Abstract
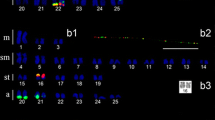
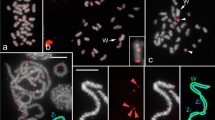
We carried out a global survey of all major types of transposable elements in Silene latifolia, a model species with sex chromosomes that are in the early stages of their evolution. A shotgun genomic library was screened with genomic DNA to isolate and characterize the most abundant elements. We found that the most common types of elements were the subtelomeric tandem repeat X-43.1 and Gypsy retrotransposons, followed by Copia retrotransposons and LINE non-LTR elements. SINE elements and DNA transposons were less abundant. We also amplified transposable elements with degenerate primers and used them to screen the library. The localization of elements by FISH revealed that most of the Copia elements were accumulated on the Y chromosome. Surprisingly, one type of Gypsy element, which was similar to Ogre elements known from legumes, was almost absent on the Y chromosome but otherwise uniformly distributed on all chromosomes. Other types of elements were ubiquitous on all chromosomes. Moreover, we isolated and characterized two new tandem repeats. One of them, STAR-C, was localized at the centromeres of all chromosomes except the Y chromosome, where it was present on the p-arm. Its variant, STAR-Y, carrying a small deletion, was specifically localized on the q-arm of the Y chromosome. The second tandem repeat, TR1, co-localized with the 45S rDNA cluster in the subtelomeres of five pairs of autosomes. FISH analysis of other Silene species revealed that some elements (e.g., Ogre-like elements) are confined to the section Elisanthe while others (e.g. Copia or Athila-like elements) are present also in more distant species. Similarly, the centromeric satellite STAR-C was conserved in the genus Silene whereas the subtelomeric satellite X-43.1 was specific for Elisanthe section. Altogether, our data provide an overview of the repetitive sequences in Silene latifolia and revealed that genomic distribution and evolutionary dynamics differ among various repetitive elements. The unique pattern of repeat distribution is found on the Y chromosome, where some elements are accumulated while other elements are conspicuously absent, which probably reflects different forces shaping the Y chromosome.
Similar content being viewed by others
References
Altschul SF, Madden TL, Schaffer AA, et al. (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25: 3389–3402.
Anzai T, Takahashi H, Fujiwara H (2001) Elimination of active Tad elements during the sexual phase of the Neurospora crassa life cycle. Fungal Genet Biol 33: 49–57.
Benson G (1999) Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res 27: 573–580.
Brodie R, Roper RL, Upton C (2004). JDotter: a Java interface to multiple dotplots generated by dotter. Bioinformatics 20: 279–281.
Buzek J, Koutnikova H, Houben A, et al. (1997) Isolation and characterization of X chromosome-derived DNA sequences from a dioecious plant Melandrium album. Chromosome Res 5: 57–65.
Charlesworth B, Charlesworth D (2000) The degeneration of Y chromosomes. Phil Trans R Soc Lond B Biol Sci 355: 1563–1572.
Charlesworth B, Sniegowski P, Stephan W (1994) The evolutionary dynamics of repetitive DNA in eukaryotes. Nature 371: 215–220.
De Keukeleire P, De Schepper S, Gielis J, Gerats T (2004) A PCR-based assay to detect hAT-like transposon sequences in plants. Chromosome Res 12: 117–123.
Dellaporta SL, Wood J, Hicks JB (1983) A plant DNA minipreparation: version II. Plant Mol Biol Rep 1: 19–21.
Downs JA, Jackson SP (1999) Involvement of DNA end-binding protein Ku in Ty element retrotransposition. Mol Cell Biol 19: 6260–6268.
Erlandsson R, Wilson JF, Paabo S (2000) Sex chromosomal transposable element accumulation and male-driven substitutional evolution in humans. Mol Biol Evol 17: 804–812.
Fawcett JA, Kawahara T, Watanabe H, Yasui Y (2006) A SINE family widely distributed in the plant kingdom and its evolutionary history. Plant Mol Biol 61: 505–514.
Feschotte C, Wessler SR (2002) Mariner-like transposases are widespread and diverse in flowering plants. Proc Natl Acad Sci U S A 99: 280–285.
Flavell JA, Dunbar E, Anderson R, Pearce SR, Hartley R, Kumar A (1992) Ty1-copia group retrotransposons are ubiquitous and heterogeneous in higher plants. Nucleic Acids Res 20: 3639–3644.
Friesen N, Brandes A, Heslop-Harrison JS (2001) Diversity, origin, and distribution of retrotransposons (gypsy and copia) in conifers. Mol Biol Evol 18: 1176–1188.
Garrido-Ramos MA, de la Herran R, Ruiz Rejon M, Ruiz Rejon C (1999) A subtelomeric satellite DNA family isolated from the genome of the dioecious plant Silene latifolia. Genome 42: 442–446.
Hobza R, Lengerova M, Cernohorska H, Rubes J, Vyskot B (2004) FAST-FISH with laser beam microdissected DOP-PCR probe distinguishes the sex chromosomes of Silene latifolia. Chromosome Res 12: 245–250.
Hobza R, Lengerova M, Svoboda J, Kubekova H, Kejnovsky E, Vyskot B (2006) An accumulation of tandem DNA repeats on the Y chromosome in Silene latifolia during early stages of sex chromosome evolution. Chromosoma 115: 376–382.
Hobza R, Kejnovsky E, Vyskot B, Widmer A (2007) The role of chromosomal rearrangements in the evolution of Silene latifolia sex chromosomes. Mol Genet Genomics 278: 633–638.
Jurka J, Kapitonov VV, Pavlicek A, Klonowski P, Kohany O, Walichiewicz J (2005) Repbase Update, a database of eukaryotic repetitive elements. Cytogenet Genome Res 110: 462–467.
Kapitonov VV, Jurka J (2005) RAG1 core and V(D)J recombination signal sequences were derived from Transib transposons. PLoS Biology 3: 998–1010.
Kazama Y, Sugiyama R, Matsunaga S et al. (2003) Organization of the KpnI family of chromosomal distal-end satellite DNAs in Silene latifolia. J Plant Res 116: 317–326.
Kazama Y, Sugiyama R, Suto Y, Uchida W, Kawano S (2006) The clustering of four subfamilies of satellite DNA at individual chromosome ends in Silene latifolia. Genome 49: 520–530.
Kejnovsky E, Kubat Z, Macas J, Hobza R, Mracek J, Vyskot B (2006a) Retand: a novel family of gypsy-like retrotransposon harboring an amplified tandem repeat. Mol Genet Genomics 276: 254–263.
Kejnovsky E, Kubat Z, Hobza R et al. (2006b) Accumulation of chloroplast DNA sequences on the Y chromosome of Silene latifolia. Genetica 128: 167–175.
Kejnovsky E, Hobza R, Kubat Z, Widmer A, Marais G, Vyskot B (2007) High intrachromosomal similarity of retrotransposon long terminal repeats: evidence for homogenization by gene conversion on plant sex chromosomes? Gene 390: 92–97.
Kidwell MG, Lisch D (2001) Perspective: transposable elements, parasitic DNA, and genome evolution. Evolution Int J Org Evolution 55: 1–24.
Kohany O, Gentles AJ, Hankus L, Jurka J (2006) Annotation, submission and screening of repetitive elements in Repbase: RepbaseSubmitter and Censor. BMC Bioinformatics 7: 474.
Kubat Z, Hobza R, Vyskot B, Kejnovsky E (2008) Microsatellite accumulation on the Y chromosome in Silene latifolia. Genome 51: 1–7.
Larkin MA, Blackshields G, Brown NP et al. (2007) ClustalW2 and ClustalX version 2. Bioinformatics 23: 2947–2948.
Lengerova M, Kejnovsky E, Hobza R, Macas J, Grant SR, Vyskot B (2004) Multicolor FISH mapping of the dioecious model plant, Silene latifolia. Theor Appl Genet 108: 1193–1199.
Lisch DR, Freeling M, Langham RJ, Choy MY (2001) Mutator transposase is widespread in the grasses. Plant Physiol 125: 1293–1303.
Liu Z, Moore PH, Ma H et al. (2004) A primitive Y chromosome in papaya marks incipient sex chromosome evolution. Nature 427: 348–352.
Macas J, Neumann P (2007) Ogre elements—a distinct group of plant Ty3/gypsy-like retrotransposons. Gene 390: 108–116.
Macas J, Pozarkova D, Navratilova A, Nouzova M, Neumann P (2000) Two new families of tandem repeats isolated from genus Vicia using genomic self-priming PCR. Mol Gen Genet 263: 741–751.
Macas J, Neumann P, Navratilova A (2007) Repetitive DNA in the pea (Pisum sativum L.) genome: comprehensive characterization using 454 sequencing and comparison to soybean and Medicago truncatula. BMC Genomics 8: 427.
Marais GAB, Nicolas M, Bergero R et al. (2008) Evidence for degeneration of the Y chromosome in the dioecious plant Silene latifolia. Current Biol 18: 1–5.
Marchler-Bauer A, Anderson JB, DeWeese-Scott C et al. (2003) CDD: a curated Entrez database of conserved domain alignments. Nucleic Acids Res 31: 383–387.
Markova M, Michu E, Vyskot B, Janousek B, Zluvova J (2007) An interspecific hybrid as a tool to study phylogenetic relationships in plants using the GISH technique. Chromosome Res 15: 1051–1059.
Matsunaga S, Kawano S, Michimoto T et al. (1999) Semi-automatic laser beam microdissection of the Y chromosome and analysis of Y chromosome DNA in a dioecious plant, Silene latifolia. Plant Cell Physiol 40: 60–68.
Matsunaga S, Yagisawa F, Yamamoto M, Uchida W, Nakao S, Kawano S (2002) LTR retrotransposons in the dioecious plant Silene latifolia. Genome 45: 745–751.
Nanda I, Kondo M, Hornung U et al. (2002) A duplicated copy of DMRT1 in the sex-determining region of the Y chromosome of the medaka, Oryzias latipes. Proc Natl Acad Sci U S A 99: 11778–11783.
Negrutiu I, Vyskot B, Barbacar N, Georgiev S, Moneger F (2001) Dioecious plants. A key to the early events of sex chromosome evolution. Plant Physiol 127: 1418–1424.
Neumann P, Pozarkova D, Macas J (2003) Highly abundant pea LTR retrotransposons Ogre is constitutively transcribed and partially spliced. Plant Mol Biol 53: 399–410.
Neumann P, Koblizkova A, Navratilova A, Macas J (2006) Significant expansion of Vicia pannonica genome size mediated by amplification of a single type of giant retroelement. Genetics 173: 1047–1056.
Nicolas M, Marais G, Hykelova V et al. (2005) A gradual process of recombination restriction in the evolutionary history of the sex chromosomes in dioecious plants. PLoS Biol. 3: 47–56.
Noma K, Ohtsubo E, Ohtsubo H (1999) Non-LTR retrotransposons (LINEs) as ubiquitous components of plant genomes. Mol Gen Genet 261: 71–79.
Okada S, Sone T, Fujisawa M et al. (2001) The Y chromosome in the liverwort Marchantia polymorpha has accumulated unique repeat sequences harboring a male-specific gene. Proc Natl Acad Sci U S A 98: 9454–9459.
Peichel CL, Ross JA, Matson CK et al. (2004) The master sex-determination locus in threespine sticklebacks is on nascent Y chromosome. Curr Biol 14: 1416–1424.
Pelissier T, Tutois S, Deragon JM, Tourmente S, Genestier S, Picard G (1995) Athila, a new retroelement from Arabidopsis thaliana. Plant Mol Biol 29: 441–452.
Presting GG, Malysheva L, Fuchs J, Schubert I (1998) A Ty3/GYPSY retrotransposon-like sequence localizes to the centromeric regions of cereal chromosomes. Plant J 16: 721–728.
Sakamoto K, Ohmido N, Fukui K, Kamada H, Satoh S (2000) Site-specific accumulation of LINE-like retrotransposon in a sex chromosome of the dioecious plant Cannabis sativa. Plant Mol Biol 44: 723–732.
Sanger F, Nicklen D, Coulson AR (1977) DNA sequencing with chain terminating inhibitors. Proc Natl Acad Sci USA 74: 5463–5467.
SanMiguel P, Tikhonov A, Jin YK et al. (1996) Nested retrotransposons in the intergenic regions of the maize genome. Science 273: 765–769.
Shibata F, Hizume M, Kuroki Y (1999) Chromosome painting of Y chromosomes and isolation of a Y chromosome-specific repetitive sequence in the dioecious plant Rumex acetosa. Chromosoma 108: 266–270.
Shibata F, Hizume M, Kuroki Y (2000) Differentiation and the polymorphic nature of the Y chromosome revealed by repetitive sequences in the dioecious plant, Rumex acetosa. Chromosome Res 8: 229–236.
Siroky J, Lysak MA, Dolezel J, Kejnovsky E, Vyskot B (2001) Heterogeneity of rDNA distribution and genome size in Silene spp. Chromosome Res 9: 387–393.
Staginnus Ch, Huettel B, Desel Ch, Schmidt T, Kahl G (2001) A PCR-based assay to detect En/Spm-like transposon sequences in plants. Chromosome Res 9: 591–605.
Steinemann S, Steinemann M (2005) Y chromosomes: born to be destroyed. BioEssays 27: 1076–1083.
Suoniemi A, Tanskanen J, Schulman AH (1998) Gypsy-like retrotransposons are widespread in the plant kingdom. Plant J 13: 699–705.
Sykorova E, Fajkus J, Mikako I, Fukui K (2001) Transition between two forms of heterochromatin at plant subtelomeres. Chromosome Res 9: 309–323.
Vershinin AV, Druka A, Alkhimova AG, Kleinhofs A, Heslop-Harrison JS (2002) LINEs and gypsy-like retrotransposons in Hordeum species. Plant Mol Biol 49: 1–14.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
Clones used for FISH experiments. (DOC 27.5 kb)
Rights and permissions
About this article
Cite this article
Cermak, T., Kubat, Z., Hobza, R. et al. Survey of repetitive sequences in Silene latifolia with respect to their distribution on sex chromosomes. Chromosome Res 16, 961–976 (2008). https://doi.org/10.1007/s10577-008-1254-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-008-1254-2