MES-1, SPC-A1, H661) were evaluated for the dasatinib sensitivity in vitro. Cells were cultivated in 96-well plates (Corning) at a seeding density of 5,000 cells per well in the presence of dasatinib (Dalian Meilun Biotech Co.; Dalian, China). The experiments were conducted on 96-well plates with eight compound concentrations (0 nM, 7 nM, 0.029 μM, 0.117 μM, 0.469 μM, 1.88 μM, 7.5 μM, 30 μM). Four wells per concentration were tested. Sensitivity of the cell lines to dasatinib was determined using a CCK-8 assay after 96 hours of treatment.
K-Ras mutation detection
Genomic DNA was extracted from all cell lines. The primers used to amplify the K-Ras gene exons 2, 3 and 4 are listed in Table S5. The PCR products were purified from a 2.0% agarose gel and isolated products were subjected to direct sequencing using the same primers.
Statistical analysis
Data were expressed as median and interquartile range (IQR); statistical differences in miRNA expression levels between two groups were calculated using the nonparametric Wilcoxon Mann Whitney U test. The Kruskal-Wallis H test was used on several independent samples. A paired t-test was used for paired samples. Pearson's correlation was used to evaluate the correlation coefficient of the two groups. The significance of tumor volume increases and cell line growth curves were evaluated by univariate test of repeated measures ANOVA. A chi-square test was used to compare proportional data. The survival curves were estimated by the Kaplan-Meier method and the resulting curves were compared using the log-rank test. Statistical analyses were performed with SPSS 17.0 software (SPSS Inc.; Chicago, IL, USA). The level of significance was defined as a p value of less than 0.05.
References
Jemal, A. et al. Global cancer statistics. CA Cancer J Clin 61, 69–90 (2011).
Siegel, R., Naishadham, D. & Jemal, A. Cancer statistics, 2013. CA Cancer J Clin 63, 11–30 (2013).
Govindan, R. et al. Changing epidemiology of small-cell lung cancer in the United States over the last 30 years: analysis of the surveillance, epidemiologic and end results database. J Clin Oncol 24, 4539–4544 (2006).
Seike, M. et al. MiR-21 is an EGFR-regulated anti-apoptotic factor in lung cancer in never-smokers. Proc Natl Acad Sci U S A 106, 12085–12090 (2009).
Garofalo, M. et al. miR-221&222 regulate TRAIL resistance and enhance tumorigenicity through PTEN and TIMP3 downregulation. Cancer Cell 16, 498–509 (2009).
Skrzypski, M., Dziadziuszko, R. & Jassem, J. MicroRNA in lung cancer diagnostics and treatment. Mutat Res 717, 25–31 (2011).
Catuogno, S. et al. miR-34c may protect lung cancer cells from paclitaxel-induced apoptosis. Oncogene 32, 341–351 (2013).
Bandi, N. & Vassella, E. miR-34a and miR-15a/16 are co-regulated in non-small cell lung cancer and control cell cycle progression in a synergistic and Rb-dependent manner. Mol Cancer 10, 55 (2011).
Corney, D. C., Flesken-Nikitin, A., Godwin, A. K., Wang, W. & Nikitin, A. Y. MicroRNA-34b and MicroRNA-34c are targets of p53 and cooperate in control of cell proliferation and adhesion-independent growth. Cancer Res 67, 8433–8438 (2007).
Stark, M. S. et al. Characterization of the Melanoma miRNAome by Deep Sequencing. PLoS One 5, e9685 (2010).
Dannemann, M., Nickel, B., Lizano, E., Burbano, H. A. & Kelso, J. Annotation of primate miRNAs by high throughput sequencing of small RNA libraries. BMC Genomics 13, 116 (2012).
Persson, H. et al. Identification of new microRNAs in paired normal and tumor breast tissue suggests a dual role for the ERBB2/Her2 gene. Cancer Res 71, 78–86 (2011).
Janowska-Wieczorek, A. et al. Bcr-abl-positive cells secrete angiogenic factors including matrix metalloproteinases and stimulate angiogenesis in vivo in Matrigel implants. Leukemia 16, 1160–1166 (2002).
Nakamura, Y., Grumont, R. J. & Gerondakis, S. NF-kappaB1 can inhibit v-Abl-induced lymphoid transformation by functioning as a negative regulator of cyclin D1 expression. Mol Cell Biol 22, 5563–5574 (2002).
Mayerhofer, M., Valent, P., Sperr, W. R., Griffin, J. D. & Sillaber, C. BCR/ABL induces expression of vascular endothelial growth factor and its transcriptional activator, hypoxia inducible factor-1alpha, through a pathway involving phosphoinositide 3-kinase and the mammalian target of rapamycin. Blood 100, 3767–3775 (2002).
Noy, P., Gaston, K. & Jayaraman, P. S. Dasatinib inhibits leukaemic cell survival by decreasing PRH/Hhex phosphorylation resulting in increased repression of VEGF signalling genes. Leuk Res 36, 1434–1437 (2012).
Renshaw, M. W., Lewis, J. M. & Schwartz, M. A. The c-Abl tyrosine kinase contributes to the transient activation of MAP kinase in cells plated on fibronectin. Oncogene 19, 3216–3219 (2000).
Scholzen, T. & Gerdes, J. The Ki-67 protein: from the known and the unknown. J Cell Physiol 182, 311–322 (2000).
Sirvent, A., Benistant, C. & Roche, S. Cytoplasmic signalling by the c-Abl tyrosine kinase in normal and cancer cells. Biol Cell 100, 617–631 (2008).
Ganguly, S. S. & Plattner, R. Activation of abl family kinases in solid tumors. Genes Cancer 3, 414–425 (2012).
Srinivasan, D. & Plattner, R. Activation of Abl tyrosine kinases promotes invasion of aggressive breast cancer cells. Cancer Res 66, 5648–5655 (2006).
Lin, J. et al. Oncogenic activation of c-Abl in non-small cell lung cancer cells lacking FUS1 expression: inhibition of c-Abl by the tumor suppressor gene product Fus1. Oncogene 26, 6989–6996 (2007).
Suh, Y. et al. Claudin-1 induces epithelial-mesenchymal transition through activation of the c-Abl-ERK signaling pathway in human liver cells. Oncogene 32, 4873–4882 (2013).
Singer, C. F. et al. Expression of tyrosine kinases in human malignancies as potential targets for kinase-specific inhibitors. Endocr Relat Cancer 11, 861–869 (2004).
Plattner, R., Kadlec, L., DeMali, K. A., Kazlauskas, A. & Pendergast, A. M. c-Abl is activated by growth factors and Src family kinases and has a role in the cellular response to PDGF. Genes Dev 13, 2400–2411 (1999).
Tanos, B. & Pendergast, A. M. Abl tyrosine kinase regulates endocytosis of the epidermal growth factor receptor. J Biol Chem 281, 32714–32723 (2006).
Pendergast, A. M. et al. BCR-ABL-induced oncogenesis is mediated by direct interaction with the SH2 domain of the GRB-2 adaptor protein. Cell 75, 175–185 (1993).
Renshaw, M. W., Lea-Chou, E. & Wang, J. Y. Rac is required for v-Abl tyrosine kinase to activate mitogenesis. Curr Biol 6, 76–83 (1996).
Talpaz, M. et al. Dasatinib in imatinib-resistant Philadelphia chromosome-positive leukemias. N Engl J Med 354, 2531–2541 (2006).
Johnson, F. M. et al. Phase II study of dasatinib in patients with advanced non-small-cell lung cancer. J Clin Oncol 28, 4609–4615 (2010).
Johnson, F. M., Saigal, B., Talpaz, M. & Donato, N. J. Dasatinib (BMS-354825) tyrosine kinase inhibitor suppresses invasion and induces cell cycle arrest and apoptosis of head and neck squamous cell carcinoma and non-small cell lung cancer cells. Clin Cancer Res 11, 6924–6932 (2005).
Nagaraj, N. S., Smith, J. J., Revetta, F., Washington, M. K. & Merchant, N. B. Targeted inhibition of SRC kinase signaling attenuates pancreatic tumorigenesis. Mol Cancer Ther 9, 2322–2332 (2010).
Xiao, J., Lin, H., Luo, X. & Wang, Z. miR-605 joins p53 network to form a p53:miR-605:Mdm2 positive feedback loop in response to stress. EMBO J 30, 5021 (2011).
Zhang, J. F. et al. Primate-specific microRNA-637 inhibits tumorigenesis in hepatocellular carcinoma by disrupting signal transducer and activator of transcription 3 signaling. Hepatology 54, 2137–2148 (2011).
Perdomo, C. et al. MicroRNA 4423 is a primate-specific regulator of airway epithelial cell differentiation and lung carcinogenesis. Proc Natl Acad Sci U S A 110, 18946–51 (2013).
Mor, E. & Shomron, N. Species-specific microRNA regulation influences phenotypic variability: perspectives on species-specific microRNA regulation. Bioessays 35, 881–888 (2013).
Soomro, I. N., Holmes, J. & Whimster, W. F. Predicting prognosis in lung cancer: use of proliferation marker, Ki67 monoclonal antibody. J Pak Med Assoc 48, 66–69 (1998).
Acknowledgements
This work was funded by National Natural Science Foundation of China (No. 81200034), Science and Technology Commission of Shanghai Municipality (No. 10411956100, No. 12ZR1426100, No. 09411960500, No. 10411967600, No. 2010Y094, 12nm0500800) and Wu Jieping Medical Foundation (No. 320.6720.10003).We are extremely grateful to Mr. Junjie Zhu, Mr. Liang Tan, Mr. Xiaojun Yang, Mr. Jie Zhang for experimental instruction and to Mrs. Meiling Zhu for providing patient information.
Author information
Authors and Affiliations
Contributions
Y.S., P.Z., C.C. W.G. and B.S. conceived the experiments. Y.S., B.S. and H.Z. wrote the paper. L.H., Y.X. and H.X. performed the experiments. Q.D. helped to do the experiments. Z.X. performed Statistical analysis. W.G. gave scientific advice. All authors contributed to the analysis of this manuscript and reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Sun, Y., Chen, C., Zhang, P. et al. Reduced miR-3127-5p expression promotes NSCLC proliferation/invasion and contributes to dasatinib sensitivity via the c-Abl/Ras/ERK pathway. Sci Rep 4, 6527 (2014). https://doi.org/10.1038/srep06527
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06527
- Springer Nature Limited