Abstract
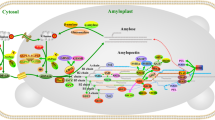
Gene duplication and divergence are important evolutionary processes. It has been suggested that a whole genome duplication (WGD) event occurred in the Gramineae, predating its divergence, and a second WGD occurred in maize during its evolution. In this study we compared the fate of the genes involved in the core pathway of starch biosynthesis following the ancient and second WGDs in maize and rice. In total, thirty starch synthesis genes were detected in the maize genome, which covered all the starch synthesis gene families encoded by 27 genes in rice. All of these genes, except ZmGBSSIIb and ZmBEIII, are anchored within large-scale synteny blocks of rice and maize chromosomes. Previous findings and our results indicate that two of the current copies of many starch synthesis genes (including AGPL, AGPS, GBSS, SSII, SSIII, and BEII) probably arose from the ancient WGD in the Gramineae and are still present in the maize and rice genome. Furthermore, two copies of at least six genes (AGPS1, SSIIb, SSIIIb, GBSSII, BEI, and ISA3) appear to have been retained in the maize genome after its second WGD, although complete coding regions were only detected among the duplicate sets of AGPS1, SSIIb, and SSIIIb. The expression patterns of the remaining duplicate sets of starch synthesis genes (AGPL1/2, AGPS1/2, SSIIa/b, SSIIIa/b, GBSSI/II, and BEIIa/b) differ in their expression and could be classified into two groups in maize. The first group is mainly expressed in the endosperm, whereas the second is expressed in other organs and the early endosperm development. The four duplicate sets of ZmGBSSII, ZmSSIIb, ZmSSIIIb and AGPS1, which arose from the second WGD diverged in gene structure and/or expression patterns in maize. These results indicated that some duplicated starch synthesis genes were remained, whereas others diverged in gene structure and/or expression pattern in maize. For most of the duplicated genes, one of the copies has disappeared in the maize genome after the WGD and the subsequent “diploidization”.



Similar content being viewed by others
References
Beatty MK, Rahman A, Cao H, Woodman W, Lee M, Myers AM, James MG (1999) Purification and molecular genetic characterization of ZPU1, a pullulanase-type starch-debranching enzyme from maize. Plant Physiol 119:255–266
Brunner S, Fengler K, Morgante M, Tingey S, Rafalski A (2005) Evolution of DNA sequence nonhomologies among maize inbreds. Plant Cell 17:343–360
Dian WM, Jiang HW, Wu P (2005) Evolution and expression analysis of starch synthase III and IV in rice. J Exp Bot 56:623–632
Dinges JR, Colleoni C, James MG, Myers AM (2003) Mutational analysis of the pullulanase-type debranching enzyme of maize indicates multiple functions in starch metabolism. Plant Cell 15:666–680
Fisher DK, Kim KN, Gao M, Boyer CD, Guiltinan MJ (1995) A cDNA encoding starch branching enzyme I from maize endosperm. Plant Physiol 108:1313–1314
Force A, Lynch M, Pickett FB, Amores A, Yan YL, Postlethwait J (1999) Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151:1531–1545
Fujita N, Yoshida M, Asakura N, Ohdan T, Miyao A, Hirochika H, Nakamura Y (2006) Function and characterization of starch synthase I using mutants in rice. Plant Physiol 140:1070–1084
Fujita N, Yoshida M, Kondo T, Saito K, Utsumi Y, Tokunaga T, Nishi A, Satoh H, Park JH, Jane JL, Miyao A, Hirochika H, Nakamura Y (2007) Characterization of SSIIIa-deficient mutants of rice: the function of SSIIIa and pleiotropic effects by SSIIIa deficiency in the rice endosperm. Plant Physiol 144:2009–2023
Gao M, Fisher DK, Kim KN, Shannon JC, Guiltinan MJ (1997) Independent genetic control of maize starch-branching enzymes IIa and IIb. Plant Physiol 114:69–78
Gao M, Wanat J, Stinard PS, James MG, Myers AM (1998) Characterization of dull1, a maize gene coding for a novel starch synthase. Plant Cell 10:399–412
Gaut BS, Doebley JF (1997) DNA sequence evidence for the segmental allotetraploid origin of maize. Proc Natl Acad Sci 94:6809–6814
Georgelis N, Braun EL, Hannah LC (2008) Duplications and functional divergence of ADP-glucose pyrophosphorylase genes in plants. BMC Evol Biol 8:232
Giroux MJ, Hannah LC (1994) ADP-glucose pyrophosphorylase in shrunken-2 and brittle-2 mutants of maize. Mol Gen Genet 243:400–408
Han Y, Sun FJ, Rosales-Mendoza S, Korban SS (2007) Three orthologs in rice, Arabidopsis, and Populus encoding starch branching enzymes (SBEs) are different from other SBE gene families in plants. Gene 15:123–130
Harn C, Knight M, Ramakrishnan A, Guan H, Keeling PL, Wasserman BP (1998) Isolation and characterization of the zSSIIa and zSSIIb starch synthase cDNA clones from maize endosperm. Plant Mol Biol 37:639–649
James MG, Robertson DS, Myers AM (1995) Characterization of the maize gene sugary1, a determinant of starch composition in kernels. Plant Cell 7:417–429
James GM, Denyer K, Myers AM (2008) Starch synthesis in the cereal endosperm. Curr Opin Plant Biol 6:215–222
Kawagoe Y, Kubo A, Satoh H, Takaiwa F, Nakamura Y (2005) Roles of isoamylase and ADP-glucose pyrophosphorylase in starch granule synthesis in rice endosperm. Plant J 42:164–174
Knight ME, Harn C, Lilley CE, Guan H, Singletary GW, MuForster C, Wasserman BP, Keeling PL (1998) Molecular cloning of starch synthase I from maize (W64) endosperm and expression in Escherichia coli. Plant J 14:613–622
Lee SK, Yu SK, Wang H, Han M, Eom JS, Kang HG, Han Y, Choi SB, Cho MH, Seong HB, Gynheung A, Hahn TR, Okita TW, Jeon JS (2007) Identification of the ADP-glucose pyrophosphorylase isoforms essential for starch synthesis in the leaf and seed endosperm of rice (Oryza sativa L.). Plant Mol Biol 65:531–546
Nakamura Y, Umemoto T, Takahata Y, Komae K, Amano E, Satoh H (1996) Changes in structure of starch and enzyme activities affected by sugary mutations in developing rice endosperm. Possible role of starch debranching enzyme (R-enzyme) in amylopectin biosynthesis. Physiol Plant 97:491–498
Nishi A, Nakamura Y, Tanaka N, Satoh H (2001) Biochemical and genetic analysis of the effects of amylose-extender mutation in rice endosperm. Plant Physiol 127:459–472
Ohdan T, Francisco PBJR, Sawada T, Hirose T, Terao T, Satoh H, Nakamura Y (2005) Expression profiling of genes involved in starch synthesis in sink and source organs of rice. J Exp Bot 56:3229–3244
Paterson AH, Bowers JE, Chapman BA (2004) Ancient polyploidization predating divergence of the cereals, and its consequences for comparative genomics. Proc Natl Acad Sci 101:9903–9908
Rahman A, Wong K, Jane J, Myers AM, James MG (1998) Characterization of SU1 isoamylase, a determinant of storage starch structure in maize. Plant Physiol 117:425–435
Rösti S, Denyer K (2007) Two paralogous genes encoding small subunits of ADP-glucose pyrophosphorylase in maize, Bt2 and L2, replace the single alternatively spliced gene found in other cereal species. J Mol Evol 65:316–327
Ryoo N, Yu C, Park CS, Baik MY, Park IM, Cho MH, Bhoo SH, An G, Hahn TR, Jeon JS (2007) Knockout of a starch synthase gene OsSSIIIa/Flo5 causes white-core floury endosperm in rice (Oryza sativa L.). Plant Cell Rep 26:1083–1095
Sano Y (1984) Differential regulation of waxy gene expression in rice endosperm. Theor Appl Genet 68:467–473
Satoh H, Nishi A, Yamashita K, Takemoto Y, Tanaka Y, Hosaka Y, Sakurai A, Fujita N, Nakamura Y (2003) Starch-branching enzyme I-deficient mutation specifically affects the structure and properties of starch in rice endosperm. Plant Physiol 133:1111–1121
Satoh H, Shibahara K, Tokunaga T, Nishi A, Tasaki M, Hwang SK, Okita TW, Kaneko N, Fujita N, Yoshida M, Hosaka Y, Sato A, Utsumi Y, Ohdan T, Nakamura Y (2008) Mutation of the plastidial alpha-glucan phosphorylase gene in rice affects the synthesis and structure of starch in the endosperm. Plant Cell 20:1833–1849
Shure M, Wessler S, Fedoroff N (1983) Molecular identification and isolation of the waxy locus in maize. Cell 35:225–233
Song R, Messing J (2003) Gene expression of a gene family in maize based on noncollinear haplotypes. Proc Natl Acad Sci 100:9055–9060
Stinard PS, Robertson DS, Schnable PS (1993) Genetic isolation, cloning, and analysis of a mutator-induced, dominant antimorph of the maize amylose extender1 locus. Plant Cell 5:1555–1566
Sullivan TD, Kaneko Y (1995) The maize brittle 1 gene encodes amyloplast membrane polypeptides. Planta 196:477–484
Takeda Y, Preiss J (1993) Structures of B90 (sugary) and W64A (normal) maize starches. Carbohydr Res 240:265–275
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4 Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Umemoto T, Yano M, Satoh H, Shomura A, Nakamura Y (2002) Mapping of a gene responsible for the difference in amylopectin structure between japonica-type and indica-type rice varieties. Theor Appl Genet 104:1–8
Wu Y, Zhu Z, Ma L, Chen M (2008) The preferential retention of starch synthesis genes reveals the impact of whole-genome duplication on grass evolution. Mol Biol Evol 25:1003–1006
Yan H, Jiang H, Pan X, Li M, Chen Y, Wu G (2009) The gene encoding starch synthase IIc exists in maize and wheat. Plant Sci 176:51–57
Yano M, Okuno K, Kawakami J, Satoh H, Omura T (1985) High amylose mutants of rice, Oryza sativa L. Theor Appl Genet 69:253–257
Yu Y, Mu HH, Wasserman BP, Carman GM (2001) Identification of the maize amyloplast stromal 112-kD protein as a plastidic starch phosphorylase. Plant Physiol 125:351–359
Zhang X, Colleoni C, Ratushna V, Sirghie-Colleoni M, James MG, Myers AM (2004) Molecular characterization demonstrates that the Zea mays gene sugary2 codes for the starch synthase isoform SSIIa. Plant Mol Biol 54:865–879
Acknowledgments
This research was supported by the Key Innovation Program of the Chinese Academy of Sciences (KSCX2-YW-G-027-3), the CAS ‘100 Talents’ program and the CAS/SAFEA International Partnership Program for Creative Research Teams.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by J. Dubcovsky.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yan, HB., Pan, XX., Jiang, HW. et al. Comparison of the starch synthesis genes between maize and rice: copies, chromosome location and expression divergence. Theor Appl Genet 119, 815–825 (2009). https://doi.org/10.1007/s00122-009-1091-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-009-1091-5