Abstract
Infectious diseases caused by pathogenic microorganisms such as viruses and bacteria pose a great threat to human health. Although a significant progress has been obtained in the diagnosis and prevention of infectious diseases, it still remains challenging to develop rapid and cost-effective detection approaches and overcome the side effects of therapeutic agents and pathogen resistance. Functional nucleic acids (FNAs), especially the most widely used aptamers and DNAzymes, hold the advantages of high stability and flexible design, which make them ideal molecular recognition tools for bacteria and viruses, as well as potential therapeutic drugs for infectious diseases. This review summarizes important advances in the selection and detection of bacterial- and virus-associated FNAs, along with their potential prevention ability of infectious disease in recent years. Finally, the challenges and future development directions are concluded.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Infectious diseases caused by viruses, bacteria, and other pathogenic microorganisms are important causes of human morbidity and mortality. Pathogenic microorganisms can gain infectivity through transmission between food, water sources, or hosts. Therefore, effective and reliable diagnosis and therapy methods are in urgent need for infectious diseases.
The process of microscopic identification includes isolation of pathogenic microorganisms from clinical specimens, culture, observation, and genome sequencing. It usually takes long time to cultivate pathogenic microorganisms, and some pathogenic microorganisms are not easy to profile, which leads to misjudgment of the diagnosis result. After acquiring the genetic characteristics of the pathogenic microorganism, microscopic observation, PCR, and immunology methods can be applied to diagnose infectious diseases [1]. PCR-based methods are usually one of the gold standards for the diagnosis of infectious diseases, which enable an accurate detection of the corresponding DNA/RNA fragments of the pathogenic genome, but they require specialized instruments and conditions and experienced personnel. Moreover, immunology methods including the agglutination test, enzyme-linked immunosorbent assay (ELISA), and Western blot analysis use specific antibodies to detect surface proteins, carbohydrates of pathogenic microorganisms, or IgG and IgM. However, antibodies are usually expensive and have poor thermal stability. Therefore, it is necessary to develop an accessible, stable, highly specific, and sensitive recognition molecule for the detection and treatment of pathogenic microorganisms.
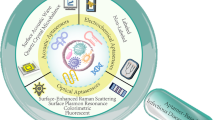
Functional nucleic acids (FNAs) refer to a class of nucleic acid molecules with target binding or catalytic properties. The most commonly used FNAs are aptamers and DNAzymes (Fig. 1), which are DNA or RNA molecules selected from in vitro screening and confirmed with high binding affinity and selectivity against targets [4]. Compared with antibodies, FNAs promise several advantages. First, FNAs have lower molecular weights with better penetration for in vivo applications. Second, as nuclei acid molecules, FNAs can be massively produced by solid-phase synthesis, which is cost-effective with no batch effects, and feasible chemical modification can easily be incorporated. Third, the low immunogenicity of FNAs outweighs antibodies and is crucial for immunology-related tests. Fourth, as the selection of FNAs is routine and the library contains large numbers of random sequences, various FNAs against targets can be enriched with different functions. Taken together, with these unique merits, FNAs have been broadly used in the detection and treatment of pathogenic microorganism–caused diseases. This paper first introduces different strategies for the selection of aptamers and DNAzymes. Moreover, corresponding applications for FNAs in the diagnosis and treatment of bacterial- and virus-caused infectious diseases are summarized (Scheme 1). Finally, we share the overview of challenges and the growing trend of novel FNAs against bacteria and viruses along with their potential applications in disease diagnosis.
FNA selection for pathogenic microorganisms
Aptamers
Aptamers are synthetic small oligonucleotides with high binding affinity and specific affinity against targets (Fig. 1a) [5]. Compared with antibodies, aptamers are smaller, have easer of synthesis and modification, and have better thermal stability. Aptamers have been broadly used in in vitro diagnosis and therapy. The targets for aptamer selection against pathogenic microorganisms include whole bacteria, viruses, and protein markers. Most aptamers were obtained by Systematic Evolution of Ligands by Exponential Enrichment (SELEX), which was first developed in 1990 [6, 7]. The workflow of SELEX usually includes the following steps: a random library of approximately 1013–1015 oligonucleotide sequences is incubated with the target (whole cells, cell debris, or biomarkers) and the binding sequences are collected for amplification followed by increased selection pressure or negative selection to exclude nonspecific binding aptamers. Although a large number of aptamers have been obtained by SELEX, improved strategies are needed for enhancing the affinity and specificity of the aptamers with reduced workload of screening. Basically, aptamer selection can be divided into two categories: selecting aptamers against known biomarkers and targeted selection against whole cells or virus, both of which will be introduced in the following sections. Finally, aptamers selected for the detection of bacteria and viruses are summed up in Table 1 and Table 2, respectively.
Identifying aptamers against biomarkers of pathogenic microorganisms
Biomarker-based SELEX uses known biomarkers of bacteria/viruses or bacterial/viral toxin as SELEX targets. For example, aptamers against specific biomarkers on bacteria/viruses, such as lipopolysaccharides (components of the membranes of Gram-negative bacteria) [21], hepatitis B surface antigen (HBsAg) [26], SARS-CoV spike protein [32], and proteins related with infection and assembly, such as hemagglutinin of the H9N2 avian influenza virus (adhering the sialic acid receptors on surfaces of host cells) [35] and HCV core protein (participating in virus assembly) [30], were obtained. In order to improve selection efficiency, an innovative selection technology based on microfluidic technology [36] and artificial intelligence [36, 37] has been developed. Hong et al. [38] proposed a microfluidic chip based on magnetic separation, which combined the cleaning process, operation of micromagnetic beads, and real-time evaluation of the screening effect. Only three rounds of screening were required to obtain highly selective aptamers for GP and NP proteins. Song et al. [2] used ACE2-based competition and machine learning algorithms to select high affinity aptamers for SARS-CoV-2 RBD. The pre-enriching ssDNA library was sequenced and scored by the enrichment of the motif/substructure, the tolerance of the aptamer family, the ability of the motif/substructure, the size of the aptamer family, and the stability of the total secondary structure. Two high-affinity aptamers that compete with ACE2 were selected. Combined with the machine learning technology, this novel strategy improved the efficiency of screening candidate aptamers and avoided the tedious process of manually selecting aptamers.
In addition, toxins secreted by bacteria are also targets for detecting bacterial infections and neutralizing toxicity. In this case, aptamers against anthrax lethal factor secreted by Bacillus anthracis [39] and staphylococcal enterotoxin type A (SEA) secreted by Staphylococcus aureus [20] were evolved. Overall, biomarker-based SELEX methods can generate highly specific aptamers. With defined biomarkers, creative strategies can be introduced to evolve aptamers with desired characteristics. Moreover, the designed aptamers can be easily implemented with downstream analysis, such as nucleic acid amplification for biosensing. Since this technique is limited to known biomarkers, approaches for evolving aptamers against unknown targets are needed to expand the applications of aptamers.
Bacterial/virus-based SELEXs
The whole-bacteria/virus SELEX technology incorporates whole bacteria/virus with an ssDNA library by identifying oligonucleotide chains and complex markers on the bacterial surface, which does not require verification of special markers on the surface. DNA aptamers against infectious pathogens including Haemophilus influenzae type b [11], Mycobacterium tuberculosis [40], Escherichia coli (E. coli) [9, 17], Streptococcus (Streptococcus pyogenes [12], Streptococcus pneumoniae [15], Staphylococcus aureus [18, 41], Staphylococcus epidermidis [42]), and Vibrio (Vibrio parahaemolyticus [43], Vibrio vulnificus [10], Vibrio alginolyticus [14]) were selected. Moreover, DNA aptamers for nonpathogenic bacteria such as Bifidobacterium [13] and Vibrio fischeri [16] were selected. For virus-based SELEXs, DNA aptamers targeting H1N1 virus [25] and RNA aptamers targeting hepatitis C virus [34], dengue virus (DENV) [31], vaccinia virus [44, 45], etc. were selected. However, due to the large diversity of biomarkers and surface area of the whole cell, large amounts of nonspecific adsorption and low-affinity sequences hinder the selecting process, resulting in redundant selection rounds and aptamers with low specificity. To tackle this problem, Song et al. [19] poured a functionalized GO (PC-GO) material composed of polyethylene glycol (PEG) and chitosan (CTS) to the ssDNA library to dissolve nonspecific binding and low-affinity aptamers. The ssDNA sequences were amplified by the complementary ring mediated rolling circle amplification (CRM-RCA) method, and the RCA products can be directly put into the next round, which improved the enrichment efficiency and shortens the enrichment time.
DNAzymes
Similar to aptamers, DNAzymes are single-stranded DNA molecules selected from random DNA libraries in vitro, but with an added catalytic activity (Fig. 1b–c) [46]. The most common DNAzymes are RNA-cleaving DNAzymes and G-quadruplex DNAzymes (G4-DNAzymes). RNA-cleaving DNAzymes usually split the RNA-containing substrate into two fragments with a high catalytic efficiency. G4-DNAzymes [47] are highly ordered nucleic acids rich in guanines with a peroxidase-like catalytic activity. Due to Hoogsteen hydrogen bonding, guanines can form planar G-tetrads stabilized by monovalent cations. DNAzymes were first obtained by “in vitro selection” in 1994, which were confirmed to have an RNA-cleaving activity. The procedures of DNAzyme selection are as follows: First, the target is incubated with a pool of random DNAzymes that contained a single RNA linkage and the activity sequences are cleaved and release the remained sequences, which were collected and amplified for the next round. After sequencing, the candidate sequences are selected. The target of DNAzymes can be RNA [48], mixture derived from the bacteria [49], potential cell target mixture [50], and crude extracellular matrix (CEM) of bacteria [3]. For instance, Li’s group [51] developed a selection method that incubated a DNA library with a bacterial culture supernatant for both counter selection and positive selection. The method did not require pre-determined biomarkers, and the selected DNAzymes were highly sensitive to bacteria and could respond to potential targets in the sample matrix (e.g., serum or lake water). Using this method, a series of bacterial-responsive DNAzymes were screened out, such as E. coli–activated RFD-EC1 [51], Clostridium difficile–activated RFD-CD1 [49], Klebsiella pneumoniae–activated RFD-KP6 [52], and Helicobacter pylori–activated DHp3T4 [53]. Moreover, the DNAzymes can be combined with specifically designed probes for the highly sensitive detection of bacteria. However, the target and specificity of these DNAzymes are unknown, and the detection performances of complex samples are fuzzy.
In summary, both aptamers and DNAzymes were selected from ssDNA libraries, where random ssDNA libraries and ssDNA libraries with RNA sequences were used for the selection of aptamers and DNAzymes, respectively. Besides, aptamers were evolved by amplifying the sequences with a strong binding affinity of the target. On the contrary, the sequences with cleavage activity activated by the target were acquired as DNAzymes. Overall, the selection of aptamers was focused on the binding affinity, while the evolution of DNAzymes was based on the cleavage activity or peroxidase activity. The obtained aptamers and DNAzymes can be employed for different strategies. For instance, the structure change of aptamers after interaction with the target was usually employed in biosensors. And the catalytic cleaving activity or peroxide activity of DNAzymes was commonly used for signal amplification. However, the applications of DNAzymes were limited due to the uncertainty of their targets.
Applications of FNAs for infectious diseases
Aptamers
Pathogenic bacteria can cause severe problems in public health [1]. Therefore, detection of specific pathogen plays an important role in human health and the environment to avoid bacterial-derived diseases and potential epidemics [54]. Ideal bacterial sensors are expected to be rapid and sensitive, where DNAzyme [49], antibodies [55], and aptamers [56] have been successfully incorporated. With advantages of ease of production and modification, flexibility, and stability, aptamers have been widely adopted in detection of bacteria and therapy for bacterial-induced diseases. A variety of aptamers of desired targets are selected from the SELEX process [57]. Besides, bacterial or viral contamination often appears in the process of antibody production [58]. On the contrary, the chemical production of aptamers can avoid these problems in the biological process [58, 59]. Moreover, the incorporation of nucleic acid amplification can largely enhance the detection sensitivity of aptamer-based strategies [60, 61]. Overall, the unique advantages of aptamers make them ideal tools for biosensing, and the latest applications of aptamer-based strategies for detection of infectious diseases will be introduced in the following sections.
Bacterial detection
Ligand-responsive biosensors
To obtain highly sensitive and rapid detection of pathogen bacteria, aptamers have been combined with ligand-responsive biosensing devices, like quartz crystal microbalance (QCM) [62] and multichannel series piezoelectric quartz crystal (MSPQC) [63]. QCM is a sensor that can measure the quality with high sensitivity. Specifically, the quality change of the crystal surface will result in a positive change in the resonance frequency of the quartz crystal. Therefore, when aptamers are modified on the crystal surface, the target can be precisely detected according to the mass change caused by the bound targets. MSPQC reflects slight changes of the interaction between targets and aptamers by varied electrical signals. For example, He’s group developed a H37Rv sensor platform [63] with MSPQC for the specific and rapid detection of M. tuberculosis (Fig. 2a). The H37Rv aptamer was obtained by the whole-cell SELEX, and the specifically designed probe complementary to the specific H37RV aptamer was modified with the Au-IDE electrode by the Au-S interaction. Single-walled carbon nanotubes (SWCNTs) were bound to aptamers through π-π stacking. The interaction of the target H37Rv with its aptamers led to a frequency shift response, where the changed electrical signals were detected by the MSPQC. This system can distinguish H37Rv from the Bacillus Calmette-Guerin vaccine (BCG) and Mycobacterium smegmatis (M. smegmatis), and down to 100 CFU mL−1 M. tuberculosis could be detected within 70 min.
Schematic of a the electrochemical detection for H37Rv by the Au-IED MSPQC system [63], b a SERS platform for the detection of S. aureus [64], c a SERS platform for the multiplex analysis of S. aureus and E. coli [65], and d a fluorescence strategy for the detection of Staphylococcus aureus (S. aureus) [66]
Surface-enhanced Raman scattering
Different from conventional Raman spectra, the surface-enhanced Raman scattering (SERS) technology is based on plasmon resonances on illuminated metallic substrates and can achieve a 104~106-fold signal amplification. For SERS-based bacterial biosensors, SERS signals were substantially enhanced by the interaction with aptamers with the corresponding target; thus, pathogens of interest could be identified directly and quantified sensitively through the SERS spectrum. For instance, a SERS-based biosensor was developed for the detection of Staphylococcus aureus (S. aureus), which is a common food-borne pathogen widely existing in the natural environment and can produce enterotoxins. The aptamers coated with AgNPs bound with S. aureus, and generated enhanced SERS signals, achieving linear analysis of S. aureus ranging from 101 to 107 CFU/mL, with the LOD of 1.5 CFU/mL within 40 min [64] (Fig. 2b). Aside from single bacterial analysis, multiple bacteria detection is vital for practical applications. A dual-recognition platform was proposed to distinguish two bacteria by magnetic enrichment and SERS tags for specific targets (Fig. 2c) [65]. SERS tags were labeled with S. aureus– or E. coli–specific aptamers to form sandwich structures. By establishing the linear relationship between the SERS signal intensity and the logarithm of bacterial concentrations, the developed biosensor could distinguish both bacteria specifically within 1.5 h, where the bacterial capture efficiency of S. aureus was 88.89% and 74.96% for E. coli, and the limit of detection was 50 cells mL−1 for E. coli and 20 cells mL−1 for S. aureus. Despite the high sensitivity, the reliability of quantitation of SERS-based approaches still remains controversial.
Electrochemical methods
Electrochemical detection relies on the changes of flow currents on the electrochemical material surface [67]. The DNA sequence can be folded into various 3D structures to bind to specific targets, and the DNA-modified surface was stable for a long time. Therefore, DNA-nanostructured electrochemical detection technologies have been developed for bacterial analysis with stringent specificity and high stability [68].
Nanoengineered materials are often used as a signaling hybridization medium in electrochemical biosensors [69]. Metal-organic frameworks (MOFs) have complementary or synergistic effects with the addition of metal ions and have been applied in electrochemical biosensors. In Shahrokhian’s work [70], they designed a nanoengineered MOF material based on an E. coli aptamer. The binding of the bacteria and aptamers could be monitored and quantified by analyzing resultant pulse voltammetry (DPV) signals, where a linear dependence on the logarithm of the E. coli O157:H7 concentration ranging from 2.1 × 101 to 2.1 × 107 CFU mL−1 was established, with the detection of limit of 2 CFU mL−1 and the limit of quantitation (LOQ) of 21 CFU mL−1.
Visual detection
Visual detection of pathogens can translate complex signals from a series of biochemistry reactions into the ones that can be observed directly by naked eyes, such as color changes [16], therefore eliminating the need for complex instruments for signal readout and saving the cost [71]. Specifically, aptamer-based methods provide a simple, precise, and universal way for a rapid screening of pathogens [72], and serve as powerful tools for point-of-care testing (POCT).
Lee et al. [72] introduced a magnetic-composite membrane system to achieve an automated detection of Acinetobacter baumannii (AB). They developed the whole nitrocellulose (NC) membrane–based dual-aptamer assay on a microfluidic chip, where the existence of bacteria could simply be determined by the color intensity of the test line, which is useful in point-of-care bacterial diagnostics. Yi and Li’s team developed an aptamer-based sensor for a rapid and sensitive detection of Mycobacterium tuberculosis (M. tuberculosis), which caused almost 1.5 million people’s death in the world [73]. Specifically, they used an app to acquire the results, which is applicable for ordinary people. The pictures of bacteria were taken and the number of bacteria was given by the app via special algorithm. With a special aptamer linked to the direct or indirect enzyme, the whole assay is designed for complex specimens. This system achieved a low quantitation limit of 104 CFU mL−1 with a linear response. The whole process costs only 5 h and significantly saves time compared with the traditional bacterial culture way which takes 3–5 weeks.
Fluorescence biosensors
Since fluorophores can be easily modified with aptamers, aptamer-based fluorescent methods usually harness aptamer-modified fluorophores for fluorescence signal transduction based on the binding of the target and aptamers, with the characteristics of rapidness and high sensitivity [74]. A variety of fluorophores with different properties can be incorporated, thereby boosting the applications of aptamer-based fluorescent sensors especially in food safety [75] and public health [76]. For example, Song et al. employed an aptamer-based strategy for the quantification of S. aureus using the fluorescence resonance energy transfer (FRET) principle (Fig. 2d) [66]. The Van-functionalized gold nanoclusters modified by aptamer served as the energy receptor. Within 30 min of incubation time, a linear response with the concentration of pathogen ranging from 2*101 to 108 CFU mL−1 was obtained, with the LOD down to 10 CFU mL−1. The strategy was also applicable for real samples, where the recovery of S. aureus rose from 99.00 to 109.75%, and relative standard derivations (RSDs) were controlled in less than 4%. It is noted that fluorescence interference should be taken into account and the combination of different fluorophores matters especially when multiplex analysis is needed.
In summary, high specificity and sensitivity are the basic requirements of detection of bacteria. Combining the flexibility of aptamers and the innovation of instruments, detection methods including SERS, electrochemistry, and fluorescence enable an accurate and specific analysis of bacteria and hold the potential for clinical use. However, the relative high cost of equipment may hinder their widespread applications. On the contrary, visualized approaches where signal outputs can be directly read by naked eyes eliminate the need for complex instruments; thus, it is suitable for POCT. Meanwhile, its sensitivity is sometimes sacrificed by the ease-of-use and short analytical time. Therefore, it is critical to strike the balance between sensitivity, specificity, analytical time, cost, and operation threshold.
Therapy for bacterial-induced diseases
Aside from bacterial detection, aptamers can also be applied for the inhibition of bacterial activities by binding specific sites and inhibiting interaction [39], binding viral enzymes and reducing activities of bacteria [77], and blocking immune escape and activating immune cells [40]. Biofilms are organized groups of bacteria, attached to the surface of inanimate or living objects, and wrapped by bacterial extracellular macromolecules. Therefore, biofilm bacteria are highly resistant to host immune defense mechanisms and antibiotics [78]. Pseudomonas aeruginosa (P. aeruginosa) is a universal microorganism that can survive under extreme conditions. It causes diseases both in plants and in humans, such as immune disorders, cystic fibrosis, and severe burns. And P. aeruginosa can form strong biofilms, which consist of numerous extracellular polymeric substances of bacterial communities, and causes some serious diseases [79]. P. aeruginosa aptamer inhibited bacterial infections by targeting biofilm. Therefore, the P. aeruginosa aptamer can be used in biofilm treatment [80]. Ciprofloxacin (CPX) and single-walled carbon nanotubes (SWNTs) are the most common nanomaterials with a strong antibiofilm activity. Combination of aptamer-CPX-SWNTs showed the potential of drug delivery and biofilm formation overmaster [80].
Moreover, Kim’s group reported that a lethal factor (LF) aptamer could cure Bacillus anthracis infection by neutralizing LF protease toxicity [39]. The value of IC50 was 15 ± 1.5 μM and with 85% cell viability, proving that the aptamer was a powerful neutralizing reagent to inhibit anthrax. Additionally, Zhang’s team selected an aptamer against Mycobacterium tuberculosis, which can cause tuberculosis (TB), the second lethal infectious disease in the world [40]. The highly specific aptamer can induce self-protective and activate T cells to produce IFN-γ after it bound with H37RV (a strain of Mycobacterium tuberculosis). With the aptamer treatment, the number of bacteria in the spleen of mice significantly decreased, and less pulmonary alveolar fusion presented in their lungs, indicating the inhibitory effect of the aptamer on TB.
Virus biosensors engineered with aptamers
Electrochemical methods
Electrochemical aptamer–based sensors can be employed with a minimal loss of sensor signals [81]. Since aptamers are electrically charged, the electrochemical responses of aptamers upon the target binding can be used to detect viral markers. Ruslinda et al. [82] used the RNA Tat aptamer probe to detect the HIV-1 Tat protein based on the changed charge density when the HIV-1 Tat protein bound to the RNA aptamer. Li et al. [83] assembled gold nanoparticles (GNPs), polypyrrole nanowires (PPNWs), and multi-walled carbon nanotubes (MWNT) and on the electrode to increase the effective surface area, electrocatalytic activity, and conductivity of the modified electrode, followed by aptamer modification. The hybridization of the aptamer with the target H5N1 gene sequence produced electrochemical signals; therefore, the avian influenza virus was sensitively detected. In addition, Li et al. exploited an aptamer-based enzyme catalysis system with bare interdigitated electrodes to detect avian influenza virus [84]. By constructing sandwich structures on the surface, Lee’s team illustrated an aerometric biosensing platform based on a screen-printed carbon electrode (SPCE) modified with gold nanoparticles to monitor avian influenza virus [85] (Fig. 3a), which detected spiked H5N1 from human serum samples sensitively and specifically. In addition to avian influenza virus, Han et al. combined a hybrid nanomaterial–modified electrode with a selected DNA aptamer, which enabled an accurate and highly selective detection of AIV (avian influenza virus) H5N1 gene sequences [89].
Visual detection
With the advantages of simple operation, rapidness, and independence of complex instruments, lateral flow arrays (LFAs) have become one of the most popular methods for the visual detection of virus in laboratory, community, and home scenarios [90]. Kim et al. [86] adopted lateral stream strips with aptamers to form sandwich structures for the detection of H5N2 virus (Fig. 3b). To overcome the cross-reactivity of antibodies and the compromised binding kinetics of aptamers, Li et al. [91] constructed an influenza virus recognition lateral flow assay equipped with the aptamer and antibody that can distinguish specific virus strains from different subtypes.
Fluorescence
With the advantages of high specificity and excellent sensitivity, aptamer-based fluorescent methods enable signal amplification and allow selective detection. Real-time detection and imaging of the virus is important for tracing virus behavior. However, conventional strategies using protein labeling face a major challenge in virus visualization, in which fluorescent tags may interfere with the activity of target proteins and cause damage to cells due to the size effect. To solve this problem, Zhang’s team developed a strategy combing specific aptamer and molecular beacons (MBs) to achieve protein imaging and HIV recognition [92]. It is expected that imaging of different proteins in living cells could be achieved by introducing different aptamer beacons. Coronaviruses have caused large-scale pandemics in the past decades. Ahn et al. [32] selected aptamers that specifically bind to the N protein of severe acute respiratory syndrome (SARS) coronavirus, and constructed an aptamer-based nanoarray chip to capture SARS coronavirus followed by incorporation of stained fluorescent antibody to detect SARS coronavirus. Recently, the outbreak of COVID-19 caused by coronavirus (SARS-CoV-2 or 2019-nCoV) has posed a great threat to human health globally [93, 94]. As effective recognition and detection probes, aptamers that could recognize SARS coronavirus were obtained and applied for a rapid detection of COVID-19 [2].
Surface plasmon resonance–based sensors
Surface plasmon resonance (SPR) plays an important role in drug discovery and small molecule detection. Different from fluorescent methods or electrochemical assays, SPR-based sensors based on interaction between biological molecules are label-free and have been developed for the rapid detection of virus, such as AIV H5N1 [87], norovirus capsid protein VP1 [27], and H5Nx viruses [23], and NS5B viral protein of HCV (hepatitis C virus) [95]. Li et al. [87] developed an SPR sensor to detect avian influenza virus (AIV) H5N1 (Fig. 3c). They installed the sensor in a flow cell, calibrated in deionized water, and initialized in the air. Within 1.5 h, the whole process was completed, achieving a LOD of 0.128 HAU (one hemagglutination unit) in poultry swab samples. The result showed the potential of the SPR aptasensor in the poultry industry.
Others
Other methods such as chemiluminescence immunoassay (CLIA) and SERS have also been applied for the detection of virus. CLIA is based on the chemiluminescence assay where highly specific immune response occurs for various targets [96], such as viruses or bacteria. Colorimetric sensors reveal the presence of the virus by the color change. In the comparison of other analytical methods, colorimetric sensors have extraordinary superiorities in units of small molecular testing [97]. For instance, Zeng’s group selected the aptamer against Zika virus, which is associated with meningoencephalitis and Guillain-Barré syndrome (GBS) and threatens human survival. They designed an aptamer-based ELISA system to recognize Zika NS1 protein [98]. As for the detection of SARS-CoV, Cho et al. [29] separated the N protein of SARS-CoV by SDS-PAGE, where the 5′-biotinylated aptamer was used to detect the N protein followed by SA-HRP and ECL signal output. Moreover, to detect the infections titer of the hepatitis C virus (HCV), Jang et al. [88] described a system based on enzyme-linked apto-sorbent assay (ELASA), where the aptamers against HCV E2 were employed to measure infectious HCV particles (Fig. 3d). As for SERS-based platforms, Dluhy et al. identified the SERS spectra for the aptamer-nucleoprotein complex to monitor the influenza virus with signal amplification [99].
Therapy for virus-induced diseases
Aptamers can cure diseases caused by virus by preventing the virus from invading cells, inhibiting the proteins or enzymes involved in viral replication [100], or stimulating the immune system [101]. Due to the easy modification of aptamers, therapeutic molecules, for example, small interfering RNA (siRNA), can be conjugated to aptamers for antiviral therapy [102]. Typical examples will be discussed in the following section.
Inhibition viral fusion with the target cell
Viruses infect cells by binding to the component on the surface of target cells via the ligands on the surface of viruses. One of the most direct and effect antiviral therapies is preventing the fusion of viruses to target cells using aptamers [103]. A variety of oligonucleotide aptamers have been selected from pools and shown to be able to inhibit the fusion of virus to host cells, including the HA protein–specific aptamers [28, 104, 105], the hepatitis C virus envelope protein (E1E2) aptamers [106], and Herpes simplex virus type 1 (HSV-1) [100] aptamers. Aside from DNA aptamers, RNA aptamers against hemagglutinins (HAs) of avian influenza (HPAI) H5 and H7 viruses [107] were also found to inhibit viral entry efficiently.
Inhibition of viral replication cycle
Inhibition of viral replication cycle can block virus proliferation. Aptamers have been shown to have antiviral effects by inhibiting enzymes or other proteins participating in viral replication, transcription, and translation [103]. Aptamers have been confirmed to be able to inhibit the activity of endonuclease [108, 109], RNA polymerase [34, 110], reverse transcriptase (HIV-1RT) [111, 112], and methyl transferase activity of dengue virus [31]. The aptamers of Ebola virus can block the competitive binding of dsRNA to VP35 protein of Ebola virus and destroy the interaction of evp35 nucleoprotein (NP) [113]. Moreover, the aptamer against HBV core protein could inhibit the assembly of the nucleocapsid, resulting in reducing extracellular HBV DNA synthesis [114]. Hepatitis C virus aptamers disrupted the localization of the core protein via lipid droplets and NS5A, holding back the core protein from binding to viral RNA [30].
Delivery of antiviral molecule drugs
Since aptamers are easy to be modified, they can be easily modified with antiviral molecules and serve as carriers. In addition, aptamers have low immunogenicity and can reduce side effects. At present, the most commonly used method is to combine aptamers with siRNAs to transport siRNAs to target tissues which knock out the target mRNA, and effectively inhibit HIV-1 replication. Zhou et al. [115] conjugated anti-gp120 aptamer with anti-HIV-1 siRNAs to target mRNA of HIV-1 tat/rev protein. Zhu et al. [102] modified the anti-CD4 DNA aptamer with siRNAs against the mRNA of HIV-1 protease. Both methods showed an inhibitory effect on viral infectivity.
Although studies have shown that aptamers had great potential as antiviral drugs, it is still necessary to develop more stable aptamer screening technology, modification methods, and efficient targeted transfer methods to evolve aptamers with higher affinity or desired properties. Moreover, the pharmacokinetics of aptamers in the circulatory system also needs attention.
DNAzymes
Bacterial biosensors engineered with DNAzymes
Visual detection
Due to the catalytic activity and ease of modification, it is easy to couple signal conversion molecules with DNAzymes. By introducing different combinations of substrates and DNAzymes, visual signal conversion was realized during the catalytic reaction. For example, Tram et al. [116] combined urea hydrolase with DNAzyme that specifically cleaved bacterial-specific RNA on magnetic beads. Urea was hydrolyzed by urease to convert bacterial signals into pH signals, providing a simple, low-cost, and easy-to-promote method for bacterial detection (Fig. 4a). Liu et al. [120] integrated PCR and g-quadruple DNAzyme which mimic the horseradish peroxidase to initiate a colorimetric reaction in the presence of DNA of L. monocytogenes. Court et al. [121] conjugated urease to DNAzyme targeting Helicobacter pylori (HP) via streptavidin-biotin interaction. RNA cleavage reaction releases urease upon adding HP which was then added to urea and phenol red for colorimetric reaction.
Fluorescence biosensors
Due to its RNA-cleaving activity, DNAzymes can induce the switch of specially designed probes and trigger fluorescent signal changes [117]. Typically, DNAzymes are coupled with RNA modified with a fluorescent group and a quencher, where the fluorescence is quenched in the absence of the target. After the introduction of the target, it will induce the cleavage of the RNA strand and keep the quencher away from the fluorescent group, thereby producing fluorescent signals (Fig. 4b). Based on this strategy, detection of Escherichia coli [51] [50, 117, 122], Klebsiella pneumoniae [52], and Vibrio anguillarum [3] has been realized with high specificity [123]. Cao et al. [124] described a turn-off strategy where the MBs modified with fluorophores and quenchers were complementary to RNA sequences and displayed fluorescent signals. However, the target could trigger DNAzymes to cleave RNA sequences, which were therefore not able to hybridize with the MBs. As a result, the fluorescent signals were quenched.
Other methods
Apart from visual detection and fluorescent methods, nucleic acid amplification has also been incorporated for bacterial detection. Bengtson et al. [125] developed a binary deoxyribose sensor called TB-DzT that combined multiplex PCR and single-nucleotide polymorphism (SNP) for the identification of resistant mutants of Mycobacterium tuberculosis (M. tuberculosis). He’s group [118] proposed a DNAzyme-integrated localized surface plasmon resonance (LSPR) sensing strategy to detect E. coli, where the DNAzyme-induced RNA cleavage triggered the HCR signal amplification reaction. Glucose oxidase was modified to the amplificated chain to catalyze glucose, followed by oxidizing silver TNPS to produce a plasma resonance response (Fig. 4c). Li’s group [119] proposed a DNAzyme feedback amplification (DFA) strategy, which used primers and complementary sequences for RNA cleavage. Subsequently, the cleaved RNAs served as substrates and were furthered amplified by rolling circle amplification (RCA), leading to exponential amplification, and the sensitivity was 3–6 orders of magnitude higher than conventional methods (Fig. 4d). Collectively, the integration with nucleic acid amplification can significantly enhance the sensitivity of bacterial detection.
Virus biosensors engineered with DNAzymes
Visual detection
G-quadruplex-hemin DNAzyme has a similar function to HRP, which catalyzes the oxidation of colorless 2,2-azinobis(3-ethylbenzothiozoline)-6-sulfonicacid (ABTS2−) to the green ABTS radical by H2O2. This principle has been applied for the detection of HBV gene [126], DNA related to HIV [127], and dengue virus (DENV) [128]. Yin et al. [129] constructed a visual biosensor with polystyrene (PS) electrospun nanofibrous membrane as a basement to enhance the DNAzyme catalysis efficiency. Except for the detection of HIV DNA, it can also distinguish single-base and double-base mismatch with high selectivity and sensitivity (Fig. 5a). James et al. [134] detected the dengue virus by coupling the specific RNA sequence of the virus to the nanogold using DNAzyme, which caused the aggregation of the nanogold, resulting in the color of the solution changing from red to colorless.
Fluorescence biosensors
The specific recognition of DNAzymes and the target RNA can produce fluorescence signal changes and realize the detection of virus. Kim et al. [130] reported on a nanosized graphene oxide (nGO)–based DNAzyme delivery system, in which the fluorescent DNAzymes were doped into GO and delivered to cells. In the presence of the hepatitis C virus miRNA, the DNAzyme hybridized with the miRNA and detached from the GO, thereby generating fluorescence signals (Fig. 5b). Xiang et al. [135] used Mg(II)-dependent DNAzyme to cleave the substrate and release G-quadruplex, which bound to thioflavin to produce fluorescence. Du et al. [131] developed the HsCR strategy in which DNAzymes were amplified in the form of hairpin DNA strands and tDNase was synthesized by polymerase by RCA. With these two strategies, a homogeneous target-induced cascaded amplification strategy was developed for the simultaneous detection of enterovirus 71 and coxsackievirus B3 (Fig. 5c).
Electrochemical methods
A common strategy for the electrochemical detection of viruses based on DNAzyme is to construct a recognition probe of the target DNA on the electrode. When the target exists, the probe is reassembled into a G-quadruplex DNAzyme. When mixed with hemin and hydrogen peroxide, 3.3′,5.5′-tetramethylbenzidine sulfate (TMB) will be oxidized, resulting in the change of electrorheological signals, which can be revealed by voltammograms [132, 133]. Yu et al. [132] designed two DNA sequences that blocked each other (Fig. 5d). The target triggered isothermal exponential amplification (EXPAR) and HCR reaction, and the polymerase-initiated polymerization and cleavage enzyme can regenerate target DNA. At the same time, the released DNA sequence self-assembled into G-quadruplex. In the presence of hemin, TMB was oxidized to generate electrochemical signals to detect avian influenza A (H7N9); hemin/G-quadruplex DNAzyme can also oxidize methylene blue (MB) in the presence of hydrogen peroxide. Utilizing this characteristic, electrochemical detection of influenza virus subtype H7N9 [132] and hepatitis B virus surface antigen (HBsAg) [136, 137] has been developed.
DNAzyme-based treatment strategies for diseases caused by viruses
DNAzymes can effectively cleave RNA and hold the advantages such as flexible design and independence of cell mechanisms. It can be effectively used in the body without expensive chemical modification and is used to down-regulate a series of important therapeutic genes [138]. DNAzymes were found to be able to inhibit the expression of HBV s- and e-genes [139], inhibit hepatitis B virus X gene expression [140], suppress the transcription and expression of F gene in respiratory syncytial virus (RSV) [141], target M2 gene of influenza A virus [142], and cleave the RNA sequence of the 3′-NCR of JEV genome [8]. Hong et al. [143] constructed nanocarrier chitosan-g-stearic acid (CSO-SA) beads on chitosan for the intracellular delivery of HBV-specific DNAzyme, which targeted cytoplasm, resulting in the enhancement of inhibition of HBV virus.
Conclusions and future directions
Pathogenic microorganism–caused infectious diseases such as viruses and bacteria pose a great threat to human health. Although a significant progress has been achieved in the diagnosis and treatment of infectious diseases, it still remains challenging to develop rapid and cost-effective detection methods and overcome the side effects of therapeutic reagents and resistance of pathogens. FNAs have high affinity, little batch effects, low immunogenicity, and ease of modification, thus providing powerful tools for the development of new diagnostic and therapeutic agents. This article reviews the selection of nucleic acid aptamers and DNAzymes and their latest development in the detection of bacteria and viruses and the treatment of diseases. The SELEX technology of aptamers and the screening technology of DNAzyme enabled a flexible evolution of the desired FNA towards whole bacteria, whole viruses, bacterial and viral markers, and virus’s DNA and RNA. Moreover, aptamers and DNAzymes have been proven as potential tools for antiviral therapy due to the ability to inhibit the recognition of viruses and cells and the proliferation of viruses. However, the application of FNAs requires a comprehensive evaluation of its characteristics, including the stability, equilibrium dissociation constant Kd, and half-inhibitory concentration IC50 in different application scenarios. On the other side, the aptamers selected with whole cell and whole bacteria as targets cannot guarantee high affinity to real samples due to the unknown targets. The side effects of DNA enzymes with unknown cleavage sites in the treatment of infectious diseases are also unknown. Therefore, it is necessary to study the targets and activation mechanism of aptamers and DNAzymes carefully before clinical use.
References
Davydova A, Vorobjeva M, Pyshnyi D, Altman S, Vlassov V, Venyaminova A. Aptamers against pathogenic microorganisms. Crit Rev Microbiol. 2016;42:847–65.
Song Y, Song J, Wei X, Huang M, Sun M, Zhu L, et al. Discovery of aptamers targeting the receptor-binding domain of the SARS-CoV-2 spike glycoprotein. Anal Chem. 2020;92:9895–900.
Gu L, Yan W, Wu H, Fan S, Ren W, Wang S, et al. Selection of DNAzymes for sensing aquatic bacteria: Vibrio anguillarum. Anal Chem. 2019;91:7887–93.
McConnell EM, Cozma I, Morrison D, Li Y. Biosensors made of synthetic functional nucleic acids toward better human health. Anal Chem. 2020;92:327–44.
Bayat P, Nosrati R, Alibolandi M, Rafatpanah H, Abnous K, Khedri M, et al. SELEX methods on the road to protein targeting with nucleic acid aptamers. Biochimie. 2018;154:132–55.
Ellington AD, Szostak JW. INVITRO selection of RNA molecules that bind specific ligands. Nature. 1990;346:818–22.
Tuerk C, Gold L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science. 1990;249:505–10.
Appaiahgari MB, Vrati S. DNAzyme-mediated inhibition of Japanese encephalitis virus replication in mouse brain. Mol Ther. 2007;15:1593–9.
Amraee M, Oloomi M, Yavari A, Bouzari S. DNA aptamer identification and characterization for E-coli O157 detection using cell based SELEX method. Anal Biochem. 2017;536:36–44.
Yan W, Gu L, Liu S, Ren W, Lyu M, Wang S. Identification of a highly specific DNA aptamer for Vibrio vulnificus using systematic evolution of ligands by exponential enrichment coupled with asymmetric PCR. J Fish Dis. 2018;41:1821–9.
Bitaraf FS, Rasooli I, Gargari SLM. DNA aptamers for the detection of Haemophilus influenzae type b by cell SELEX. Eur J Clin Microbiol Infect Dis. 2016;35:503–10.
Hamula CLA, Peng H, Wang Z, Tyrrell GJ, Li X-F, Le XC. An improved SELEX technique for selection of DNA aptamers binding to M-type 11 of Streptococcus pyogenes. Methods. 2016;97:51–7.
Hu L, Wang L, Lu W, Zhai Q, Fan D, Liu X, et al. Selection, identification and application of DNA aptamers for the detection of Bifidobacterium breve. RSC Adv. 2017;7:11672–9.
Yu Q, Liu M, Su H, Xiao H, Wu S, Qin X, et al. Selection and characterization of ssDNA aptamers specifically recognizing pathogenic Vibrio alginolyticus. J Fish Dis. 2019;42:851–8.
Bayrac AT, Donmez SI. Selection of DNA aptamers to Streptococcus pneumonia and fabrication of graphene oxide based fluorescent assay. Anal Biochem. 2018;556:91–8.
Shin W-R, Sekhon SS, Rhee S-K, Ko JH, Ahn J-Y, Min J, et al. Aptamer-based paper strip sensor for detecting Vibrio fischeri. ACS Comb Sci. 2018;20:261–8.
Zou Y, Duan N, Wu S, Shen M, Wang Z. Selection, identification, and binding mechanism studies of an ssDNA aptamer targeted to different stages of E-coli O157:H7. J Agric Food Chem. 2018;66:5677–82.
Ramlal S, Mondal B, Lavu PS, Bhavanashri N, Kingston J. Capture and detection of Staphylococcus aureus with dual labeled aptamers to cell surface components. Int J Food Microbiol. 2018;265:74–83.
Song S, Wang X, Xu K, Li Q, Ning L, Yang X. Selection of highly specific aptamers to Vibrio parahaemolyticus using cell-SELEX powered by functionalized graphene oxide and rolling circle amplification. Anal Chim Acta. 2019;1052:153–62.
Sedighian H, Halabian R, Amani J, Heiat M, Amin M, Fooladi AAI. Staggered target SELEX, a novel approach to isolate non-cross-reactive aptamer for detection of SEA by apta-qPCR. J Biotechnol. 2018;286:45–55.
Ye H, Duan N, Wu S, Tan G, Gu H, Li J, et al. Orientation selection of broad-spectrum aptamers against lipopolysaccharides based on capture-SELEX by using magnetic nanoparticles. Microchim Acta. 2017;184:4235–42.
Park J-W, Lee SJ, Choi E-J, Kim J, Song J-Y, Gu MB. An ultra-sensitive detection of a whole virus using dual aptamers developed by immobilization-free screening. Biosens Bioelectron. 2014;51:324–9.
Van-Thuan N, Seo HB, Kim BC, Kim SK, Song C-S, Gu MB. Highly sensitive sandwich-type SPR based detection of whole H5Nx viruses using a pair of aptamers. Biosens Bioelectron. 2016;86:293–300.
Lai YT, DeStefano JJ. DNA aptamers to human immunodeficiency virus reverse transcriptase selected by a primer-free SELEX method: characterization and comparison with other aptamers. Nucleic Acid Ther. 2012;22:162–76.
Bai C, Lu Z, Jiang H, Yang Z, Liu X, Ding H, et al. Aptamer selection and application in multivalent binding-based electrical impedance detection of inactivated H1N1 virus. Biosens Bioelectron. 2018;110:162–7.
Xi ZJ, Huang RR, Li ZY, He NY, Wang T, Su EB, et al. Selection of HBsAg-specific DNA aptamers based on carboxylated magnetic nanoparticles and their application in the rapid and simple detection of hepatitis B virus infection. ACS Appl Mater Interfaces. 2015;7:11215–23.
Beier R, Pahlke C, Quenzel P, Henseleit A, Boschke E, Cuniberti G, et al. Selection of a DNA aptamer against norovirus capsid protein VP1. FEMS Microbiol Lett. 2014;351:162–9.
Cheng C, Dong J, Yao L, Chen A, Jia R, Huan L, et al. Potent inhibition of human influenza H5N1 virus by oligonucleotides derived by SELEX. Biochem Biophys Res Commun. 2008;366:670–4.
Cho SJ, Woo HM, Kim KS, Oh JW, Jeong YJ. Novel system for detecting SARS coronavirus nucleocapsid protein using an ssDNA aptamer. J Biosci Bioeng. 2011;112:535–40.
Shi S, Yu X, Gao Y, Xue B, Wu X, Wang X, et al. Inhibition of hepatitis C virus production by aptamers against the core protein. J Virol. 2014;88:1990–9.
Jung JI, Han SR, Lee S-W. Development of RNA aptamer that inhibits methyltransferase activity of dengue virus. Biotechnol Lett. 2018;40:315–24.
Ahn DG, Jeon IJ, Kim JD, Song MS, Han SR, Lee SW, et al. RNA aptamer-based sensitive detection of SARS coronavirus nucleocapsid protein. Analyst. 2009;134:1896–901.
Gabriela Valencia-Resendiz D, Palomino-Vizcaino G, Virginia Tapia-Vieyra J, Luisa Benitez-Hess M, Gabriela Leija-Montoya A, Marat Alvarez-Salas L. Inhibition of human papillomavirus type 16 infection using an RNA aptamer. Nucleic Acid Ther. 2018;28:97–105.
Biroccio A, Hamm J, Incitti I, De Francesco R, Tomei L. Selection of RNA aptamers that are specific and high-affinity ligands of the hepatitis C virus RNA-dependent RNA polymerase. J Virol. 2002;76:3688–96.
Choi SK, Lee C, Lee KS, Choe S-Y, Mo IP, Seong RH, et al. DNA aptamers against the receptor binding region of hemagglutinin prevent avian influenza viral infection. Mol Cell. 2011;32:527–33.
Lou X, Qian J, Xiao Y, Viel L, Gerdon AE, Lagally ET, et al. Micromagnetic selection of aptamers in microfluidic channels. PNAS. 2009;106:2989–94.
Song J, Zheng Y, Huang M, Wu L, Wang W, Zhu Z, et al. A sequential multidimensional analysis algorithm for aptamer identification based on structure analysis and machine learning. Anal Chem. 2020;92:3307–14.
Hong SL, Xiang MQ, Tang M, Pang DW, Zhang ZL. Ebola virus aptamers: from highly efficient selection to application on magnetism-controlled chips. Anal Chem. 2019;91:3367–73.
Lahousse M, Park H-C, Lee S-C, Ha N-R, Jung I-P, Schlesinger SR, et al. Inhibition of anthrax lethal factor by ssDNA aptamers. Arch Biochem Biophys. 2018;646:16–23.
Chen F, Zhou J, Luo F, Mohammed A-B, Zhang X-L. Aptamer from whole-bacterium SELEX as new therapeutic reagent against virulent Mycobacterium tuberculosis. Biochem Biophys Res Commun. 2007;357:743–8.
Cao X, Li S, Chen L, Ding H, Xu H, Huang Y, et al. Combining use of a panel of ssDNA aptamers in the detection of Staphylococcus aureus. Nucleic Acids Res. 2009;37:4621–8.
Kaur SJ, Gilman V, Duong M, Asher DM, Gregori L. Rapid selection of single-stranded DNA aptamers binding Staphylococcus epidermidis in platelet concentrates. Biotechniques. 2018;65:331–8.
Ahn J-Y, Lee K-A, Lee M-J, Sekhon SS, Rhee S-K, Cho S-J, et al. Surface plasmon resonance aptamer biosensor for discriminating pathogenic bacteria Vibrio parahaemolyticus. J Nanosci Nanotechnol. 2018;18:1599–605.
Nitsche A, Kurth A, Dunkhorst A, Paenke O, Sielaff H, Junge W, et al. One-step selection of Vaccinia virus-binding DNA aptamers by MonoLEX. BMC Biotechnol. 2007;7:48.
Tang Z, Parekh P, Turner P, Moyer RW, Tan W. Generating aptamers for recognition of virus-infected cells. Clin Chem. 2009;55:813–22.
Morrison D, Rothenbroker M, Li YF. DNAzymes: selected for applications. Small Methods. 2018;2:1700319.
Burge S, Parkinson GN, Hazel P, Todd AK, Neidle S. Quadruplex DNA: sequence, topology and structure. Nucleic Acids Res. 2006;34:5402–15.
Kim KS, Choi WH, Gong SJ, Oh S, Kim JH, Kim DE. Efficient target site selection for an RNA-cleaving DNAzyme through combinatorial library screening. Bull Kor Chem Soc. 2006;27:657–62.
Shen Z, Wu Z, Chang D, Zhang W, Kha T, Lee C, et al. A catalytic DNA activated by a specific strain of bacterial pathogen. Angew Chem Int Ed. 2016;55:2431–4.
Zhang W, Feng Q, Chang D, Tram K, Li Y. In vitro selection of RNA-cleaving DNAzymes for bacterial detection. Methods. 2016;106:66–75.
Ali MM, Aguirre SD, Lazim H, Li Y. Fluorogenic DNAzyme probes as bacterial indicators. Angew Chem Int Ed. 2011;50:3751–4.
Ali MM, Slepenkin A, Peterson E, Zhao W. A simple DNAzyme-based fluorescent assay for Klebsiella pneumoniae. Chembiochem. 2019;20:906–10.
Ali MM, Wolfe M, Tram K, Gu J, Filipe CDM, Li Y, et al. A DNAzyme-based colorimetric paper sensor for Helicobacter pylori. Angew Chem Int Ed Eng. 2019;58:9907–11.
O'Sullivan CK. Aptasensors - the future of biosensing. Anal Bioanal Chem. 2002;372:44–8.
Masdor NA, Altintas Z, Tothill IE. Sensitive detection of Campylobacter jejuni using nanoparticles enhanced QCM sensor. Biosens Bioelectron. 2016;78:328–36.
Xu Y, Wang H, Luan C, Liu Y, Chen B, Zhao Y. Aptamer-based hydrogel barcodes for the capture and detection of multiple types of pathogenic bacteria. Biosens Bioelectron. 2018;100:404–10.
Stoltenburg R, Reinemann C, Strehlitz B. SELEX-A (r)evolutionary method to generate high-affinity nucleic acid ligands. Biosens Bioelectron. 2007;24:381–403.
Kulbachinskiy AV. Methods for selection of aptamers to protein targets. Biochemistry-Moscow. 2007;72:1505–18.
Gopinath SCB. Methods developed for SELEX. Anal Bioanal Chem. 2007;387:171–82.
Torres-Chavolla E, Alocilja EC. Aptasensors for detection of microbial and viral pathogens. Biosens Bioelectron. 2009;24:3175–82.
Hamula CLA, Zhang HQ, Li F, Wang ZX, Le XC, Li XF. Selection and analytical applications of aptamers binding microbial pathogens. Trac-Trends in Anal Chem. 2011;30:1587–97.
Wang L, Wang R, Chen F, Jiang T, Wang H, Slavik M, et al. QCM-based aptamer selection and detection of Salmonella typhimurium. Food Chem. 2017;221:776–82.
Zhang X, Feng Y, Yao Q, He F. Selection of a new Mycobacterium tuberculosis H37Rv aptamer and its application in the construction of a SWCNT/aptamer/Au-IDE MSPQC H37Rv sensor. Biosens Bioelectron. 2017;98:261–6.
Gao W, Li B, Yao R, Li Z, Wang X, Dong X, et al. Intuitive label-free SERS detection of bacteria using aptamer-based in situ silver nanoparticles synthesis. Anal Chem. 2017;89:9836–42.
Zhang C, Wang C, Xiao R, Tang L, Huang J, Wu D, et al. Sensitive and specific detection of clinical bacteria via vancomycin-modified Fe3O4@Au nanoparticles and aptamer-functionalized SERS tags. J Mater Chem B. 2018;6:3751–61.
Yu M, Wang H, Fu F, Li L, Li J, Li G, et al. Dual-recognition forster resonance energy transfer based platform for one-step sensitive detection of pathogenic bacteria using fluorescent vancomycin-gold nanoclusters and aptamer-gold nanoparticles. Anal Chem. 2017;89:4085–90.
Han D, Yan Y, Wang J, Zhao M, Duan X, Kong L, et al. An enzyme-free electrochemiluminesce aptasensor for the rapid detection of Staphylococcus aureus by the quenching effect of MoS2-PtNPs-vancomycin to S2O82-/O-2 system. Sensors Actuators B Chem. 2019;288:586–93.
Jo N, Kim B, Lee S-M, Oh J, Park IH, Lim KJ, et al. Aptamer-functionalized capacitance sensors for real-time monitoring of bacterial growth and antibiotic susceptibility. Biosens Bioelectron. 2018;102:164–70.
Hasan MR, Pulingam T, Appaturi JN, Zifruddin AN, Teh SJ, Lim TW, et al. Carbon nanotube-based aptasensor for sensitive electrochemical detection of whole-cell Salmonella. Anal Biochem. 2018;554:34–43.
Shahrokhian S, Ranjbar S. Aptamer immobilization on amino-functionalized metal-organic frameworks: an ultrasensitive platform for the electrochemical diagnostic of Escherichia coli O157:H7. Analyst. 2018;143:3191–201.
Bayrac C, Eyidogan F, Oktem HA. DNA aptamer-based colorimetric detection platform for Salmonella Enteritidis. Biosens Bioelectron. 2017;98:22–8.
Wu J-H, Wang C-H, Ma Y-D, Lee B. A nitrocellulose membrane-based integrated microfluidic system for bacterial detection utilizing magnetic-composite membrane microdevices and bacteria-specific aptamers. Lab Chip. 2018;18:1633–40.
Li L, Liu Z, Zhang H, Yue W, Li C-W, Yi C. A point-of-need enzyme linked aptamer assay for Mycobacterium tuberculosis detection using a smartphone. Sensors Actuators B Chem. 2018;254:337–46.
Song M-S, Sekhon SS, Shin W-R, Kim HC, Min J, Ahn J-Y, et al. Detecting and discriminating Shigella sonnei using an aptamer-based fluorescent biosensor platform. Molecules. 2017;22:825.
Bayramoglu G, Ozalp VC, Dincbal U, Arica MY. Fast and sensitive detection of Salmonella in milk samples using aptamer-functionalized magnetic silica solid phase and MCM-41-aptamer gate system. ACS Biomater Sci Eng. 2018;4:1437–44.
Yan W, Gu L, Ren W, Ma X, Qin M, Lyu M, et al. Recognition of Helicobacter pylori by protein-targeting aptamers. Helicobacter. 2019;24:e12577.
Shum KT, Tanner JA. Differential inhibitory activities and stabilisation of DNA aptamers against the SARS coronavirus helicase. Chembiochem. 2008;9:3037–45.
Flemming HC, Wuertz S. Bacteria and archaea on Earth and their abundance in biofilms. Nat Rev Microbiol. 2019;17:247–60.
Chen Z, Wang Z, Ren J, Qu X. Enzyme mimicry for combating bacteria and biofilms. Acc Chem Res. 2018;51:789–99.
Wang S, Mao B, Wu M, Liang J, Deng L. Influence of aptamer-targeted antibiofilm agents for treatment of Pseudomonas aeruginosa biofilms. Anton Leeuw Int J Gen Mol Microbiol. 2018;111:199–208.
Liu J, Cao Z, Lu Y. Functional nucleic acid sensors. Chem Rev. 2009;109:1948–98.
Rahim Ruslinda A, Tanabe K, Ibori S, Wang X, Kawarada H. Effects of diamond-FET-based RNA aptamer sensing for detection of real sample of HIV-1 Tat protein. Biosens Bioelectron. 2013;40:277–82.
Liu X, Cheng Z, Fan H, Ai S, Han R. Electrochemical detection of avian influenza virus H5N1 gene sequence using a DNA aptamer immobilized onto a hybrid nanomaterial-modified electrode. Electrochim Acta. 2011;56:6266–70.
Fu Y, Callaway Z, Lum J, Wang R, Lin J, Li Y. Exploiting enzyme catalysis in ultra-low ion strength media for impedance biosensing of avian influenza virus using a bare interdigitated electrode. Anal Chem. 2014;86:1965–71.
Diba FS, Kim S, Lee HJ. Amperometric bioaffinity sensing platform for avian influenza virus proteins with aptamer modified gold nanoparticles on carbon chips. Biosens Bioelectron. 2015;72:355–61.
Kim SH, Lee J, Lee BH, Song C-S, Gu MB. Specific detection of avian influenza H5N2 whole virus particles on lateral flow strips using a pair of sandwich-type aptamers. Biosens Bioelectron. 2019;134:123–9.
Bai H, Wang R, Hargis B, Lu H, Li Y. A SPR aptasensor for detection of avian influenza virus H5N1. Sensors. 2012;12:12506–18.
Park JH, Jee MH, Kwon OS, Keum SJ, Jang SK. Infectivity of hepatitis C virus correlates with the amount of envelope protein E2: development of a new aptamer-based assay system suitable for measuring the infectious titer of HCV. Virology. 2013;439:13–22.
Lum J, Wang R, Hargis B, Tung S, Bottje W, Lu H, et al. An impedance aptasensor with microfluidic chips for specific detection of H5N1 avian influenza virus. Sensors. 2015;15:18565–78.
Liu RD, McConnell EM, Li JX, Li YF. Advances in functional nucleic acid based paper sensors. J Mater Chem B. 2020;8:3213–30.
Le TT, Chang P, Benton DJ, McCauley JW, Iqbal M, Cass AEG. Dual recognition element lateral flow assay toward multiplex strain specific influenza virus detection. Anal Chem. 2017;89:6781–6.
Liang Y, Zhang Z, Wei H, Hu Q, Deng J, Guo D, et al. Aptamer beacons for visualization of endogenous protein HIV-1 reverse transcriptase in living cells. Biosens Bioelectron. 2011;28:270–6.
Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579:270–3.
Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020;367:1260–3.
Roh C, Kim SE, Jo S-K. Label free inhibitor screening of hepatitis C virus (HCV) NS5B viral protein using RNA oligonucleotide. Sensors. 2011;11:6685–96.
Wang C, Wu J, Zong C, Xu J, Ju H-X. Chemiluminescent immunoassay and its applications. Chin J Anal Chem. 2012;40:3–10.
Zhao L, Sun L, Chu X. Chemiluminescence immunoassay. TrAC Trend Anal Chem. 2009;28:404–15.
Lee KH, Zeng H. Aptamer-based ELISA assay for highly specific and sensitive detection of Zika NS1 protein. Anal Chem. 2017;89:12743–8.
Negri P, Chen G, Kage A, Nitsche A, Naumann D, Xu B, et al. Direct optical detection of viral nucleoprotein binding to an anti-influenza aptamer. Anal Chem. 2012;84:5501–8.
Yadavalli T, Agelidis A, Jaishankar D, Mangano K, Thakkar N, Penmetcha K, et al. Targeting herpes simplex virus-1 gD by a DNA aptamer can be an effective new strategy to curb viral infection. Mol Ther-Nucl Acids. 2017;9:365–78.
Hwang SY, Sun HY, Lee KH, Oh BH, Cha YJ, Kim BH, et al. 5'-Triphosphate-RNA-independent activation of RIG-I via RNA aptamer with enhanced antiviral activity. Nucleic Acids Res. 2012;40:2724–33.
Zhu Q, Shibata T, Kabashima T, Kai M. Inhibition of HIV-1 protease expression in T cells owing to DNA aptamer-mediated specific delivery of siRNA. Eur J Med Chem. 2012;56:396–9.
Wandtke T, Wozniak J, Kopinski P. Aptamers in diagnostics and treatment of viral infections. Viruses-Basel. 2015;7:751–80.
Jeon SH, Kayhan B, Ben-Yedidia T, Arnon R. A DNA aptamer prevents influenza infection by blocking the receptor binding region of the viral hemagglutinin. J Biol Chem. 2004;279:48410–9.
Park SY, Kim S, Yoon H, Kim KB, Kalme SS, Oh S, et al. Selection of an antiviral RNA aptamer against hemagglutinin of the subtype H5 avian influenza virus. Nucleic Acid Ther. 2011;21:395–402.
Yang D, Meng X, Yu Q, Xu L, Long Y, Liu B, et al. Inhibition of hepatitis C virus infection by DNA aptamer against envelope protein. Antimicrob Agents Chemother. 2013;57:4937–44.
Suenaga E, Kumar PK. An aptamer that binds efficiently to the hemagglutinins of highly pathogenic avian influenza viruses (H5N1 and H7N7) and inhibits hemagglutinin-glycan interactions. Acta Biomater. 2014;10:1314–23.
Yuan S, Zhang N, Singh K, Shuai H, Chu H, Zhou J, et al. Cross-protection of influenza A virus infection by a DNA aptamer targeting the PA endonuclease domain. Antimicrob Agents Chemother. 2015;59:4082–93.
Jang KJ, Lee NR, Yeo WS, Jeong YJ, Kim DE. Isolation of inhibitory RNA aptamers against severe acute respiratory syndrome (SARS) coronavirus NTPase/Helicase. Biochem Biophys Res Commun. 2008;366:738–44.
Bellecave P, Cazenave C, Rurni J, Staedel C, Cosnefroy O, Andreola M-L, et al. Inhibition of hepatitis C virus (HCV) RNA polymerase by DNA aptamers: mechanism of inhibition of in vitro RNA synthesis and effect on HCV-infected cells. Antimicrob Agents Chemother. 2008;52:2097–110.
Ditzler MA, Bose D, Shkriabai N, Marchand B, Sarafianos SG, Kvaratskhelia M, et al. Broad-spectrum aptamer inhibitors of HIV reverse transcriptase closely mimic natural substrates. Nucleic Acids Res. 2011;39:8237–47.
Shiang Y-C, Ou C-M, Chen S-J, Ou T-Y, Lin H-J, Huang C-C, et al. Highly efficient inhibition of human immunodeficiency virus type 1 reverse transcriptase by aptamers functionalized gold nanoparticles. Nanoscale. 2013;5:2756–64.
Binning JM, Wang T, Luthra P, Shabman RS, Borek DM, Liu G, et al. Development of RNA aptamers targeting Ebola virus VP35. Biochemistry. 2013;52:8406–19.
Zhang ZW, Zhang J, Pei XY, Zhang Q, Lu B, Zhang XJ, et al. An aptamer targets HBV core protein and suppresses HBV replication in HepG2.2.15 cells. Int J Mol Med. 2014;34:1423–9.
Zhou J, Neff CP, Swiderski P, Li H, Smith DD, Aboellail T, et al. Functional in vivo delivery of multiplexed anti-HIV-1 siRNAs via a chemically synthesized aptamer with a sticky bridge. Mol Ther. 2013;21:192–200.
Tram K, Kanda P, Salena BJ, Huan S, Li Y. Translating bacterial detection by DNAzymes into a litmus test. Angew Chem Int Ed. 2014;53:12799–802.
Yousefi H, Ali MM, Su H-M, Filipe CDM, Didar TF. Sentinel wraps: real-time monitoring of food contamination by printing DNAzyme probes on food packaging. ACS Nano. 2018;12:3287–94.
Yu F, Li Y, Li M, Tang L, He J-J. DNAzyme-integrated plasmonic nanosensor for bacterial sample-to-answer detection. Biosens Bioelectron. 2017;89:880–5.
Liu M, Zhang Q, Chang D, Gu J, Brennan JD, Li Y. A DNAzyme feedback amplification strategy for biosensing. Angew Chem Int Ed. 2017;56:6142–6.
Liu Z, Yao C, Yang C, Wang Y, Wan S, Huang J. Development of DNAzyme-based PCR signal cascade amplification for visual detection of Listeria monocytogenes in food. Anal Biochem. 2018;553:7–11.
Court CM, Hou S, Winograd P, Segel NH, Li QW, Zhu Y, et al. A novel multimarker assay for the phenotypic profiling of circulating tumor cells in hepatocellular carcinoma. Liver Transpl. 2018;24:946–60.
Aguirre SD, Ali MM, Salena BJ, Li Y. A sensitive DNA enzyme-based fluorescent assay for bacterial detection. Biomolecules. 2013;3:563–77.
Zaouri N, Cui Z, Peinetti AS, Lu Y, Hong P-Y. DNAzyme-based biosensor as a rapid and accurate verification tool to complement simultaneous enzyme-based media for E. coli detection. Environ Sci: Water Res Technol. 2019;5:2260–8.
Cao T, Wang Y, Zhao L-L, Wang Y, Tao Y, Heyman JA, et al. A simple mix-and-read bacteria detection system based on a DNAzyme and a molecular beacon. Chem Commun. 2019;55:7358–61.
Bengtson HN, Homolka S, Niemann S, Reis AJ, da Silva PE, Gerasimova YV, et al. Multiplex detection of extensively drug resistant tuberculosis using binary deoxyribozyme sensors. Biosens Bioelectron. 2017;94:176–83.
Li YB, Liu S, Deng QJ, Ling LS. A sensitive colorimetric DNA biosensor for specific detection of the HBV gene based on silver-coated glass slide and G-quadruplex-hemin DNAzyme. J Med Virol. 2018;90:699–705.
Wang XQ, Huang ZJ, Chen JM, Luo ZW, Xu Y, Duan YX. A colorimetric sensing platform based on site-specific endonuclease IV-aided signal amplification for the detection of DNA related to the human immunodeficiency virus. Anal Methods. 2019;11:2190–6.
Ida J, Kuzuya A, Choong YS, Lim TS. An intermolecular-split G-quadruplex DNAzyme sensor for dengue virus detection. RSC Adv. 2020;10:33040–51.
Yin CY, Wu YY, Li XL, Niu JJ, Lei J, Ding XD, et al. Highly selective, naked-eye, and trace discrimination between perfect-match and mismatch sequences using a plasmonic nanoplatform. Anal Chem. 2018;90:7371–6.
Kim S, Ryoo S-R, Na H-K, Kim Y-K, Choi B-S, Lee Y, et al. Deoxyribozyme-loaded nano-graphene oxide for simultaneous sensing and silencing of the hepatitis C virus gene in liver cells. Chem Commun. 2013;49:8241–3.
Du MY, Mao GB, Tian SB, Liu YC, Zheng J, Ke XL, et al. Target-induced cascade amplification for homogeneous virus detection. Anal Chem. 2019;91:15099–106.
Yu YY, Chen ZG, Jian WS, Sun DP, Zhang BB, Li XC, et al. Ultrasensitive electrochemical detection of avian influenza A (H7N9) virus DNA based on isothermal exponential amplification coupled with hybridization chain reaction of DNAzyme nanowires. Biosens Bioelectron. 2015;64:566–71.
Shi LJ, Yu YY, Chen ZG, Zhang L, He SJ, Shi QJ, et al. A label-free hemin/G-quadruplex DNAzyme biosensor developed on electrochemically modified electrodes for detection of a HBV DNA segment. RSC Adv. 2015;5:11541–8.
Carter JR, Balaraman V, Kucharski CA, Fraser TS, Fraser MJ. A novel dengue virus detection method that couples DNAzyme and gold nanoparticle approaches. Virol J. 2013;10:201.
Xiang L, Zhang F, Chen CY, Cai CQ. A general scheme for fluorometric detection of multiple oligonucleotides by using RNA-cleaving DNAzymes: application to the determination of microRNA-141 and H5N1 virus DNA. Microchim Acta. 2019;186:511.
Alizadeh N, Hallaj R, Salimi A. A highly sensitive electrochemical immunosensor for hepatitis B virus surface antigen detection based on Hemin/G-quadruplex horseradish peroxidase-mimicking DNAzyme-signal amplification. Biosens Bioelectron. 2017;94:184–92.
Alizadeh N, Hallaj R, Salimi A. Dual amplified electrochemical immunosensor for hepatitis B virus surface antigen detection using hemin/G-quadruplex immobilized onto Fe3O4-AuNPs or (hemin-amino-rGO-Au) nanohybrid. Electroanalysis. 2018;30:402–14.
Fokina AA, Stetsenko DA, Francois JC. DNA enzymes as potential therapeutics: towards clinical application of 10-23 DNAzymes. Expert Opin Biol Ther. 2015;15:689–711.
Wo JE, Wu XL, Zhou LF, Yao HP, Chen LW, Dennin RH. Effective inhibition of expression of hepatitis B virus genes by DNAzymes. World J Gastroenterol. 2005;11:3504–7.
Hou W, Ni Q, Wo JN, Li MW, Liu KZ, Chen LW, et al. Inhibition of hepatitis B virus X gene expression by 10-23 DNAzymes. Antivir Res. 2006;72:190–6.
Xie YY, Zhao XD, Jiang LP, Liu HL, Wang LJ, Fang P, et al. Inhibition of respiratory syncytial virus in cultured cells by nucleocapsid gene targeted deoxyribozyme (DNAzyme). Antivir Res. 2006;71:31–41.
Kumar B, Rajput R, Pati DR, Khanna M. Potent intracellular knock-down of influenza a virus M2 gene transcript by DNAzymes considerably reduces viral replication in host cells. Mol Biotechnol. 2015;57:836–45.
Hong Y, Mao DS, Wu R, Gao Z, Meng TT, Wang RR, et al. Hepatitis B virus S gene therapy with 10-23 DNAzyme delivered by chitosan-g-stearic acid micelles. RSC Adv. 2019;9:15196–204.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Additional information
Published in the topical collection Analytical Chemistry for Infectious Disease Detection and Prevention with guest editors Chaoyong Yang and XiuJun (James) Li.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhu, L., Ling, J., Zhu, Z. et al. Selection and applications of functional nucleic acids for infectious disease detection and prevention. Anal Bioanal Chem 413, 4563–4579 (2021). https://doi.org/10.1007/s00216-020-03124-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-020-03124-3