Abstract
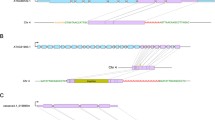
Structural variability of Tvv1, a grapevine retrotransposon Ty1 copia-like family, was investigated within the grape genome and the canonical sequence of Tvv1 determined. Then, two remarkable elements, Tvv1-Δ3001 and Tvv1-Δ3640, which had suffered large deletions 3,001 bp and 3,460 bp in length of their coding sequences were compared to the canonical copy. In both deleted elements, the deletion breakpoint was characterized by a stretch 13 bp-long in Tvv1-Δ3001 and 11 bp-long in Tvv1-Δ3640 found duplicated in the canonical copy at each bound of the deleted regions. Tvv1-Δ3001 and Tvv1-Δ3460 were both shown to be unique copies fixed at a single locus in the grapevine genome. Their presence was very variable in a set of 58 varieties and wild vines. These elements have most likely been dispersed through natural intermixing after their initial insertion whose chronology was estimated. The model that we propose to explain the structure of Tvv1-Δ3001 and Tvv1-Δ3640, implies illegitimate recombination involving template switching between two RNA molecules co-packaged in the VLP prior to the integration of the deleted daughter copy into the host genome.



Similar content being viewed by others
References
Anonymous (1995) Catalogue des variétés et clones de vigne cultivés en France, ENTAV, Ed, Le Grau du Roi. France, p 357
Boeke JD, Corces VG (1989) Transcription and reverse transcription of retrotransposons. Ann Rev Microbiol 43:403–434
Boeke JD, Styles C, Fink R (1986) Saccharomyces cerevisiae SPT3 gene is required for transposition and transpositional recombination of chromosomal Ty elements. Mol Cell Biol 6:3575–3581
Bowers J, Meredith C (1997) The parentage of a classic wine grape, Cabernet Sauvignon. Nature Genet 16:84–87
Bronner A (2003) Muscats et variétés muscatées. Inventaire et synonymie universels, des origines à nos jours, Avenir-Oenologie, INRA Editions/Oenoplurimedia, p 222
Casacuberta JM, Grandbastien M-A (1993) Characterization of LTR sequences involved in the protoplast specific expression of the tobacco Tnt1 retrotransposon. Nucl Acids Res 21:2087–2093
Casacuberta JM, Vernhettes S, Grandbastien M-A (1995) Sequence variability within the tobacco retrotransposon Tnt1 population. EMBO J 14:2670–2678
Feng Y-X, Moore SP, Garfinkel DJ, Rein A (2000) The genomic RNA in Ty1 virus-like particles is dimeric. J Virol 74:10819–10821
Feschotte C, Jiang N, Wessler SR (2002) Plant transposable elements: where genetics meets genomics. Nature Rev Genet 3:329–341
Flavell AJ, Dunbar E, Anderson R, Pearce SR, Hartley R, Kumar A (1992) Ty1-copia group retrotransposons are ubiquitous and heterogeneous in higher plants. Nucl Acids Res 20:3639–3644
Galet P (1988) Cépages et vignobles de France. Les vignes américaines, Edition, p 553
Gaut BS, Morton BR, McCaig BC, Clegg MT (1996) Substitution rate comparisons between grasses and palms: Synonymous rate differences at the nuclear gene Adh parallel rate differences at the plastid gene rbcL. Proc Natl Acad Sci USA 93:10274–10279
Gorbunova V, Levy AA (1997) Non-homologous DNA end joining in plant cells is associated with deletions and filler DNA insertions. Nucl Acids Res 25:4650–4657
Hu W, Temin HM (1990) Genetic consequences of packaging two RNA genomes in one retroviral particle: pseudodiploidy and high rate of genetic recombination. Proc Natl Acad Sci USA 87:1556–1560
Jordan IK, McDonald JF (1998) Evidence for the role of recombination in the regulatory evolution of Saccharomyces cerevisiae Ty elements. J Mol Evol 47:14–20
Jordan IK, McDonald JF (1999) Tempo and mode of Ty element evolution in Saccharomyces cerevisiae. Genetics 151:1341–1351
Kalendar R, Vicient CM, Peleg O, Amamthawat-Jonsson K, Bolshoy A, Schulman A (2004) Large retrotransposon derivatives: abundant, conserved by non autonomous retroelements of barley and related genomes. Genetics 166:1437–1450
Kirik A, Salomon S, Puchta H (2000) Species-specific double-strand break repair and genome evolution in plants. EMBO J 19:5562–5566
Kobayashi S, Goto-Yamamoto N, Hirochika H (2004) Retrotransposon-induced mutations in grape skin color. Science 304:982
Lamoureux D, Bernole A, Clainche IL, Tual S, Thareau V, Paillard S, Legeai F, Dossat C, Wincker P, Oswald M, Merdinoglu D, Vignault C, Delrot S, Caboche M, Chalhoub B, Adam-Blondon A-F (2006) Anchoring of a large set of markers onto a BAC library for the development of a draft physical map of the grapevine genome. Theor Appl Genet 113:344–356
Marillonnet S, Wessler SR (1998) Extreme structural heterogeneity among the members of a maize retrotransposon family. Genetics 150:1245–1256
Nagy PD, Bujarski JJ (1995) Efficient system of homologous RNA recombination in brome mosaic virus: sequence and structure requirements and accuracy of crossovers. J Virol 69:131–140
Nymark-McMahon MH, Beliakova-Bethell NS, Darlix J-L, Le Grice SFJ, Sandmeyer SB (2002) Ty3 integrase is required for initiation of reverse transcription. J Virol 76:2804–2816
Pelsy F (2007) Untranslated leader region polymorphism of Tvv1, a retrotransposon family, is a novel marker useful for analyzing genetic diversity and relatedness in the genus Vitis. Theor Appl Genet 116:15–27
Pelsy F, Merdinoglu D (2002) Complete sequence of Tvv1, a family of Ty1 copia-like retrotransposons of Vitis vinifera L., reconstituted by chromosome walking. Theor Appl Genet 105:614–621
Preston BD (1996) Error-prone retrotransposition: Rime of the ancient mutators. Proc Natl Acad Sci USA 93:7427–7431
Sabot F, Sourdille P, Chantret N, Bernard M (2006) Morgane, a new LTR retrotransposon group, and its subfamilies in wheats. Genetica 128:439–447
SanMiguel P, Gaut BS, Tikhonov A, Nakajima Y, Bennetzen JL (1998) The paleontology of intergene retrotransposons of maize. Nat Genet 20:43–45
Shirasu K, Schulman AH, Lahaye T, Schulze-Lefert P (2000) A contiguous 66-kb barley DNA sequence provides evidence for reversible genome expansion. Genome Res 10:908–915
Suoniemi A, Tanskanen J, Schulman AH (1998) Gypsy-like retrotransposons are widespread in the plant kingdom. Plant J 13:699–705
Temin H (1993) Retrovirus variation and reverse transcription: abnormal strand transfers result in retrovirus genetic variation. Proc Natl Acad Sci USA 90:6900–6903
The French–Italian Public Consortium for Grapevine Genome Characterization (2007) The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449(7161):463–467
Verriès C, Bès C, This P, Tesnière C (2000) Cloning and characterization of Vine-1, a LTR- retrotransposon-like element in Vitis vinifera L., and other Vitis species. Genome 43:366–376
Vicient CM, Kalendar R, Schulman AH (2005) Variability, recombination, and mosaic evolution of the barley BARE-1 retrotransposon. J Mol Evol 61:275–291
Vicient CM, Suoniemi A, Anamthamat-Jonsson K, Tanskanen J, Beharav A, Nevo E, Schulman AH (1999) Retrotransposon BARE-1 and its role in genome evolution in the genus Hordeum. Plant Cell 11:1769–1784
Voytas DF, Cumming MP, Konieczny A, Ausubel FM, Rodermel SR (1992) Copia-like retrotransposons are ubiquitous among plants. Proc Natl Acad Sci USA 89:7124–7128
Acknowledgments
The authors thank Dr. Eva Zyprian and Dr. Reinhardt Töpfer from the Federal Centre for Breeding Research on Cultivated Plant, Institute for Grapevine Breeding Geilweilerhof (Siebeldingen, Germany) for kindly providing the partial sequence of the BAC clone A1-16, Riccardo Velasco from the Istituto Agrario San Michele all’Adige (Italy) for the pinot sequence (VV78X150312.9) and Anne Françoise Adam-Blondon (INRA-URGV, France) for the BAC clone 3E11. We thank Lucie Bevilacqua-Schehrer and Cindy Ponama for their valuable technical assistance, Vincent Dumas and the experimental domain staff for grapevine accessions, Malek Alioua (ULP-Strasbourg, France) for sequencing, and Pere Mestre for careful revision of the English. Cedric Moisy is a doctoral fellow supported by the Conseil Régional d’Alsace. All experiments described in this study comply with the current laws of France.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. Gebhardt.
Rights and permissions
About this article
Cite this article
Moisy, C., Blanc, S., Merdinoglu, D. et al. Structural variability of Tvv1 grapevine retrotransposons can be caused by illegitimate recombination. Theor Appl Genet 116, 671–682 (2008). https://doi.org/10.1007/s00122-007-0700-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-007-0700-4