Abstract
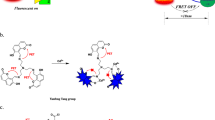
The hairpin probe is characterized by a higher fluorescence quenching efficiency as compared with the linear one under the conditions of real-time PCR, which leads to a lower background level of fluorescence and, consequently, a higher signal to noise ratio during real-time PCR. An experimental comparison of the fluorescence-quenching efficiency of two oligonucleotide probes in different conformations (hairpin in a molecular beacon format and linear in a TaqMan format) was made. There is a difference in the interaction between the quencher and fluorophore for the probes of different conformations. For a linear probe, quenching occurs through the mechanism of inductive-resonance energy transfer (Förster resonance energy transfer, FRET), while that for a hairpin probe quenching occurs through contact quenching through a closer arrangement of the fluorophore and quencher, but a resonant energy transfer according to the Förster mechanism is also possible. It was demonstrated that the absorption spectrum for the linear probe almost coincides with the absorption spectrum of an oligonucleotide representing a probe without a quencher, which indicates a dynamic (Förster) mechanism of energy transfer. On the contrary, the absorption spectra for the hairpin probe and oligonucleotide representing the probe without a quencher differ significantly, which indicates a contact mechanism of energy transfer between the fluorophore and fluorescence quencher. The fluorescence spectra of the probes and their complexes with the oligonucleotide complementary to the linear probe (and the loop of the hairpin probe) and the amplicon (200 bp in length containing a DNA target for the probes) allowed for comparison of these two probes by comparing the energy migration radii, the efficiency of donor fluorescence quenching. The energy migration radius R calculated by the experimental data was 32.4 Å for the hairpin probe and 47.3 Å for the linear probe.




Similar content being viewed by others
REFERENCES
Demchenko, A.P., Introduction in Fluorescence Sensing, Berlin: Springer Science and Business Media, 2009.
Dexter, D.L., A theory of sensitized luminescence in solids, J. Chem. Phys., 1953, vol. 21, pp. 836–850. https://doi.org/10.1063/1.1699044
Forster, T., Zwischenmolekulare energiewanderung und fluoreszenz, Ann. Phys. (Leipzig), 1948, vol. 2, p. 55. https://doi.org/10.1002/andp.19484370105
Hadjinicolaou, A.V., Demetriou, V.L., Hezka, J., et al., Use of molecular beacons and multi-allelic real-time PCR for detection of and discrimination between virulent Bacillus anthracis and other Bacillus isolates, J. Microbiol. Methods, 2009, vol. 78, pp. 45–53. https://doi.org/10.1016/j.mimet.2009.04.005
Howell, W.M., Jobs, I.M., and Brookes, A.J., iFRET: An improved fluorescence system for DNA-melting analysis, Genome Res., 2002, vol. 12, pp. 1401–1407. http://www.genome.org/cgi/doi/10.1101/gr.297202
Hunyadi, S. and Murphy, C., Tunable one-dimensional silver-silica nanopeapod architectures, J. Phys. Chem. B., 2006, vol. 110, pp. 7226–7231. https://doi.org/10.1021/jp0603076
Inokuti, M. and Hirayama, F., Influence of energy transfer by the exchange mechanism on donor luminescence, J. Chem. Phys., 1965, vol. 43, p. 1978. https://doi.org/10.1063/1.1697063
Josefsen, M.H., Löfström, C., Sommer, H.M., et al., Diagnostic PCR: Comparative sensitivity of four probe chemistries, Mol. Cell. Probes, 2009, vol. 23, pp. 201–203. https://doi.org/10.1016/j.mcp.2009.02.03
Kapanidis, A. and Weiss, S., Fluorescent probes and bioconjugation chemistries for single molecule fluorescence analysis of biomolecules, J. Chem. Phys., 2002, vol. 117, pp. 10953–10964. https://doi.org/10.1063/1.1521158
Krasnoperov, L.N., Marras, S.A., Kozlov, M., et al., Luminescent probes for ultrasensitive detection of nucleic acids, Bioconjugate Chem., 2010, vol. 21, pp. 319–327. https://doi.org/10.1021/bc900403n
Lakowicz, J.R., Principles of Fluorescence Spectroscopy, New York: Springer-Verlag, 2007.
Latorra, D., Arar, K., and Hurley, J.M., Design considerations and effects of LNA in PCR primers, Mol. Cell. Probes, 1986, vol. 17, pp. 253–259. https://doi.org/10.1016/s0890-8508(03)00062-8
Le Reste, L., Hohlbein, J., Gryte, K., et al., Characterization of dark quencher chromophores as nonfluorescent acceptors for single-molecule FRET, Biophys. J., 2012, vol. 102, pp. 2658–2668. https://doi.org/10.1016/j.bpj.2012.04.028
Limanskaya, O.Yu., Murtazaeva, L.Î., and Limanskii, A.P., Species specific detection of causative agent of anthrax, Biotechnology, 2012, vol. 5, no. 1, pp. 92–99.
Ma, C., Liu, H., Wu, K., et al., An exonuclease I-based quencher-free fluorescent method using DNA hairpin probes for rapid detection of microRNA, Sensors, 2017, vol. 17, p. 760. https://doi.org/10.3390/s17040760
Marras, S., Interactive fluorophore and quencher pairs for labeling fluorescent nucleic acid hybridization probes, Mol. Biotechnol., 2008, vol. 38, pp. 247–255. https://doi.org/10.1007/s12033-007-9012-9
Marras, S., Kramer, F., and Tyagi, S., Efficiencies of fluorescence resonance energy transfer and contact-mediated quenching in oligonucleotide probes, Nucleic Acids Res., 2002, vol. 30, p. e122. https://doi.org/10.1093/nar/gnf121
Marras, S.A., Kramer, F.R., and Tyagi, S., Genotyping SNPs with molecular beacons, Methods Mol. Biol., 2003, vol. 212, pp. 111–128. https://doi.org/10.1385/1-59259-327-5:111
Miura, M., Tanigawa, C., Fujii, Y., et al., Comparison of six commercially-available DNA polymerases for direct PCR, Rev. Inst. Med. Trop. Sao Paulo, 2013, vol. 55, pp. 401–406. https://doi.org/10.1590/S0036-46652013000600005
Munoz, C., Talquenca, S.G., and Volpe, M.L., Tetra primer ARMS-PCR for identification of SNP in β-tubulin of Botrytis cinerea, responsible of resistance to benzimidazole, J. Microbiol. Methods, 2009, vol. 78, pp. 245–246. https://doi.org/10.1016/j.mimet.2009.06.007
Parsons, B.L., McKinzie, P.B., and Heflich, R.H., Allele-specific competitive blocker-PCR detection of rare base substitution, Methods Mol. Biol., 2005, vol. 291, pp. 235–245. https://doi.org/10.1385/1-59259-840-4:235
Sjoback, R., Nygren, J., and Kubista, M., Absorption and fluorescence properties of fluorescein, Spectrochim. Acta, Part A, 1995, vol. 51, pp. L7–L21.
Takayamaa, I., Nakauchia, M., Takahashia, H., et al., Development of real-time fluorescent reverse transcription loop-mediated isothermal amplification assay with quenching primer for influenza virus and respiratory syncytial virus, J. Virol. Methods, 2019, vol. 267, pp. 53–58. https://doi.org/10.1016/j.jviromet.2019.02.010
Tyagi, S. and Kramer, F.R., Molecular beacons: probes that fluoresce upon hybridization, Nat. Biotechnol., 1996, vol. 14, pp. 303–308. https://doi.org/10.1038/nbt0396-303
Vester, B. and Wengel, J., LNA (locked nucleic acid): high-affinity targeting of complementary RNA and DNA, Biochemistry, 2004, vol. 43, pp. 13233–13241. https://doi.org/10.1021/bi0485732
Wang, R.-H., Liu, L.-M., Zhao, J.-L., et al., A new method for SNP typing based on allele specific PCR, Fa Yi Xue Za Zhi, 2008, vol. 24, pp. 189–193.
Wu, D.Y., Ugozzoli, L., Pal, B.K., and Wallace, R.B., Allele-specific enzymatic amplification of β-globin genomic DNA for diagnosis of sickle cell anemia, Proc. Natl. Acad. Sci. U. S. A., 1989, vol. 86, pp. 2757–2760. https://doi.org/10.1073/pnas.86.8.2757
Yaku, H., Yukimasa, T., Nakano, S., et al., Design of allele-specific primers and detection of the human ABO genotyping to avoid the pseudopositive problems, Electrophoresis, 2008, vol. 29, pp. 4130–4140. https://doi.org/10.1002/elps.200800097
Zimmers, Z.A., Adam, N.M., Gabella, W.E., et al., Fluorophore-quencher interactions effect on hybridization characteristics of complementary oligonucleotides, Anal. Methods, 2019, vol. 11, pp. 2862–2867. https://doi.org/10.1039/c9ay00584f
Zuker, M., Mfold web server for nucleic acid folding and hybridization prediction, Nucl. Acids Res., 2003, vol. 31, pp. 3406–3415. https://doi.org/10.1093/nar/gkg595
ACKNOWLEDGMENTS
We are grateful to L.A. Murtazaeva for the assistance in performing spectral studies and to Doctor in Physics and Mathematics, Professor of Karazin Kharkiv National University G.P. Gorbenko.
Funding
This study was supported by Doctor in Biology O.Yu. Limanskaya and Doctor in Biology O.P. Limanskii without the external funding.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare that they have no conflicts of interest. This article does not contain any studies involving human participants and animals as objects.
About this article
Cite this article
Limanskaya, O.Y., Limanskii, O.P. Intramolecular Interactions in the Fluorophore–Quencher System in Linear and Hairpin Probes for Real-Time PCR. Cytol. Genet. 57, 134–141 (2023). https://doi.org/10.3103/S009545272302007X
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.3103/S009545272302007X