Abstract
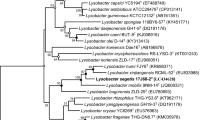
A novel proteobacterial strain designated SYSU H10001T was isolated from a soil sample collected from plateau meadow in Hongyuan county, Sichuan province, south-western China. The taxonomic position of the strain was investigated using a polyphasic approach. On the basis of 16S rRNA gene sequence similarities and phylogenetic analysis, strain SYSU H10001T was most closely related to Lysobacter soli KCTC 22011T (98.6%, sequence similarity) and Lysobacter panacisoli JCM 19212T (98.2%). The prediction result of secondary metabolites based on genome shown that the strain SYSU H10001T contained 3 clusters of bacteriocins, 1 cluster of non-ribosomal peptide synthetase, 1 cluster of type 1 polyketide synthase and 1 cluster of arylpolyene. In addition, the major isoprenoid quinone was Q-8 and the major fatty acids were identified as iso-C15:0, iso-C17:0 and Summed feature 9. The polar lipids contained diphosphatidylglycerol, phosphatidylglycerol, phosphatidylethanolamine, and three unidentified phospholipids. The genomic DNA G + C content of strain SYSU H10001T was 66.5% (genome). On the basis of phenotypic, genotypic and phylogenetic data, strain SYSU H10001T represents a novel species of the genus Lysobacter, for which the name Lysobacter prati sp. nov. is proposed. The type strain is SYSU H10001T (= KCTC 72062T = CGMCC 1.16662T).


Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Athalye M, Goodfellow M, Lacey J, White RP (1985) Numerical classification of Actinomadura and Nocardiopsis. Int J Syst Bacteriol 35:86–98
Bligh EG, Dyer WJ (1959) A rapid method of total lipid extraction and purification. Can J Biochem Physiol 37:911–917
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY, Medema MH, Weber T (2019) antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. https://doi.org/10.1093/nar/gkz310
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Chen W, Zhao YL, Cheng J, Zhou XK, Salam N, Fang BZ, Li QQ, Hozzein WN, Li WJ (2016) Lysobacter cavernae sp. nov., a novel bacterium isolated from a cave sample. Antonie Van Leeuwenhoek 109:1047–1053
Chhetri G, Kim J, Kim I, Seo T (2019) Lysobacter caseinilyticus, sp nov, a casein hydrolyzing bacterium isolated from sea water. Antonie Van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01267-7
Choi JH, Seok JH, Cha JH, Cha CJ (2014) Lysobacter panacisoli sp. nov., isolated from ginseng soil. Int J Syst Evol Microbiol 64:2193–2197
Christensen P, Cook FD (1978) Lysobacter, a new genus of nonfruiting, gliding bacteria with a high base ratio. Int J Syst Evol Microbiol 28:367–393
Collins MD, Jones D (1980) Lipids in the classification and identification of coryneform bacteria containing peptidoglycan based on 2,4-diaminobutyric acid. J Appl Bacteriol 48:459–470
Collins MD, Pirouz T, Goodfellow M, Minnikin DE (1977) Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol 100:221–230
De Bruijn I, Cheng X, de Jager V, Expósito RG, Watrous J, Patel N, Postma J, Dorrestein PC, Kobayashi D, Raaijmakers JM (2015) Comparative genomics and metabolic profiling of the genus Lysobacter. BMC Genom 16:991
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Biol 20:406–416
Fukuda W, Kimura T, Araki S, Miyoshi Y, Atomi H, Imanaka T (2013) Lysobacter oligotrophicus sp. nov., isolated from an Antarctic freshwater lake in Antarctica. Int J Syst Evol Microbiol 63:3313–3318
Gordon RE, Barnett DA, Handerhan JE, Pang CHN (1974) Nocardia coeliaca, Nocardia autotrophica, and the nocardin strain. Int J Syst Bacteriol 24:54–63
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM (2007) DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91
Harrison PG, Strulo B (2000) SPADES—a process algebra for discrete event simulation. J Logic Comput 10:3–42
Hernández I, Fernàndez C (2017) Draft genome sequence and assembly of a Lysobacter enzymogenes strain with biological control activity against root knot nematodes. Genome Announc 5:e00271–17
Hyatt D, Chen GL, Locascio PF, Land ML, Larimer FW (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11:119
Jiang Y, Li QY, Chen X, Jiang CL (2016). Isolation and cultivation methods of Actinobacteria. In: Dhanasekaran D (ed) Actinobacteria-basics and biotechnological applications, pp 39–57, (Rijeka: InTech)
Kim I, Choi J, Chhetri G, Seo T (2019) Lysobacter helvus sp. nov. and Lysobacter xanthus sp. nov., isolated from soil in South Korea. Antonie Van Leeuwenhoek 112:1253–1262
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kimura M (1985) The neutral theory of molecular evolution. Cambridge University Press, Cambridge
Kroppenstedt RM (1982) Separation of bacterial menaquinones by HPLC using reverse phase (RP18) and a silver loaded ion exchanger as stationary phases. J Liq Chromatogr 5:2359–2367
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Letunic I, Bork P (2019) Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Res 47:256–259
Li WJ, Xu P, Schumann P, Zhang YQ, Pukall R, Xu LH, Stackebrandt E, Jiang CL (2007) Georgenia ruanii sp. nov., a novel actinobacterium isolated from forest soil in Yunnan (China), and emended description of the genus Georgenia. Int J Syst Evol Microbiol 57:1424–1428
Lin SY, Hameed A, Wen CZ, Liu YC, Hsu YH, Lai WA, Young CC (2015) Lysobacter lycopersici sp. nov., isolated from tomato plant Solanum lycopersicum. Antonie Van Leeuwenhoek 107:1261–1270
Luo Y, Dong HQ, Zhou M, Huang YL, Zhang H, He WL, Sheng HM, An LZ (2019) Lysobacter psychrotolerans sp. nov., isolated from soil in the Tianshan Mountains, Xinjiang, China. Int J Syst Evol Microbiol 69:926–931
Minnikin DE, Collins MD, Goodfellow M (1979) Fatty acid and polar lipid composition in the classification of Cellulomonas, Oerskovia and related taxa. J Appl Bacteriol 47:87–95
Minnikin D, O’Donnell A, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Appl Bacteriol 2:233–241
Nie GX, Ming H, Li S, Zhou EM, Cheng J, Tang X, Feng HG, Tang SK, Li WJ (2012) Amycolatopsis dongchuanensis sp. nov., a novel actinobacterium isolated from dry-hot valley in Yunnan, south-west China. Int J Syst Evol Microbiol 62:2650–2656
Panthee S, Hamamoto H, Paudel A, Sekimizu K (2016) Lysobacter, species: a potential source of novel antibiotics. Arch Microbiol 198(9):839–845
Park JH, Kim R, Aslam Z, Jeon CO, Chung YR (2008) Lysobacter capsici sp. nov. with antimicrobial activity, isolated from the rhizosphere of pepper, and emended description of the genus Lysobacter. Int J Syst Evol Microbiol 58:387–392
Pridham TG, Gottlieb G (1948) The utilization of carbon compounds by some Actinomycetales as an aid for species determination. J Bacteriol 56:107–114
Reasoner DJ, Geldreich EE (1985) A new medium for the enumeration and subculture of bacteria from potable water. Appl Environ Microbiol 49:1–7
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106:19126–19131
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids, MIDI technical note 101. Microbial ID Inc, Newark
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340
Siddiqi MZ, Im WT (2016) Lysobacter hankyongensis sp. nov., isolated from activated sludge and Lysobacter sediminicola sp. nov., isolated from freshwater sediment. Int J Syst Evol Microbiol 66:212–218
Srinivasan S, Kim MK, Sathiyaraj G, Kim HB, Kim YJ, Yang DC (2010) Lysobacter soli sp. nov., isolated from soil of a ginseng field. Int J Syst Evol Microbiol 60:1543–1547
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
Tamaoka J, Katayama-Fujimura Y, Kuraishi H (1983) Analysis of bacterial menaquinone mixtures by high performance liquid chromatography. J Appl Bacteriol 54:31–36
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Wei DQ, Yu TT, Yao JC, Zhou EM, Song ZQ, Yin YR, Ming H, Tang SK, Li WJ (2012) Lysobacter thermophilus sp. nov., isolated from a geothermal soil sample in Tengchong, south-west China. Antonie Van Leeuwenhoek 102:643–651
Weon HY, Kim BY, Kim MK, Yoo SH, Kwon SW, Go SJ, Stackebrandt E (2007) Lysobacter niabensis sp. nov. and Lysobacter niastensis sp. nov., isolated from greenhouse soils in Korea. Int J Syst Evol Microbiol 57:548–551
Williams ST, Goodfellow M, Alderson G (1989) Genus Streptomyces Waksman and Henrici 1943, 339AL. In: Williams ST, Sharpe ME, Holt JG (eds) Bergey’s manual of systematic bacteriology, vol 4. Baltimore, Williams & Willkins, pp 2453–2492
Wu M, Scott AJ (2012) Phylogenomic analysis of bacterial and archaeal sequences with AMPHORA2. Bioinformatics 28:1033–1034
Xiao M, Zhou XK, Chen X, Duan YQ, Alkhalifah MDH, Im WT, Hozzein WN, Chen W, Li WJ (2019) Lysobacter tabacisoli sp. nov., isolated from rhizosphere soil of Nicotiana tabacum L. Int J Syst Evol Microbiol 69:1875–1880
Xie YX, Wright S, Shen YM, Du LC (2012) Bioactive natural products from Lysobacter. Nat Prod Rep 29:1277–1287
Xu P, Li WJ, Tang SK, Zhang YQ, Chen GZ, Chen HH, Xu LH, Jiang CL (2005) Naxibacter alkalitolerans gen. nov., sp. nov., a novel member of the family ‘Oxalobacteraceae’ isolated from China. Int J Syst Evol Microbiol 55:1149–1153
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017a) Introducing EzBioCloud: a taxonomically united database of 16S rRNA and whole genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Yoon SH, Ha SM, Lim JM, Kwon SJ, Chun J (2017b) A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek 110:1281–1286
Zhang XJ, Yao Q, Wang YH, Yang SZ, Feng GD, Zhu HH (2019) Lysobacter silvisoli sp. nov., isolated from forest soil. Int J Syst Evol Microbiol 69:93–98
Acknowledgements
The authors are grateful to Prof. Jung-Sook Lee (KCTC, Korea) and Prof. Takuji Kudo (JCM, Japan) for kindly providing the experimental control strains. This research was supported by the National Key R&D Program of China (2017YFD0200503), Fundamental research funds for the central universities, SYSU (No. 19lgpy173) and National Fundamental Fund Project Subsidy Funds of Personnel Training of China (No. J1310025). W-J Li was also supported by Guangdong Province Higher Vocational Colleges and Schools Pearl River Scholar Funded Scheme (2014).
Author information
Authors and Affiliations
Contributions
BZF and WJL designed research and project outline. BZF, ZXK, and LL performed isolation, deposition, and identification. YGX, JYJ and XTZ performed genome analysis. BZF, MW and WJL drafted the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All the authors have declared no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fang, BZ., Xie, YG., Zhou, XK. et al. Lysobacter prati sp. nov., isolated from a plateau meadow sample. Antonie van Leeuwenhoek 113, 763–772 (2020). https://doi.org/10.1007/s10482-020-01386-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-020-01386-6