Abstract
Enantiopure alcohols have received much attention due to their widespread use as pharmaceutical intermediates. In the asymmetric biosynthesis of enantiopure alcohols, the excellent performance of carbonyl reductase makes it be the best choice as the biocatalysts. In this work, an alkali-tolerant carbonyl reductase (BsCR, encoded by yueD) from Bacillus subtilis (strain 168) was obtained through gene mining, and successfully heterologously expressed in Escherichia coli with pET-32a. BsCR showed excellent alkali resistance and even can keep more than 70% of its peak activity after incubation in Tris–HCl buffer at pH 9.0 for 40 h. The Michaelis constants and maximal velocity of the BsCR to NADPH (A) and ethyl 4-chloroacetoacetate (B) are \(K_{m}^{A}\) = 5.390 × 10−2 mmol/L, \(K_{m}^{B}\) = 1.855 mmol/L, and \(V_{max}\) = 147.3 μmol·min−1·mg−1, respectively. Applying the E. coli BL21(DE3)/pET-32a-yueD to catalyze asymmetric reduction of ethyl 4-chloroacetoacetate and acetophenone, the yield of S-CHBE reached 89.9% and S-1-phenyl ethanol reached 66.7%, and e.e. of both products reached more than 99%. This work provides a novel CR for asymmetric reduction.
Graphic Abstract
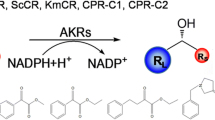
A carbonyl reductase (BsCR) and its gene were identified through gene mining, and overexpressed in Escherichia coli BL21(DE3) for whole-cell biocatalytic asymmetric reduction







Similar content being viewed by others
References
Xu P, Zheng GW, Du PX, Zong MH, Lou WY (2016) ACS Sustain Chem Eng 4:371–386
Wei P, Liang J, Cheng J, Zong MH, Lou WY (2016) Microb Cell Fact 15:5
Schmid A, Dordick JS, Hauer B, Kiener A, Wubbolts M, Witholt B (2001) Nature 409:258–268
Bian GL, Yang SW, Huang HY, Zong H, Song L (2016) Sens Actuators, B 231:129–134
Ward TJ, Ward KD (2012) Anal Chem 84:626–635
Bredikhina ZA, Kurenkov AV, Krivolapov DB, Bredikhin A (2015) Tetrahedron Asymmetry 26:577–583
Bai DY, He JY, Ouyang B, Huang J, Wang P (2017) Prog Chem 29:491–501
Zheng YG, Yin HH, Yu DF, Chen X, Tang XL, Zhang XJ, Xue YP, Wang YJ, Liu ZQ (2017) Appl Microbiol Biotechnol 101:987–1001
Itoh N (2014) Appl Microbiol Biotechnol 98:3889–3904
Malatkova P, Wsol V (2015) Drug Metab Rev 46:96–123
Shi SM, Di L (2017) Expert Opin Drug Metab Toxicol 13:859–870
Zhang RZ, Xu Y, Xiao R (2015) Biotechnol Adv 33:1671–1684
Wei P, Gao JX, Zheng GW, Wu H, Zong MH, Lou WY (2016) J Biotechnol 230:54–62
Liu ZQ, Dong SC, Yin HH, Xue YP, Tang XL, Zhang XJ, He JY, Zheng YG (2017) Bioresour Technol 229:26–32
Wang YJ, Ying BB, Shen W, Zheng RC, Zheng YG (2017) Enzyme Microb Technol 107:32–40
Wang YJ, Shen W, Luo X, Liu ZQ, Zheng YG (2017) Biotechnol Prog 33:1235–1242
Fukuda Y, Sakuraba H, Araki T, Ohshima T, Kazunari Y (2016) Enzyme Microb Technol 91:17–25
Xie Y, Xu JH, Xu Y (2010) Bioresour Technol 101:1054–1059
Ni Y, Li CX, Ma HM, Zhang J, Xu JH (2011) Appl Microbiol Biotechnol 89:1111–1118
Samuel N, Bao T, Zhang X, Yang T, Xu M, Li X, Komera I, Philibert T, Rao Z (2017) J Chem Technol Biotechnol 92:2477–2487
Sheng X, Yan M, Xu L, Wei M (2016) J Mol Catal B 130:14–18
Forlana G, Nocek B, Chakravarthy S, Joachimiak A (2017) Front Microbiol 8:1442
Cui Z, Zhang J, Fan X, Zheng G, Chang H, Wei W (2017) J Biotechnol 243:1–9
Chen X, Mei T, Cui Y, Chen Q, Liu X, Feng J, Wu Q, Zhu D (2015) ChemistryOpen 4:483–488
Luo W, Deng XX, Gong ZW, Yang ZH (2016) Asia-Pac J Chem Eng 11:2408–2411
Luo W, Deng XX, Huo J, Ruan T, Gong ZW, Yan JB, Yang ZH, Quan C, Cui ZF (2018) Catal Lett 148:1714–1722
Ni Y, Li CX, Wang LJ, Zhang J, Xu JH (2011) Org Biomol Chem 9:5463–5468
Maruyama R, Nishizawa M, Itoi Y, Ito S, Inoue M (2010) Biotechnol Bioeng 75:630–633
Qin YL, Ruan T, Hou HS, Hou YL, Yang ZH, Quan C (2019) Catal Lett 149:610–618
Cao H, Mi L, Ye Q, Zang G, Yan M, Wang Y, Zhang Y, Li X, Xu L, Xiong J, Ouyang P, Ying H (2011) Bioresour Technol 102:1733–1739
Wang YJ, Ying BB, Min C, Shen W, Liu ZQ, Zheng YG (2017) World J Microbiol Biotechnol 33:144
Wang LJ, Li CX, Ni Y, Zhang J, Liu X, Xu JH (2011) Bioresour Technol 102:7023–7028
Acknowledgments
The authors acknowledge all the financial supports for this research by the National Natural Science Foundation of China (Grant No. 21376184), Foundation from the Educational Commission of Hubei Province of China (Grant No. D20121108), the National Key Research and Development Project (2017YFF0205803, Ministry of Science and Technology of China), and the National Institute of Metrology of China (21-AKY1615).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Luo, W., Du, HJ., Bonku, E.M. et al. An Alkali-tolerant Carbonyl Reductase from Bacillus subtilis by Gene Mining: Identification and Application. Catal Lett 149, 2973–2983 (2019). https://doi.org/10.1007/s10562-019-02873-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10562-019-02873-w