Abstract
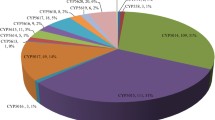
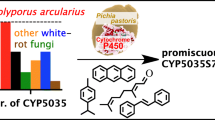
We explored the molecular diversity and functional capabilities of cytochrome P450 monooxygenases (P450s) from the brown-rot basidiomycete Postia placenta. Using bioinformatic and experimental data, we found 250 genes of P450s in the whole genome, including 60 putative allelic variants. Phylogenetic analysis revealed the presence of 42 families, including 18 novel families. Comparative phylogenetic analysis of P450s from P. placenta and the white-rot basidiomycete Phanerochaete chrysosporium suggested that vigorous gene duplication and molecular evolution occurred after speciation of basidiomycetes. Among the 250 gene models, 184 were isolated as full-length cDNA and transformed into Saccharomyces cerevisiae to construct a functional library in which recombinant P450s were co-expressed with yeast NADPH-P450 oxidoreductase. Using this library, the catalytic potentials of P450s against a wide variety of compounds were investigated. A functionomic survey allowed the discovery of novel catalytic properties of P. placenta P450s. The phylogenetic diversity of the CYP53 family in P. placenta was clear, and CYP53D2 is capable of converting stilbene derivatives. This is the first report of this peculiar function of the CYP53 family. Our increased understanding of the molecular and functional diversity of P450s in this fungus will facilitate comprehension of metabolic diversity in basidiomycetes and has future biotechnology applications.




Similar content being viewed by others
References
Chang MCY, Eachus RA, Trieu W, Ro D-K, Keasling JD (2007) Engineering Escherichia coli for production of functionalized terpenoids using plant P450s. Nat Chem Biol 3:274–277
Chigu NL, Hirosue S, Nakamura C, Teramoto H, Ichinose H, Wariishi H (2010) Cytochrome P450 monooxygenases involved in anthracene metabolism by the white-rot basidiomycete Phanerochaete chrysosporium. Appl Microbiol Biotechnol 87:1907–1916
Eriksson K-EL, Blanchette RA, Ander P (1990) Microbial and enzymatic degradation of wood and wood components. Springer, Berlin
Faber BW, Van Gorcom RFM, Duine JA (2001) Purification and characterization of benzoate-para-hydroxylase, a cytochrome P450 (CYP53A1), from Aspergillus niger. Arch Biochem Biophys 394:245–254
Felsenstein J (1989) PHYLIP—phylogeny inference package (version 3.2). Cladistics 5:164–166
Gillam EMJ (2008) Engineering cytochrome P450 enzymes. Chem Res Toxicol 21:220–231
Gold MH, Wariishi H, Valli K (1989) Extracellular peroxidases involved in lignin degradation by the white rot basidiomycete Phanerochaete chrysosporium. In: Whitaker JR, Sonnet PE (eds) Biocatalysis in agricultural biotechnology, ASC symposium series 389. American Chemical Society, Washington D.C, pp 127–140
Grogan G (2011) Cytochromes P450: exploiting diversity and enabling application as biocatalysts. Curr Opin Chem Biol 15:241–248
Guengerich FP (2002) Cytochrome p450 enzymes in the generation of commercial products. Nat Rev Drug Discov 1:359–366
Hammel KE, Moen MA (1991) Depolymerization of a synthetic lignin in vitro by lignin peroxidase. Enzyme Microb Technol 13:15–18
Hirokawa T, Boon-Chieng S, Mitaku S (1998) SOSUI: classification and secondary structure prediction system for membrane proteins. Bioinformatics 14:378–379
Hirosue S, Tazaki M, Hiratsuka N, Yanai S, Kabumoto H, Shinkyo R et al (2011) Insight into functional diversity of cytochrome P450 in the white-rot basidiomycete Phanerochaete chrysosporium: involvement of versatile monooxygenase. Biochem Biophys Res Commun 407:118–123
Ichinose H, Wariishi H, Tanaka H (1999) Biotransformation of recalcitrant 4-methyldibenzothiophene to water-extractable products using lignin-degrading basidiomycete Coriolus versicolor. Biotechnol Prog 15:706–714
Ichinose H, Wariishi H, Tanaka H (2002) Identification and characterization of novel cytochrome P450 genes from the white-rot basidiomycetes Coriolus versicolor. Appl Microbiol Biotechnol 58:97–105
Ingelman-Sundberg M (2001) Implications of polymorphic cytochrome p450-dependent drug metabolism for drug development. Drug Metab Dispos 29:570–573
Jiang H, Morgan JA (2004) Optimization of an in vivo plant P450 monooxygenase system in Saccharomyces cerevisiae. Biotechnol Bioeng 85:130–137
Kelly DE, Krasevec N, Mullins J, Nelson DR (2009) The CYPome (Cytochrome P450 complement) of Aspergillus nidulans. Fungal Genet Biol 46:S53–S61
Kirk TK, Farrell RL (1987) Enzymatic combustion—the microbial-degradation of lignin. Annu Rev Microbiol 41:465–505
Kirk TK, Schultz E, Connors WJ, Lorenz LF, Zeikus JG (1978) Influence of culture parameters on lignin metabolism by Phanerochaete chrysosporium. Arch Microbiol 117:277–285
Lamb DC, Skaug T, Song HL, Jackson CJ, Podust LM, Waterman MR et al (2002) The cytochrome P450 complement (CYPome) of Streptomyces coelicolor A3(2). J Biol Chem 277:24000–24005
Martinez D, Larrondo LF, Putnam N, Gelpke MD, Huang K, Chapman J et al (2004) Genome sequence of the lignocellulose degrading fungus Phanerochaete chrysosporium strain RP78. Nat Biotechnol 22:695–700
Martinez D, Challacombe J, Morgenstern I, Hibbett D, Schmoll M, Kubicek CP et al (2009) Genome, transcriptome, and secretome analysis of wood decay fungus Postia placenta supports unique mechanisms of lignocellulose conversion. Proc Natl Acad Sci USA 106:1954–1959
Matsuzaki F, Wariishi H (2005) Molecular characterization of cytochrome P450 catalyzing hydroxylation of benzoates from the white-rot fungus Phanerochaete chrysosporium. Biochem Biophys Res Commun 334:1184–1190
Matsuzaki F, Shimizu M, Wariishi H (2008) Proteomic and metabolomic analyses of the white-rot fungus Phanerochaete chrysosporium exposed to exogenous benzoic acid. J Proteome Res 7:2342–2350
Murakami H, Yabusaki Y, Sakaki T, Shibata M, Ohkawa H (1990) Expression of cloned yeast NADPH-cytochrome P450 reductase gene in Saccharomyces cerevisiae. J Biochem 108:859–865
Nazir KHMNH, Ichinose H, Wariishi H (2010) Molecular characterization and isolation of cytochrome P450 genes from the filamentous fungus Aspergillus oryzae. Arch Microbiol 192:395–408
Nazir KHMNH, Ichinose H, Wariishi H (2011) Construction and application of a functional library of cytochrome P450 monooxygenases from the filamentous fungus Aspergillus oryzae. Appl Environ Microbiol 77:3147–3150
Nelson DR (2009) The cytochrome p450 homepage. Hum Genomics 4:59–65
Nelson DR, Koymans L, Kamataki T, Stegeman JJ, Feyereisen R, Waxman DJ et al (1996) P450 superfamily: update on new sequences, gene mapping, accession numbers and nomenclature. Pharmacogenetics 6:1–42
Niemenmaa O, Uusi-Rauva A, Hatakka A (2008) Demethoxylation of O14CH3-labelled lignin model compounds by the brown-rot fungi Gloeophyllum trabeum and Poria (Postia) placenta. Biodegradation 19:555–565
Omura T, Sato R (1964) The carbon monoxide-binding pigment of liver microsomes, II. Solubilization, purification, and properties. J Biol Chem 239:2379–2385
Ortiz de Montellano PR (2005) Cytochrome P450: structure, mechanism, and biochemistry, 3rd ed. Kluwer Academic/Plenum Publishers, New York
Park J, Park B, Jung K, Jang S, Yu K, Choi J et al (2008) CFGP: a web-based, comparative fungal genomics platform. Nucleic Acids Res 36:D562–D571
Podobnik B, Stojan J, Lah L, Krasevec N, Seliskar M, Rizner TL et al (2008) CYP53A15 of Cochliobolus lunatus, a target for natural antifungal compounds. J Med Chem 51:3480–3486
Rimando AM, Suh N (2008) Biological/chemopreventive activity of stilbenes and their effect on colon cancer. Planta Med 74:1635–1643
Ro DK, Paradise EM, Ouellet M, Fisher KJ, Newman KL, Ndungu JM et al (2006) Production of the antimalarial drug precursor artemisinic acid in engineered yeast. Nature 440:940–943
Roupe KA, Remsberg CM, Yáñez JA, Davies NM (2006) Pharmacometrics of stilbenes: seguing towards the clinic. Curr Clin Pharmacol 1:81–101
Sabbadin F, Hyde R, Robin A, Hilgarth EM, Delenne M, Flitsch S et al (2010) LICRED: a versatile drop-in vector for rapid generation of redox-self-sufficient cytochrome P450s. Chembiochem 11:987–994
Sakaki T, Akiyoshi-Shibata M, Yabusaki Y, Ohkawa H (1992) Organella-targeted expression of rat liver cytochrome P450c27 in yeast: genetically engineered alteration of mitochondrial P450 into a microsomal form creates a novel functional electron transport chain. J Biol Chem 267:16497–16502
Sakaki T, Shinkyo R, Takita T, Ohta M, Inouye K (2002) Biodegradation of polychlorinated dibenzo-p-dioxins by recombinant yeast expressing rat CYP1A subfamily. Arch Biochem Biophys 410:91–98
Sambrook J, Russel DW (2001) Extraction, purification, and analysis of mRNA from eukaryotic cells. In: Argentine J (ed) Molecular cloning: a laboratory manual, 3rd edn, vol 1. Cold spring harbor laboratory press, New York
Sato S, Feltus FA, Iyer P, Tien M (2009) The first genome-level transcriptome of the wood-degrading fungus Phanerochaete chrysosporium grown on red oak. Curr Genet 55:273–286
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Urlacher VB, Eiben S (2006) Cytochrome P450 monooxygenases: perspectives for synthetic application. Trends Biotechnol 24:324–330
Vanden Wymelenberg A, Gaskell J, Mozuch M, Sabat G, Ralph J, Skyba O et al (2010) Comparative transcriptome and secretome analysis of wood decay fungi Postia placenta and Phanerochaete chrysosporium. Appl Environ Microbiol 76:3599–3610
Wei D, Houtman CJ, Kapich AN, Hunt CG, Cullen D, Hammel KE (2010) Laccase and its role in production of extracellular reactive oxygen species during wood decay by the brown rot basidiomycete Postia placenta. Appl Environ Microbiol 76:2091–2097
Yadav JS, Doddapaneni H, Subramanian V (2006) P450ome of the white rot fungus Phanerochaete chrysosporium: structure, evolution and regulation of expression of genomic P450 clusters. Biochem Soc Trans 34:1165–1169
Yelle DJ, Wei D, Ralph J, Hammel KE (2011) Multidimensional NMR analysis reveals truncated lignin structures in wood decayed by the brown rot basidiomycete Postia placenta. Environ Microbiol 13:1091–1100
Yu A, Kneller BM, Rettie AE, Haining RL (2002) Expression, purification, biochemical characterization, and comparative function of human cytochrome P450 2D6.1, 2D6.2, 2D6.10, and 2D6.17 allelic isoforms. J Pharmacol Exp Ther 303:1291–1300
Acknowledgments
We are deeply grateful to Dr. Osamu Gotoh (Kyoto University, Japan) for technical support of our bioinformatic gene prediction and to Dr. David Nelson (University of Tennessee, USA) for assistance with the CYP naming. This research was supported in part by a Grant-in-Aid (no. 21688013) for Young Scientists (A) from the Ministry of Education, Culture, Sports, Science and Technology, Japan (to H.I.).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Axel Brakhage.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ide, M., Ichinose, H. & Wariishi, H. Molecular identification and functional characterization of cytochrome P450 monooxygenases from the brown-rot basidiomycete Postia placenta . Arch Microbiol 194, 243–253 (2012). https://doi.org/10.1007/s00203-011-0753-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-011-0753-2