Abstract
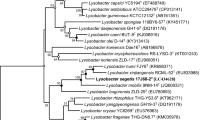
Two bacterial strains, designated D10T and U8T, were isolated from soil samples from the Dong-angyeong cave and Geommeolle wharf sea-coast, Udo-Island, Jeju, South Korea. Both novel bacterial strains are yellow-pigmented, Gram-stain negative, motile by means of monotrichous flagella, short rod shaped and strictly aerobic. A phylogenetic tree was reconstructed based on their 16S rRNA gene sequences, which indicated that these two strains belong to the genus Lysobacter within the family Xanthomonadaceae. Strain D10T showed high 16S rRNA gene sequence similarities with Lysobacter humi FJY8T (99.0%), Lysobacter xinjiangensis RCML-52T (98.9%) and Lysobacter mobilis 9NM-14T (97.2%), whereas strain U8T showed high sequence similarities to L. mobilis 9NM-14T (97.9%), L. xinjiangensis RCML-52T (97.8%), L. humi FJY8T (97.5%) and Lysobacter bugurensis ZLD-29T (97.1%). The 16S rRNA gene sequence similarity between D10T and U8T was 97.0%. Strain D10T showed low DNA–DNA relatedness to U8T (57.7 ± 3.4%), L. humi FJY8T (48.8 ± 4.3%), L. xinjiangensis RCML-52T (60.1 ± 2.4%) and L. mobilis 9NM-14T (55.9 ± 1.9%). The level of DNA–DNA relatedness for strain U8T with respect to D10T, L. mobilis 9NM-14T, L. xinjiangensis RCML-52T, L. humi FJY8T, and L. bugurensis ZLD-29T was 55.5 ± 0.5%, 54.5 ± 2.1%, 58.1 ± 0.8%, and 51.9 ± 3.4%, respectively. The major polar lipids for both strains were identified as diphosphatidylglycerol, phosphatidylethanolamine and phosphatidylglycerol. The major cellular fatty acids for both strains were identified as iso-C15:0, iso-C16:0 and summed feature 9 (iso-C17:1 ω9c/C16:0 10-methyl), and ubiquinone (Q-8) as the only isoprenoid quinone for both strains. The DNA G + C contents of the strains D10T and U8T were determined to be 70.2 mol% and 70.6 mol%. On the basis of phenotypic, genotypic, chemotaxonomic, and phylogenetic analysis, both strains D10T and U8T represent a novel species in the genus Lysobacter, for which the names Lysobacter helvus sp. nov. and Lysobacter xanthus sp. nov. are proposed, respectively. The type strain of L. helvus is D10T (= KCTC 62111T = JCM 32364T) and the type strain of L. xanthus is U8T (= KCTC 62112T = JCM 32365T).

Similar content being viewed by others
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman J (eds) (1995) Short protocols in molecular biology: a compendium of methods from current protocols in molecular biology, 3rd edn. Wiley, New York
Buck JD (1982) Nonstaining (KOH) method for determination of Gram reactions of marine bacteria. Appl Environ Microbiol 44:992–993
Christensen P, Cook FD (1978) Lysobacter, a new genus of nonfruiting, gliding bacteria with a high ratio. Int J Syst Evol Microbiol 28:367–393
Collins MD, Jones D (1981) Distribution of isoprenoid quinone structural types in bacteria and their taxonomic implication. Microbiol Rev 45:316–354
De Ley J, Cattoir H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Fautz E, Reichenbach H (1980) A simple test for flexirubin type pigments. FEMS Microbiol Lett 8:87–91
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fukuda W, Kimura T, Araki S, Miyoshi Y, Atomi H, Imanaka T (2013) Lysobacter oligotrophicus sp. nov., isolated from an Antarctic freshwater lake in Antarctica. Int J Syst Evol Microbiol 63:3313–3318
Gillis M, Ley JD, Cleene MD (1970) The determination of molecular weight of bacterial genome DNA from renaturation rates. Eur J Biochem 12:143–153
Gonzalez JM, Saiz-Jimenez C (2002) A fluorimetric method for the estimation of G + C mol% content in microorganisms by thermal denaturation temperature. Environ Microbiol 4:770–773
Hall T (1997) BioEdit. Biological sequence alignment editor for Win 95/98/NT/2 K/XP. Ibis Therapeutics, Carlsbad
Hiraishi A, Ueda Y, Ishihara J, Mori T (1996) Comparative lipoquinone analysis of influent sewage and activated sludge by high-performance liquid chromatography and photodiode array detection. J Gen Appl Microbiol 42:457–469
Jeong SE, Lee HJ, Jeon CO (2016) Lysobacter aestuarii sp. nov., isolated from estuary sediment. Int J Syst Evol Microbiol 66:1346
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Komagata K, Suzuki KI (1987) Lipid and cell-wall analysis in bacterial systematics. Methods Microbiol 19:161–205
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Kuykendall LD, Roy MA, O’Neill JJ, Devine TE (1988) Fatty acids, antibiotic resistance and deoxyribonucleic acid homology groups of Bradyrhizobium japonicum. Int J Syst Evol Microbiol 38:358–361
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Lee D, Jang JH, Cha S, Seo T (2017) Lysobacter humi sp. nov., isolated from soil. Int J Syst Evol Microbiol 67:951–955
Lin SY, Hameed A, Wen CZ, Liu YC, Hsu YH, Lai WA, Young CC (2014) Lysobacter lycopersici sp. nov., isolated from tomato plant Solanum lycopersicum. Antonie Van Leeuwenhoek 107:1261–1270
Loveland-Curtze J, Miteva VI, Brenchley JE, Vanya IM, Jean EB (2011) Evaluation of a new fluorimetric DNA–DNA hybridization method. Can J Microbiol 57:250–255
Luo Y, Dong H, Zhou M, Huang Y, Zhang H, He W, Sheng H, An L (2019) Lysobacter psychrotolerans sp. nov., isolated from soil in the Tianshan Mountains, Xinjiang. China. Int J Syst Evol Microbiol 2019:69
Margesin R, Zhang DC, Albuquerque L, Froufe HJC, Egas C, da Costa MS (2018) Lysobacter silvestris sp. nov., isolated from alpine forest soil, and reclassification of Luteimonas tolerans as Lysobacter tolerans comb. nov. Int J Syst Evol Microbiol 68:1571–1577
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Ngo HT, Won K, Du J, Son HM, Park Y, Kook M, Kim KY, Jin FX, Yi TH (2015) Lysobacter terrae sp. nov. isolated from Aglaia odorata rhizosphere soil. Int J Syst Evol Microbiol 65:587
Reichenbach H (2006) The Genus Lysobacter. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (eds) The prokaryotes. Springer, New York
Rosselló-Móra R, Trujillo ME, Sutcliffe IC (2017) Introducing a digital protologue: timely move towards a database-driven systematics of archaea and bacteria. Syst Appl Microbiol 40:121–122
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Siddiqi MZ, Im WT (2016) Lysobacter hankyongensis sp. nov., isolated from activated sludge and Lysobacter sediminicola sp. nov., isolated from fresh water sediment. Int J Syst Evol Microbiol 66:212
Singh H, Du J, Ngo HT, Won K, Yang JE, Kim KY, Yi TH (2015a) Lysobacter fragariae sp. nov. and Lysobacter rhizosphaerae sp. nov. isolated from rhizosphere of strawberry plant. Antonie Van Leeuwenhoek 107:1437–1444
Singh H, Won K, Du J, Yang J-E, Akter S, Kim K-Y, Yi T-H (2015b) Lysobacter agri sp. nov., a bacterium isolated from soil. Antonie Van Leeuwenhoek 108:553–561
Srinivasan S, Kim MK, Sathiyaraj G, Kim HB, Kim YJ, Yang DC (2010) Lysobacter soli sp. nov., isolated from soil of a ginseng field. Int J Syst Evol Microbiol 60:1543–1547
Stackebrandt E, Goebel BM (1994) Taxonomic note: a place for DNA–DNA reassociation and 16s rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Evol Microbiol 44:846–849
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O et al (1987) International Committee on Systematic Bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Evol Microbiol 37:463–464
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Xiao M, Xing KZ, Xing C, Yan QD, Dalal HMA, Wan TI, Wael NH, Wei C, Wen JL (2018) Lysobacter tabacisoli sp. nov., isolated from rhizosphere soil of Nicotiana tabacum L. Int J Syst Evol Microbiol 2018:68
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Zhang L, Bai J, Wang Y, Wu GL, Dai J, Fang CX (2011) Lysobacter korlensis sp. nov. and Lysobacter bugurensis sp. nov., isolated from soil. Int J Syst Evol Microbiol 61:2259–2265
Zhang XJ, Yao Q, Wang YH, Yang SZ, Feng GD, Zhu HH (2018) Lysobacter silvisoli sp. nov., isolated from forest soil. Int J Syst Evol Microbiol 69:93–98
Acknowledgements
This research was supported by a National Research Foundation of Korea (NRF) grant by the Korean government (MIST) (NRF-2017R1A2B4009448).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kim, I., Choi, J., Chhetri, G. et al. Lysobacter helvus sp. nov. and Lysobacter xanthus sp. nov., isolated from Soil in South Korea. Antonie van Leeuwenhoek 112, 1253–1262 (2019). https://doi.org/10.1007/s10482-019-01256-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-019-01256-w