Abstract
Context
Evidence for effects of habitat loss and fragmentation on the viability of temperate forest herb populations in agricultural landscapes is so far based on population genetic studies of single species in single landscapes. However, forest herbs differ in their life histories, and landscapes have different environments, structures and histories, making generalizations difficult.
Objectives
We compare the response of three slow-colonizing forest herbs to habitat loss and fragmentation and set this in relation to differences in life-history traits, in particular their mating system and associated pollinators.
Methods
We analysed the herbs’ landscape-scale population genetic structure based on microsatellite markers from replicate forest fragments across seven European agricultural landscapes.
Results
All species responded to reductions in population size with a decrease in allelic richness and an increase in genetic differentiation among populations. Genetic differentiation also increased with enhanced spatial isolation. In addition, each species showed unique responses. Heterozygosity in the self-compatible Oxalis acetosella was reduced in smaller populations. The genetic diversity of Anemone nemorosa, whose main pollinators are less mobile, decreased with increasing spatial isolation, but not that of the bumblebee-pollinated Polygonatum multiflorum.
Conclusions
Our study indicates that habitat loss and fragmentation compromise the long-term viability of slow-colonizing forest herbs despite their ability to persist for many decades by clonal propagation. The distinct responses of the three species studied within the same landscapes confirm the need of multi-species approaches. The mobility of associated pollinators should be considered an important determinant of forest herbs’ sensitivity to habitat loss and fragmentation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Large parts of the Earth are covered by human-modified, agricultural landscapes, in which only small remnants of (semi-)natural habitats are left (Kennedy et al. 2019). Habitat loss and fragmentation are, therefore, considered among the most important threats to biodiversity (Wilson et al. 2016). In the temperate zones of the Northern Hemisphere, agricultural landscapes are dominant (Kennedy et al. 2019). Many organisms are not adapted to live in small populations restricted to isolated habitat fragments. Typical temperate forest herbs, for instance, evolved during times when forests were much more common and connected than today (Honnay et al. 2005) and therefore exhibit traits associated with long-term stable conditions, such as a high age of first flowering, the production of few and heavy seeds and the absence of long-distance seed dispersal mechanisms (Whigham 2004). These traits limit the species’ ability to colonize new habitat patches (Verheyen et al. 2003) and their populations nowadays are often spatially isolated (Jamoneau et al. 2012; Naaf and Kolk 2015).
Small, isolated populations are increasingly prone to local extinction due to environmental, demographic and genetic stochasticity (Shaffer 1981). Theoretically, genetic diversity in small and isolated populations is eroded through random genetic drift and enhanced inbreeding (Young et al. 1996), reducing both the populations’ fitness and their potential to adapt to environmental changes (Jump et al. 2009). Empirically, however, forest herbs often appear to be able to survive in small, spatially isolated populations for many decades (Vellend et al. 2006; Valdés et al. 2015). Many forest specialist herbs are long-lived and able to propagate vegetatively, thus exhibiting long generation times. This delays fragmentation effects through genetic drift and results in extinction debts (Honnay et al. 2005). Thus, it remains unclear how susceptible temperate forest herbs are to detrimental effects of habitat loss and fragmentation in the long term.
Population genetics offers tools to answer this question by assessing both genetic diversity within and genetic connectivity among spatially isolated populations. There are also some population genetic studies for temperate forest herbs (see below). Interestingly, the majority of these focus on species relying solely on sexual reproduction, despite the fact that most temperate forest herbs are clonal (Whigham 2004). A reduced genetic diversity in smaller, more isolated forest herb populations compared to larger, better connected populations has been found in a number of case studies (Vellend 2004; Jacquemyn et al. 2006; Kolb and Durka 2013; Schlaepfer et al. 2018). However, this pattern is not ubiquitous: the genetic diversity of some self-compatible forest herbs appears to be unaffected by habitat loss and fragmentation (Culley et al. 2007; Toma et al. 2015). High levels of among-population genetic differentiation independent of among-population spatial distance (absence of isolation-by-distance) were found in some case studies (Jacquemyn et al. 2006; Schmidt et al. 2009; Zomlefer et al. 2018; Toczydlowski & Waller 2019), indicating that drift dominates over gene flow and populations are genetically isolated. But also low levels of genetic differentiation among fragmented populations have been observed, when drift was either compensated by gene flow (Tomimatsu and Ohara 2003; Jacquemyn et al. 2009) or impeded by very limited sexual reproduction (Honnay et al. 2006).
Almost all of the above-mentioned studies focus on a single species in a single landscape or region (but see Schmidt et al. 2009 for a multi-landscape study). Any generalization is therefore difficult (Segelbacher et al. 2010). One way to draw general conclusions are meta-analyses across various plant species. These have shown that the susceptibility of plant populations to negative effects of habitat loss and fragmentation varies with their life-history traits (Leimu et al. 2006; Duminil et al. 2007; Honnay and Jacquemyn 2007; Aguilar et al. 2008; Gonzalez et al. 2020). The decrease in genetic diversity in response to habitat fragmentation is more pronounced in predominantly outcrossing species than in selfing species (Leimu et al. 2006; Honnay and Jacquemyn 2007; Aguilar et al. 2008). Populations of species with gravity-dispersed seeds show generally a higher genetic differentiation than populations of species with wind- or animal-dispersed seeds (Duminil et al. 2007). The loss of genetic diversity through drift also depends on the number of generations elapsed since fragmentation took place (Young et al. 1996; Aguilar et al. 2008). We should therefore assume that long-lived clonal plants with limited seedling recruitment, such as many forest herbs, lose genetic diversity more slowly than annuals and short-lived plants (Honnay et al. 2005).
An important element of the mating strategy that has been neglected so far in meta-analyses on genetic effects of habitat fragmentation is the pollinator group associated with the species. The contribution of pollen flow to gene flow can be essential in plants (Auffret et al. 2017). Therefore, pollinator mobility may affect susceptibility to fragmentation effects (Breed et al. 2015). On the one hand, the contribution of pollen flow to total gene flow among forest herb populations is generally considered low (Honnay et al. 2005) because most pollinator species do not travel far enough to cross the agricultural matrix between spatially isolated herb populations (Wilcock and Neiland 2002). Pollinator visits are perceived to be relatively rare in forest herbs, and habitat loss and fragmentation are assumed to limit pollination service even more because small and sparse populations appear less attractive for pollinators (Kwak et al. 1998; Kolb 2008). On the other hand, some pollinator groups, including bumblebees and some solitary bees, have foraging distances of as much as several hundred meters (Gathmann and Tscharntke 2002; Redhead et al. 2016). These groups are also important pollinators for many temperate forest herbs (e.g., Phyteuma spp.: Kolb 2008, Polygonatum spp.: Hasegawa and Kudo 2005, or Stachys sylvatica: Fussell and Corbet 1991). Sensitivity to habitat loss and fragmentation in these species might therefore be lower than expected.
Moreover, the above-mentioned meta-analyses cannot account for various interactions between life-history traits and landscape context, which finally determine the regional population genetic structure (Aparicio et al. 2012). Also, accounting for time since fragmentation is difficult in meta-analyses (Schlaepfer et al. 2018). A powerful approach would therefore be to compare the population genetic structure of multiple species with contrasting traits within the same landscape (Bolliger et al. 2014). Such studies are, however, extremely rare and have so far not dealt with temperate forest herbs.
Here, we compared the population genetic structure of three slow-colonizing, clonal forest herb species: Anemone nemorosa L., Oxalis acetosella L. and Polygonatum multiflorum (L.) All. (Fig. 1). These species differ in their mating strategy, which comprises in our understanding not only the mating system s. str. (outcrossing vs. selfing), but also the associated pollinator group. To enhance the generality of our results, we compared the three species over six to seven agricultural landscapes across temperate Europe from North France to Central Sweden and Estonia. Our objective was to assess the species’ sensitivity to habitat loss and fragmentation (i.e., the degree by which genetic diversity decreases and among-population genetic differentiation increases in response to a reduced population size and an increased spatial isolation). We tested the following hypotheses: (i) All three species show general responses to habitat loss and fragmentation that are consistent across landscapes. (ii) As a regular selfing species, we expect O. acetosella to be less sensitive to habitat loss and fragmentation than the mostly outcrossing A. nemorosa and P. multiflorum. (iii) Given its pollination by far-flying bumblebees, we expect P. multiflorum to be less sensitive to habitat loss and fragmentation than A. nemorosa, which is associated with less mobile pollinators.
Methods
Studied species
The selected forest herbs (Fig. 1) are common species in European temperate deciduous forests. They have a similar life history in that they are all known as slow-colonizing forest specialists (Verheyen et al. 2003; Schmidt et al. 2014), concurrently flower in spring (Klotz et al. 2002), are pollinated by insects (Klotz et al. 2002) and propagate vegetatively, besides regular seedling recruitment (Holderegger et al. 1998; Berg 2002; Kosiński 2012). They all have a relatively broad ecological amplitude in terms of soil conditions and frequently co-occur at mesic, slightly to moderately acidic and moderately nutrient-rich sites (Leuschner and Ellenberg 2017).
However, the three species differ in their mating strategy (Table 1). While O. acetosella has a mixed mating system and produces most of its seeds from cleistogamous flowers (Berg and Redbo-Torstensson 1998), A. nemorosa and P. multiflorum are considered mainly and strictly self-incompatible, respectively (Müller et al. 2000; Kosiński 2012). The two outcrossing species differ in their associated pollinators. Anemone nemorosa is pollinated by several groups, including flies, bees, small beetles and thrips (Shirreffs 1985), however, according to standardized pollinator observations (unpublished data, see Appendix S1 in the Online Resource), the most important groups are hoverflies and solitary bees. The most important hoverfly pollinators of A. nemorosa, such as Melanostoma scalare or Syrphus torvus, show a clear preference for deciduous forest (Speight et al. 2016) and are therefore expected to cross the agricultural matrix between forest patches rarely. The maximum foraging distances of solitary bees do not exceed 600 m (Gathmann and Tscharntke 2002). In contrast, P. multiflorum is pollinated by long-tongued bumblebees (Hasegawa and Kudo 2005; Kosiński 2012), which are generally more mobile than solitary bees (Redhead et al. 2016). Our observations confirmed that Bombus pascuorum and B. pratorum with maximum forage distances of > 1800 m (Redhead et al. 2016) and > 670 m (Knight et al. 2005), respectively, are the most important pollinators of P. multiflorum (Appendix S1).
Apart from their mating strategy, the three species also differ in some other traits such as ploidy level and age of first flowering, which might affect their response to habitat loss and fragmentation (Table 1). In contrast to O. acetosella and P. multiflorum that are diploid, A. nemorosa is an aneuploid tetraploid with 2n = 30 = 4x−2 throughout Central Europe (Baumberger 1971). Hexaploid individuals have been reported from southern Sweden (Bernström 1946). Since our raw data did not show any evidence for differences in ploidy among regions and more than four allele peaks were only exceptionally observed across all regions, we treated A. nemorosa as tetraploid.
Study design and population fragmentation
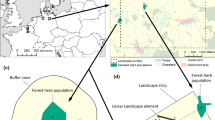
We studied the landscape-scale population genetic structure of the three plant species in seven (A. nemorosa) and six (O. acetosella and P. multiflorum) 5 × 5 km2 landscape windows, which cover a geographical gradient across temperate Europe from North France, over Belgium, West Germany, East Germany and South Sweden up to Central Sweden and Estonia (Fig. 2). Polygonatum multiflorum is fairly rare in Central Sweden and did not occur within the landscape window. Oxalis acetosella had too few occurrences in the Belgian landscape window to be included. The landscape windows have previously been extensively studied to quantify the plant species diversity in all forest fragments (Valdés et al. 2015; Vanneste et al. 2019). Therefore, the locations of all forest herb populations in these landscapes are well documented. All landscape windows represent agricultural landscapes, in which forest fragments are embedded in an agricultural matrix. Forest cover ranges between 6% and 28% (Fig. 2; Table S3).
Seven 5 × 5 km2 landscape windows used to study the regional population genetic structure of three forest herb species: a North France, b Belgium, c West Germany, d East Germany, e South Sweden, f Central Sweden and g Estonia. Landscape window locations across temperate Europe are shown in the top-left panel. Locations of the surveyed populations within each landscape window are indicated by the initial of the corresponding species name: Anemone nemorosa, Oxalis acetosella, Polygonatum multiflorum
In each landscape window, we aimed at selecting six populations for each species. For O. acetosella, only five populations were available in France and Central Sweden (Table S6). Thus, the total number of surveyed populations amounted to 42, 34 and 36 for A. nemorosa, O. acetosella and P. multiflorum, respectively. We defined a population as a spatially distinct group of shoots > 100 m apart from other shoots. Typically, populations covered the whole forest patch, but were in some cases restricted to certain parts of a forest patch if habitat conditions were heterogeneous. We selected the populations within landscape windows according to the following criteria: (a) the corresponding forest patches should be older than 200 years, as assessed from old maps (Valdés et al. 2015) to avoid potential founder effects, which can occur in young populations; (b) the populations had to represent the full range of available population sizes; (c) they should be maximally distributed across the entire landscape window; (d) the populations of all three species should preferably be in the same forest patches; and (e) more than one population within the same forest patch was only accepted if these populations were separated by > 200 m and if no other populations were available in other forest patches.
For each selected population, we estimated census population size and its degree of spatial isolation. Population size was either determined by counting all flowering shoots during the time of full flowering or, if this was not feasible, estimated as the product of population area and population density (i.e., the mean number of flowering shoots per square meter). If the population area did not correspond to the forest patch area, it was demarcated in the field by marking the outermost shoots of the population with a GPS device and afterwards measured in GIS. To estimate population density, we counted 40 flowering shoots along each of five randomly placed transects of 2 m width and measured transect length (Appendix S4). For O. acetosella, we took only the easily visible chasmogamous flowers into account, which are produced in spring.
The degree of spatial isolation was measured based on the connectivity of a population to all other populations in each landscape window (including those we did not survey) using the incidence function model derived from Hanski (1994): \(C_{i} = \sum\limits_{{j \ne i}} {A_{j}^{b} \cdot e^{{ - \alpha d_{{ij}} }} }\), where Ci is the connectivity of population i, Aj is the population size of population j, dij is the edge-to-edge distance between populations i and j, and α and b are calibration parameters. The parameter α scales the effect of distance to dispersal (i.e., with a larger α, migration between habitats becomes more difficult). We estimated 1/α for each species as the mean nearest-neighbour distance among all populations in a landscape window, averaged across landscape windows. This resulted in α values for A. nemorosa, O. acetosella and P. multiflorum of 3.65 km−1, 3.16 km−1 and 3.59 km−1, respectively. The parameter b was set to 0.5 as recommended by Moilanen and Nieminen (2002). Population size of non-surveyed populations was estimated based on data from an earlier study (Appendix S5; Valdés et al. 2015). Population areas and edge-to-edge distances among populations were calculated using ArcGIS 10.6.1 (ESRI 2017).
Sampling, DNA extraction and genotyping
From each plant population, we randomly collected leaf material from up to 20 flowering, healthy individuals spread across the population for DNA extraction. A minimum distance of 10 m between selected plants was set to avoid sampling of clones. Fewer than 20 samples per population were available in 29.5% of the populations (Tables S6 and S7.1) either due to genotyping failures or due to a very small population size. Leaf samples were dried and stored using silica gel. In total, 2,159 leaf samples were used in subsequent genetic analyses. Total genomic DNA was extracted with the innuPREP Plant DNA Kit (Analytik Jena AG, Germany). We genotyped our samples based on sets of microsatellite markers (Appendix S8) that either had been developed for congeneric species (A. nemorosa and P. multiflorum) or were newly developed for O. acetosella by AllGenetics & Biology SL (Spain) on demand. The applied marker sets comprised six, nine and six markers with a total number of 102, 61 and 149 alleles for A. nemorosa, O. acetosella and P. multiflorum, respectively (Tables S8.1-S3). The corresponding primers were multiplexed and amplified with PCR following different protocols (Appendix S8). Fragment analysis was performed on a 3730XL DNA analyzer (Applied Biosystems, USA) by Macrogen Europe (Amsterdam, Netherlands) with GeneScan 500 LIZ (Applied Biosystems) for A. nemorosa and P. multiflorum and GeneScan 350 ROX (Applied Biosystems) for O. acetosella as an internal size standard. Alleles were manually scored using GeneMapper 5 (Applied Biosystems). For a stratified random subsample of 10% from all landscapes, we repeated genotyping to estimate the multilocus genotyping error rate, which turned out to be 4.8%, 2.8% and 5.7% for A. nemorosa, O. acetosella and P. multiflorum, respectively. The number of repeated multilocus genotypes (MLG) was low in A. nemorosa (10 out of 814 samples) and P. multiflorum (44 out of 669 samples), reflecting our efforts to avoid sampling of clones. In O. acetosella, the number of repeated MLG was higher (144 out of 663), likely as a result of the lower degree of polymorphism in the applied marker set (Appendix S9). Nevertheless, the repeated MLG were randomly distributed across the regions (Appendix S9) and 519 unique MLG remained for our population genetic analyses. For further analyses, we excluded all repeated MLG.
For all three species, we tested for linkage disequilibrium using the function pair.ia in the R package poppr 2.8.3 (Kamvar et al. 2014). Although significant in some pairs of loci, the standardized index of association was always ≤ 0.12 in all three species, indicating that linkage among loci was negligible.
While the calculation of allele frequencies is straightforward in diploid species, the distribution of alleles in partially heterozygous polyploids is ambiguous (Meirmans et al. 2018). Therefore, we used the function deSilvaFreq from the R package polysat 1.7–4 (Clark and Jasieniuk 2011) to estimate allele frequencies according to De Silva et al. (2005) assuming a selfing rate of 0.06 (Müller et al. 2000) and polysomic inheritance (Stehlik and Holderegger 2000) for A. nemorosa.
Genetic diversity and differentiation
We calculated several measures of genetic diversity and differentiation. These are mostly not directly comparable among species, given their different ploidy and the varying level of polymorphism in the applied marker sets. However, the aim of this study was to compare species in terms of their relative responses to population size and connectivity (Sect. 2.5) rather than in terms of absolute values. To quantify genetic diversity within populations, we calculated allelic richness (Ar), unbiased expected heterozygosity (He), observed heterozygosity (Ho) and the inbreeding coefficient F = 1−Ho/He for each population. Allelic richness is only comparable among populations when based on the same sample size. However, nine populations had genotype numbers < 10 (Table S7.1). Therefore, rarefying on the minimum sample size would mean losing a lot of information and to down-weight rare alleles. Furthermore, low genotype numbers were not due to limited sampling effort, but very small population sizes (i.e., more genotypes could not be found in the field). As a trade-off, we therefore rarefied allelic richness based on the mean sample size across populations corresponding to 19, 15 and 17 samples in A. nemorosa, O. acetosella and P. multiflorum, respectively. Heterozygosity measures were calculated using standard formulas in the diploid species (Nei and Chesser 1983). For A. nemorosa, we calculated unbiased He using Eq. 8 in Hardy (2016) and Ho using the concept of gametic homozygosity (Moody et al. 1993). The latter is the observed homozygosity when drawing random diploid gametes from individuals, and thus comparable among ploidy levels. The probability Φ of choosing two alleles that are identical in state from a given tetraploid sample depends on the allele dosage information, which was missing. Therefore, we assembled all possible unambiguous genotypes for each phenotype and averaged Φ across these genotypes using the genotype probabilities as weights. Unambiguous genotype probabilities were calculated using code from the function meandistance.matrix2 of the polysat package (Appendix S10).
To quantify genetic differentiation among populations, we used two measures, G’’ST (Meirmans and Hedrick 2011) and DPS (Bowcock et al. 1994). While G’’ST is based on heterozygosity (like the traditional FST or GST) and is the recommended genetic differentiation measure with microsatellite markers (Meirmans and Hedrick 2011), DPS is equal to 1 minus the proportion of shared alleles and therefore facilitates an intuitive interpretation. We also tested two other commonly used differentiation measures, Dest (Jost 2008) and ρ (Ronfort et al. 1998), which were, however, highly correlated (r ≥ 0.92) with G’’ST and gave very similar results (Appendix S15). To calculate G’’ST, we used the original formula for diploid species based on the average (HS) and total expected heterozygosity (HT). For A. nemorosa, we used formulas for HS and HT analogous to Eqs. 15 and 16, respectively, in Nei and Chesser (1983), but replaced the harmonic mean of the number of samples in the populations by the harmonic mean of the number of genomes in the populations to allow for polyploidy (Clark and Jasieniuk 2011). We calculated both global statistics (G’’ST) for each landscape window (tested with permutation tests) and pairwise statistics (G’’ST and DPS) for each population pair within each landscape window. In addition, we calculated for each population a site-level differentiation value as the average pairwise differentiation of a focal population to all other populations in the landscape.
Data analysis
In a first step, we tested for differences in population size, connectivity, genetic diversity, geographic distance and genetic differentiation among species using stratified two-samples permutation tests as implemented in the R package permute 0.1 (Ottoboni 2020). Permutation was done independently within landscape windows. Next, we used linear mixed models (LMMs) to model genetic diversity and site-level genetic differentiation as a function of population size, connectivity and their interaction with species identity, using forest patch ID nested within landscape window ID as random intercept terms. Collinearity of population size and connectivity was low as indicated by a generalized variance-inflation factor < 3.0 for all predictors in all models. In order to compare specific responses to the degree of population fragmentation among species regardless of absolute differences in species-level genetic diversity and differentiation, we standardized all quantitative variables to mean = 0 and s.d. = 1 separately for each species. Prior to standardization, all variables were Box-Cox transformed (Box and Cox 1964) to make them approach normality. We fitted separate models for each genetic diversity and differentiation variable. All LMMs were fitted with the R function lme (Pinheiro et al. 2019).
Finally, we used two approaches to test for isolation-by-distance. First, we regressed pairwise genetic differentiation against geographic distance and its interaction with species using maximum-likelihood population-effects models (MLPE, Clarke et al. 2002). These models take the dependence of two pairwise distances involving a common population into account and were fitted using the function corMLPE (Pope 2020) to define a correlation structure within the lme function. Landscape window was included as a random intercept term. A positive relationship between genetic differentiation and geographic distance is often found only over a limited extent of the spatial range, usually reflecting gene dispersal distances (Slatkin 1993). Therefore, we used an autocorrelation analysis as a second approach to test for isolation-by-distance. We produced Mantel correlograms (Borcard and Legendre 2012) separately for each species, but with data from all landscape windows combined, to test, up to which distance nearby populations are genetically more similar to each other than more distant populations. For this purpose, we used the R function eco.cormantel (Roser et al. 2017) and restricted permutations to occur within landscape windows only. Whether significant spatial autocorrelation is found at a certain distance depends largely on the defined distance interval and thus the number of population pairs included in the test. Therefore, we tested a sequence of distance intervals from 600 m to 2000 m to find the interval that yielded the highest Mantel statistic rM at the first distance lag that was significant according to a permutation test with 9999 permutations. All calculations and analyses were done in R 3.6.2 (R Core Team 2019).
Results
Differences in population attributes among species
Census population size and connectivity were much higher in A. nemorosa than in P. multiflorum, with O. acetosella taking an intermediate position. The range of values covered several orders of magnitude in all three species (Fig. 3a, b; Table S7.2). Genetic diversity measures were generally higher in A. nemorosa and P. multiflorum than in O. acetosella (Fig. 3c-e), reflecting the different degree of polymorphism in the applied marker sets. Differences between expected and observed heterozygosity were highest in A. nemorosa, for which the inbreeding coefficient was highest (Fig. 3f) and significantly positive (i.e., indicating inbreeding) in 41 out of 42 populations (Fig. S11a). For O. acetosella, the F-value was significantly positive in 21 out of 34 populations (Fig. S11b), whereas for P. multiflorum, F-values were mostly negative, indicating outbreeding (significant in 10 out of 36 populations; Fig. S11c). The geographic distance among surveyed populations was similar between species (Fig. 3g) as intended by our sampling design. The degree of pairwise genetic differentiation, however, differed among species (Fig. 3h, i). In general, genetic differentiation was highest in P. multiflorum, while the ranking of A. nemorosa and O. acetosella depended on the applied differentiation measure. Global G’’ST statistics were generally significant, indicating population genetic structure, for all species and all landscapes except for A. nemorosa in France (Tables S12).
Differences in population attributes among the studied species Anemone nemorosa (An), Oxalis acetosella (Ox) and Polygonatum multiflorum (Po): a population size, b connectivity, c allelic richness (Ar), d expected heterozygosity (He), e observed heterozygosity (Ho), f inbreeding coefficient (F), g geographic distance, h and i measures of pairwise genetic differentiation (link level). Boxplots are standard boxplots. Significant differences (P ≤ 0.05 after controlling the false-discovery rate) according to pairwise stratified two-samples permutation tests (cf. Table S7.2) are indicated with lower-case letters
Effects of population size and connectivity on genetic diversity and site-level differentiation
Allelic richness increased significantly with population size in all three species (Table 2; Fig. 4a). This effect corresponded to a loss of allelic richness between the largest and smallest populations studied of 20%, 41% and 44% for A. nemorosa, O. acetosella and P. multiflorum, respectively. In O. acetosella, also expected and observed heterozygosity increased significantly with population size (Fig. 4b, c). The loss of heterozygosity between the largest and smallest population amounted to 60%. In A. nemorosa and P. multiflorum, heterozygosity was unaffected or weakly affected (only He) by population size, respectively. The inbreeding coefficient was unrelated to population size in all three species. Site-level genetic differentiation decreased with increasing population size for all species and differentiation measures (Table 2).
Partial effects of population size (a–c) and connectivity (d–f) on measures of genetic diversity and site-level differentiation as resulting from linear mixed models (cf. Table 2). All variables are scaled in standard deviation units. Shown are regression lines and 95% confidence bands (only for significant slopes). Lower-case letters denote significant differences (α = 0.1) in slopes between species. See Fig. S13 for plots with partial residuals
Connectivity only affected the genetic diversity of A. nemorosa. Specifically, allelic richness, expected heterozygosity and the inbreeding coefficient showed a positive response to connectivity (Table 2; Fig. 4d, e). The loss of allelic richness between the least and most spatially isolated population amounted to 30%. In all three species, a high connectivity reduced site-level genetic differentiation (Table 2). This effect was significantly stronger in A. nemorosa than in P. multiflorum (Fig. 4f).
In general, the amount of variation explained by population size and connectivity, as expressed by the marginal R2, was moderate to low (8–31%; Table 2). The conditional R2 was generally much higher (20–77%), indicating differences in genetic diversity and differentiation among forest patches and landscapes.
Effects of geographic distance on genetic differentiation
Geographic distance had no significant effect on pairwise genetic differentiation in MLPE models (Tables S14). However, spatial autocorrelation analysis revealed for all species that genetic differentiation at the first distance lag was significantly lower than expected by chance, indicating isolation-by-distance (Fig. 5). The lag distance yielding the highest Mantel statistic (rM) was lowest in O. acetosella (700 m), intermediate in A. nemorosa (1300–1400 m) and highest in P. multiflorum (1600 m). The rM-value at the first distance lag was highest for O. acetosella (0.20–0.21), intermediate for A. nemorosa (0.12–0.17) and lowest for P. multiflorum (0.06–0.07).
Mantel correlograms showing the spatial autocorrelation of genetic differentiation measured by the Mantel statistic rM in relation to geographic distance. Marks represent the mean distance for the corresponding distance lag. The distance intervals differ for the species and/or the genetic differentiation metrics and amount to 1400 m (G’’ST in Anemone nemorosa), 1300 m (DPS in A. nemorosa), 700 m (Oxalis acetosella) and 1600 m (Polygonatum multiflorum). A maximum of four lags was used to test for isolation-by-distance
Discussion
Our study demonstrates that clonal, slow-colonizing forest herbs are in fact sensitive to habitat loss and fragmentation. All three species responded to reductions in population size with a decrease in allelic richness and an increase in genetic differentiation among populations. Genetic differentiation also increased with enhanced spatial isolation. For the first time, we show this by using simultaneous population genetic analysis of three forest herbs across multiple landscapes. At the same time, however, our results reveal unique responses for each species, reflecting differences in life-history traits.
Differences in the inbreeding coefficient between species
While most genetic variables were not comparable among species in absolute terms, the relative difference between observed and expected heterozygosity, as expressed by the inbreeding coefficient, should be independent of allele numbers and thus comparable among species. Surprisingly, the mainly outcrossing A. nemorosa (median 0.29) rather than the regularly selfing O. acetosella (0.20) showed the highest F-values (Fig. 3f). In a study on 20 Central European populations of A. nemorosa, F-values were lower (mean 0.21) and significantly > 0 in only half of the populations (Stehlik and Holderegger 2000). Although A. nemorosa is not strictly self-incompatible, seed set is strongly reduced when open pollination is excluded (Müller et al. 2000). We think, therefore, that mostly other factors than occasional selfing caused the homozygosity excess, in particular biparental inbreeding (Stehlik and Holderegger 2000), null alleles and double reduction (i.e., the erroneous inheritance of twice the same gene copy during meiosis; Hardy 2016). In fact, the null allele frequencies in our A. nemorosa samples (inherently estimated by the function deSilvaFreq, Sect. 2.5) were relatively high, amounting to 0.237 ± 0.183 (grand mean across loci and populations ± s.d.). An observation that supports the idea that null alleles and double reduction contributed to the homozygosity excess in A. nemorosa is the unexpected positive relationship between the inbreeding index and connectivity (Table 2). It likely results from the fact that expected heterozygosity increased with connectivity, while observed heterozygosity remained low independent of connectivity. This pattern is unlikely to be caused by uni- or biparental inbreeding, which are usually enhanced in more isolated populations (Young et al. 1996).
The range of F-values in O. acetosella was large compared to A. nemorosa and values at the upper extreme exceeded those in A. nemorosa, indicating that levels of inbreeding can be higher in O. acetosella than in A. nemorosa. However, average F-values were unexpectedly low compared to other selfing forest herbs (Geum urbanum: 0.92 [Vandepitte et al. 2007], Viola grypoceras: 0.87 [Toma et al. 2015]). In their thorough study on the recruitment strategy in O. acetosella, Berg and Redbo-Torstensson (1998, 2000) found that the contribution of seedlings originating from cross-fertilized flowers to the adult plant population can be substantial, amounting to 30% on average though varying greatly between years and sites. This number might be even underestimated considering that their study was conducted near Uppsala in Sweden, which is located further north than any of our landscape windows. Thus, a lower pollinator availability might have enhanced the relative importance of selfing in their study sites (Moeller et al. 2017), while our results suggest that O. acetosella is mostly outcrossing.
In P. multiflorum, F-values close to zero were expected, given its self-incompatibility (Kosiński 2012). The tendency towards negative values (Fig. 3f; Fig. S11c) indicates that sexual reproduction occurs more often between individuals from different populations than between nearby individuals (outbreeding). A mechanism that could explain this pattern is the fact that bumblebees, on which the pollination of P. multiflorum flowers depends, prefer to fly across the open landscape rather than through dense forest (Kamm et al. 2010).
Does the mating system matter?
Allelic richness was positively related to population size in all three species (Table 2; Fig. 4a). In contrast to our second hypothesis, this relationship was not more pronounced in the mainly outcrossing species. Meta-analyses have shown a stronger positive relationship between population size and the number of alleles in self-incompatible than in self-compatible species (Leimu et al. 2006; Honnay and Jacquemyn 2007; Aguilar et al. 2008). As discussed above, the high outcrossing rate observed in the self-compatible O. acetosella explains the lack of this pattern in our results. In terms of heterozygosity, O. acetosella showed the strongest relationship with population size (Fig. 4b, c). It was the only species, in which observed heterozygosity was significantly reduced in smaller populations. Although not unexpected (Frankham 1996), a positive relationship between observed heterozygosity and population size has to our knowledge not been reported in any other study on fragmented forest herb populations. The most obvious mechanism that could lead to decreased heterozygosity in small populations is selfing or biparental inbreeding (Young et al. 1996). However, the inbreeding index was not related to population size in O. acetosella (Table 2). Genetic drift can also increase homozygosity and might occur faster in O. acetosella than in A. nemorosa and P. multiflorum given its shorter generation time. Although all three species exhibit strong clonal growth, O. acetosella relies more on sexual reproduction than the other two species. Seedling production in O. acetosella may exceed ramet production (Berg 2002). Both seedlings that have survived their first winter as well as new ramets may already flower in their first year as adults (Berg 2002). Therefore, the generation time is probably much shorter in O. acetosella than in A. nemorosa and P. multiflorum, which reach their generative phase mostly after ten or more years (Shirreffs 1985; Kosiński 2015).
In accordance with our second hypothesis, genetic diversity (Ar and He) decreased with increasing spatial isolation in the mainly outcrossing A. nemorosa, but not in the self-compatible O. acetosella (Table 2; Fig. 4d). However, given the low selfing signal in O. acetosella (and the apparent high inbreeding signal in A. nemorosa), we cannot be sure that this distinct response is due to the originally assumed difference in mating system. Furthermore, there was no effect for the other outcrossing species, P. multiflorum (Table 2). The different response between A. nemorosa and O. acetosella indicates that the inflow of new alleles is reduced in more isolated A. nemorosa populations, but not much impeded in more isolated O. acetosella populations. In plants, gene flow takes place through seed or pollen dispersal. However, both dispersal pathways do not clearly differ between the two species. Both species are strongly dispersal-limited (Table 1) and have similarly low colonization capacities (Verheyen et al. 2003). According to anecdotal knowledge (Knuth et al. 1898; Redbo-Torstensson and Berg 1995), O. acetosella is pollinated by a similar range of pollinators as A. nemorosa, including flies, beetles, thrips, bees and bumblebees. Moreover, further results of our analysis argue against a higher seed or pollen dispersal distance in O. acetosella than in A. nemorosa. First, in both species, site-level genetic differentiation increased with decreasing population size and decreasing connectivity at similar rates (Table 2; Fig. 4f). Second, the Mantel correlograms indicated that isolation-by-distance is effective over a shorter distance in O. acetosella than in A. nemorosa (Fig. 5). We therefore conclude that the answer to the question, whether and why genetic diversity is more sensitive to spatial isolation in A. nemorosa than in O. acetosella, requires further investigation. So far, the difference in mating system seems to be the best explanation despite the unexpectedly high and low inbreeding coefficients in A. nemorosa and O. acetosella, respectively.
Does the mobility of the associated pollinators matter?
The higher mobility of the pollinators associated with P. multiflorum compared to A. nemorosa is reflected in some of our results. For A. nemorosa, the spatial proximity to other populations was important to maintain high levels of genetic diversity, whereas for P. multiflorum this was not the case (Table 2; Fig. 4d). Also, the degree of genetic differentiation was stronger determined by connectivity in A. nemorosa than in P. multiflorum (Table 2; Fig. 4f). The isolation-by-distance signal was stronger in A. nemorosa (higher rM) and restricted to a lower distance compared to P. multiflorum (Fig. 5). These results are in line with our assumption that the main pollinators of A. nemorosa (i.e., hoverflies and solitary bees), either prefer deciduous forest as habitat (Appendix S1; Speight et al. 2016), or cover shorter forage distances than the bumblebee pollinators of P. multiflorum (Gathmann and Tscharntke 2002; Redhead et al. 2016). Alternatively, a higher seed dispersal potential in P. multiflorum than in A. nemorosa would explain the weaker response of P. multiflorum to increased spatial isolation. The fleshy berries of P. multiflorum appear to be destined for endozoochorous dispersal. They are, however, toxic and in fact hardly dispersed by birds or large mammals (Ehrlén and Eriksson 1993; Schaumann and Heinken 2002). Also, both plant species show a strong affinity to ancient forests, indicating their low colonization capacity (Graae 2000; Schmidt et al. 2014; Naaf and Kolk 2015). We think, therefore, that the higher mobility of the bumblebees associated with P. multiflorum is a more likely explanation for the observed patterns than a higher seed dispersal potential.
Other results were not in line with our expectations. In terms of genetic diversity, both species were similarly sensitive to habitat loss (i.e., a reduced population size; Table 2; Fig. 4a, b). This suggests that pollen-mediated gene flow between populations via bumblebees is not able to compensate for genetic erosion in small populations of P. multiflorum. This is also reflected in a relatively high level of genetic differentiation among P. multiflorum populations (Fig. 3h, i). Small populations of P. multiflorum may have difficulties to attract bumblebees and thus to sustain pollen flow. The flowering shoot density in P. multiflorum is relatively low compared to, for instance, that in A. nemorosa. Although a single P. multiflorum shoot may have more than 20 flowers, only three of these flowers are open at the same time (Kosiński 2012). In another self-incompatible forest herb that is pollinated by bumblebees, Phyteuma spicatum, seed production was significantly reduced in smaller compared to larger populations, but not in more isolated populations (Kolb 2005). Furthermore, pollen limitation in small populations of plants pollinated by bumblebees has been frequently demonstrated through differences in seed or fruit set between hand-pollinated and open-pollinated patches (Kolb 2005; Sih and Baltus 1987; Kwak et al. 1998).
The relatively weak response of genetic diversity in A. nemorosa to a reduced population size (i.e., not significantly stronger than in P. multiflorum as hypothesized) may have two reasons. First, A. nemorosa is tetraploid, which means that genetic erosion through drift happens more slowly (Meirmans et al. 2018). Second, although the main pollinator groups associated with A. nemorosa may be less mobile than those associated with P. multiflorum, they might be attracted at higher rates. Even small populations of A. nemorosa build carpets of conspicuous flowers, of which each offers pollen for approximately one week (Shirriffs 1985). Moreover, A. nemorosa offers pollen early in the spring, when few other floral resources are available.
General patterns across temperate Europe
Population size and connectivity as well as geographic distance explained only small to moderate proportions of the variation in genetic response variables. The conditional R2 was always much higher than the marginal R2 (Table 2), indicating that considerable amounts of variation in genetic response variables could be explained by the different landscapes. Although all landscapes represent agricultural landscapes of temperate Europe and in this sense were regarded as replicates in this study, we know from previous research that our landscapes differ in many respects including the macroclimate (Valdés et al. 2015), the degree of forest cover and fragmentation (Vanneste et al. 2019), the composition and configuration of the agricultural matrix (Vanneste et al. 2019), the position within the species’ distribution range (Reinecke et al. 2016) and the species’ ecological behaviour (De Frenne et al. 2009; Reinecke et al. 2016). While all these factors might directly or indirectly affect the species’ population genetic structure, analysing their influence and relative importance would require larger N and is out of scope in this study. We want to stress, however, that despite these differences among landscapes, we were able to find general patterns of species’ responses to habitat loss and fragmentation (including differences among species) that were consistent across landscapes. In this regard, our study extends and goes beyond population genetic case studies that focus on a single species in a single landscape (Segelbacher et al. 2010).
Conclusions
From our results, we draw three main conclusions. First, despite their long life spans and ability to delay fragmentation effects through clonal propagation (Honnay et al. 2005), all three forest herb species are sensitive to habitat loss and fragmentation. This will compromise their long-term viability, particularly in small and isolated populations. Second, each species has its own life-history, which led to distinct responses to habitat loss and fragmentation. Results for a single species may be difficult to assess without reference to other species with known differences in life-history traits studied within similar landscapes. Multi-species comparisons across replicate populations in multiple landscapes are thus needed to assess the generality of patterns found in independent case studies. Third, our results indicate that the mobility of associated pollinators may strongly affect forest herbs' sensitivity to habitat loss and fragmentation beyond the commonly considered effects of mating system and clonality.
Data availability
The microsatellite allele tables for all species and populations as well as population locations are available on DRYAD (https://doi.org/10.5061/dryad.tb2rbp00k).
References
Aguilar R, Quesada M, Ashworth L, Herrerias-Diego Y, Lobo J (2008) Genetic consequences of habitat fragmentation in plant populations: susceptible signals in plant traits and methodological approaches. Mol Ecol 17(24):5177–5188
Aparicio A, Hampe A, Fernandez-Carrillo L, Albaladejo RG (2012) Fragmentation and comparative genetic structure of four mediterranean woody species: complex interactions between life history traits and the landscape context. Divers Distrib 18(3):226–235
Auffret AG, Rico Y, Bullock JM, Hooftman DAP, Pakeman RJ, Soons MB, Suárez-Esteban A, Traveset A, Wagner HH, Cousins SAO (2017) Plant functional connectivity – integrating landscape structure and effective dispersal. J Ecol 105(6):1648–1656
Baumberger H (1971) Chromosomenzahlbestimmungen und Karyotypanalysen bei den Gattungen Anemone, Hepatica und Pulsatilla. Ber Schweiz Bot Ges 80:17–95
Berg H (2002) Population dynamics in Oxalis acetosella: the significance of sexual reproduction in a clonal, cleistogamous forest herb. Ecography 25(2):233–243
Berg H, Redbo-Torstensson P (1998) Cleistogamy as a bet-hedging strategy in Oxalis acetosella, a perennial herb. J Ecol 86(3):491–500
Berg H, Redbo-Torstensson P (2000) Offspring performance in Oxalis acetosella, a cleistogamous perennial herb. Plant Biol 2(6):638–645
Bernström P (1946) Chromosome numbers in Anemone nemorosa and A. ranunculoides. Hereditas 32(4):514–520
Bolliger J, Lander T, Balkenhol N (2014) Landscape genetics since 2003: status, challenges and future directions. Landsc Ecol 29(3):361–366
Borcard D, Legendre P (2012) Is the Mantel correlogram powerful enough to be useful in ecological analysis? a simulation study. Ecology 93(6):1473–1481
Bowcock AM, Ruizlinares A, Tomfohrde J, Minch E, Kidd JR, Cavallisforza LL (1994) High resolution of human evolutionary trees with polymorphic microsatellites. Nature 368(6470):455–457
Box GEP, Cox DR (1964) An analysis of transformations. J Roy Stat Soc Ser B Stat Methodol 26(2):211–252
Breed MF, Ottewell KM, Gardner MG, Marklund MHK, Dormontt EE, Lowe AJ (2015) Mating patterns and pollinator mobility are critical traits in forest fragmentation genetics. Heredity 115(2):108–114
Clark LV, Jasieniuk M (2011) polysat: an R package for polyploid microsatellite analysis. Mol Ecol Resour 11(3):562–566
Clarke RT, Rothery P, Raybould AF (2002) Confidence limits for regression relationships between distance matrices: Estimating gene flow with distance. J Agric Biol Env Stat 7(3):361–372
Culley TM, Sbita SJ, Wick A (2007) Population genetic effects of urban habitat fragmentation in the perennial herb Viola pubescens (Violaceae) using ISSR markers. Ann Bot 100(1):91–100
De Frenne P, Kolb A, Verheyen K, Brunet J, Chabrerie O, Decocq G, Diekmann M, Eriksson O, Heinken T, Hermy M, Jõgar Ü, Stanton S, Quataert P, Zindel R, Zobel M, Graae BJ (2009) Unravelling the effects of temperature, latitude and local environment on the reproduction of forest herbs. Glob Ecol Biogeogr 18(6):641–651
De Silva HN, Hall AJ, Rikkerink E, McNeilage MA, Fraser LG (2005) Estimation of allele frequencies in polyploids under certain patterns of inheritance. Heredity 95(4):327–334
Duminil J, Fineschi S, Hampe A, Jordano P, Salvini D, Vendramin GG, Petit RJ (2007) Can population genetic structure be predicted from life-history traits? Am Nat 169(5):662–672
Ehrlén J, Eriksson O (1993) Toxicity in fleshy fruits: a non-adaptive trait? Oikos 66(1):107–113
ESRI (2017) ArcGIS 10.6.1. Esri Inc., Redlands, CA, USA
Frankham R (1996) Relationship of genetic variation to population size in wildlife. Conserv Biol 10(6):1500–1508
Fussell M, Corbet SA (1991) Forage for bumble bees and honey bees in farmland: a case study. J Apicult Res 30(2):87–97
Gathmann A, Tscharntke T (2002) Foraging ranges of solitary bees. J Anim Ecol 71(5):757–764
Gonzalez AV, Gomez-Silva V, Ramirez MJ, Fonturbel FE (2020) Meta-analysis of the differential effects of habitat fragmentation and degradation on plant genetic diversity. Conserv Biol 34(3):711–720
Graae BJ (2000) The effect of landscape fragmentation and forest continuity on forest floor species in two regions of Denmark. J Veg Sci 11(6):881–892
Hanski I (1994) A practical model of metapopulation dynamics. J Anim Ecol 63(1):151–162
Hardy OJ (2016) Population genetics of autopolyploids under a mixed mating model and the estimation of selfing rate. Mol Ecol Resour 16(1):103–117
Hasegawa T, Kudo G (2005) Comparisons of growth schedule, reproductive property and allocation pattern among three rhizomatous Polygonatum species with reference to their habitat types. Plant Species Biol 20(1):23–32
Holderegger R, Stehlik I, Schneller JJ (1998) Estimation of the relative importance of sexual and vegetative reproduction in the clonal woodland herb Anemone nemorosa. Oecologia 117(1–2):105–107
Honnay O, Jacquemyn H (2007) Susceptibility of common and rare plant species to the genetic consequences of habitat fragmentation. Conserv Biol 21(3):823–831
Honnay O, Jacquemyn H, Bossuyt B, Hermy M (2005) Forest fragmentation effects on patch occupancy and population viability of herbaceous plant species. New Phytol 166(3):723–736
Honnay O, Jacquemyn H, Roldan-Ruiz I, Hermy M (2006) Consequences of prolonged clonal growth on local and regional genetic structure and fruiting success of the forest perennial Maianthemum bifolium. Oikos 112(1):21–30
Jacquemyn H, Brys R, Honnay O, Hermy M, Roldan-Ruiz I (2006) Sexual reproduction, clonal diversity and genetic differentiation in patchily distributed populations of the temperate forest herb Paris quadrifolia (Trilliaceae). Oecologia 147(3):434–444
Jacquemyn H, Brys R, Adriaens D, Honnay O, Roldan-Ruiz I (2009) Effects of population size and forest management on genetic diversity and structure of the tuberous orchid Orchis mascula. Conserv Genet 10(1):161–168
Jamoneau A, Chabrerie O, Closset-Kopp D, Decocq G (2012) Fragmentation alters beta-diversity patterns of habitat specialists within forest metacommunities. Ecography 35(2):124–133
Jost L (2008) G(ST) and its relatives do not measure differentiation. Mol Ecol 17(18):4015–4026
Jump AS, Marchant R, Peñuelas J (2009) Environmental change and the option value of genetic diversity. Trends Plant Sci 14(1):51–58
Kamm U, Gugerli F, Rotach P, Edwards P, Holderegger R (2010) Open areas in a landscape enhance pollen-mediated gene flow of a tree species: evidence from northern Switzerland. Landsc Ecol 25(6):903–911
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:14
Kennedy CM, Oakleaf JR, Theobald DM, Baruch-Mordo S, Kiesecker J (2019) Managing the middle: a shift in conservation priorities based on the global human modification gradient. Glob Change Biol 25(3):811–826
Klotz S, Kühn I, Durka W (2002) BIOLFLOR - Eine Datenbank mit biologisch-ökologischen Merkmalen zur Flora von Deutschland. Bundesamt für Naturschutz, Bonn
Knight ME, Martin AP, Bishop S, Osborne JL, Hale RJ, Sanderson RA, Goulson D (2005) An interspecific comparison of foraging range and nest density of four bumblebee (Bombus) species. Mol Ecol 14(6):1811–1820
Knuth PEOW, Appel O, Loew E, Müller H (1898) Handbuch der Blütenbiologie, vol 2. W. Engelmann, Leipzig
Kolb A (2005) Reduced reproductive success and offspring survival in fragmented populations of the forest herb Phyteuma spicatum. J Ecol 93(6):1226–1237
Kolb A (2008) Habitat fragmentation reduces plant fitness by disturbing pollination and modifying response to herbivory. Biol Conserv 141(10):2540–2549
Kolb A, Durka W (2013) Reduced genetic variation mainly affects early rather than late life-cycle stages. Biol Conserv 159:367–374
Kosiński I (2012) Generative reproduction dynamics in populations of the perennial herb Polygonatum multiflorum (Asparagaceae). Ann Bot Fenn 49(4):217–228
Kosiński I (2015) Ontogenetic development and maturity of individuals of Polygonatum multiflorum. Flora 216:1–5
Kwak MM, Velterop O, van Andel J (1998) Pollen and gene flow in fragmented habitats. Appl Veg Sci 1(1):37–54
Leimu R, Mutikainen P, Koricheva J, Fischer M (2006) How general are positive relationships between plant population size, fitness and genetic variation? J Ecol 94(5):942–952
Leuschner C, Ellenberg H (2017) Ecology of Central European forests: vegetation ecology of Central Europe. Springer International Publishing, Cham
Meirmans PG, Hedrick PW (2011) Assessing population structure: F-ST and related measures. Mol Ecol Resour 11(1):5–18
Meirmans PG, Liu S, van Tienderen PH (2018) The analysis of polyploid genetic data. J Heredity 109(3):283–296
Moeller DA, Briscoe Runquist RD, Moe AM, Geber MA, Goodwillie C, Cheptou P-O, Eckert CG, Elle E, Johnston MO, Kalisz S, Ree RH, Sargent RD, Vallejo-Marin M, Winn AA (2017) Global biogeography of mating system variation in seed plants. Ecol Lett 20(3):375–384
Moilanen A, Nieminen M (2002) Simple connectivity measures in spatial ecology. Ecology 83(4):1131–1145
Moody ME, Mueller LD, Soltis DE (1993) Genetic-variation and random drift in autotetraploid populations. Genetics 134(2):649–657
Müller N, Schneller JJ, Holderegger R (2000) Variation in breeding system among populations of the common woodland herb Anemone nemorosa (Ranunculaceae). Plant Syst Evol 221(1–2):69–76
Naaf T, Kolk J (2015) Colonization credit of post-agricultural forest patches in NE Germany remains 130–230 years after reforestation. Biol Conserv 182:155–163
Nei M, Chesser RK (1983) Estimation of fixation indexes and gene diversities. Ann Hum Genet 47:253–259
Ottoboni K (2020) permuter: permutation tests. R package version 0.1.
Pinheiro J, Bates D, DebRoy S, Sarkar D, Team RC (2019) nlme: Linear and nonlinear mixed effects models. R package version 3.1–143.
Pope N (2020) corMLPE: a correlation structure for symmetric relational data. R package version 0.0.2.
Redbo-Torstensson P, Berg H (1995) Seasonal cleistogamy - a conditional strategy to provide reproductive assurance. Acta Bot Neerl 44(3):247–256
Redhead JW, Dreier S, Bourke AFG, Heard MS, Jordan WC, Sumner S, Wang J, Carvell C (2016) Effects of habitat composition and landscape structure on worker foraging distances of five bumble bee species. Ecol Appl 26(3):726–739
Reinecke J, Wulf M, Baeten L, Brunet J, Decocq G, De Frenne P, Diekmann M, Graae BJ, Heinken T, Hermy M, Jamoneau A, Lenoir J, Plue J, Orczewska A, Van Calster H, Verheyen K, Naaf T (2016) Acido– and neutrophilic temperate forest plants display distinct shifts in ecological pH niche across north–western Europe. Ecography 39(12):1164–1175
Ronfort JL, Jenczewski E, Bataillon T, Rousset F (1998) Analysis of population structure in autotetraploid species. Genetics 150(2):921–930
Roser LG, Ferreyra LI, Saidman BO, Vilardi JC (2017) EcoGenetics: an R package for the management and exploratory analysis of spatial data in landscape genetics. Mol Ecol Resour 17(6):e241–e250
Schaumann F, Heinken T (2002) Endozoochorous seed dispersal by martens (Martes foina, M. martes) in two woodland habitats. Flora 197(5):370–378
Schlaepfer DR, Braschler B, Rusterholz HP, Baur B (2018) Genetic effects of anthropogenic habitat fragmentation on remnant animal and plant populations: a meta-analysis. Ecosphere 9(10):17
Schmidt M, Mölder A, Schönfelder E, Engel F, Schmiedel I, Culmsee H (2014) Determining ancient woodland indicator plants for practical use: a new approach developed in northwest Germany. For Ecol Manag 330:228–239
Schmidt T, Arens P, Smulders MJM, Billeter R, Liira J, Augenstein I, Durka W (2009) Effects of landscape structure on genetic diversity of Geum urbanum L. populations in agricultural landscapes. Flora 204(7):549–559
Segelbacher G, Cushman S, Epperson B, Fortin M-J, Francois O, Hardy OJ, Holderegger R, Taberlet P, Waits LP, Manel S (2010) Applications of landscape genetics in conservation biology: concepts and challenges. Conserv Genet 11(2):375–385
Shaffer ML (1981) Minimum population sizes for species conservation. Bioscience 31(2):131–134
Shirreffs DA (1985) Biological flora of the British isles – Anemone nemorosa L. J Ecol 73(3):1005–1020
Sih A, Baltus MS (1987) Patch size, pollinator behaviour, and pollinator limitation in catnip. Ecology 68(6):1679–1690
Slatkin M (1993) Isolation by distance in equilibrium and non-equilibrium populations. Evolution 47(1):264–279
Speight MCD, Castella E, Sarthou JP (2016) StN 2016. In Speight MCD, Castella E, Sarthou JP, Vanappelghem C (eds), Syrph the Net on CD, Issue 11. Syrph the Net Publications, Dublin
Stehlik I, Holderegger R (2000) Spatial genetic structure and clonal diversity of Anemone nemorosa in late successional deciduous woodlands of Central Europe. J Ecol 88(3):424–435
Team RC (2019) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Toczydlowski RH, Waller DM (2019) Drift happens: molecular genetic diversity and differentiation among populations of jewelweed (Impatiens capensis Meerb.) reflect fragmentation of floodplain forests. Mol Ecol 28(10):2459–2475
Toma Y, Imanishi J, Yokogawa M, Hashimoto H, Imanishi A, Morimoto Y, Hatanaka Y, Isagi Y, Shibata S (2015) Factors affecting the genetic diversity of a perennial herb Viola grypoceras A. Gray var. grypoceras in urban fragmented forests. Landsc Ecol 30(8):1435–1447
Tomimatsu H, Ohara M (2003) Genetic diversity and local population structure of fragmented populations of Trillium camschatcense (Trilliaceae). Biol Conserv 109(2):249–258
Valdés A, Lenoir J, Gallet-Moron E, Andrieu E, Brunet J, Chabrerie O, Closset-Kopp D, Cousins SAO, Deconchat M, De Frenne P, De Smedt P, Diekmann M, Hansen K, Hermy M, Kolb A, Liira J, Lindgren J, Naaf T, Paal T, Prokofieva I, Scherer-Lorenzen M, Wulf M, Verheyen K, Decocq G (2015) The contribution of patch-scale conditions is greater than that of macroclimate in explaining local plant diversity in fragmented forests across Europe. Glob Ecol Biogeogr 24(9):1094–1105
Vandepitte K, Jacquemyn H, Roldan-Ruiz I, Honnay O (2007) Landscape genetics of the self-compatible forest herb Geum urbanum: effects of habitat age, fragmentation and local environment. Mol Ecol 16(19):4171–4179
Vanneste T, Valdés A, Verheyen K, Perring MP, Bernhardt-Römermann M, Andrieu E, Brunet J, Cousins SAO, Deconchat M, De Smedt P, Diekmann M, Ehrmann S, Heinken T, Hermy M, Kolb A, Lenoir J, Liira J, Naaf T, Paal T, Wulf M, Decocq G, De Frenne P (2019) Functional trait variation of forest understorey plant communities across Europe. Basic Appl Ecol 34:1–14
Vellend M (2004) Parallel effects of land-use history on species diversity and genetic diversity of forest herbs. Ecology 85(11):3043–3055
Vellend M, Verheyen K, Jacquemyn H, Kolb A, Van Calster H, Peterken G, Hermy M (2006) Extinction debt of forest plants persists for more than a century following habitat fragmentation. Ecology 87(3):542–548
Verheyen K, Honnay O, Motzkin G, Hermy M, Foster DR (2003) Response of forest plant species to land-use change: a life-history trait-based approach. J Ecol 91(4):563–577
Whigham DE (2004) Ecology of woodland herbs in temperate deciduous forests. Annu Rev Ecol Evol Syst 35:583–621
Wilcock C, Neiland R (2002) Pollination failure in plants: why it happens and when it matters. Trends Plant Sci 7(6):270–277
Wilson MC, Chen X-Y, Corlett RT, Didham RK, Ding P, Holt RD, Holyoak M, Hu G, Hughes AC, Jiang L, Laurance WF, Liu J, Pimm SL, Robinson SK, Russo SE, Si X, Wilcove DS, Wu J, Yu M (2016) Habitat fragmentation and biodiversity conservation: key findings and future challenges. Landsc Ecol 31(2):219–227
Young A, Boyle T, Brown T (1996) The population genetic consequences of habitat fragmentation for plants. Trends Ecol Evol 11(10):413–418
Zomlefer WB, Comer JR, Lucardi RD, Hamrick JL, Allison JR (2018) Distribution and genetic diversity of the rare plant Veratrum woodii (Liliales: Melanthiaceae) in Georgia: A preliminary study with AFLP fingerprint data. Syst Botany 43(4):858–869
Acknowledgements
We thank Eva Kleibusch (Senckenberg German Entomological Institute, SDEI) and Ute Jahn (ZALF) for technical assistance during molecular lab work and GIS work, respectively. Basic research work for this study relied on data from the smallFOREST geodatabase (https://www.u-picardie.fr/smallforest/uk/).
Funding
Open Access funding enabled and organized by Projekt DEAL. This research was primarily funded by the German Research Foundation (DFG) (Research Grants NA 1067/2-1, HO 4742/2-1 and KR 5060/1-1). This includes the research work of JTF, KK, SH, SIJH and TN. CM, PDF and PV were supported by the European Research Council (ERC Starting Grant FORMICA no. 757833, 2018). JaanL was supported by the European Union through the European Regional Development Fund (the Centre of Excellence, EcolChange). SG received funding from the Research Foundation Flanders (FWO) (project G0H1517N). SAOC was supported by the Bolin Centre for Climate Research. TV was funded by the Special Research Fund (BOF) from Ghent University (grant number 01N02817).
Author information
Authors and Affiliations
Contributions
TN, SIJH and KK conceived and designed the study. All authors were involved in site selection, field work and sampling. JTF, SH, TN and KK performed the molecular lab work and did the allele calling. TN analyzed the data and wrote the first draft of the manuscript with contributions of JTF, SIJH, SH and KK. All authors contributed to revisions.
Corresponding author
Ethics declarations
Conflict of interest
We have no conflict of interest to declare.
Ethical approval
We have no ethical concerns to declare.
Consent to participate
All authors contributed to this study and this manuscript of their own accord.
Consent for publication
All authors agree with the content of this manuscript and its publication in Landscape Ecology.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Naaf, T., Feigs, J.T., Huang, S. et al. Sensitivity to habitat fragmentation across European landscapes in three temperate forest herbs. Landscape Ecol 36, 2831–2848 (2021). https://doi.org/10.1007/s10980-021-01292-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10980-021-01292-w