Abstract
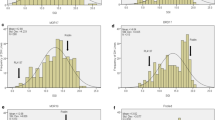
Pre-harvest sprouting (PHS) in developing wheat (Triticum aestivum L.) spikes is stimulated by cool and wet weather and leads to a decline in grain quality. A low level of harvest-time seed dormancy is a major factor for PHS, which generally is a larger problem in white-grained as compared to red-grained wheat. We have in this study analyzed seed dormancy levels at the 92nd Zadok growth stage of spike development in a doubled-haploid (DH) white wheat population and associated variation for the trait with regions on the wheat genome. The phenotypic data was generated by growing the parent lines Argent (non-dormant) and W98616 (dormant) and 151 lines of the DH population in the field during 2002 and 2003, at two locations each year, followed by assessment of harvest-time seed dormancy by germination tests. A genetic map of 2681 cM was constructed for the population upon genotyping 90 DH lines using 361 SSR, 292 AFLP, 252 DArT and 10 EST markers. Single marker analysis of the 90 genotyped lines associated regions on chromosomes 1A, 2B, 3A, 4A, 5B, 6B, and 7A with seed dormancy in at least two out of the four trials. All seven putative quantitative trait loci (QTLs) were contributed by alleles of the dormant parent, W98616. The strongest QTLs positioned on chromosomes 1A, 3A, 4A and 7A were confirmed by interval mapping and markers at these loci have potential use in marker-assisted selection of PHS resistant white-grained wheat.









Similar content being viewed by others
Abbreviations
- AFLP:
-
Amplified Fragment Length Polymorphism
- DArT:
-
Diversity Array Technology
- DH:
-
Doubled-haploid
- EST:
-
Expressed sequence tag
- PHS:
-
Pre-harvest sprouting
- QTL:
-
Quantitative trait locus
- SSR:
-
Simple sequence repeats
References
Akbari M, Wenzl P, Caig V, Carling J, Xia L, Yang S, Uszynski G, Mohler V, Lehmensiek A, Kuchel H, Hayden MJ, Howes N, Sharp P, Vaughn P, Rathmell B, Huttner E, Kilian A (2006) Diversity arrays technology (DArT) for high-throughput profiling of the hexaploid wheat genome. Theor Appl Genet 113:1409–1420
Anderson JA, Sorrells ME, Tanksley SD (1993) RFLP analysis of genomic regions associated with resistance to preharvest sprouting in wheat. Crop Sci 33:453–459
Båga M, Chodaparambil SV, Limin AE, Pecar M, Fowler DB, Chibbar RN (2007) Identification of quantitative trait loci and associated candidate genes for low-temperature tolerance in cold-hardy winter wheat. Funct Integr Genomics 7:53–68
Bailey PC, McKibbin RS, Lenton JR, Holdsworth MJ, Flintham JE, Gale MD (1999) Genetic map locations for orthologous Vp1 genes in wheat and rice. Theor Appl Genet 98:281–284
Bewley JD (1997) Seed germination and dormancy. Plant Cell 9:1055–1066
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Eujayl I, Sorrells ME, Baum M, Wolters P, Powell W (2002) Isolation of EST-derived microsatellite markers for genotyping the A and B genomes of wheat. Theor Appl Genet 104:399–407
Faris JD, Laddomada B, Gill BS (1998) Molecular mapping of segregation distortion loci in Aegilops tauschii. Genetics 149:319–327
Flintham J, Adlam R, Bassoi M, Holdsworth M, Gale M (2002) Mapping genes for resistance to sprouting damage in wheat. Euphytica 126:39–45
Groos C, Gay G, Perretant MR, Gervais L, Bernard M, Dedryver F, Charmet G (2002) Study of the relationship between pre-harvest sprouting and grain color by quantitative trait loci analysis in a white × red grain bread-wheat cross. Theor Appl Genet 104:39–47
Himi E, Noda K (2005) Red grain colour gene (R) of wheat is a Myb-type transcription factor. Euphytica 143:239–242
Himi E, Mares DJ, Yanagisawa A, Noda K (2002) Effect of grain colour gene (R) on grain dormancy and sensitivity of the embryo to abscisic acid (ABA) in wheat. J Exp Bot 53:1569–1574
Holdsworth MJ, Bentsink L, Soppe WJJ (2008) Molecular networks regulating Arabidopsis seed maturation, after-ripening, dormancy and germination. New Phytol 179:33–54
Howell KA, Narsai R, Carroll A, Ivanova A, Lohse M, Usadel B, Millar AH, Whelan J (2009) Mapping metabolic and transcript temporal switches during germination in rice highlights specific transcription factors and the role of RNA instability in the germination process. Plant Physiol 149:961–980
Hucl P, Matus-Cádiz M, Hanson K, Perron C, Tyler R (2005) Development of hard white bread wheat with sprouting resistance. ADF project report (19990102). http://www.agriculture.gov.sk.ca/Projects. pp 67
Kammholz SJ, Campbell AW, Sutherland MW, Hollamby GJ, Martin PJ, Eastwood RF, Barclay I, Wilson RE, Brennan PS, Sheppard JA (2001) Establishment and characterisation of wheat genetic mapping populations. Aust J Agric Res 52:1079–1088
Kato K, Nakamura W, Tabiki T, Miura H, Sawada S (2001) Detection of loci controlling seed dormancy on group 4 chromosomes of wheat and comparative mapping with rice and barley genomes. Theor Appl Genet 102:980–985
Knox RE, Clarke FR, Clarke JM, Fox SL (2005) Genetic analysis of pre-harvest sprouting in a durum wheat cross. Euphytica 143:261–264
Kulwal PL, Kumar N, Gaur A, Khurana P, Khurana JP, Tyagi AK, Balyan HS, Gupta PK (2005) Mapping of a major QTL for pre-harvest sprouting tolerance on chromosome 3A in bread wheat. Theor Appl Genet 111:1052–1059
Lagudah ES, Appels R, Brown AHD, McNeil D (1991) The molecular-genetic analysis of Triticum tauschii, the D-genome donor to hexaploid wheat. Genome 34:375–386
Laurie DA, Bennett MD (1988) The production of haploid wheat plants from wheat × maize crosses. Theor Appl Genet 76:393–397
Li C, Ni P, Francki M, Hunter A, Zhang Y, Schibeci D, Li H, Tarr A, Wang J, Cakir M, Yu J, Bellgard M, Lance R, Appels R (2004) Genes controlling seed dormancy and pre-harvest sprouting in a rice-wheat-barley comparison. Funct Integr Genomics 4:84–93
Lyttle TW (1991) Segregation distorters. Annu Rev Genet 25:511–557
Mares D, Mrva K, Cheong J, Williams K, Watson B, Storlie E, Sutherland M, Zou Y (2005) A QTL located on chromosome 4A associated with dormancy in white- and red-grained wheats of diverse origin. Theor Appl Genet 111:1357–1364
McKibbin RS, Wilkinson MD, Bailey PC, Flintham JE, Andrew LM, Lazzeri PA, Gale MD, Lenton JR, Holdsworth MJ (2002) Transcripts of Vp-1 homologues are misspliced in modern wheat and ancestral species. Proc Natl Acad Sci USA 99:10203–10208
Mori M, Uchino N, Chono M, Kato K, Miura H (2005) Mapping QTLs for grain dormancy on wheat chromosome 3A and the group 4 chromosomes, and their combined effect. Theor Appl Genet 110:1315–1323
Nakamura S, Komatsuda T, Miura H (2007) Mapping diploid wheat homologues of Arabidopsis seed ABA signaling genes and QTLs for seed dormancy. Theor Appl Genet 114:1129–1139
Nicot N, Chiquet V, Gandon B, Amilhat L, Legeai F, Leroy P, Bernard M, Sourdille P (2004) Study of simple sequence repeat (SSR) markers from wheat expressed sequence tags (ESTs). Theor Appl Genet 109:800–805
Noda K, Matsuura T, Maekawa M, Taketa S (2002) Chromosomes responsible for sensitivity of embryo to abscisic acid and dormancy in wheat. Euphytica 123:203–209
Ogbonnaya FC, Imtiaz M, DePauw RM (2007) Haplotype diversity of preharvest sprouting QTLs in wheat. Genome 50:107–118
Ogbonnaya FC, Imtiaz M, Ye G, Hearnden PR, Hernandez E, Eastwood RF, van Ginkel M, Shorter SC, Winchester JM (2008) Genetic and QTL analyses of seed dormancy and preharvest sprouting resistance in wheat germplasm CN10955. Theor Appl Genet 116:891–902
Ranford JC, Bryce JH, Morris PC (2002) PM19, a barley (Hordeum vulgare L.) gene encoding a putative plasma membrane protein, is expressed during embryo development and dormancy. J Exp Bot 53:147–148
Röder MS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149:2007–2023
Singh K, Ghai M, Garg M, Chhuneja P, Kaur P, Schnurbusch T, Keller B, Dhaliwal HS (2007) An integrated molecular linkage map of diploid wheat based on a Triticum boeoticum × T. monococcum RIL population. Theor Appl Genet 115:301–312
Singh R, Matus-Cádiz MA, Båga M, Hucl P, Chibbar RN (2008) Comparison of different methods for phenotyping pre-harvest sprouting in white-grained wheat. Cereal Chem 85:238–242
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Sourdille P, Tavaud M, Charmet G, Bernard M (2001) Transferability of wheat microsatellites to diploid Triticeae species carrying the A, B and D genomes. Theor Appl Genet 103:346–352
Sreenivasulu N, Usadel B, Winter A, Radchuk V, Scholz U, Stein N, Weschke W, Strickert M, Close TJ, Stitt M, Graner A, Wobus U (2008) Barley grain maturation and germination: metabolic pathway and regulatory network commonalities and differences highlighted by new MapMan/PageMan profiling tools. Plant Physiol 146:1738–1758
Tondelli A, Francia E, Barabaschi D, Aprile A, Skinner JS, Stockinger EJ, Stanca AM, Pecchioni N (2005) Mapping regulatory genes as candidates for cold and drought stress tolerance in barley. Theor Appl Genet 112:445–454
Torada A, Ikeguchi S, Koike M (2005) Mapping and validation of PCR-based markers associated with a major QTL for seed dormancy in wheat. Euphytica 143:251–255
van Ooijen JW (2004) MapQTL®5, software for the mapping of quantitative trait loci in experimental populations. Kyazma BV, Wageningen, The Netherlands
van Ooijen JW, Voorrips RE (2001) Joinmap®3.0, software for the calculation of genetic linkage maps. Plant Research International, Wageningen, The Netherlands
Vardi A (1973) Introgression between different ploidy levels in the wheat group. In: Sears ER, Sears LMS (eds) Proceedings of 4th international wheat genetics symposium. University Missouri Press, Columbia, USA, pp 131–141
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Hornes M, Friters A, Pot J, Paleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–4414
Walker-Simmons M (1987) ABA levels and sensitivity in developing wheat embryos of sprouting resistant and susceptible cultivars. Plant Physiol 84:61–66
Wilson ID, Barker GLA, Lu C, Coghill JA, Beswick RW, Lenton JR, Edwards KJ (2005) Alteration of the embryo transcriptome of hexaploid winter wheat (Triticum aestivum cv. Mercia) during maturation and germination. Funct Integr Genomics 5:144–154
Yu K, Park SJ, Poysa V (2000) Marker assisted selection of common beans for resistance to common bacterial blight: efficacy and economics. Plant Breed 119:411–415
Zadoks JC, Chang TT, Konzak CF (1974) A decimal code for the growth stages of cereals. Weed Res 14:415–421
Zanetti S, Winzeler M, Keller M, Keller B, Messmer M (2000) Genetic analysis of pre-harvest sprouting resistance in a wheat × spelt cross. Crop Sci 40:1406–1417
Acknowledgements
Canada Research Chairs Program, Canada Foundation for Innovation, Saskatchewan Innovation Fund and Saskatchewan Agriculture and Food are gratefully acknowledged for financial support for this research. R. Singh is also grateful to the Indian Council of Agricultural Research (ICAR), New Delhi, India, for granting study leave for Ph.D.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Singh, R., Matus-Cádiz, M., Båga, M. et al. Identification of genomic regions associated with seed dormancy in white-grained wheat. Euphytica 174, 391–408 (2010). https://doi.org/10.1007/s10681-010-0137-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-010-0137-8