Abstract
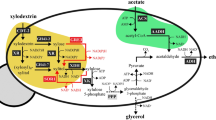
The important platform chemicals ethylene glycol and glycolic acid were produced via the oxidative D-xylose pathway in the yeast Saccharomyces cerevisiae. The expression of genes encoding D-xylose dehydrogenase (XylB) and D-xylonate dehydratase (XylD) from Caulobacter crescentus and YagE or YjhH aldolase and aldehyde dehydrogenase AldA from Escherichia coli enabled glycolic acid production from D-xylose up to 150 mg/L. In strains expressing only xylB and xylD, 29 mg/L 2-keto-3-deoxyxylonic acid [(S)-4,5-dihydroxy-2-oxopentanoic acid] (2K3DXA) was produced and D-xylonic acid accumulated to ca. 9 g/L. A significant amount of D-xylonic acid (ca. 14%) was converted to 3-deoxypentonic acid (3DPA), and also, 3,4-dihydroxybutyric acid was formed. 2K3DXA was further converted to glycolaldehyde when genes encoding by either YagE or YjhH aldolase from E. coli were expressed. Reduction of glycolaldehyde to ethylene glycol by an endogenous aldo-keto reductase activity resulted further in accumulation of ethylene glycol of 14 mg/L. The possibility of simultaneous production of lactic and glycolic acids was evaluated by expression of gene encoding lactate dehydrogenase ldhL from Lactobacillus helveticus together with aldA. Interestingly, this increased the accumulation of glycolic acid to 1 g/L. The D-xylonate dehydratase activity in yeast was notably low, possibly due to inefficient Fe–S cluster synthesis in the yeast cytosol, and leading to D-xylonic acid accumulation. The dehydratase activity was significantly improved by targeting its expression to mitochondria or by altering the Fe–S cluster metabolism of the cells with FRA2 deletion.







Similar content being viewed by others
References
Alkim C, Cam Y, Trichez D, Auriol C, Spina L, Vax A, Bartolo F, Besse P, François JM, Walther T (2015) Optimization of ethylene glycol production from (D)-xylose via a synthetic pathway implemented in Escherichia coli. Microb Cell Factories 14:127. https://doi.org/10.1186/s12934-015-0312-7
Alkim C, Trichez D, Cam Y, Spina L, François JM, Walther T (2016) The synthetic xylulose-1 phosphate pathway increases production of glycolic acid from xylose-rich sugar mixtures. Biotechnol Biofuels 9:201. https://doi.org/10.1186/s13068-016-0610-2
Andberg M, Aro-Kärkkäinen N, Carlson P, Oja M, Bozonnet S, Toivari M, Hakulinen N, O’Donohue M, Penttilä M, Koivula A (2016) Characterization and mutagenesis of two novel iron–sulphur cluster pentonate dehydratases. Appl Microbiol Biotechnol 100:7549–7563. https://doi.org/10.1007/s00253-016-7530-8
Archer RM, Royer SF, Mahy W, Winn CL, Danson MJ, Bull SD (2013) Syntheses of 2-keto-3-deoxy-D-xylonate and 2-keto-3-deoxy-L-arabinonate as stereochemical probes for demonstrating the metabolic promiscuity of sulfolobus solfataricus towards D-xylose and L-arabinose. Chem Eur J 19:2895–2902. https://doi.org/10.1002/chem.201203489
Babilas P, Knie U, Abels C (2012) Cosmetic and dermatologic use of alpha hydroxy acids. J Dtsch Dermatol Ges 10:488–491
Baudot A, Odagescu V (2004) Thermal properties of ethylene glycol aqueous solutions. Cryobiology 48:283–294. https://doi.org/10.1016/j.cryobiol.2004.02.003
Badia J, Gimenez R, Baldomá L, Barnes E, Fessner WD, Aguilar J (1991) L-lyxose metabolism employs the Lrhamnose pathway in mutant cells of Escherichia coli adapted to grow on L-lyxose. J Bacteriol 173:5144–5150
Benisch F, Boles E (2014) The bacterial Entner–Doudoroff pathway does not replace glycolysis in Saccharomyces cerevisiae due to the lack of activity of iron–sulfur cluster enzyme 6-phosphogluconate dehydratase. J Biotechnol 171:45–55. https://doi.org/10.1016/j.jbiotec.2013.11.025
Buchanan CL, Connaris H, Danson MJ, Reeve CD, Hough DW (1999) An extremely thermostable aldolase from Sulfolobus solfataricus with specificity for non-phosphorylated substrates. Biochemical J 343:563–570
Cabulong RB, Valdehuesa KNG, Ramos KRM, Nisola GM, Lee W-K, Lee CR, Chung W-J (2017) Enhanced yield of ethylene glycol production from D-xylose by pathway optimization in Escherichia coli. Enz Microb Technol 97:11–20. https://doi.org/10.1016/j.enzmictec.2016.10.020
Cam Y, Alkim C, Trichez D, Trebosc V, Vax A, Bartolo F, Besse P, François JM, Walther T (2016) Engineering of a synthetic metabolic pathway for the assimilation of (D)-xylose into value-added chemicals. ACS Synth Biol 5:607–618. https://doi.org/10.1021/acssynbio.5b00103
Cao Y, Niu W, Guo J, Xian M, Liu H (2015) Biotechnological production of 1,2,4-butanetriol: An efficient process to synthesize energetic material precursor from renewable biomass. Sci Rep 5:18149. https://doi.org/10.1038/srep18149
Carlsen S, Ajikumar PK, Formenti LR, Zhou K, Phon TH, Nielsen ML, Lantz AE, Kielland-Brandt MC, Stephanopoulos G (2013) Heterologous expression and characterization of bacterial 2-C-methyl-d-erythritol-4-phosphate pathway in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 97:5753–5769. https://doi.org/10.1007/s00253-013-4877-y
Chen R, Dou J (2016) Biofuels and bio-based chemicals from lignocellulose: metabolic engineering strategies in strain development. Biotechnol Lett 38:213–221. https://doi.org/10.1007/s10529-015-1976-0
Choi SY, Park SJ, Kim WJ, Yang JE, Lee H, Shin J, Lee SY (2016) One-step fermentative production of poly(lactate-co-glycolate) from carbohydrates in Escherichia coli. Nature Biotechnol 34:435–440. https://doi.org/10.1038/nbt.3485
Choi SY, Kim WJ, Yu SJ, Park SJ, Im SG, Lee SY (2017) Engineering the xylose-catabolizing Dahms pathway for production of poly(D-lactate-co-glycolate) and poly(D-lactate-co-glycolate-co-D-2-hydroxybutyrate) in Escherichia coli. Microb Biotechnol. https://doi.org/10.1111/1751-7915.12721
Chomvong K, Bauer S, Benjamin DI, Li X, Nomura DK, Cate JHD (2016) Bypassing the pentose phosphate pathway: towards modular utilization of xylose. PLoS One 11:e0158111. https://doi.org/10.1371/journal.pone.0158111
Christianson TW, Sikorski RS, Dante M, Shero JH, Hieter P (1992) Multifunctional yeast high-copy-number shuttle vectors. Gene 110:119–122
Dahms A (1974) 3-Deoxy-D-pentulosonic acid aldolase and its role in a new pathway of D-xylose degradation. Biochem Biophys Res Commun 60:1433–1439
Flint D, Brian JP, Ye RW (2012) Activity of Fe-S cluster requiring proteins. Patent US2012/0064561A
Frost JW, Niu W (2008) Microbial synthesis of D-1,2,4-butanetriol. Patent WO 2008/091288 A2
Gädda TM, Pirttimaa MM, Koivistoinen O, Richard P, Penttilä M, Harlin A (2013) The industrial potential of bio-based glycolic acid and polyglycolic acid. Appita Magazine 67:352
Gietz RD, Sugino A (1988) New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene 74:527–534
Harmsen P, Hackmann M (2013) Green building blocks for biobased plastics. Biobased processes and market development in “Green raw material” series. International Energy Agency (IEA) Bioenergy Task 42 on biorefinery. International Energy Agency, Paris, France, ISBN 978-94-6173-610-9
Hummel M, Leppikallio M, Heikkinen S, Niemelä K, Sixta H (2010) Acidity and lactonization of xylonic acid: a nuclear magnetic resonance study. J Carbohydr Chem 29:416–428. https://doi.org/10.1080/07328303.2011.567424
Irena/Iea-Etsap (2013) Production of bio-ethylene: Technology brief 9. IEA-ETSAP Int Renew Energy Agency, Paris, France. https://www.irena.org/DocumentDownloads/Publications/IRENA-ETSAP%20Tech%20Brief%20I13%20Production_of_Bio-ethylene.pdf
Jayakody LN, Horie K, Hayashi N, Kitagaki H (2013) Engineering redox cofactor utilization for detoxification of glycolaldehyde, a key inhibitor of bioethanol production, in yeast Saccharomyces cerevisiae. Appl Microbiol Biotechnol 97:6589–6600. https://doi.org/10.1007/s00253-013-4997-4
Johnson DC, Dean DR, Smith AD, Johnson MK (2005) Structure, function, and formation of biological iron-sulfur clusters. Annu Rev Biochem 74:247–281. https://doi.org/10.1146/annurev.biochem.74.082803.133518
Koivistoinen OM, Kuivanen J, Barth D, Turkia H, Pitkänen J-P, Penttilä M, Richard P (2013) Glycolic acid production in the engineered yeasts Saccharomyces cerevisiae and Kluyveromyces lactis. Microb Cell Factories 12:1–16. https://doi.org/10.1186/1475-2859-12-82
Kosinski AM, Brugnano JL, Seal BL, Knight FC, Panitch A (2012) Synthesis and characterization of a poly(lactic-co-glycolic acid) core + poly(N-isopropylacrylamide) shell nanoparticle system. Biomatter 2:195–201. https://doi.org/10.4161/biom.22494; 10.4161/biom.22494
Kumánovics A, Chen OS, Li L, Bagley D, Adkins EM, Lin H, Dingra NN, Outten CE, Keller G, Winge D, Ward DM, Kaplan J (2008) Identification of FRA1 and FRA2 as genes involved in regulating the yeast iron regulon in response to decreased mitochondrial iron-sulfur cluster synthesis. J Biol Chem 283:10276–10286
Lee JY, Kang CD, Lee SH, Park YK, Cho KM (2015) Engineering cellular redox balance in Saccharomyces cerevisiae for improved production of L-lactic acid. Biotechnol Bioeng 112:751–758. https://doi.org/10.1002/bit.25488
Lill R (2009) Function and biogenesis of iron–sulphur proteins. Nature 460:831–838. https://doi.org/10.1038/nature08301
Liu H, Valdehuesa KN, Nisola GM, Ramos KR, Chung WJ (2012) High yield production of D-xylonic acid from D-xylose using engineered Escherichia coli. Bioresour Technol 115:244–248. https://doi.org/10.1016/j.biortech.2011.08.065;%2010.1016/j.biortech.2011.08.065
Liu H, Ramos KRM, Valdehuesa KNG, Nisola GM, Lee W-KK, Chung W-JJ (2013) Biosynthesis of ethylene glycol in Escherichia coli. Appl Microbiol Biotechnol 97:3409–3417. https://doi.org/10.1007/s00253-012-4618-7
Meijnen J-P, de Winde HJ, Ruijssenaars HJ (2009) Establishment of oxidative D-xylose metabolism in Pseudomonas putida S12. Appl Envinron Microbiol 75:2784–2791
Miller C, Fosmer A, Rush B, McMullin T, Beacom D, Suominen P (2011) Bio-based chemicals | industrial production of lactic acid. In: Moo-Young M (ed) Comprehensive Biotechnology, 2nd edn. Elsevier, Amsterdam, pp 179–188 eBook ISBN: 9780080885049
Milne N, Wahl SA, van Maris AJA, Pronk JT, Daran JM (2016) Excessive by-product formation: a key contributor to low isobutanol yields of engineered Saccharomyces cerevisiae strains. Metab Eng Commun 3:39–51. https://doi.org/10.1016/j.meteno.2016.01.002
Niu W, Molefe MN, Frost JW (2003) Microbial synthesis of the energetic material precursor 1,2,4-butanetriol. J Am Chem Soc 125:12998–12999. https://doi.org/10.1021/ja036391+
Nygård Y, Toivari MH, Penttilä M, Ruohonen L, Wiebe MG (2011) Bioconversion of D-xylose to D-xylonate with Kluyveromyces lactis. Metab Eng 13:383–391. https://doi.org/10.1016/j.ymben.2011.04.001; 10.1016/j.ymben.2011.04.001
Nygård Y, Maaheimo H, Mojzita D, Toivari M, Wiebe M, Resnekov O, Pesce CG, Ruohonen L, Penttilä M (2014) Single cell and in vivo analyses elucidate the effect of xylC lactonase during production of D-xylonate in Saccharomyces cerevisiae. Metab Eng 25:238–247. https://doi.org/10.1016/j.ymben.2014.07.005
Partow S, Siewers V, Daviet L, Schalk M, Nielsen J (2012) Reconstruction and evaluation of the synthetic bacterial MEP pathway in Saccharomyces cerevisiae. PLoS One 7:e52498. https://doi.org/10.1371/journal.pone.0052498
Pereira B, Li Z-J, De Mey M, Lim CG, Zhang H, Hoeltgen C, Stephanopoulos G (2016) Efficient utilization of pentoses for bioproduction of the renewable two-carbon compounds ethylene glycol and glycolate. Metab Eng 34:80–87. https://doi.org/10.1016/j.ymben.2015.12.004
Petersson G (1970) Mass spectrometry of aldonic and deoxyaldonic acids as trimethylsilyl derivatives. Tetrahedron 26:3413–3428. https://doi.org/10.1016/S0040-4020(01)92918-7
Petersson G (1977) Retention data in GLC analysis; carbohydrate-related hydroxy carboxylic and dicarboxylic acids as trimethylsilyl derivatives. J Chromatogr Sci 15(7):245–255. https://doi.org/10.1093/chromsci/15.7.245
Robertson G (2012) Food packaging: principles and practice, 3rd edn. CRC Press, Boca Raton
Salusjärvi L, Pitkänen J-P, Aristidou A, Ruohonen L, Penttilä M (2006) Transcription analysis of recombinant Saccharomyces cerevisiae reveals novel responses to xylose. Appl Biochem Biotechnol 128:237–261. https://doi.org/10.1385/ABAB:128:3:237
Salusjärvi L, Kankainen M, Soliymani R, Pitkänen J-P, Penttilä M, Ruohonen L (2008) Regulation of xylose metabolism in recombinant Saccharomyces cerevisiae. Microb Cell Factories 7:18. https://doi.org/10.1186/1475-2859-7-18
Sherman F, Fink G, Hicks JB (1983) Methods in Yeast Genetics. A Laboratory Manual. Cold Springs Harbor Laboratory, Cold Springs Harbor
Sridhara S, Wu TT (1969) Purification and properties of lactaldehyde dehydrogenase from Escherichia coli. J Biol Chem 244:5233–5238
Steensma HY, Ter Linde JJM (2001) Plasmids with the Cre-recombinase and the dominantnat marker, suitable for use in prototrophic strains of Saccharomyces cerevisiae and Kluyveromyces lactis. Yeast 18:469–472
Stephens C, Christen B, Fuchs T, Sundaram V, Watanabe K, Jenal U (2007) Genetic analysis of a novel pathway for D-xylose metabolism in Caulobacter crescentus. J Bacteriol 189:2181–2185
Suzuki M (1985) Analytical aspects of endogenous sugar-derived compounds: special emphasis on inductive agents to hunger or satiety. GC-MS News 13:41–44
Tai Y-S, Xiong M, Jambunathan P, Wang J, Wang J, Stapleton C, Zhang K (2016) Engineering nonphosphorylative metabolism to generate lignocellulose-derived products. Nat Chem Biol 12:247–253. https://doi.org/10.1038/nchembio.2020
Toivari MH, Ruohonen L, Richard P, Penttilä M, Wiebe MG (2010) Saccharomyces cerevisiae engineered to produce D-xylonate. Appl Microbiol Biotechnol 88:751–760. https://doi.org/10.1007/s00253-010-2787-9
Toivari M, Nygård Y, Kumpula EP, Vehkomäki ML, Bencina M, Valkonen M, Maaheimo H, Andberg M, Koivula A, Ruohonen L, Penttilä M, Wiebe MG (2012) Metabolic engineering of Saccharomyces cerevisiae for bioconversion of D-xylose to D-xylonate. Metab Eng 14:427–436. https://doi.org/10.1016/j.ymben.2012.03.002
Toivari M, Vehkomäki M-L-L, Nygård Y, Penttilä M, Ruohonen L, Wiebe MG (2013) Low pH D-xylonate production with Pichia kudriavzevii. Bioresour Technol 133:555–562. https://doi.org/10.1016/j.biortech.2013.01.157
Turkia H, Siren H, Pitkänen JP, Wiebe M, Penttilä M (2010) Capillary electrophoresis for the monitoring of carboxylic acid production by Gluconobacter oxydans. J Chromatogr 1217:1537–1542. https://doi.org/10.1016/j.chroma.2009.12.075;%2010.1016/j.chroma.2009.12.075
Wang J, Shen X, Jain R, Wang J, Yuan Q, Yan J (2017) Establishing a novel biosynthetic pathway for the production of 3,4-dihydroxybutyric acid from xylose in Escherichia coli. Metab Eng 41:39–45. https://doi.org/10.1016/j.ymben.2017.03.003
Weimberg R (1961) Pentose oxidation by Pseudomonas fragi. J Biol Chem 236:629–635
Winston F, Dollard C, Ricupero-Hovasse SL (1995) Construction of a set of convenient Saccharomyces cerevisiae strains that are isogenic to S288C. Yeast 11:53–55
Woods RA, Gietz RD (2001) High-efficiency transformation of plasmid DNA into yeast. Methods Mol Biol 177:85–97
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mpl8 and pUC19 vectors. Gene 33:103–119
Zhang N, Wang J, Zhang Y, Gao H (2016) Metabolic pathway optimization for biosynthesis of 1,2,4-butanetriol from xylose by engineered Escherichia coli. Enz Microb Technol 93:51–58. https://doi.org/10.1016/j.enzmictec.2016.07.007
Acknowledgements
Technical assistance of Mari Helanterä and Merja Helanterä is gratefully acknowledged. Heidi Turkia is acknowledged for the CE analytics.
Funding
This study was financially supported by the Academy of Finland through the Centre of Excellence in White Biotechnology—Green Chemistry (grant 118573) and the Pentoval project (grant 129174) and by the Finnish Funding Agency for Innovation (TEKES) (Living Factories project 562/31/2014).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Salusjärvi, L., Toivari, M., Vehkomäki, ML. et al. Production of ethylene glycol or glycolic acid from D-xylose in Saccharomyces cerevisiae . Appl Microbiol Biotechnol 101, 8151–8163 (2017). https://doi.org/10.1007/s00253-017-8547-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8547-3