Abstract
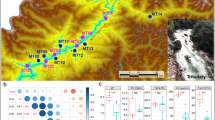
The gastrointestinal microbial community plays a crucial role in host health, immunity, protection, development and provides nutrients to the host. The rising human-induced pollution and heavy metal contamination in all aquatic systems globally has led us to explore the gut microbial diversity of two exotic invasive fish Cyprinus carpio (Linnaeus, 1858) and Oreochromis niloticus (Linnaeus,1857) from river Yamuna, India. These fishes are aquatic bioindicators with high demographic resilience. Exploring these associations would pave the way for addressing problems that inhabitant fishes are facing due to the increasing pollution load in the River Yamuna. Based on 16S rRNA gene amplicon sequencing, our results deliver comparative information on the gut microbiome of these fishes and highlight connotations between the microbiome of gut and water samples. The gut of C. carpio and O. niloticus was dominated by phyla Proteobacteria whereas Bacteroidetes dominated the water sample. Microbial communities showed predicted roles such as pathogenicity (Escherichia-Shigella, Aeromonas veronii, Vibrio cholerae, Streptococcus iniae, Flavobacterium columnare, Klebsiella pneumoniae, Mycobacterium sp.), probiotic applications (Bacillus velezensis, Lactobacillus plantarum, Enterococcus faecalis, Bifidobacterium longum, Lactococcus lactis, Leuconostoc falkenbergense) and involvement in sewage and organic matter decomposition (Nitrosomonas sp., Methanosaeta harundinacea, Dechloromonas agitata, Thauera humireducens, Zoogloea ramigera). Heavy metal degrading members (Leucobacter chromiireducens, Pseudomonas fluorescens, P. aeruginosa, Klebsiella pneumoniae, and Micrococcus luteus) were detected in gut microbiome samples thus supporting the notion that fish shapes its gut microbiota with changing ecology. Functional profiling showed that microbial communities are specialized in metabolic functions thus reflecting the dietary profile of these invasive fishes.





Similar content being viewed by others
Data availability
The raw sequencing read data have been deposited in the National Centre for Biotechnology Information (NCBI) under the project Accession Number PRJNA809116.
References
Ahmed W, Hughes B, Harwood VJ (2016) Current status of marker genes of Bacteroides and related taxa for identifying sewage pollution in environmental waters. Water J 8(6):231. https://doi.org/10.3390/w8060231
Allameh KS, Ringø E, Yusoff MF, Daud MH, Ideris A (2014) Properties of Enterococcus faecalis, a new probiotic bacterium isolated from the intestine of snakehead fish (Channa striatus Bloch). Afr J Microbiol Res 8(22):2215–2222. https://doi.org/10.5897/AJMR2013.5830
Bentley R, Meganathan R (1982) Biosynthesis of vitamin K (menaquinone) in bacteria. Microbiol. Rev 46(3):241–280
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37(8):852–857. https://doi.org/10.1038/s41587-019-0209-9
Cebeci A, Gürakan C (2003) Properties of potential probiotic Lactobacillus plantarum strains. Food Microbiol 20(5):511–518. https://doi.org/10.1016/S0740-0020(02)00174-0
Claus SP, Guillou H, Ellero-Simatos S (2016) The gut microbiota: a major player in the toxicity of environmental pollutants? NPJ Biofilms Microbiomes 2:16003. https://doi.org/10.1038/npjbiofilms.2016.3
Douglas GM, Maffei VJ, Zaneveld J, Yurgel SN, Brown JR, Taylor CM, Langille MG (2020) PICRUSt2: an improved and customizable approach for metagenome inference. BioRxiv. https://doi.org/10.1101/672295
Dwivedi AC, Tiwari A, Mayank P (2018) Environmental pollution supports to constancy and invader potential of Cyprinus carpio and Oreochromis niloticus from the Ganga river. India Int J Poultry Sci 2(1):1–7. https://doi.org/10.15226/2578-1898/2/2/00113
Fakhar A, Gul B, Gurmani AR, Khan SM, Ali S, Sultan T et al (2020) Heavy metal remediation and resistance mechanism of Aeromonas, Bacillus, and Pseudomonas: a review. Crit Rev Environ Sci Technol. https://doi.org/10.1080/10643389.2020.1863112
IUCN/SSC Invasive Species Specialist Group (ISSG) (2014) Global Invasive Species Database version 2014-2 http://193.206.192.138/gisd/ (Accessed 10 October 2014)
Jaber SM, Al-Mayahi FSA (2020) Screening and characterization of Pseudomonas aeruginosa resistant for heavy metal from surface sediment of Euphrates river Iraq biochem. Cell Arch 20(2):5203–5210
Jami M, Ghanbari M, Kneifel W, Domig KJ (2015) Phylogenetic diversity and biological activity of culturable Actinobacteria isolated from freshwater fish gut microbiota. Microbiol Res 175:6–15. https://doi.org/10.1016/j.micres.2015.01.009
Jampasri K, Pokethitiyook P, Poolpak T, Kruatrachue M, Ounjai P, Kumsopa A (2020) Bacteria-assisted phytoremediation of fuel oil and lead co-contaminated soil in the salt-stressed condition by chromolaena odorata and Micrococcus luteus. Int J Phytoremediation 22(3):333. https://doi.org/10.1080/15226514.2016.1183568
Johny TK, Puthusseri RM, Bhat SG (2021) Metagenomic landscape of taxonomy, metabolic potential and resistome of Sardinella longiceps gut microbiome. Arch Microbiol 204(1):87. https://doi.org/10.1007/s00203-021-02675-y
Kaevska M, Videnska P, Sedlar K, Slana I (2016) Seasonal changes in microbial community composition in river water studied using 454-pyrosequencing. Springerplus 5:1–8. https://doi.org/10.1186/s40064-016-2043-6
Khalid F, Khalid A, Fu Y, Hu Q, Zheng Y, Khan S, Wang Z (2021) Potential of Bacillus velezensis as a probiotic in animal feed: a review. J Microbiol 59(7):627–633. https://doi.org/10.1007/s12275-021-1161-1
Khurana H, Sharma M, Bharti M, Singh DN, Negi RK (2021) Gut milieu shapes the bacterial communities of invasive silver carp. Genomics 113(2):815–826. https://doi.org/10.1016/j.ygeno.2021.01.013
Kim PS, Shin NR, Lee JB, Kim MS, Whon TW, Hyun DW, Bae JW (2021) Host habitat is the major determinant of the gut microbiome of fish. Microbiome 9(1):1–6. https://doi.org/10.1186/s40168-021-01113-x
Koushlesh SM, Sajina AM, Roshith CM (2021) Ichthyofaunal diversity of the major Indian rivers: A review. J Inland Fish Soc India 53(1&2):22–35
Krossøy C, Waagbø R, Ørnsrud R (2011) Vitamin K fish nutrition. Aquac Nutr 17(6):585–594. https://doi.org/10.1111/j.1365-2095.2011.00904.x
Mittal P, Prasoodanan PKV, Dhakan DB, Kumar S, Sharma VK (2019) Metagenome of a polluted river reveals a reservoir of metabolic and antibiotic resistance genes. Environ Microbiome 14(1):1–12. https://doi.org/10.1186/s40793-019-0345-3
Mohan AK, Martis S, Chiplunkar S, Kamath S, Goveas LC, Rao CV (2019) Heavy Metal Tolerance of Klebsiella pneumoniae Kpn555 Isolated from Coffee Pulp Waste. Borneo J Res Sci Technol 9(2):101–106
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glockner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41(D1):D590–D596. https://doi.org/10.1093/nar/gks1219
Rajeev AC, Sahu N, Arvind K, Deori M, Grace T, Dev SA et al (2021) Exploring prevalence of potential pathogens and fecal indicators in geographically distinct river systems through comparative metagenomics. Environ Pollut 282:117003. https://doi.org/10.1016/j.envpol.2021.117003
Restivo VE, Kidd KA, Surette MG, Servos MR, Wilson JY (2021) Rainbow darter (Etheostomacaeruleum) from a river impacted by municipal wastewater effluents have altered gut content microbiomes. Sci Total Environ 751:141724. https://doi.org/10.1016/j.scitotenv.2020.141724
Sharma S, Sundaram CS, Luthra PM, Singh Y, Sirdeshmukh R, Gade WN (2006) Role of proteins in resistance mechanism of Pseudomonas fluorescens against heavy metal induced stress with proteomics approach. J Biotechnol 126(3):374–382. https://doi.org/10.1016/j.jbiotec.2006.04.032
Sharma R, Singh NS, Singh DK (2020) Impact of heavy metal contamination and seasonal variations on enzyme’s activity of Yamuna river soil in Delhi and NCR. Appl Water Sci 10(3):1–8. https://doi.org/10.1007/s13201-020-1166-7
Sodhi KK, Kumar M, Singh DK (2021) Assessing the bacterial diversity and functional profiles of the river Yamuna using illumina MiSeq sequencing. Arch Microbiol 203:367–375. https://doi.org/10.1007/s00203-020-02045-0
Sun H, He X, Ye L, Zhang XX, Wu B, Ren H (2017) Diversity, abundance, and possible sources of fecal bacteria in the Yangtze river. Appl Microbiol and Biotechnol 101:2143–2152. https://doi.org/10.1007/s00253-016-7998-2
Talwar C, Nagar S, Lal R, Negi RK (2018) Fish gut microbiome: current approaches and future perspectives. Indian J Microbiol 58(4):397–414. https://doi.org/10.1007/s12088-018-0760-y
Thomas J, Thanigaivel S, Vijayakumar S, Acharya K, Shinge D, Seelan JTS, Chandrasekaran N (2014) Pathogenecity of Pseudomonas aeruginosa in Oreochromis mossambicus and treatment using lime oil nanoemulsion. Colloids Surf B Biointerfaces 116:372–377. https://doi.org/10.1016/j.colsurfb.2014.01.019
Toranzo AE, Magariños B, Romalde JL (2005) A review of the main bacterial fish diseases in mariculture systems. Aquac Res 246(1–4):37–61. https://doi.org/10.1016/j.aquaculture.2005.01.002
Tyagi A, Singh B (2017) Microbial diversity in rohu fish gut and inland saline aquaculture sediment and variations associated with next-generation sequencing of 16S rRNA gene. J Fish Life Sci 2:8
Wagner M, Loy A, Nogueira R, Purkhold U, Lee N, Daims H (2002) Microbial community composition and function in wastewater treatment plants. Antonie Van Leeuwenhoek 81(1):665–680. https://doi.org/10.1023/A:1020586312170
WHO (2011) Guidelines for drinking-water quality 4th edn. Geneva: World Health Organization (WHO)
Xue X, Jia J, Yue X, Guan Y, Zhu L, Wang Z (2021) River contamination shapes the microbiome and antibiotic resistance in sharpbelly (Hemiculterleucisculus). Environ Pollut 268:115796. https://doi.org/10.1016/j.envpol.2020.115796
Zhang L, Shen T, Cheng Y, Zhao T, Li L, Qi P (2020) Temporal and spatial variations in the bacterial community composition in lake Bosten, a large, brackish lake in China. Sci Rep 10:1–10. https://doi.org/10.1038/s41598-019-57238-5
Zhu W, Yang Z, Ma Z, Chai L (2008) Reduction of high concentrations of chromate by Leucobacter sp. CRB1 isolated from Changsha, China. World J Microbiol Biotechnol 24:991–996. https://doi.org/10.1007/s11274-007-9564-7
Acknowledgements
MB and SN thank the Council of Scientific and Industrial Research (CSIR) for providing doctoral fellowships.
Funding
This work was supported by funds from the Institute of Eminence (IoE), University Grants Commission-Special Assistance Programme (UGC-SAP), and Application of Microorganisms in Agriculture and Allied Sectors (AMAAS).
Author information
Authors and Affiliations
Contributions
RKN conceived the idea. MB wrote the manuscript, MB and SN performed the analysis. SN, HK helped shape the manuscript. RKN critically reviewed the manuscript and improved it. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interests
The authors declare no conflict of interest.
Ethical approval
No special permission was required for this study.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bharti, M., Nagar, S., Khurana, H. et al. Metagenomic insights to understand the role of polluted river Yamuna in shaping the gut microbial communities of two invasive fish species. Arch Microbiol 204, 509 (2022). https://doi.org/10.1007/s00203-022-03127-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-022-03127-x