Abstract
Key message
We found two loci on chromosomes 2BS and 6AL that significantly contribute to stripe rust resistance in current European winter wheat germplasm.
Abstract
Stripe or yellow rust, caused by the fungus Puccinia striiformis Westend f. sp. tritici, is one of the most destructive wheat diseases. Sustainable management of wheat stripe rust can be achieved through the deployment of rust resistant cultivars. To detect effective resistance loci for use in breeding programs, an association mapping panel of 230 winter wheat cultivars and breeding lines from Northern and Central Europe was employed. Genotyping with the Illumina® iSelect® 25 K Infinium® single nucleotide polymorphism (SNP) genotyping array yielded 8812 polymorphic markers. Structure analysis revealed two subpopulations with 92 Austrian breeding lines and cultivars, which were separated from the other 138 genotypes from Germany, Norway, Sweden, Denmark, Poland, and Switzerland. Genome-wide association study for adult plant stripe rust resistance identified 12 SNP markers on six wheat chromosomes which showed consistent effects over several testing environments. Among these, two marker loci on chromosomes 2BS (RAC875_c1226_652) and 6AL (Tdurum_contig29607_413) were highly predictive in three independent validation populations of 1065, 1001, and 175 breeding lines. Lines with the resistant haplotype at both loci were nearly free of stipe rust symptoms. By using mixed linear models with those markers as fixed effects, we could increase predictive ability in the three populations by 0.13–0.46 compared to a standard genomic best linear unbiased prediction approach. The obtained results facilitate an efficient selection for stripe rust resistance against the current pathogen population in the Northern and Central European winter wheat gene pool.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stripe or yellow rust is a major disease in bread wheat (Triticum aestivum L.) production under temperate climatic conditions or at high altitudes with a significant impact on grain yield and end-use quality characteristics (Singh et al. 2016). It is caused by Puccinia striiformis Westend f. sp. tritici (Pst), which is a biotrophic and heteroecious fungus. The pathogen is a common fungal disease of cereals and grasses, and various Berberis species can serve as alternate hosts (Hovmøller et al. 2011). Depending on various factors such as disease duration, infection stage, speed of disease development, susceptibility of cultivars, and favorable climatic conditions, yield reductions range from 10 to 40% (13% on average, Laidig et al. 2021) and can be as high as 100% if infection occurs at the seedling stage and persists until maturity (Afzal et al. 2007; Pradhan et al. 2020). Although stripe rust can be controlled with fungicides, economical, sustainable and environmentally sound management of this disease is needed. Breeding of resistant cultivars can be facilitated through identification, introduction, and subsequent selection of effective rust resistance genes during breeding cycles in wheat.
Fungal disease resistance genes in crop plants can generally be divided into all-stage resistance (ASR) genes and adult plant resistance (APR) genes. ASR genes are effective against avirulent pathotypes at all growth stages of the plant, are also referred to as race-specific, are inherited qualitatively, and can be overcome by new races. APR genes, on the other hand, express resistance only at post-seedling stages, are considered race-nonspecific, are inherited quantitatively, and tend to be durable (Bariana 2003; Rosewarne et al. 2013; Zetzsche et al. 2019). Because resistance levels in this group can be highly affected by temperature, it also includes high-temperature adult-plant (HTAP) resistance genes (Chen 2013). Since strong selection pressure is exerted on the pathogen to become virulent against a single ASR gene and thus survive, the use of a combination or pyramiding of APR genes with effective ASR genes is recommended to achieve commercially acceptable levels of resistance (Bariana 2003; Gessese et al. 2021).
The use of fungicides and the cultivation of resistant varieties have prevented devastating epidemics in Europe in the past, while genetic resistance is still the most effective and sustainable approach. In Europe, stripe rust occurred infrequently due to the use of a few key resistance genes with long-term efficacy such as Yr1, Yr2, Yr3, Yr4, Yr6, Yr9, Yr15, Yr17, Yr25, and Yr32 (Hovmøller 2007; Hovmøller et al. 2016). In 2011, "Warrior," a new virulent stripe rust strain from the near-Himalayan region (Hovmøller et al. 2016), emerged simultaneously in several European countries and spread rapidly across much of the continent. According to monitoring by the Julius Kühn Institute (JKI, Germany), the Pst race "Warrior (-)" dominates the European yellow rust population, followed by the races "Triticale2015" and the original "Warrior," as well as a new race "PstS15," which was first discovered in 2020 (Flath et al. 2021). It seems that only a few resistance genes, including Yr5, Yr10, Yr15, and Yr27, are still effective against these races in European wheat (K. Flath 2022, JKI, personal communication).
The chromosomal positions of several stripe rust resistance genes and quantitative trait loci (QTL) were determined using classical mapping approaches (Tsomin et al. 1990; Bariana and McIntosh 1993; Michelmore et al. 1991; Xu et al. 2008; Edae et al. 2016; Gessese et al. 2021) and, more recently, using genome-wide association studies (GWAS) (Bouvet et al. 2021; Rollar et al. 2021). To complement traditional QTL mapping, GWAS combined with high-density SNP genotyping have been successfully used as powerful tools for discovery of stripe rust resistance loci in a global collection of winter wheat accessions (Bulli et al. 2016), spring wheat landraces (Kankwatsa et al. 2017), diverse Indian spring wheat cultivars (Kumar et al. 2020), US winter wheat cultivars and breeding lines (Mu et al. 2020), elite wheats of the International Maize and Wheat Improvement Center (CIMMYT) (Juliana et al. 2018), European winter wheat (Miedaner et al. 2019), and modern Chinese wheat (Jia et al. 2020).
Based on the “Catalogue of Gene Symbols for Wheat” available to date (https://wheat.pw.usda.gov/GG3/WGC), 85 formally named and more than 300 tentatively named genes or QTL located on different wheat chromosomes have been reported for stripe rust resistance (Maccaferri et al. 2015; Bulli et al. 2016; McIntosh et al. 2017; Mu et al. 2020). Of these, only a few genes, namely Yr5/YrSP and Yr7 (Marchal et al. 2018), Yr15 (Klymiuk et al. 2018), Yr10 (Liu et al. 2014), Yr18 (Krattinger et al. 2009), Yr27 (Athiyannan et al. 2022), Yr36 (Fu et al. 2009), Yr46 (Moore et al. 2015), and YrAS2388R (Zhang et al. 2019) have been cloned and functionally characterized. Some chromosomes, such as 1B, 2A, 2B, and 7B, harbor a substantial number of genes or QTL that confer different types and degrees of resistance to stripe rust (Maccaferri et al. 2015). Although some race-nonspecific resistance genes are effective and durable against the new Pst races (Abou-Zeid and Mourad 2021), virulent Pst races have emerged against most Yr genes, rendering them ineffective (Maccaferri et al. 2015). Thus, continued efforts are needed to characterize new resistance sources for maintaining or increasing resistance levels. Compared to wild relatives of wheat, identification and characterization of resistance genes from landraces and cultivated genotypes is more preferable due to the absence of undesirable agronomic traits and chromosomal linkage drags in the latter (Burt et al. 2014; Gessese et al. 2021). The selection and combination of resistance genes already present in the advanced breeding pool allows for a faster development of new resistant varieties.
Classical phenotypic selection for Pst resistance is resource demanding, and its success strongly depends on environmental factors. The prediction of breeding values using molecular markers has become a promising approach to facilitate genomic selection. Marker-based prediction of breeding values can be categorized into marker-assisted selection (MAS), which makes use of a preselected set of markers associated with important resistance genes, and genomic selection (GS), which is based on genome-wide marker information. Previous studies have shown the potential of both MAS and GS for the prediction of Pst resistance, but comparisons of predictive abilities of the two approaches do not yet allow a clear conclusion about the optimal prediction method (Juliana et al. 2017; Muleta et al. 2017; Beukert et al. 2020b).
Compared to previous studies (Miedaner et al. 2019; Beukert et al. 2020a; Bouvet et al. 2021; Rollar et al. 2021) that used European winter wheat materials released in a broader time period, the present work identified QTL for stripe rust resistance in a collection of 230 current Central and Northern European winter wheat, examined the effects of these QTL in practical breeding programs and suggested prediction models for breeding cultivars with improved stripe rust resistance. Selection and combination of alleles already present in the advanced breeding pool is essential for improving durable resistance to stripe rust and for developing new wheat varieties. Specifically, the objectives of our study were to (1) perform GWAS in modern wheat germplasm using disease responses to current Pst populations assessed in field trials, (2) identify the sources of effective resistance alleles and associated QTL for use in breeding programs, (3) validate QTL in breeding materials, (4) compare the resistance loci identified in this study with previously reported Yr genes and QTL. Our results will help to understand the genetic basis of stripe rust resistance in Northern and Central European winter wheat and facilitate improvement of stripe rust resistance through MAS and GS.
Materials and methods
Plant materials and stripe rust assessment
With the aim to capture wide genetic variability across Europe, we selected a population of 230 winter wheat cultivars and breeding lines for the GWAS. Using the breeder´s knowledge and the coefficient of determination algorithm (Akdemir et al. 2021), we selected genotypes from Germany, Austria, Norway, Sweden, Denmark, Poland, and Switzerland comprising 157, 50, 14, 4, 3, 1, and 1 genotype(s), respectively (Table S1).
The 230 genotypes of the association panel were evaluated for stripe rust resistance at the adult plant stage in field experiments. Each entry was sown in two-row microplots 0.5–1.5 m long and 0.17–0.3 m wide with approximately 50 grains per row in October each year. Field trials were carried out at Lemgo and Lenglern in Germany, and in Tulln and Reichersberg in Austria in 2020 and 2021 (Table S2). We used non-replicated trials in favor of multiple sites, except for Tulln, where a randomized complete block trial with two replicates was conducted. To validate the results of this study, using same sowing conditions, two F6 populations of 1065 and 1001 German breeding lines from two consecutive breeding cycles were evaluated in Lemgo in 2020 and 2021, respectively, and another independent population of 175 breeding lines was evaluated in Lenglern in 2021. Field responses to either natural or artificial inoculation with Pst isolates (Table S1) were recorded between plant heading (Zadoks 50) and grain filling (Zadoks 80) stages when most flag leaves of susceptible controls had disease severity of at least 50%. Disease severity as a percentage of infected leaf area was scored 3–5 times in June using the modified Cobb scale (Peterson et al. 1948) and means for the recorded scores for stripe rust were used in the analyses.
Statistical analysis
Implementation of the coefficient of determination algorithm for selection of the genotypes was done in R package "TrainSel" (R Core Team, 2017). After testing several transformation methods to meet the assumption of normality of residuals, the arcsine square root transformation method (Maccaferri et al. 2015) was chosen to prepare phenotypic data for association analysis. Analyses of variance (ANOVA) and estimation of adjusted means across environments were conducted using "PROC GLM" of the SAS statistical package v.9.4 (SAS Institute, Cary, USA) by applying the following model:
where yij is the phenotypic value of genotype i in trial j, µ is the overall mean, gi is the fixed effect of genotype i, t is the random effect of trial/environment j and eij is the random error, which was confounded with genotype × environment effects. The heritability coefficient was estimated according to the formula
where \(\sigma_{{\text{G}}}^{2}\) is the genotypic variance, \(\sigma_{{\text{e}}}^{2}\) is the residual variance, and T is the number of trials. Variance components for estimating h2 were derived from the above-mentioned model assuming a random genotypic effect.
SNP genotyping
Genomic DNA of each genotype belonging to the association panel and validation populations was extracted from young leaf tissue according to the procedure of Plaschke et al. (1995). Genotyping was performed using the 25 K Infinium iSelect array (TraitGenetics, Seeland OT Gatersleben, Germany), which is an extension of 20 K array (Arif et al. 2021) plus 5000 Axiom array markers and candidate genes. SNP markers on the 25 K array were selected from the 90 K Infinium array, the 35 K Axiom wheat breeder array, and two proprietary TraitGenetics wheat arrays (135 K Axiom and 12 K Infinium). These markers were also selected based on the analysis of more than 2500 wheat cultivars, accessions and breeding lines according to the quality of performance on each analysis platform, their polymorphism information content (LD to flanking SNPs, MAF) and their position on the wheat reference genome (Dr. Jörg Plieske, TraitGenetics, personal communication). The physical position of the markers was determined by BLAST and e-value cut off for the highest probability match using the published genome sequence of Chinese Spring (IWGSC RefSeq v1.0). SNPs were filtered to avoid producing redundant genotyping information. The monomorphic SNPs and those with more than 10% missing values and minor allele frequency of less than 5% were excluded from further analysis using the "synbreed" package (Wimmer et al. 2012) in R (R Core Team 2017). Heterozygous marker signals were treated as missing data. Chromosomal positions of these SNPs were obtained from the 90 K consensus map (Wang et al. 2014).
Population structure
The genetic structure of the 230 genotypes was determined using the Bayesian clustering program STRUCTURE (Pritchard et al. 2000). The output of STRUCTURE was analyzed in STRUCTURE HARVESTER (Earl and Vonholdt 2012) to determine the possible number of subpopulations using the ΔK ad hoc statistic. The best K value representing the optimal number of clusters in the populations was estimated as ΔK based on the rate of change of log-likelihood of data between successive values, as described by Evanno et al. (2005). Population structure was also analyzed by principal component analysis (PCA) to distinguish different groups using the "PROC PCA" in the SAS statistical package.
Association mapping
Linkage disequilibrium (LD) between markers was estimated for the association mapping panel using observed versus expected allele frequencies in TASSEL 5.2.78 (Bradbury et al. 2007). LD decay was measured as the distance at which the average R2 value between pairwise SNPs fell to half its maximum value. Both Bayesian clustering method of STRUCTURE and PCA revealed a population structure in the panel. Marker-trait association in each environment and across environments were carried out using a mixed linear model that accounts for population structure (Q) and kinship matrix (K). The model can describe as follows:
where y is the vector of observations, β is a vector containing fixed effects for genetic markers and population structure (Q), u is a vector of random additive genetic effects from multiple background QTL with u ~ N(0, \(\sigma_{{\text{G}}}^{2}\) K), X and Z are the design matrices, and e is a vector of random residuals with e ~ N(0, \(\sigma_{{\text{G}}}^{2}\) I). To provide the adjusted p-values, the false discovery rate (FDR) was calculated, using a threshold of < 5%, with the "q-value" package in R (R Core Team, 2017). To evaluate the performance of the models and appropriate thresholds, QQ plots drawn by TASSEL were inspected. Associations of SNP markers with stripe rust severity in each environment and across environments are represented as Manhattan plots. The physical position of significantly associated markers across environments was compared to previously published Yr genes and QTL using the “Catalogue of Gene Symbols for Wheat” (https://wheat.pw.usda.gov/GG3/WGC) and an integrated map for chromosomal positions of loci associated with reactions to Pst constructed by Bulli et al. (2016) and Maccaferri et al. (2015).
Validation of associated markers
To detect allelic effect of associated SNP markers identified by GWAS in validation populations, the Student’s t-test was applied for evaluating statistically significant differences (P ≤ 0.05) between means of two allelic groups belonging to each locus of interest for disease severity.
Putative candidate gene identification
Candidate genes for validated loci were identified by blasting the sequences of the markers on the corresponding chromosomes of the International Wheat Genome Sequencing Consortium RefSeq v2.1 (https://urgi.versailles.inrae.fr/blast_iwgsc/?dbgroup=wheat_iwgsc_refseq_v2.1_chromosomes&program=blastn; Zhu et al., 2021) to retrieve the gene identifiers within a window of 2 Mb (1 Mb upstream and downstream) from the peak of each targeted marker. Gene ontology terms were obtained from EnsmblPlants using the biomaRt package (Durinck et al. 2009).
Prediction of stripe rust resistance
The predictive ability of different linear regression models was evaluated using adjusted phenotypic mean values after arcsine square root transformation as response variable. Only a common set of SNP markers that remained after filtering in the GWAS panel and the three validation panels was considered for prediction. Linear regression using the ordinary least squares (OLS) method was performed using the "stats" package (R Core Team 2017). Following the matrix notation of the GWAS model shown above, the OLS model was fitted considering only fixed marker effects (β) and a random error (e). Genomic best linear unbiased prediction (GBLUP) models were fitted with rrBLUP (Endelman 2011) applying a model similar to the GWAS model, with the only exception of β, which only contained fixed marker effects without effects of population structure. To avoid collinearity due to linkage among detected markers, we discarded markers with a variance inflation factor > 5, from the design matrix X using the R package “car” (Fox and Weisberg 2019). Predictive ability was defined as the correlation between estimated breeding values and observed phenotypic values. Predictive ability was first estimated within the GWAS panel using a five-fold cross-validation with 200 replications (Ntraining = 184, Ntest = 46). The predictive ability of these models was also evaluated in the three independent validation populations using the complete GWAS panel for model training.
Results
Phenotypic evaluation for stripe rust
Depending on the environmental conditions and the Pst pathotypes present in the different field trials, a wide variation was observed for the severity of stripe rust in the panel in each environment (ranging from 0 to 87.5%). The four breeding lines TS085, TS175, TS185, and TS220 and three cultivars TS140 (Gentleman), TS173 (Tobak) and TS195 (Sinatra) from Germany and the cultivar TS027 (Mariboss) from Norway were completely resistant in different field trials and across environments (Table S1). The cultivars TS012 (Rida) from Norway and TS014 (Akteur) from Germany were highly susceptible in this panel (Table S1). Stripe rust severity was left skewed in each experiment and for means across experiments (Table 1; Fig. S1.A). Significant (P < 0.05) and positive correlations ranging from 0.57 to 0.76 were observed between all pairs of test environments (Table 2), with the highest correlation (0.76) between Lemgo 20 and Lemgo 21, while the lowest correlation (0.57) was found between Lenglern 21 and Reichersberg 21.. Normality of residuals was achieved by applying the arcsine square root transformation method (Fig. S1.B). ANOVA revealed significant differences (P < 0.01) among the genotypes. The estimate of genetic variance (\(\sigma_{{\text{G}}}^{2}\) = 0.026) contributed to a high broad-sense heritability for stripe rust resistance (h2 = 0.87; \(\sigma_{{\text{T}}}^{2}\) = 0.017; \(\sigma_{{\text{e}}}^{2}\) = 0.019).
Genotyping, population structure and LD decay
After marker filtering, 8812 informative and polymorphic SNP markers (Table S3) with an average minor allele frequency of 0.26 were used for analyzing population structure, LD, and genetic association. Of these, 3390, 3977, and 1445 markers belonged to subgenomes A, B, and D, respectively, with most markers (683) found on chromosome 5B and the fewest markers (70) on chromosome 4D. All SNPs were physically anchored to the reference sequence of Chinese Spring wheat.
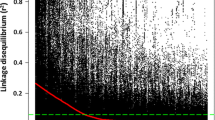
Two subpopulations were detected in the winter wheat panel of 230 genotypes (Fig. S2). The number of subpopulations (K) was estimated based on the rate of change of the log-likelihood of the data between successive K values. In the plot of K versus ΔK, a reduction in the slope was observed at ΔK = 2 (Fig. S2A). Therefore, the panel was divided into two subgroups based on the corresponding population membership coefficients (Q) of the individuals, with subgroup 1 containing 92 genotypes mainly from Austria and subgroup 2 containing 138 genotypes mainly from Germany Norway, Sweden, Denmark, Poland, and Switzerland (Fig. S2B). PCA also classified the panel into two subpopulations (Fig. 1). PC1 and PC2 accounted for 20% and 10% of the total marker-based variation, respectively. In the LD analysis (Fig. S3), patterns between significantly associated markers were determined using the squared Pearson correlation coefficient (R2) between SNP markers as a function of physical map position between markers. As shown in the scatter plot (Fig. S3), the strength of LD due to linkage decreases as the physical distance between SNPs in the genome increases. LD decay reached R2 = 0.2 at 200 Mb.
GWAS and candidate genes for stripe rust resistance
GWAS using a mixed linear model identified 67 different loci on 12 wheat chromosomes, namely 1A, 2A, 2B, 3A, 3B, 4B, 5B, 5D, 6A, 7A, 7B, and 7D which were significantly associated with stripe rust disease severity assessed in at least one trial (Table 3; Fig. S2). The QQ plots evaluating the performance of the mixed linear models indicated a high corrective effect of the GWAS model (Fig. S4). The percentage of explained phenotypic variance (R2) of the associated markers ranged from 5 to 11%, while the effect size of those markers ranged from − 0.37 to 0.28 (Table 3). The strongest association (R2 = 11%, p value = 0.0001) was found for the SNP markers Jagger_c1423_102 and GENE-4167_145 on chromosome 6A in Lemgo 21 (Table S4). Resistant allele G at marker locus IAAV1743 and susceptible allele G at marker locus RAC875_c1226_652 both on chromosome 2B showed the maximum effects (0.37 and 0.28, respectively) on stripe rust severity identified in Lemgo 20 and Reichersberg 21 field trials, respectively (Table S4). Of the 67 markers, 12 SNP markers on chromosomes 2B, 4B, and 6A, 7A, 7B, and 7D displayed significant associations for means across environments (Table 4). Four SNP markers on chromosomes 2B (Jagger_c6853_60), 7A (BS00093016_51), 7B (AX-95154562), and 7D (AX-94720261) were significantly associated solely with the average stripe rust severity across all field trials (Table 3). In addition, eight SNP markers were found to be common (Table S3; Table 4) in at least two environments on chromosomes 2B (RAC875_c1226_652, IAAV1743, Ra_c6266_136), 4B (AX-94684920), and 6A (Tdurum_contig29607_413, Jagger_c1423_102, GENE-4167_145, BS00040814_51).
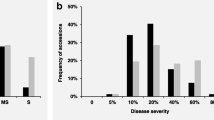
Notably, the association of only two markers, namely RAC875_c1226_652 on chromosome 2B and Tdurum_contig29607_413 on chromosome 6A, with stripe rust resistance were validated by Student’s t-test (P ≤ 0.001) in three validation populations of 1065, 1001, and 175 breeding lines assessed for disease severity in Lemgo, Germany, in 2020 and 2021 and in Lenglern, Germany, in 2021. For the marker RAC875_c1226_652, the G allele and for the marker Tdurum_contig29607_413, the T allele contributed to a 10–18% and 8–17% reduction in stripe rust disease severity in the validation populations, respectively (Fig. 2). Lines in the validation that harbored the resistance improving alleles at both markers (Fig. 2), showed a highly to completely resistant phenotype (0–1.6% stripe rust severity) compared to haplotypes with the “susceptible” alleles (10–35% stripe rust severity). Interestingly, only one breeding line (TS128) in the GWAS panel possessed this allele combination.
Effects of allelic combination of the markers located on chromosomes 2B (RAC875_c1226_652, A and G alleles) and 6A (Tdurum_contig29607_413, C and T alleles) on disease severity (%) in validation populations of a 1065 and b 1001 breeding lines evaluated in Lemgo in 2020 and 2021, respectively, and c 175 breeding lines evaluated in Lenglern in 2021. The more susceptible alleles are shown in yellow. The number of lines in each group is presented at the top of each box plot
To further investigate the validated loci, sequences of the two SNP markers RAC875_c1226_652 and Tdurum_contig29607_413 were aligned to the physical map of the reference genome to search for annotated genes. As a result, 20 to 68 putative candidate genes (Table S5) were identified in a 2-Mb physical interval around the peak markers on chromosomes 2B and 6A, respectively. The genes TraesCS2B02G182800 and TraesCS6A02G399600, which were directly tagged by these markers, encode a putative disease resistance protein RGA4 (LOC123044110), BST_chr2B_nlr_143 and the E3 ubiquitin protein ligase RHB1A (LOC123129052), respectively, and are known to be involved in disease resistance and defense mechanisms in wheat.
Finally, the corresponding physical interval of the associated loci for means across environments identified in the present study were compared with previously reported Yr genes and QTL (Table S6). The physical interval of the compared QTL showed overlap with eighteen QTL and six genes including Yr27, Yr5, Yr44, YrSp, Yr62, and Yr18.
Predicting resistance to stripe rust
The potential of MAS and GS to select for Pst resistant wheat was evaluated using (1) 6846 genome-wide SNP markers that were common between all four data sets, (2) the set of 12 markers identified in the present GWAS based on transformed means across environments, and (3) the two associated markers that were also significant in the three validation panels. Two of the 12 markers (Ra_c6266_136 and GENE-4167_145) were excluded from all fixed effect matrices to avoid collinearity resulting from linkage. Five-fold cross-validation of ordinary least squares (OLS) and GBLUP models within the GWAS panel confirmed the association between the remaining ten markers and Pst resistance, whereas a reduced model with only RAC875_c1226_652 and Tdurum_contig29607_413 as predictors resulted in a lower predictive ability (Table 5). In this population, the highest predictive ability was achieved using a GBLUP model with all ten detected markers included as fixed effects. Predicting breeding values in three independent validation panels showed that OLS and GBLUP including the QTL-linked markers as covariates outperformed the standard GBLUP model. Contrary to the cross-validation within the GWAS data set, the estimated breeding values of the three validation panels were more accurate with only RAC875_c1226_652 and Tdurum_contig29607_413 as fixed effects compared to models including all ten associated markers as predictors. In these independent data sets, the highest predictive ability was achieved using either OLS or GBLUP treating the two above-mentioned markers as fixed effects. With these models, we could increase predictive ability in the three populations by 0.13–0.46 compared to the standard GBLUP approach.
Discussion
Host plant resistance is generally the most ecological and economic strategy for stripe rust disease control. Integration of additional resistance alleles into the genetic background of already resistant cultivars is important for improving the durability of stripe rust resistance in wheat breeding. Understanding the genetic basis of stripe rust resistance could facilitate the transfer of existing or new resistance alleles into high-yielding and regionally adapted bread wheat lines.
In the present study, the frequency distribution of the percentage of disease severity assessed in the field trials was right skewed with a large proportion of resistant lines (Fig. S1.A), suggesting that many effective APR and possibly race-nonspecific alleles may be present in this panel. Similar results were obtained when the disease severity of stripe rust was assessed in a European winter wheat diversity panel of 158 old and new wheat cultivars (Miedaner et al. 2020) and in 419 pre-breeding lines developed at CIMMYT (Ledesma-Ramírez et al. 2019). All these studies indicated that this type of resistance is of high priority in breeding programs: it is established through the targeted crossing of resistant lines followed by high selection pressure in early generations. In addition, the broad-sense heritability coefficient (h2 = 0.87) obtained in this study suggests that phenotypic variation in stripe rust severity was mainly due to genotypic effects. High broad-sense heritability values have been reported in previous studies by Ling et al. (2012), Beukert et al. (2020a), and Abou-Zeid and Mourad (2021).
GWAS identified 67 QTL for stripe rust resistance in a panel of 230 winter wheat cultivars and breeding lines. Among all marker loci identified in five environments, only eight SNPs were found significant in at least two environments, therefore could be considered as stable QTL. Four SNPs were found significantly associated with stripe rust resistance for means across environments, but not in individual environments. Among these, two QTL appeared particularly attractive for stripe rust resistance breeding, namely those predicted by markers RAC875_c1226_652 (chromosome 2B) and Tdurum_contig29607_413 (chromosome 6A). Lines in the validation populations combining the favorable alleles at these loci displayed a near immune phenotype.
The results of the present study revealed QTL close to the genomic regions of the stripe rust resistance genes Yr5/YrSP (Macer 1966; Murphy et al. 2009; Feng et al. 2015) and Yr44 (Sui et al. 2009) on chromosome 2B, Yr62 (Lu et al. 2014) on chromosome 4B, and Yr18 (Singh 1992) on chromosome 7D near the SNP markers IAAV1743, AX-94684920, and AX-94720261, respectively. Yr5 provides a high level of resistance to stripe rust and originates from spring spelt Triticum spelta var. album (Macer 1966). It is allelic to YrSP and paralogous to Yr7, both of which have been overcome by several Pst isolates. A QTL (qYr.A) in the same genomic region was previously reported by Losert et al. (2017) in a diverse set of 919 triticale genotypes from the private and public breeding sectors in Europe. In this region, five QTL controlling stripe rust resistance (Table S6) have been reported in common wheat (Guo et al. 2008; Jighly et al. 2015; Ando et al. 2018; Ren et al. 2012; Tehseen et al. 2021).
We found a QTL on chromosome 4B in the region of the Yr62 gene. Lu et al. (2014) found a QTL on chromosome 4BL (QYrPI192252.wgp-4BL) in a mapping population of 150 F5 recombinant inbred lines (derived from a cross between PI 192252 and “Avocet susceptible”) which explained 40–60% of the total phenotypic variation of the relative area under the stripe rust disease progress curve and was inherited as a single gene. The gene, named Yr62, provides a high level of HTAP resistance and was located proximal to Yr50, transferred from T. intermedium into wheat. Jia et al. (2020) and Naruoka et al. (2015) also reported two QTL for stripe resistance in a similar region on chromosome 4BL.
We discovered a QTL in the Lr34/Yr18 region on chromosome 7DS. Cloning of Lr34/Yr18 has shown that the gene encodes an adenosine triphosphate-binding cassette transporter that resembles a pleiotropic drug resistance transporter (Krattinger et al. 2009). Lr34/Yr18 is active at the adult plant stage and shows moderate but durable resistance to stripe and leaf rust (Spielmeyer et al. 2005). It appears that the gene is relatively common in German cultivars (Zetzsche et al. 2020). However, further investigations are needed to confirm whether the QTL identified in the present study on chromosomes 2BL, 4BL, and 7DS are allelic to or distinct from Yr5/YrSP, Yr44, Yr62, and Yr18, respectively.
Our study also detected four loci associated with stripe rust resistance on chromosomes 2BS, 6AL, 7AL, and 7BL (Table 4), which could correspond to QTL previously reported by Prins et al. (2011), Vazquez et al. (2012), Rosewarne et al. (2012), Agenbag et al. (2014), Miedaner et al. (2019), Beukert et al. (2020a), Jia et al. (2020), and Rollar et al. (2021). Of these, QTL on chromosomes 2BS and 6AL were identified in German plant materials including a winter wheat diversity panel (Miedaner et al. 2019), a hybrid wheat panel (Beukert et al. 2020a), and a multiparental population (Rollar et al. 2021). Interestingly, these two QTL were not reported in the worldwide collections of hexaploid spring (Maccaferri et al. 2015) and winter wheat (Bulli et al. 2016), suggesting that they were specifically enriched through European breeding activities.
On chromosome 2B, the putative QTL associated with RAC875_c1226_652 was in a region that referred to BST_chr2B_nlr_143 and disease resistance protein RGA4. In this region, a QTL (QYr.sgi-2B.1) was previously mapped by Agenbag et al. (2014) near the marker IWB52095, located at 157.694 Mbp of the wheat physical map (IWGSC Refseq v.1). More recently, the QTL was cloned in bread wheat as a major factor for the race-specific disease resistance gene Yr27, which encodes an intracellular immune receptor (Athiyannan et al. 2022). Yr27 is allelic to leaf rust resistance gene Lr13 with 97% sequence identity. The predicted coding sequence of the gene with a length of 3219 base pairs encodes a protein of 1072 amino acids with an N-terminal coiled-coil domain, a central NB-ARC domain, and a carboxy-terminal leucine-rich repeat (LRR) domain (Athiyannan et al. 2022). In rice, resistance to Magnaporthe oryzae is mediated by a pair of interacting nucleotide-binding site leucine-rich repeat domain-containing immune sensors, RGA4 and RGA5 (Césari et al. 2014). RGA4 mediates cell death but is repressed by RGA5. The repressor is neutralized by binding pathogen-derived proteins to the dimer. RGA4 and RGA5 interact through their CC domains to form homo- and heterocomplexes. In addition, BLAST searches using the sequence of another SNP marker on chromosome 6AL (Tdurum_contig29607_413) yielded direct hits for a gene annotated as E3 ubiquitin ligase protein, which is a module that controls innate immunity and programmed cell death in plants and strongly contributes to promoting antimicrobial defense while preventing autoimmunity (You et al. 2016). However, the mechanisms contributing to this immune homeostasis are poorly understood. Understanding the mode of action between candidate genes and their effects on disease resistance can help in the development of functional and predictive markers to detect resistant. It also provides new insights into the genetic mechanisms controlling stripe rust resistance in wheat and lays the foundation for characterization, cloning, and manipulation of genes in future studies. Since most of the genes/QTL identified in this study are already present in adapted varieties and/or elite breeding lines, pyramiding of these genes for breeding new resistant varieties seems relatively straightforward.
The markers identified in the present study showed great potential to facilitate an efficient selection for Pst resistance. OLS and GBLUP models that included RAC875_c1226_652 and Tdurum_contig29607_413 as fixed effects allowed a more accurate prediction of breeding values in the three independent panels compared to a GBLUP model with only the intercept as fixed effect. More complex models with all ten QTL-linked markers as fixed effects were only superior within the GWAS panel but yielded comparably low predictive abilities in the independent validation panels, which can be explained by overfitting. The value of preselected markers for the prediction of Pst resistance was evaluated in previous studies, but the comparison with genome-wide markers did not yield consistent results (Juliana et al. 2017; Muleta et al. 2017; Beukert et al. 2020b). These differing outcomes could be attributed to the size and composition of the training set, the genetic architecture of Pst resistance in the analyzed populations, and the prevalent pathotypes. Nevertheless, the present study demonstrates that, for the Northern and Central European winter wheat gene pool and current pathogen races, the validated QTL on chromosomes 2B and 6A and associated markers appear highly attractive to facilitate selection of Pst resistant cultivars. Since GBLUP with fixed effects for RAC875_c1226_652 and Tdurum_contig29607_413 was superior to OLS, we recommend including these markers in GS models to enrich quantitative, minor-effect alleles for a more sustainable resistance in combination with large-effect alleles.
Overall, our study identified resistant genotypes and the potential source of effective resistance alleles and associated QTL that could be used to improve stripe rust resistance levels in the current Northern and Central European breeding materials, particularly through genomic approaches.
Data availability
All datasets generated for this study are included in the article/Supplementary Material.
Abbreviations
- ANOVA:
-
Analysis of variance
- APR:
-
Adult plant resistance
- ASR:
-
All-stage resistance
- CIMMYT:
-
International Maize and Wheat Improvement Center
- FDR:
-
False discovery rate
- GBLUP:
-
Genomic best linear unbiased prediction
- GS:
-
Genomic selection
- GWAS:
-
Genome-wide association study
- HTAP:
-
High-temperature adult-plant
- LD:
-
Linkage disequilibrium
- LRR:
-
Leucine-rich repeat
- MAS:
-
Marker-assisted selection
- OLS:
-
Ordinary least squares
- PCA:
-
Principle component analysis
- Pst :
-
Puccinia striiformis
- QTL:
-
Quantitative trait loci
- SNP:
-
Single nucleotide polymorphism
References
Abou-Zeid MA, Mourad AMI (2021) Genomic regions associated with stripe rust resistance against the Egyptian race revealed by genome-wide association study. BMC Plant Biol 21:42. https://doi.org/10.1186/s12870-020-02813-6
Afzal SN, Haque MI, Ahmedani MS, Bashir S, Rattu AR (2007) Assessment of yield losses caused by Puccinia striiformis triggering stripe rust in the most common wheat varieties. Pak J Bot 39:2127–2134
Agenbag GM, Pretorius ZA, Boyd LA, Bender CM, MacCormack R et al (2014) High-resolution mapping and new marker development for adult plant stripe rust resistance QTL in the wheat cultivar Kariega. Mol Breed 34:2005–2020. https://doi.org/10.1007/s11032-014-0158-4
Akdemir D, Rio S, Isidro y Sánchez J (2021) TrainSel: an R package for selection of training populations. Front Genet 12:655287. https://doi.org/10.3389/fgene.2021.655287
Ando K, Rynearson S, Muleta KT, Gedamu J, Girma B et al (2018) Genome-wide associations for multiple pest resistances in a Northwestern United States elite spring wheat panel. PLoS ONE 13:e0191305. https://doi.org/10.1371/journal.pone.0191305
Arif MAR, Shokat S, Plieske J, Ganal M, Lohwasser U et al (2021) A SNP-based genetic dissection of versatile traits in bread wheat (Triticum aestivum L.). Plant J 108(4):960–976. https://doi.org/10.1111/tpj.15407
Athiyannan N, Abrouk M, Boshoff WHP, Cauet S, Rodde N et al (2022) Long-read genome sequencing of bread wheat facilitates disease resistance gene cloning. Nat Genet 54:227–231. https://doi.org/10.1038/s41588-022-01022-1
Bariana HS (2003) Breeding for disease resistance. In: Murray BG, Murphy DJ (eds) Encyclopedia of applied plant sciences. Academic Press, Oxford, pp 244–253
Bariana HS, McIntosh RA (1993) Cytogenetic studies in wheat. XV. Location of rust resistance genes in VPM1 and their genetic linkage with other disease resistance genes in chromosome 2A. Genome 36:476–482. https://doi.org/10.1139/g93-065
Beukert U, Liu G, Thorwarth P, Boeven PHG, Longin CFH et al (2020a) The potential of hybrid breeding to enhance leaf rust and stripe rust resistance in wheat. Theor Appl Genet 133:2171–2181. https://doi.org/10.1007/s00122-020-03588-y
Beukert U, Thorwarth P, Zhao Y, Longin CFH, Serfling A et al (2020b) Comparing the potential of marker-assisted selection and genomic prediction for improving rust resistance in hybrid wheat. Front Plant Sci 11:594113. https://doi.org/10.3389/fpls.2020.594113
Bouvet L, Percival-Alwyn L, Berry S, Fenwick P, Campos Mantello C et al (2021) Wheat genetic loci conferring resistance to stripe rust in the face of genetically diverse races of the fungus Puccinia striiformis f. sp. tritici. Theor Appl Genet 135:301–319. https://doi.org/10.1007/s00122-021-03967-z
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y et al (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635. https://doi.org/10.1093/bioinformatics/btm308
Bulli P, Zhang J, Chao S, Chen X, Pumphrey M (2016) Genetic architecture of resistance to stripe rust in a global winter wheat germplasm collection. G3 6:2237–2253. https://doi.org/10.1534/g3.116.028407
Burt C, Griffe LL, Ridolfini AP, Orford S, Griffiths S et al (2014) Mining the Watkins collection of wheat landraces for novel sources of eyespot resistance. Plant Pathol 63:1241–1250. https://doi.org/10.1111/ppa.12221
Césari S, Kanzaki H, Fujiwara T, Bernoux M, Chalvon V et al (2014) The NB-LRR proteins RGA4 and RGA5 interact functionally and physically to confer disease resistance. EMBO J 33:1941–1959. https://doi.org/10.15252/embj.201487923
Chen XM (2013) High-temperature adult-plant resistance, key for sustainable control of stripe rust. Am J Plant Sci 4:608–627. https://doi.org/10.4236/ajps.2013.43080
Durinck S, Spellman PT, Birney E, Huber W (2009) Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package. Nat Protocols 4:1184–1191. https://doi.org/10.1038/nprot.2009.97
Earl DA, Vonholdt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Edae EA, Olivera PD, Jin Y, Poland JA, Rouse MN (2016) Genotype-by-sequencing facilitates genetic mapping of a stem rust resistance locus in Aegilops umbellulata, a wild relative of cultivated wheat. BMC Genomics 17:1039. https://doi.org/10.1186/s12864-016-3370-2
Endelman JB (2011) Ridge regression and other kernels for genomic selection with R package rrBLUP. Plant Genome 4:250–255. https://doi.org/10.3835/plantgenome2011.08.0024
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Feng JY, Wang MN, Chen XM, See DR, Zheng YL et al (2015) Molecular mapping of YrSP and its relationship with other genes for stripe rust resistance in wheat chromosome 2BL. Phytopathology 105:1206–1213. https://doi.org/10.1094/PHYTO-03-15-0060-R
Flath K, Schulz P, Klocke B (2021) RustWatch – das erste Frühwarnsystem für Getreideroste in Europa. In: Julius Kühn-Institut (Hrsg) 62. Deutsche Pflanzenschutztagung: Gesunde Pflanzen in Verantwortung für unsere Welt. 21.–23. September 2021, Julius-Kühn-Archiv 467, Quedlinburg, pp 211–212
Fox J, Weisberg S (2019) An R companion to applied regression, 3rd edn. Sage, Thousand Oaks
Fu D, Uauy C, Distelfeld A, Blechl A, Epstein L et al (2009) A kinase-START gene confers temperature-dependent resistance to wheat stripe rust. Science 323:1357–1360. https://doi.org/10.1126/science.1166289
Gessese M, Miah H, Bansal U, Bariana H (2021) Genetics of stripe rust resistance in a common wheat landrace Aus27492 and its transfer to modern wheat cultivars. Can J Plant Pathol 43:256–262. https://doi.org/10.1080/07060661.2021.1979657
Guo Q, Zhang ZJ, Xu YB, Li GH, Feng J et al (2008) Quantitative trait loci for high-temperature adult-plant and slow-rusting resistance to Puccinia striiformis f. sp. tritici in wheat cultivars. Phytopathology 98:803–809. https://doi.org/10.1094/PHYTO-98-7-0803
Hovmøller MS (2007) Sources of seedling and adult plant resistance to Puccinia striiformis f. sp. tritici in European wheats. Plant Breed 126:225–233. https://doi.org/10.1111/j.1439-0523.2007.01369.x
Hovmøller MS, Sørensen CK, Walter S, Justesen AF (2011) Diversity of Puccinia striiformis on cereals and grasses. Annu Rev Phytopathol 49:197–217. https://doi.org/10.1146/annurev-phyto-072910-095230
Hovmøller MS, Walter S, Bayles RA, Hubbard A, Flath K et al (2016) Replacement of the European wheat yellow rust population by new races from the centre of diversity in the near-Himalayan region. Plant Pathol 65:402–411. https://doi.org/10.1111/ppa.12433
Jia M, Yang L, Zhang W, Rosewarne G, Li J et al (2020) Genome-wide association analysis of stripe rust resistance in modern Chinese wheat. BMC Plant Biol 20:491. https://doi.org/10.1186/s12870-020-02693-w
Jighly A, Oyiga BC, Makdis F, Nazari K, Youssef O et al (2015) Genome-wide DArT and SNP scan for QTL associated with resistance to stripe rust (Puccinia striiformis f. sp. tritici) in elite ICARDA wheat (Triticum aestivum L.) germplasm. Theor Appl Genet 128:1277–1295. https://doi.org/10.1007/s00122-015-2504-2
Juliana P, Singh RP, Singh PK, Crossa J, Huerta-Espino J et al (2017) Genomic and pedigree-based prediction for leaf, stem, and stripe rust resistance in wheat. Theor Appl Genet 130:1415–1430. https://doi.org/10.1007/s00122-017-2897-1
Kankwatsa P, Singh D, Thomson PC, Ebrahiem M, Babiker EM et al (2017) Characterization and genome-wide association mapping of resistance to leaf rust, stem rust and stripe rust in a geographically diverse collection of spring wheat landraces. Mol Breed 37:113. https://doi.org/10.1007/s11032-017-0707-8
Klymiuk V, Yaniv E, Huang L, Raats D, Fatiukha A et al (2018) Cloning of the wheat Yr15 resistance gene sheds light on the plant tandem kinase pseudo-kinase family. Nat Commun 9:3735. https://doi.org/10.1038/s41467-018-06138-9
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Huerta-Espino J et al (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363. https://doi.org/10.1126/science.1166453
Kumar D, Kumar A, Chhokar V, Gangwar OP, Bhardwaj SC et al (2020) Genome-wide association studies in diverse spring wheat panel for stripe, stem, and leaf rust resistance. Front Plant Sci 11:748. https://doi.org/10.3389/fpls.2020.00748
Laidig F, Feike T, Hadasch SRD, Klocke B et al (2021) Breeding progress of disease resistance and impact of disease severity under natural infections in winter wheat variety trials. Theor Appl Genet 134:1281–1302. https://doi.org/10.1007/s00122-020-03728-4
Ledesma-Ramírez L, Solís-Moya E, Iturriaga G, Sehgal D, Reyes Valdes MH et al (2019) GWAS to identify genetic loci for resistance to yellow rust in wheat pre-breeding lines derived from diverse exotic crosses. Front Plant Sci 10:1390. https://doi.org/10.3389/fpls.2019.01390
Ling W, Xian-chun X, You-liang Z, Zheng-yu Z, Hua-zhong Z et al (2012) QTL mapping for adult-plant resistance to stripe rust in a common wheat RIL population derived from Chuanmai 32/Chuanyu 12. J Integr Agric 11:1775–1782. https://doi.org/10.1016/S2095-3119(12)60182-8
Liu W, Frick M, Huel R, Nykiforuk CL, Wang X et al (2014) The stripe rust resistance gene Yr10 encodes an evolutionary-conserved and unique CC-NBS-LRR sequence in wheat. Mol Plant 7:1740–1755. https://doi.org/10.1093/mp/ssu112
Losert D, Maurer HP, Leiser WL, Würschum T (2017) Defeating the Warrior: genetic architecture of triticale resistance against a novel aggressive yellow rust race. Theor Appl Genet 130:685–696. https://doi.org/10.1007/s00122-016-2843-7
Lu Y, Wang M, Chen X, See D, Chao S et al (2014) Mapping of Yr62 and a small-effect QTL for high-temperature adult-plant resistance to stripe rust in spring wheat PI 192252. Theor Appl Genet 127:1449–1459. https://doi.org/10.1007/s00122-014-2312-0
Maccaferri M, Zhang J, Bulli P, Abate Z, Chao S et al (2015) A genome-wide association study of resistance to stripe rust (Puccinia striiformis f. sp. tritici) in a worldwide collection of hexaploid spring wheat (Triticum aestivum L.). G3 5:449–465
Macer RCF (1966) The formal and monosomic genetic analysis of stripe rust (Puccinia striiformis) resistance in wheat. In: Mackey J (ed) Proceedings of the 2nd international wheat genetics symposium, Lund, Sweden. Hereditas Suppl 2:127–142
Marchal C, Zhang J, Zhang P, Fenwick P, Steuernagel B et al (2018) Bed-domain containing immune receptors confer diverse resistance spectra to yellow rust. Nat Plants 4:662–668. https://doi.org/10.1038/s41477-018-0236-4
McIntosh RA, Dubcovsky J, Rogers WJ, Morris C, Xia XC (2017) Catalogue of gene symbols for wheat. 2017 Supplement. https://shigen.nig.ac.jp/wheat/komugi/genes/macgene/supplement2017.pdf
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci 88:9828–9832. https://doi.org/10.1073/pnas.88.21.9828
Miedaner T, Akel W, Flath K, Jacobi A, Taylor M et al (2019) Molecular tracking of multiple disease resistance in a winter wheat diversity panel. Theor Appl Genet 133:419–431. https://doi.org/10.1007/s00122-019-03472-4
Moore JW, Herrera-Foessel S, Lan C, Schnippenkoetter W, Ayliffe M et al (2015) A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat Genet 47:1494–1498. https://doi.org/10.1038/ng.3439
Mu J, Liu L, Liu Y, Wang M, See DR et al (2020) Genome-wide association study and gene specific markers identified 51 genes or QTL for resistance to stripe rust in U.S. winter wheat cultivars and breeding lines. Front Plant Sci 11:998. https://doi.org/10.3389/fpls.2020.00998
Muleta KT, Bulli P, Zhang Z, Chen X, Pumphrey M (2017) Unlocking diversity in germplasm collections via genomic selection: a case study based on quantitative adult plant resistance to stripe rust in spring wheat. Plant Genome. https://doi.org/10.3835/plantgenome2016.12.0124
Murphy LR, Santra D, Kidwell K, Yan G, Chen X et al (2009) Linkage maps of wheat stripe rust resistance genes Yr5 and Yr15 for use in marker-assisted selection. Crop Sci 49:1786–1790. https://doi.org/10.2135/cropsci2008.10.0621
Naruoka Y, Garland-Campbell KA, Carter AH (2015) Genome-wide association mapping for stripe rust (Puccinia striiformis f. sp. tritici) in US Pacific Northwest winter wheat (Triticum aestivum L.). Theor Appl Genet 128:1083–1101. https://doi.org/10.1007/s00122-015-2492-2
Peterson R, Campbell A, Hannah A (1948) A diagrammatic scale for estimating rust intensity of leaves and stem of cereals. Can J Res Sect C 26:496–500
Plaschke J, Ganal MW, Röder MS (1995) Detection of genetic diversity in closely related bread wheat using microsatellite markers. Theor Appl Genet 91:1002–1007. https://doi.org/10.1007/BF00223912
Pradhan AK, Kumar S, Singh AK, Budhlakoti N, Mishra DC et al (2020) Identification of QTLs/defense genes effective at seedling stage against prevailing races of wheat stripe rust in India. Front Genet 11:572975. https://doi.org/10.3389/fgene.2020.572975
Prins R, Pretorius ZA, Bender CM, Lehmensiek A (2011) QTL mapping of stripe, leaf and stem rust resistance genes in a Kariega × Avocet S doubled haploid wheat population. Mol Breed 27:259–270. https://doi.org/10.1007/s11032-010-9428-y
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
R Core Team (2017) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Ren Y, He Z, Li J, Lillemo M, Wu L et al (2012) QTL mapping of adult-plant resistance to stripe rust in a population derived from common wheat cultivars Naxos and Shanghai 3/Catbird. Theor Appl Genet 125:1211–1221. https://doi.org/10.1007/s00122-012-1907-6
Rollar S, Geyer M, Hartl L, Mohler V, Ordon F et al (2021) Quantitative trait loci mapping of adult plant and seedling resistance to stripe rust (Puccinia striiformis Westend.) in a multiparent advanced generation intercross wheat population. Front Plant Sci 12:684671. https://doi.org/10.3389/fpls.2021.684671
Rosewarne GM, Singh RP, Huerta-Espino J, Herrera-Foessel SA, Forrest KL et al (2012) Analysis of leaf and stripe rust severities reveals pathotype changes and multiple minor QTLs associated with resistance in an Avocet × Pastor wheat population. Theor Appl Genet 124:1283–1294. https://doi.org/10.1007/s00122-012-1786-x
Rosewarne GM, Herrera-Foessel SA, Singh RP, Huerta-Espino J, Lan CX et al (2013) Quantitative trait loci of stripe rust resistance in wheat. Theor Appl Genet 126:2427–2449. https://doi.org/10.1007/s00122-013-2159-9
Singh RP (1992) Genetic association of leaf rust resistance gene Lr34 with adult plant resistance to stripe rust in bread wheat. Phytopathology 82:835–838. https://doi.org/10.1094/phyto-82-835
Singh RP, Singh PK, Rutkoski J, Hodson DP, He X et al (2016) Disease impact on wheat yield potential and prospects of genetic control. Annu Rev Phytopathol 4:303–322. https://doi.org/10.1146/annurev-phyto-080615-095835
Spielmeyer W, McIntosh RA, Kolmer J, Lagudah ES (2005) Powdery mildew resistance and Lr34/Yr18 genes for durable resistance to leaf and stripe rust cosegregate at a locus on the short arm of chromosome 7D of wheat. Theor Appl Genet 111:731–735. https://doi.org/10.1007/s00122-005-2058-9
Sui XX, Wang MN, Chen XM (2009) Molecular mapping of a stripe rust resistance gene in spring wheat cultivar Zak. Phytopathology 99:1209–1215. https://doi.org/10.1094/PHYTO-99-10-1209
Tehseen MM, Tonk FA, Tosun M, Amri A, Sansaloni CP et al (2021) Genome-wide association study of resistance to PstS2 and Warrior races of Puccinia striiformis f. sp. tritici (stripe rust) in bread wheat landraces. Plant Genome 14:e20066. https://doi.org/10.1002/tpg2.20066
Tsomin Y, Wenhua S, Kequan S (1990) Monosomic analyses of stripe rust and leaf rust resistance genes in winter wheat varieties Luquyu and Yantar. Euphytica 48:83–86
Vazquez MD, Peterson CJ, Riera-Lizarazu O, Chen X, Heesacker A et al (2012) Genetic analysis of adult plant, quantitative resistance to stripe rust in wheat cultivar “Stephens” in multi-environment trials. Theor Appl Genet 124:1–11. https://doi.org/10.1007/s00122-011-1681-x
Wang S, Wong D, Forrest K, Allen A, Chao S et al (2014) Characterization of polyploid wheat genomic diversity using a high-density 90,000 single nucleotide polymorphism array. Plant Biotechnol J 12:787–796. https://doi.org/10.1111/pbi.12183
Wimmer V, Albrecht T, Auinger HJ, Schön CC (2012) synbreed: a framework for the analysis of genomic prediction data using R. Bioinformatics 28:2086–2087. https://doi.org/10.1093/bioinformatics/bts335
Xu Y, Wang J, Crouch JH (2008) Selective genotyping and pooled DNA analysis: an innovative use of an old concept. Recognizing past achievement, meeting future needs. In: Proceedings of the 5th international crop science congress, April 3–18; Jeju, Korea
You Q, Zhai K, Yang D, Yang W, Wu J et al (2016) An E3 ubiquitin ligase-BAG protein module controls plant innate immunity and broad-spectrum disease resistance. Cell Host Microbe 20:758–769. https://doi.org/10.1016/j.chom.2016.10.023
Zetzsche H, Serfling A, Ordon F (2019) Breeding progress in seedling resistance against various races of stripe and leaf rust in European bread wheat. Crop Breed Genet Genom 1:546. https://doi.org/10.20900/cbgg20190021
Zhang C, Huang L, Zhang H, Hao Q, Lyu B et al (2019) An ancestral NB-LRR with duplicated 3′UTRs confers stripe rust resistance in wheat and barley. Nat Commun 10:4023. https://doi.org/10.1038/s41467-019-11872-9
Zhu T, Wang L, Rimbert H, Rodriguez JC, Deal KR et al (2021) Optical maps refine the bread wheat Triticum aestivum cv. Chin Spring Genome Assemb Plant J 107:303–314. https://doi.org/10.1111/tpj.15289
Acknowledgments
We are grateful to our project partners at Saatzucht Josef Breun GmbH & Co. KG for their valuable cooperation and conducting field trials in Lenglern, Germany and Dr. Theresa Albrecht for her preliminary works in selection of the genotypes. Excellent technical assistant of Sabine Schmidt, Kerstin Hoffmann and Petra Greim at LfL is appreciated. We also appreciate Jörg Plieske and Martin Ganal of TraitGenetics for providing unpublished assignments of the 25 k SNP markers to physical positions. JIS was supported by the Beatriz Galindo Program (BEAGAL18/00115) from the Ministerio de Educación y Formación Profesional of Spain and the Severo Ochoa Program for Centres of Excellence in R&D from the Agencia Estatal de Investigación of Spain, grant SEV-2016-0672 (2017–2021) to the CBGP.
Funding
Open Access funding enabled and organized by Projekt DEAL. Results have been achieved within the framework of the project "WheatSustain" under the ERA-NET Cofund SusCrop (Grant No. 771134), being part of the Joint Programming Initiative on Agriculture, Food Security and Climate Change (FACCE-JPI). This project has received funding from the European Union's Horizon 2020 research and innovation program under Grant Agreement No. 771134 and with funding from the Research Council of Norway (NFR Grant 299615), Deutsches Bundesministerium für Bildung und Forschung (031B0810), and Bundesministerium für Nachhaltigkeit und Tourismus Österreich (Forschungsprojekt Nr. 101402).
Author information
Authors and Affiliations
Contributions
FS analyzed the phenotypic data and constructed the molecular map. FS and MG performed genetic analyses and wrote the manuscript. FSch, SR, JH, HK, MS, FL, LM, and HB conducted experimental field trials for phenotyping of stripe rust resistance. VM supported genetic analyses and designing of the project. JIS and DA designed the diversity panel. HB, JIS, ML, and LH conceived the project and obtained fundings. All authors read, edited, and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
122_2022_4202_MOESM1_ESM.xlsx
Table S1 List of 230 winter wheat cultivars and breeding lines assembled from the Northern and Central European countries Germany, Austria, Norway, Sweden, Denmark, Poland, and Switzerland. Mean values of stripe rust scores (%) evaluated in field trials and SNP alleles at 12 QTL identified across environments are presented. Table S2 Information of experiments conducted in field trials for evaluating stripe rust in 230 association panel. Table S3 List of 8812 SNP markers and their chromosomal position used for GWAS analysis in the association panel of 230 genotypes. Table S4 SNP markers identified in the winter wheat panel associated with stripe rust responses were evaluated in different field trials and across environments during 2020–2021. The marker alleles associated with increased resistance are bolded. Table S5 Functional annotation of putative candidate genes within 2 Mb (1 Mb upstream and downstream) physical interval of the peaks of validated markers associated with stripe rust disease resistance on chromosomes 2B (RAC875_c1226_652) and 6A (Tdurum_contig29607_413). The genes indicated in bold were tagged directly by the markers. Table S6 Comparison between QTL identified in the present study based on means across environments and previously reported genes and QTL controlling stripe rust resistance in common wheat. (XLSX 7581 kb)
122_2022_4202_MOESM2_ESM.tif
Fig. S1. a) Frequency distribution of stripe rust disease severity (%) in different field trials and across environments.(TIF 900 kb)
122_2022_4202_MOESM3_ESM.tif
b) Residual frequency distribution of arcsine square root transformed of disease severity in different field trials and across environments. (TIF 1164 kb)
122_2022_4202_MOESM4_ESM.tif
Fig. S2 Population structure among 230 winter wheat cultivars and breeding lines. a) Determination of the number of subpopulations via the ad hoc statistic ΔK. a) Stacked bar plots of ancestry relationship of genotypes based on membership coefficient of individuals (Q). (TIF 88 kb)
122_2022_4202_MOESM5_ESM.tif
Fig. S3 Scatter plot of linkage disequilibrium (R2) versus inter-marker physical distance (Mbp), with the average R2 of increasing intervals of 0.5 Mbp physical distance plotted as a red line. (TIF 145 kb)
122_2022_4202_MOESM6_ESM.tif
Fig. S4. Manhattan plots showing the association of single nucleotide polymorphisms in the 230 genotypes (left) and quantile–quantile plot comparing the performance of the mixed linear model (right) used in the genome-wide association study for stripe rust resistance in different field trials (a-e) and across environments (f). The horizontal black line represents the genome-wide significance threshold. (TIF 1586 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shahinnia, F., Geyer, M., Schürmann, F. et al. Genome-wide association study and genomic prediction of resistance to stripe rust in current Central and Northern European winter wheat germplasm. Theor Appl Genet 135, 3583–3595 (2022). https://doi.org/10.1007/s00122-022-04202-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-022-04202-z