Abstract
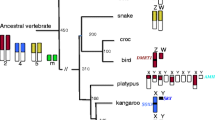
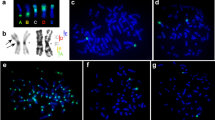
Placental (eutherian) mammals are currently classified into four superordinal clades (Afrotheria, Xenarthra, Laurasiatheria and Supraprimates) of which one, the Afrotheria (a unique lineage of African origin), is generally considered to be basal. Therefore, Afrotheria provide a pivotal evolutionary link for studying fundamental differences between the sex chromosomes of human/mouse (both representatives of Supraprimates and the index species for studies of sex chromosomes) and those of the distantly related marsupials. In this study, we use female fibroblasts to investigate classical features of X chromosome inactivation including replication timing of the X chromosomes and Barr body formation. We also examine LINE-1 accumulation on the X chromosomes of representative afrotherians and look for evidence of a pseudoautosomal region (PAR). Our results demonstrate that asynchronous replication of the X chromosomes is common to Afrotheria, as with other mammals, and Barr body formation is observed across all Placentalia, suggesting that mechanisms controlling this evolved before their radiation. Finally, we provide evidence of a PAR (which marsupials lack) and demonstrate that LINE1 is accumulated on the afrotherian and xenarthran X, although this is probably not due to transposition events in a common ancestor, but rather ongoing selection to retain recently inserted LINE1 on the X.




Similar content being viewed by others
References
Anderson LK, Reeves A, Webb LM, Ashley T (1999) Distribution of crossing over on mouse synaptonemal complexes using immunofluorescent localization of MLH1protein. Genetics 151:1569–1579
Baarends WM, Grootegoed JA (2003) Chromatin dynamics in the male meiotic prophase. Cytogenet Genome Res 103:225–234
Bailey JA, Carrel L, Chakravarti A, Eichler EE (2000) Molecular evidence for a relationship between LINE-1 elements and X chromosome inactivation: the Lyon repeat hypothesis. Proc Natl Acad Sci USA 97:6634–6639
Barr ML, Bertram G (1949) A morphological distinction between neurones of the male and female, and the behaviour of the nucleolar satellite during accelerated nucleoprotein synthesis. Nature 163:676–677
Belonogova MN, Karamysheva TV, Biltueva LS, Perepelov EA, Minina JM, Polyakov AV, Zhdanova NS, Rubtsov NB, Searle JB, Borodin PM (2006) Identification of all pachytene bivalents in the common shrew using DAPI-staining of synaptonemal complex spreads. Chromosome Res 14:673–679
Boissinot S, Entezam A, Furano AV (2001) Selection against deleterious LINE-1-containing loci in the human lineage. Mol Biol Evol 18:926–935
Brown CJ, Ballabio A, Rupert JL, Lafreniere RG, Grompe M, Tonlorenzi R, Willard HF (1991) A gene from the region of the human X inactivation centre is expressed exclusively from the inactive X chromosome. Nature 349:38–44
Carrel L, Cottle AA, Goglin KC, Willard HF (1999) A first-generation X-inactivation profile of the human X chromosome. Proc Natl Acad Sci USA 96:14440–14444
Carrel L, Park C, Tyekucheva S, Dunn J, Chiaromonte F, Makova KD (2006) Genomic environment predicts expression patterns on the human inactive X chromosome. PLoS Genet 2:e151
Carrel L, Willard HF (2005) X-inactivation profile reveals extensive variability in X-linked gene expression in females. Nature 434:400–404
Charlesworth B (1991) The evolution of sex chromosomes. Science 251:1030–1033
Dobigny G, Ozouf-Costaz C, Bonillo C, Volobouev V (2004) Viability of X-autosome translocations in mammals: an epigenomic hypothesis from a rodent case-study. Chromosoma 113:34–41
Dobson MJ, Pearlman RE, Karaiskakis A, Spyropoulos B, Moens PB (1994) Synaptonemal complex proteins: occurrence, epitope mapping and chromosome disjunction. J Cell Sci 107(Pt 10):2749–2760
Duret L, Chureau C, Samain S, Weissenbach J, Avner P (2006) The XIST RNA gene evolved in eutherians by pseudogenization of a protein-coding gene. Science 312:1653–1655
Froenicke L, Anderson KL, Wienberg J, Ashley T (2002) Male mouse recombination maps for each autosome identified by chromosome painting. Am J Hum Genet 71:1353–1368
Gartler SM, Riggs AD (1983) Mammalian X-chromosome inactivation. Annu Rev Genet 17:155–190
Gilbert CW, Muldal S, Lajtha LG, Rowley J (1962) Time-sequence of human chromosome duplication. Nature 195:869–873
Graves JAM (1967) DNA synthesis in chromosomes of cultured leucocytes from two marsupial species. Exp Cell Res 46:37–57
Graves JAM (1995) The origin and function of the mammalian Y chromosome and Y-borne genes—an evolving understanding. Bioessays 17:311–320
Grinberg MA, Sullivan MM, Benirschke K (1966) Investigation with tritiated thymidine of the relationship between the sex chromosomes, sex chromatin, and the drumstick in the cells of the female nine-banded armadillo, Dasypus novemcinctus. Cytogenetics 5:64–74
Hansen RS (2003) X inactivation-specific methylation of LINE-1 elements by DNMT3B: implications for the Lyon repeat hypothesis. Hum Mol Genet 12:2559–2567
Heard E (2005) Delving into the diversity of facultative heterochromatin: the epigenetics of the inactive X chromosome. Curr Opin Genet Dev 15:482–489
Jessberger R (2002) The many functions of SMC proteins in chromosome dynamics. Nat Rev Mol Cell Biol 3: 767–778
Johnston PG, Robinson ES (1987) X chromosome inactivation in female embryos of a marsupial mouse (Antechinus stuartii). Chromosoma 95:419–423
Kriegs JO, Churakov G, Kiefmann M, Jordan U, Brosius J, Schmitz J (2006) Retroposed elements as archives for the evolutionary history of placental mammals. PLoS Biol 4:e91
Lammers JH, Offenberg HH, van Aalderen M, Vink AC, Dietrich AJ, Heyting C (1994) The gene encoding a major component of the lateral elements of synaptonemal complexes of the rat is related to X-linked lymphocyte-regulated genes. Mol Cell Biol 14:1137–1146
Lee JT, Davidow LS, Warshawsky D (1999) Tsix, a gene antisense to XIST at the X-inactivation centre. Nat Genet 21:400–404
Lee JT, Jaenisch R (1997) Long-range cis effects of ectopic X-inactivation centres on a mouse autosome. Nature 386:275–279
Lima de Faria A, Reitalu J, Bergman S (1961) The pattern of DNA synthesis in the chromosome of man. Hereditas 47:695–704
Lynn A, Ashley T, Hassold T (2004) Variation in human meiotic recombination. Annu Rev Genomics Hum Genet 5:317–349
Lyon MF (1998) X-chromosome inactivation: a repeat hypothesis. Cytogenet Cell Genet 80:133–137
Marco E, Moens P (2003) MLH1p and MLH3p localize to precociously induced chiasmata of okadaic-acid-treated mouse spermatocytes. Genetics 165:2283–2287
McKay LM, Wrigley JM, Graves JAM (1987) Evolution of mammalian X-chromosome inactivation: sex chromatin in monotremes and marsupials. Aust J Biol Sci 40:397–404
Meuwissen RL, Offenberg HH, Dietrich AJ, Riesewijk A, van Iersel M, Heyting C (1992) A coiled-coil related protein specific for synapsed regions of meiotic prophase chromosomes. EMBO J 11:5091–5100
Mikkelsen TS, Wakefield MJ, Aken B, Amemiya CT, Chang JL, Duke S, Garber M, Gentles AJ, Goodstadt L, Heger A, Jurka J, Kamal M, Mauceli E, Searle SM, Sharpe T, Baker ML, Batzer MA, Benos PV, Belov K, Clamp M, Cook A, Cuff J, Das R, Davidow L, Deakin JE, Fazzari MJ, Glass JL, Grabherr M, Greally JM, Gu W, Hore TA, Huttley GA, Kleber M, Jirtle RL, Koina E, Lee JT, Mahony S, Marra MA, Miller RD, Nicholls RD, Oda M, Papenfuss AT, Parra ZE, Pollock DD, Ray DA, Schein JE, Speed TP, Thompson K, VandeBerg JL, Wade CM, Walker JA, Waters PD, Webber C, Weidman JR, Xie X, Zody MC; Broad Institute Genome Sequencing Platform; Broad Institute Whole Genome Assembly Team, Baldwin J, Abdouelleil A, Abdulkadir J, Abebe A, Abera B, Abreu J, Acer SC, Aftuck L, Alexander A, An P, Anderson E, Anderson S, Arachi H, Azer M, Bachantsang P, Barry A, Bayul T, Berlin A, Bessette D, Bloom T, Bloom T, Boguslavskiy L, Bonnet C, Boukhgalter B, Bourzgui I, Brown A, Cahill P, Channer S, Cheshatsang Y, Chuda L, Citroen M, Collymore A, Cooke P, Costello M, D’Aco K, Daza R, De Haan G, DeGray S, DeMaso C, Dhargay N, Dooley K, Dooley E, Doricent M, Dorje P, Dorjee K, Dupes A, Elong R, Falk J, Farina A, Faro S, Ferguson D, Fisher S, Foley CD, Franke A, Friedrich D, Gadbois L, Gearin G, Gearin CR, Giannoukos G, Goode T, Graham J, Grandbois E, Grewal S, Gyaltsen K, Hafez N, Hagos B, Hall J, Henson C, Hollinger A, Honan T, Huard MD, Hughes L, Hurhula B, Husby ME, Kamat A, Kanga B, Kashin S, Khazanovich D, Kisner P, Lance K, Lara M, Lee W, Lennon N, Letendre F, LeVine R, Lipovsky A, Liu X, Liu J, Liu S, Lokyitsang T, Lokyitsang Y, Lubonja R, Lui A, MacDonald P, Magnisalis V, Maru K, Matthews C, McCusker W, McDonough S, Mehta T, Meldrim J, Meneus L, Mihai O, Mihalev A, Mihova T, Mittelman R, Mlenga V, Montmayeur A, Mulrain L, Navidi A, Naylor J, Negash T, Nguyen T, Nguyen N, Nicol R, Norbu C, Norbu N, Novod N, O’Neill B, Osman S, Markiewicz E, Oyono OL, Patti C, Phunkhang P, Pierre F, Priest M, Raghuraman S, Rege F, Reyes R, Rise C, Rogov P, Ross K, Ryan E, Settipalli S, Shea T, Sherpa N, Shi L, Shih D, Sparrow T, Spaulding J, Stalker J, Stange-Thomann N, Stavropoulos S, Stone C, Strader C, Tesfaye S, Thomson T, Thoulutsang Y, Thoulutsang D, Topham K, Topping I, Tsamla T, Vassiliev H, Vo A, Wangchuk T, Wangdi T, Weiand M, Wilkinson J, Wilson A, Yadav S, Young G, Yu Q, Zembek L, Zhong D, Zimmer A, Zwirko Z; Broad Institute Whole Genome Assembly Team, Jaffe DB, Alvarez P, Brockman W, Butler J, Chin C, Gnerre S, MacCallum I, Graves JA, Ponting CP, Breen M, Samollow PB, Lander ES, Lindblad-Toh K (2007) Genome of the marsupial Monodelphis domestica reveals innovation in non-coding sequences. Nature 447:167–177
Morishima A, Grumbach MM, Taylor JH (1962) Asynchronous duplication of human chromosomes and the origin of sex chromatin. Proc Natl Acad Sci USA 48:756–763
Murphy WJ, Eizirik E, Johnson WE, Zhang YP, Ryder OA, O’Brien SJ (2001) Molecular phylogenetics and the origins of placental mammals. Nature 409:614–618
Murphy WJ, Pringle TH, Crider TA, Springer MS, Miller W (2007) Using genomic data to unravel the root of the placental mammal phylogeny. Genome Res 17:413–421
Nikolaev S, Montoya-Burgos JI, Margulies EH, Program NC, Rougemont J, Nyffeler B, Antonarakis SE (2007) Early history of mammals is elucidated with the ENCODE multiple species sequencing data. PLoS Genet 3:e2
Page J, Berrios S, Rufas JS, Parra MT, Sija JA, Heyting C, Fernandez-Donosoo R (2003) The meiotic pairing of X and Y chromosomes in the marsupial species Thylamys elegans is maintained by a dense plate developed from their axial elements. J Cell Sci 116:551–560
Page J, Berrios S, Parra MT, Viera A, Suja JA, Prieto I, Barbero JL, Rufas JS, Fernandez-Donoso R (2005) The program of sex chromosome pairing in meiosis is highly conserved across marsupial species: implications for sex chromosome evolution. Genetics 170:793–799
Page J, Viera A, Parra MT, Fuente RD, Suja JA, Prieto I, Barbero JL, Rufas JS, Berrios S, Fernandez-Donoso R (2006a) Involvement of synaptonemal complex proteins in sex chromosome segregation during marsupial male meiosis. PLoS Genet 2:e136
Page J, de la Fuente R, Gomez R, Calvente A, Viera A, Parra MT, Santos JL, Berrios S, Fernandez-Donoso R, Suja JA, Rufas JS (2006b) Sex chromosomes, synapsis, and cohesins: a complex affair. Chromosoma 115:250–259
Plug AW, Peters AHFM, Keegan KS, Hoekstra MF, de Boer P, Ashley T (1998) Changes in protein composition of meiotic nodules during mammalian meiosis. J Cell Sci 111:413–423
Popescu P, Hayes H, Dutrillaux B (1998) Techniques de cytogénétique animale. INRA editions, Versailles, France
Robinson TJ, Fu B, Ferguson-Smith MA, Yang F (2004) Cross-species chromosome painting in the golden mole and elephant-shrew: support for the mammalian clades Afrotheria and Afroinsectiphillia but not Afroinsectivora. Proc R Soc Lond B Biol Sci 271:1477–1484
Roig I, Liebe B, Egozcue J, Cabero L, Garcia M, Scherthan H (2004) Female-specific features of recombinational double-stranded DNA repair in relation to synapsis and telomere dynamics in human oocytes. Chromosoma 113:22–33
Ross MT, Grafham DV, Coffey AJ, Scherer S, McLay K, Muzny D, Platzer M, Howell GR, Burrows C, Bird CP, Frankish A, Lovell FL, Howe KL, Ashurst JL, Fulton RS, Sudbrak R, Wen G, Jones MC, Hurles ME, Andrews TD, Scott CE, Searle S, Ramser J, Whittaker A, Deadman R, Carter NP, Hunt SE, Chen R, Cree A, Gunaratne P, Havlak P, Hodgson A, Metzker ML, Richards S, Scott G, Steffen D, Sodergren E, Wheeler DA, Worley KC, Ainscough R, Ambrose KD, Ansari-Lari MA, Aradhya S, Ashwell RI, Babbage AK, Bagguley CL, Ballabio A, Banerjee R, Barker GE, Barlow KF, Barrett IP, Bates KN, Beare DM, Beasley H, Beasley O, Beck A, Bethel G, Blechschmidt K, Brady N, Bray-Allen S, Bridgeman AM, Brown AJ, Brown MJ, Bonnin D, Bruford EA, Buhay C, Burch P, Burford D, Burgess J, Burrill W, Burton J, Bye JM, Carder C, Carrel L, Chako J, Chapman JC, Chavez D, Chen E, Chen G, Chen Y, Chen Z, Chinault C, Ciccodicola A, Clark SY, Clarke G, Clee CM, Clegg S, Clerc-Blankenburg K, Clifford K, Cobley V, Cole CG, Conquer JS, Corby N, Connor RE, David R, Davies J, Davis C, Davis J, Delgado O, Deshazo D, Dhami P, Ding Y, Dinh H, Dodsworth S, Draper H, Dugan-Rocha S, Dunham A, Dunn M, Durbin KJ, Dutta I, Eades T, Ellwood M, Emery-Cohen A, Errington H, Evans KL, Faulkner L, Francis F, Frankland J, Fraser AE, Galgoczy P, Gilbert J, Gill R, Glockner G, Gregory SG, Gribble S, Griffiths C, Grocock R, Gu Y, Gwilliam R, Hamilton C, Hart EA, Hawes A, Heath PD, Heitmann K, Hennig S, Hernandez J, Hinzmann B, Ho S, Hoffs M, Howden PJ, Huckle EJ, Hume J, Hunt PJ, Hunt AR, Isherwood J, Jacob L, Johnson D, Jones S, de Jong PJ, Joseph SS, Keenan S, Kelly S, Kershaw JK, Khan Z, Kioschis P, Klages S, Knights AJ, Kosiura A, Kovar-Smith C, Laird GK, Langford C, Lawlor S, Leversha M, Lewis L, Liu W, Lloyd C, Lloyd DM, Loulseged H, Loveland JE, Lovell JD, Lozado R, Lu J, Lyne R, Ma J, Maheshwari M, Matthews LH, McDowall J, McLaren S, McMurray A, Meidl P, Meitinger T, Milne S, Miner G, Mistry SL, Morgan M, Morris S, Muller I, Mullikin JC, Nguyen N, Nordsiek G, Nyakatura G, O’Dell CN, Okwuonu G, Palmer S, Pandian R, Parker D, Parrish J, Pasternak S, Patel D, Pearce AV, Pearson DM, Pelan SE, Perez L, Porter KM, Ramsey Y, Reichwald K, Rhodes S, Ridler KA, Schlessinger D, Schueler MG, Sehra HK, Shaw-Smith C, Shen H, Sheridan EM, Shownkeen R, Skuce CD, Smith ML, Sotheran EC, Steingruber HE, Steward CA, Storey R, Swann RM, Swarbreck D, Tabor PE, Taudien S, Taylor T, Teague B, Thomas K, Thorpe A, Timms K, Tracey A, Trevanion S, Tromans AC, d’Urso M, Verduzco D, Villasana D, Waldron L, Wall M, Wang Q, Warren J, Warry GL, Wei X, West A, Whitehead SL, Whiteley MN, Wilkinson JE, Willey DL, Williams G, Williams L, Williamson A, Williamson H, Wilming L, Woodmansey RL, Wray PW, Yen J, Zhang J, Zhou J, Zoghbi H, Zorilla S, Buck D, Reinhardt R, Poustka A, Rosenthal A, Lehrach H, Meindl A, Minx PJ, Hillier LW, Willard HF, Wilson RK, Waterston RH, Rice CM, Vaudin M, Coulson A, Nelson DL, Weinstock G, Sulston JE, Durbin R, Hubbard T, Gibbs RA, Beck S, Rogers J, Bentley DR (2005) The DNA sequence of the human X chromosome. Nature 434:325–337
Sharman GB (1971) Late DNA replication in the paternally derived X chromosome of female kangaroos. Nature 230:231–232
Sumner AT (1972) A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res 75:304–306
Taylor JH (1960) Asynchronous duplication of chromosomes in cultured cells of Chinese hamster. J Biophys Biochem Cytol 7:455–464
Wang Z, Willard HF, Mukherjee S, Furey TS (2006) Evidence of influence of genomic DNA sequence on human X chromosome inactivation. PLoS Comput Biol 2:e113
Waters PD, Dobigny G, Pardini AT, Robinson TJ (2004) LINE-1 distribution in Afrotheria and Xenarthra: implications for understanding the evolution of LINE-1 in eutherian genomes. Chromosoma 113:137–144
Waters PD, Dobigny G, Waddell PJ, Robinson TJ (2007) Evolutionary history of LINE-1 in the major clades of placental mammals. PLoS ONE 2:e158
Yang F, Alkalaeva EZ, Perelman PL, Pardini AT, Harrison WR, O’Brien PCM, Fu B, Graphodatsky AS, Ferguson-Smith MA, Robinson TJ (2003) Reciprocal chromosome painting among human, aardvark, and elephant (superorder Afrotheria) reveals the likely eutherian ancestral karyotype. Proc Natl Acad Sci USA 100:1062–1066
Acknowledgements
SCP antibodies were kindly provided by Christa Heyting, Wageningen University, The Netherlands. P. Robles and R. Garcia from Universitat Autònoma de Barcelona are gratefully acknowledged for their technical advices. ARH is a postdoctoral fellow in the Evolutionary Genomics Group and is supported by a postdoctoral grant from the Ministry of Education and Science (MEC). Funding to TJR by a South African National Research Foundation grant is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Henikoff
Rights and permissions
About this article
Cite this article
Waters, P.D., Ruiz-Herrera, A., Dobigny, G. et al. Sex chromosomes of basal placental mammals. Chromosoma 116, 511–518 (2007). https://doi.org/10.1007/s00412-007-0116-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-007-0116-6