Abstracts
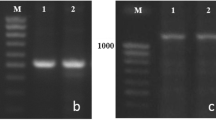
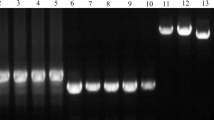
We conducted a molecular analysis of the Vrn-B1 gene in two near-isogenic lines (NILs) carrying the dominant Vrn-B1 S and Vrn-B1 Dm alleles from the Saratovskaya 29 and Diamant 2 cultivars, respectively. These lines are characterized by different times of ear emergence. PCR analysis and subsequent sequencing of the regulatory regions of Vrn-B1 revealed the full identity of the promoter region in both alleles. Simultaneously, we found significant differences in the structure of the first intron of the Vrn-B1 S allele when compared to Vrn-B1 Dm; specifically, the deletion of 0.8 kb coupled with the duplication of 0.4 kb. We suggest that these changes in intron 1 of Vrn-B1 S caused earlier ear emergence in the corresponding NIL. The unusual structure of intron 1 within the Vrn-B1 S allele was described for the first time in this study. The allele Vrn-B1 Dm was almost identical with the previously studied sequence of the Vrn-B1a allele of T. aestivum, Triple Dirk B. We designated the new Vrn-B1 S allele as Vrn-B1c. PCR analysis of the Vrn-B1 gene in 26 spring wheat cultivars of both Russian and foreign breeding revealed that 16 of them contain the Vrn-B1a allele and 6 contain the Vrn-B1c allele. Other cultivars studied contained the recessive vrn-B1 gene, except for Novosibirskaya 67. This study demonstrates that the traditional system of Vrn-1 markers does not fully encompass the allelic diversity of these genes because none of the cultivars containing the Vrn-B1c allele gave a PCR product using the previously developed set of primers for identification of the Vrn-B1 locus. We showed that the newly characterized Vrn-B1c allele is widely distributed among different genotypes of spring wheat. The findings indicate the impact of structural changes in the first intron of Vrn-1 on the vernalization response and heading time.



Similar content being viewed by others
References
Cockram J, Chiapparino E, Taylor SA, Stamati K, Donini P et al (2007) Haplotype analysis of vernalization loci in European barley germplasm reveals novel VRN-H1 alleles and a predominant winter VRN-H1/VRN-H2 multi-locus haplotype. Theor Appl Genet 115:993–1001
Distelfeld A, Li C, Dubcovsky J (2009) Regulation of flowering in temperate cereals. Curr Opin Plant Biol 12:1–7
Dubcovsky J, Lijavetzky D, Appendino L, Tranquilli G (1998) Comparative RFLP mapping of Triticum monococcum genes controlling vernalization requirement. Theor Appl Genet 97:968–975
Dubcovsky J, Loukoianov A, Fu D, Valarik M, Sanchez A, Yan L (2006) Effect of photoperiod on the regulation of wheat vernalization genes VRN-1 and VRN-2. Plant Mol Biol 60:469–480
Efremova TT, Maystrenko OI, Arbuzova VS, Laikova LI et al (2006) Effect of alien 5R(5A) chromosome substitution on ear-emergence time and winter hardiness in wheat-rye substitution lines. Euphytica 151:145–153
Efremova TT, Arbuzova VS, Leonova IN, Makhmudova K (2011) Multiple allelism in the Vrn-B1 locus of common wheat. Cereal Res Commun 39(1):12–21
Fu D, Szucs P, Yan L, Helguera M, Skinner JS, vonZitzewitz J et al (2005) Large deletions within the first intron in VRN-1 are associated with spring growth habit in barley and wheat. Mol Gen Genomics 273:54–65
Galiba G, Quarrie SA, Sutka J, Morgounov A (1995) RFLP mapping of the vernalization (Vrn-1) and frost resistance (Fr1) genes on chromosome 5A of wheat. Theor Appl Genet 90:1174–1179
Golovnina K, Kondratenko EY, Blinov AG, Goncharov NP (2010) Molecular characterization of vernalization loci VRN-1 in wild and cultivated wheats. BMC Plant Biol 10:168. doi:10.1186/1471-2229-10-168
Goncharov NP (2002) Comparative genetics of wheats and their relatives. Siberian University Publishing House (in Russian), Novosibirsk
Goncharov NP (2004) Response to vernalization in wheat: its quantitative or qualitative nature. Cereal Res Commun 32:323–330
Goncharov NP, Shitova IP (1999) The inheritance of growth habit in old local varieties and landraces of hexaploid wheat. Genetika (Russian) 35:386–392
Gorbunova V, Levy AA (1997) Non-homologous DNA end joining in plant cells is associated with deletions and filler DNA insertions. Nucleic Acids Res 25:4650–4657
Hemming MN, Peacock WJ, Dennis ES, Trevaskis B (2008) Low temperature and day length cues are integrated to regulate FLOWERING LOCUS T in barley. Plant Physiol 147:355–366
Hemming MN, Fieg S, Peacock WJ, Dennis ES, Trevaskis B (2009) Regions associated with repression of the barley (Hordeum vulgare) VERNALIZATION1 gene are not required for cold induction. Mol Genet Genomics 282:107–117
Iqbal M, Navabi A, Yang RC, Salmon DF, Spaner D (2007) Molecular characterization of vernalization response genes in Canadian spring wheat. Genome 50:511–516
Klaimi YY, Qualset CO (1974) Genetics of time of heading in wheat (Triticum aestivum L.) II. The inheritance of vernalization response. Genetics 76:119–133
Law CN, Worland AJ, Giorgi B (1975) The genetic control of ear emergence time by chromosomes 5A and 5D of wheat. Heredity 36:49–58
Lin Y, Waldman AS (2001) Promiscuous patching of broken chromosomes in mammalian cells with extrachromosomal DNA. Nucleic Acids Res 29:3975–3981
Loukoianov A, Yan L, Blechi A, Sanchez A, Dubcovsky J (2005) Regulation of VRN-1 vernalization genes in normal and transgenic polyploid wheat. Plant Physiol 138:2364–2373
Maystrenko OI (1992) Use of cytogenetic methods in studying the ontogenesis of common wheat. In: Zhakote AG (ed) Ontogenetics of higher plants. Shtiniitsa Publisher, Kishinev, pp 98–114 (In Russian)
Morgounov A, Zykin VA, Sereda GA, Urazaliev RA (2001) Siberian and North Kazakhstan wheat pool. In: Bonjean AP, Angus WJ (eds) The world wheat book, a history of wheat breeding. Lavoisier Publishing, London, pp 755–772
Oliver SN, Finnegan EJ, Dennis ES, Peacock WJ, Trevaskis B (2009) Vernalization-induced flowering in cereals is associated with changes in histone methylation at the VERNALIZATION1 gene. Proc Natl Acad Sci USA 106:8386–8391
Pidal B, Yan L, Fu D, Zhang F, Tranquilli G, Dubcovsky J (2009) The CArG-box located upstream from the transcription start of wheat vernalization gene VRN-1 is not necessary for the vernalization response. J Hered 100:355–364
Plaschke J, Ganal MW, Roeder MS (1995) Detection of genetic diversity in closely related bread wheat using microsatellite markers. Theor Appl Genet 91:1001–1007
Porter JR, Gawith M (1999) Temperatures and the growth and development of wheat: a review. Eur J Agron 10:23–26
Pugsley AT (1971) A genetic analysis of spring-winter habit of growth in wheat. Aust J Agric Res 22:21–31
Pugsley AT (1972) Additional genes inhibiting winter habit in wheat. Euphytica 21:547–552
Roberts DWA, McDonald MD (1984) Evidence for the multiplicity of alleles at Vrn1, the winter-spring habit locus in common wheat. Can J Genet Cytol (Genome) 26:191–193
Salomon S, Puchta H (1998) Capture of genomic and T-DNA sequences during double-strand break repair in somatic plant cells. EMBO J 17:6086–6095
Santra DK, Santra M, Allan RE, Campbell KG, Kidwell KK (2009) Genetic and molecular characterization of vernalization genes Vrn-A1, Vrn-B1 and Vrn-D1 in spring wheat germplasm from the pacific northwest region of the USA. Plant Breed 128:576–584
Sears ER (1966) Nullisomic-tetrasomic combinations in hexaploid wheat. In: Rilly R, Lewis KR (eds) Chromosome manipulations and plant genetics. Oliver and Boyd, Edinburgh, pp 29–45
Shumny VK, Kolchanov NA, Efimenko AV, Zakharov IK (2009) Ivan Vasilyevich Cherny. A scientist and plant breeder (23.05.1934–11.09.2009). Inf Bull VOGiS 14(4):865–874
Snape JW, Law CN, Worland AJ (1976) Chromosome variation for loci controlling ear emergence time on chromosome 5A of wheat. Heredity 37:335–340
Stelmakh AF (1993) Genetic effects of Vrn genes on heading date and agronomic traits in bread wheat. Euphytica 65:53–60
Stelmakh AF (1998) Genetic systems regulating flowering response in wheat. Euphytica 100:359–369
Thompson JD, Higgins DG, Gibson TG (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucl Acids Res 22:4673–4680
Trevaskis B, Hemming MN, Dennis ES, Peacock WJ (2007) The molecular basis of vernalization induced flowering in cereals. Trends Plant Sci 12:352–357
Tsunewaki K, Jenkins BC (1961) Monosomic and conventional gene analysis in common wheat. II. Growth habit and awnedness. Jpn J Genet 46:428–443
von Zitzewitz J, Szucs P, Dubcovsky J, Yan L, Francia E et al (2005) Molecular and structural characterization of barley vernalization genes. Plant Mol Biol 59:449–467
Wessler S, Tarpley A, Purugganan M, Spell M, Okagaki R (1990) Filler DNA is associated with spontaneous deletions in maize. Proc Natl Acad Sci USA 87:8731–8735
Yan L, Loukojanov A, Tranquillo G, Helguera M, Fahima T, Dubcovsky J (2003) Positional cloning of wheat vernalization gene VRN1. Proc Natl Acad Sci USA 100:6263–6268
Yan L, Helguera M, Kato K, Fukuyama S, Sherman J, Dubcovsky J (2004) Allelic variation at the VRN-1 promoter in polyploidy wheat. Theor Appl Genet 109:1677–1686
Yan L, Fu D, Li C, Blechl A, Tranquilli G et al (2006) The wheat and barley vernalization gene Vrn-3 is an orthologue of FT. Proc Natl Acad Sci USA 103:19581–19586
Yoshida T, Nishida H, Zhu J, Nitcher R, Distelfeld A et al (2010) Vrn-D4 is a vernalization gene located on the centromeric region of chromosome 5D in hexaploid wheat. Theor Appl Genet 120:543–552
Zhang XK, Xia XC, Xiao YG, Dubcovsky J, He ZH (2008) Allelic variation at the vernalization genes Vrn-A1, Vrn-B1, Vrn-D1 and Vrn-B3 in Chinese common wheat cultivars and their association with growth habit. Crop Sci 48:458–470
Acknowledgments
The work was supported by the Russian Foundation for Basic Research (project no. 11-04-00-178) and the “Biodiversity and Dynamics of Gene Pools” program (project no 23.28).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11032_2011_9581_MOESM1_ESM.ppt
Supplementary Fig. 1 Alignment of the promoter sequences of the Vrn-1 genes from T. aestivum genomes. The consensus forms of Vrn-A1, Vrn- B1, Vrn-D1 were constructed based on the following sequences: AY616461, AY616462, AY747601, AY616452, AY747600, AY747598, AY747599 (Vrn-A1); AY616456, AY747602, AY747603, AY747604,AY616453 (Vrn-B1); AY616454, AY616457, AY747606, AY747597, AY747605 (Vrn-D1). The base differences specific to Vrn-B1 are shown by asterisks. Pr1 and Pr2, shown by arrows, were the primers used for PCR. Numbers 1,968 indicate the first and last nucleotides in the sequence Vrn-B1 (PPT 760 kb)
11032_2011_9581_MOESM2_ESM.ppt
Supplementary Fig. 2 Hypothetical model used to explain the origin of the Vrn-B1 S (Vrn-B1c) allele from the Vrn-B1a allele during DNA replication (according Wessler et al. 1990). The region of deletion is marked by lowercase letters. The numerical positions of the deletion endpoints and the region of duplication (F) relative to the start of the first intron are shown. The F-region is boldface and underlined along with the flanking dinucleotides (A, B) and the corresponding dinucleotides at the deletion endpoints (A’, B’). Dotted lines represent newly a replicated DNA strand (PPT 148 kb)
Rights and permissions
About this article
Cite this article
Shcherban, A.B., Efremova, T.T. & Salina, E.A. Identification of a new Vrn-B1 allele using two near-isogenic wheat lines with difference in heading time. Mol Breeding 29, 675–685 (2012). https://doi.org/10.1007/s11032-011-9581-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-011-9581-y