Abstract
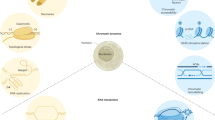
The genome integrity of all organisms is constantly threatened by replication errors and DNA damage arising from endogenous and exogenous sources. Such base pair anomalies must be accurately repaired to prevent mutagenesis and/or lethality. Thus, it is not surprising that cells have evolved multiple and partially overlapping DNA repair pathways to correct specific types of DNA errors and lesions. Great progress in unraveling these repair mechanisms at the molecular level has been made by several talented researchers, among them Tomas Lindahl, Aziz Sancar, and Paul Modrich, all three Nobel laureates in Chemistry for 2015. Much of this knowledge comes from studies performed in bacteria, yeast, and mammals and has impacted research in plant systems. Two plant features should be mentioned. Plants differ from higher eukaryotes in that they lack a reserve germline and cannot avoid environmental stresses. Therefore, plants have evolved different strategies to sustain genome fidelity through generations and continuous exposure to genotoxic stresses. These strategies include the presence of unique or multiple paralogous genes with partially overlapping DNA repair activities. Yet, in spite (or because) of these differences, plants, especially Arabidopsis thaliana, can be used as a model organism for functional studies. Some advantages of this model system are worth mentioning: short life cycle, availability of both homozygous and heterozygous lines for many genes, plant transformation techniques, tissue culture methods and reporter systems for gene expression and function studies. Here, I provide a current understanding of DNA repair genes in plants, with a special focus on A. thaliana. It is expected that this review will be a valuable resource for future functional studies in the DNA repair field, both in plants and animals.
Similar content being viewed by others
References
Cadet J, Wagner J (2013) DNA base damage by reactive oxygen species, oxidizing agents, and UV radiation. Cold Spring Harb Perspect Biol 5:a012559
Jiricny J (2013) Postreplicative mismatch repair. Cold Spring Harb Perspect Biol 5:a012633
Hu Z, Cools T, De Veylder L (2016) Mechanisms used by plants to cope with DNA damage. Annu Rev Plant Biol 67:439–462
Yoshiyama K, Sakaguchi K, Kimura S (2013) DNA damage response in plants: conserved and variable response compared to animals. Biology 2:1338–1356
Edgar B, Zielke N, Gutierrez C (2014) Endocycles: a recurrent evolutionary innovation for post-mitotic cell growth. Nat Rev Mol Cell Biol 15:197–210
The Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408:796–815
Singh S, Roy S, Choudhury S, Sengupta D (2010) DNA repair and recombination in higher plants: insights from comparative genomics of Arabidopsis and rice. BMC Genom 11:443
Spampinato C, Gómez-Casati D (2012) Research on plants for the understanding of diseases of nuclear and mitochondrial origin. J Biomed Biotech 2012:ID 836196
Biedermann S, Mooney S, Hellmann H (2011) Recognition and repair pathways of damaged DNA in higher plants. Selected Topics in DNA Repair. University of California, San Diego
Kimura S, Sakaguchi K (2006) DNA repair in plants. Chem Rev 106:753–766
Friedberg E (2015) A history of the DNA repair and mutagenesis field I. The discovery of enzymatic photoreactivation. DNA Repair (Amst) 33:35–42
Eker A, Quayle C, Chaves I, van der Horst G (2009) DNA repair in mammalian cells: direct DNA damage reversal: elegant solutions for nasty problems. Cell Mol Life Sci 66:968–980
Essen L, Klar T (2006) Light-driven DNA repair by photolyases. Cell Mol Life Sci 63:1266–1277
Sancar A (2008) Structure and function of photolyase and in vivo enzymology: 50th anniversary. J Biol Chem 283:32153–32157
Mei Q, Dvornyk V (2015) Evolutionary history of the photolyase/cryptochrome superfamily in eukaryotes. PLoS One 10:e0135940
Okafuji A, Biskup T, Hitomi K, Getzoff E, Kaiser G, Batschauer A, Bacher A, Hidema J, Teranishi M, Yamamoto K, Schleicher E, Weber S (2010) Light-induced activation of class II cyclobutane pyrimidine dimer photolyases. DNA Repair (Amst) 9:495–505
Lucas-Lledó J, Lynch M (2009) Evolution of mutation rates: phylogenomic analysis of the photolyase/cryptochrome family. Mol Biol Evol 26:1143–1153
Kiontke S, Gnau P, Haselsberger R, Batschauer A, Essen L (2014) Structural and evolutionary aspects of antenna chromophore usage by class II photolyases. J Biol Chem 289:19659–19669
Rastogi R, Richa X, Kumar A, Tyagi M, Sinha R (2010) Molecular mechanisms of ultraviolet radiation-induced DNA damage and repair. J Nucleic Acids 2010:592980
Richa Sinha R, Häder D (2015) Physiological aspects of UV-excitation of DNA. Top Curr Chem 356:203–248
Zhong D (2015) Electron transfer mechanisms of DNA repair by photolyase. Annu Rev Phys Chem 66:691–715
Mannuss A, Trapp O, Puchta H (2012) Gene regulation in response to DNA damage. Biochim Biophys Acta 1819:154–165
Ueda T, Nakamura C (2011) Ultraviolet-defense mechanisms in higher plants. Biotechnol Biotechnol Equip 25:2177–2182
Manova V, Gruszka D (2015) DNA damage and repair in plants—from models to crops. Front Plant Sci 6:art 885
Li N, Teranishi M, Yamaguchi H, Matsushita T, Watahiki M, Tsuge T, Li S, Hidema J (2015) UV-B-induced CPD photolyase gene expression is regulated by UVR8-dependent and -independent pathways in Arabidopsis. Plant Cell Physiol 56:2014–2023
Hitomi K, Arvai A, Yamamoto J, Hitomi C, Teranishi M, Hirouchi T, Yamamoto K, Iwai S, Tainer J, Hidema J, Getzoff E (2012) Eukaryotic class II cyclobutane pyrimidine dimer photolyase structure reveals basis for improved ultraviolet tolerance in plants. J Biol Chem 287:12060–12069
Hitomi K, DiTacchio L, Arvai A, Yamamoto J, Kim S, Todo T, Tainer J, Iwai S, Panda S, Getzoff E (2009) Functional motifs in the (6-4) photolyase crystal structure make a comparative framework for DNA repair photolyases and clock cryptochromes. Proc Natl Acad Sci USA 106:6962–6967
Li J, Liu Z, Tan C, Guo X, Wang L, Sancar A, Zhong D (2010) Dynamics and mechanism of repair of UV-induced (6-4) photoproduct by photolyase. Nature 466:887–890
Liu Z, Wang L, Zhong D (2015) Dynamics and mechanisms of DNA repair by photolyase. Phys Chem Chem Phys 17:11933–11949
Kim Y, Wilson DI (2012) Overview of base excision repair biochemistry. Curr Mol Pharmacol 5:3–13
Krokan H, Bjørås M (2013) Base excision repair. Cold Spring Harb Perspect Biol 5:a012583
Svilar D, Goellner E, Almeida K, Sobol R (2011) Base excision repair and lesion-dependent subpathways for repair of oxidative DNA damage. Antioxid Redox Signal 14:2491–2507
Brooks S, Adhikary S, Rubinson E, Eichman B (2013) Recent advances in the structural mechanisms of DNA glycosylases. Biochim Biophys Acta 1834:247–271
Jacobs A, Schär P (2012) DNA glycosylases: in DNA repair and beyond. Chromosoma 121:1–20
Dianov G, Hübscher U (2013) Mammalian base excision repair: the forgotten archangel. Nucleic Acids Res 41:3483–3490
Bebenek K, Pedersen L, Kunkel T (2014) Structure–function studies of DNA polymerase λ. Biochemistry 53:2781–2792
Hanssen-Bauer A, Solvang-Garten K, Sundheim O, Peña-Diaz J, Andersen S, Slupphaug G, Krokan H, Wilson DI, Akbari M, Otterlei M (2011) XRCC1coordinates disparate responses and multiprotein repair complexes depending on the nature and context of the DNA damage. Environ Mol Mutagen 52:623–635
Balestrazzi A, Confalonieri M, Macovei A, Donà M, Carbonera D (2011) Genotoxic stress and DNA repair in plants: emerging functions and tools for improving crop productivity. Plant Cell Rep 30:287–295
Roldán-Arjona T, Ariza R (2009) Repair and tolerance of oxidative DNA damage in plants. Mutat Res 681:169–179
Córdoba-Cañero D, Dubois E, Ariza R, Doutriaux M-P, Roldán-Arjona T (2010) Arabidopsis uracil DNA glycosylase (ung) is required for base excision repair of uracil and increases plant sensitivity to 5-fluorouracil. J Biol Chem 285:7475–7483
Cordoba-Cañero D, Morales-Ruiz T, Roldán-Arjona T, Ariza R (2009) Single-nucleotide and long-patch base excision repair of DNA damage in plants. Plant J 60:716–728
Córdoba-Cañero D, Roldán-Arjona T, Ariza R (2014) Arabidopsis ZDP DNA 3′-phosphatase and ARP endonuclease function in 8-oxoG repair initiated by FPG and OGG1 DNA glycosylases. Plant J 79:824–834
Duclos S, Aller P, Jaruga P, Dizdaroglu M, Wallace S, Doublié S (2012) Structural and biochemical studies of a plant formamidopyrimidine-DNA glycosylase reveal why eukaryotic Fpg glycosylases do not excise 8-oxoguanine. DNA Repair (Amst) 11:714–725
Gutman B, Niyogi K (2009) Evidence for base excision repair of oxidative DNA damage in chloroplasts of Arabidopsis thaliana. J Biol Chem 284:17006–17012
Morales-Ruiz T, Ortega-Galisteo A, Ponferrada-Marín M, Martínez-Macías M, Ariza R, Roldán-Arjona T (2006) DEMETER and REPRESSOR OF SILENCING 1 encode 5-methylcytosine DNA glycosylases. Proc Natl Acad Sci USA 103:6853–6858
Ponferrada-Marin M, Roldan-Arjona T, Ariza R (2009) ROS1 5-methylcytosine DNA glycosylase is a slow-turnover catalyst that initiates DNA demethylation in a distributive fashion. Nucleic Acids Res 37:4264–4274
Brooks S, Fischer R, Huh J, Eichman B (2014) 5-methylcytosine recognition by Arabidopsis thaliana DNA glycosylases DEMETER and DML3. Biochemistry 53:2525–2532
Parrilla-Doblas J, Ponferrada-Marin M, Roldan-Arjona T, Ariza R (2013) Early steps of active DNA demethylation initiated by ROS1 glycosylase require three putative helix-invading residues. Nucleic Acids Res 41:8654–8664
Hong S, Hashimoto H, Kow Y, Zhang X, Cheng X (2014) The carboxy-terminal domain of ROS1 is essential for 5-methylcytosine DNA glycosylase activity. J Mol Biol 426:3703–3712
Ramiro-Merina A, Ariza R, Roldán-Arjona T (2013) Molecular characterization of a putative plant homolog of MBD4 DNA glycosylase. DNA Repair (Amst) 12:890–898
Chen H, Chu P, Zhou Y, Li Y, Liu J, Ding Y, Tsang E, Jiang L, Wu K, Huang S (2012) Overexpression of AtOGG1, a DNA glycosylase/AP lyase, enhances seed longevity and abiotic stress tolerance in Arabidopsis. J Exp Bot 63:4107–4121
Nota F, Cambiagno D, Ribone P, Alvarez M (2015) Expression and function of AtMBD4L, the single gene encoding the nuclear DNA glycosylase MBD4L in Arabidopsis. Plant Sci 235:122–129
Qüesta J, Fina J, Casati P (2013) DDM1 and ROS1 have a role in UV-B induced- and oxidative DNA damage in A. thaliana. Front Plant Sci 4:420
Murphy T, Belmonte M, Shu S, Britt A, Hatteroth J (2009) Requirement for abasic endonuclease gene homologues in Arabidopsis seed development. PLoS One 4:e4297
Li Y, Córdoba-Cañero D, Qian W, Zhu X, Tang K, Zhang H, Ariza R, Roldán-Arjona T, Zhu J-K (2015) An AP endonuclease functions in active DNA demethylation and gene imprinting in Arabidopsis. PLoS Genet 11:e1004905
Cordoba-Cañero D, Roldán-Arjona T, Ariza R (2011) Arabidopsis ARP endonuclease functions in a branched base excision DNA repair pathway completed by LIG1. Plant J 68:693–702
Joldybayeva B, Prorok P, Grin I, Zharkov D, Ishenko A, Tudek B, Bissenbaev A, Saparbaev M (2014) Cloning and characterization of a wheat homologue of apurinic/apyrimidinic endonuclease Ape1L. PLoS One 9:e92963
Martínez-Macías M, Córdoba-Cañero D, Ariza R, Roldán-Arjona T (2013) The DNA repair protein XRCC1 functions in the plant DNA demethylation pathway by stimulating cytosine methylation (5-mec) excision, gap tailoring, and DNA ligation. J Biol Chem 288:5496–5505
Martínez-Macías M, Qian W, Miki D, Pontes O, Liu Y, Tang K, Liu R, Morales-Ruiz T, Ariza R, Roldán-Arjona T, Zhu J-K (2012) A DNA 3′ phosphatase functions in active DNA demethylation in Arabidopsis. Mol Cell 45:357–370
Kim H, Na S, Lee S-Y, Jeong Y-M, Hwang H-J, Hur J, Park S-H, Woo J-C, Kim S (2012) Structure–function studies of a plant tyrosyl-DNA phosphodiesterase provide novel insights into DNA repair mechanisms of Arabidopsis thaliana. Biochem J 443:49–56
Waterworth W, Kozak J, Provost C, Bray C, Angelis K, West C (2009) DNA ligase 1 deficient plants display severe growth defects and delayed repair of both DNA single and double strand breaks. BMC Plant Biol 9:79
Zhang Y, Wen C, Liu S, Zheng L, Shen B, Tao Y (2016) Shade avoidance 6 encodes an Arabidopsis flap endonuclease required for maintenance of genome integrity and development. Nucleic Acids Res 44:1271–1284
Alekseev S, Coin F (2015) Orchestral maneuvers at the damaged sites in nucleotide excision repair. Cell Mol Life Sci 72:2177–2186
Schärer O (2013) Nucleotide excision repair in eukaryotes. Cold Spring Harb Perspect Biol 5:a012609
Spivak G, Ganesan A (2014) The complex choreography of transcription-coupled repair. DNA Repair (Amst) 19:64–70
Vermeulen W, Fousteri M (2013) Mammalian transcription coupled excision repair. Cold Spring Harb Perspect Biol 5:a012625
Bedez F, Linard B, Brochet X, Ripp R, Thompson J, Moras D, Lecompte O, Poch O (2013) Functional insights into the core-TFIIH from a comparative survey. Genomics 101:178–186
Fuss J, Tainer J (2011) XPB and XPD helicases in TFIIH orchestrate DNA duplex opening and damage verification to coordinate repair with transcription and cell cycle via CAK kinase. DNA Repair (Amst) 10:697–713
Fagbemi A, Orelli B, Schärer O (2011) Regulation of endonuclease activity in human nucleotide excision repair. DNA Repair (Amst) 10:722–729
Ogi T, Limsirichaikul S, Overmeer R, Volker M, Takenaka K, Cloney R, Nakazawa Y, Nimi A, Jaspers N, Mullenders L, S Y, Fousteri M, Lehamann A (2010) Three DNA polymerases, recruited by different mechanisms, carry out NER repair synthesis in human cells. Mol Cell 37:714–727
Ganpudi A, Schroeder D (eds) (2011) UV damaged DNA repair and tolerance in plants. Selected Topics in DNA Repair, Intech, Croatia
Farmer L, Book A, Lee K, Lin Y, Fu H, Vierstra R (2010) The RAD23 family provides an essential connection between the 26S proteasome and ubiquitylated proteins in Arabidopsis. Plant Cell 22:124–142
Al Khateeb W, Schroeder D (2007) DDB2, DDB1A and DET1 exhibit complex interactions during Arabidopsis development. Genetics 176:231–242
Bernhardt A, Mooney S, Hellmann H (2010) Arabidopsis DDB1a and DDB1b are critical for embryo development. Planta 232:555–566
Zhang C, Guo H, Zhang J, Guo G, Schumaker K, Guo Y (2010) Arabidopsis cockayne syndrome A-like proteins 1A and 1B form a complex with CULLIN4 and damage DNA binding protein 1A and regulate the response to UV irradiation. Plant Cell 22:2353–2369
Biedermann S, Hellmann H (2010) The DDB1a interacting proteins ATCSA-1 and DDB2 are critical factors for UV-B tolerance and genomic integrity in Arabidopsis thaliana. Plant J 62:404–415
Molinier J, Lechner E, Dumbliauskas E, Genschik P (2008) Regulation and role of Arabidopsis CUL4-DDB1A-DDB2 in maintaining genome integrity upon UV stress. PLoS Genet 4:e1000093
Al Khateeb W, Schroeder D (2009) Overexpression of Arabidopsis damaged DNA binding protein 1A (DDB1A) enhances UV tolerance. Plant Mol Biol 70:371–383
Ganpudi A, Schroeder D (2013) Genetic interactions of Arabidopsis thaliana damaged DNA binding protein 1B (DDB1B) with DDB1A, DET1, and COP1. G3 (Bethesda) 3:493–503
Koga A, Ishibashi T, Kimura S, Uchiyama Y, Sakaguchi K (2006) Characterization of T-DNA insertion mutants and RNAi silenced plants of Arabidopsis thaliana UV-damaged DNA binding protein 2 (AtUV-DDB2). Plant Mol Biol 61:227–240
Ly V, Hatherell A, Kim E, Chan A, Belmonte M, Schroeder D (2013) Interactions between Arabidopsis DNA repair genes UVH6, DDB1A, and DDB2 during abiotic stress tolerance and floral development. Plant Sci 213:88–97
Shaked H, Avivi-Ragolsky N, Levy A (2006) Involvement of the Arabidopsis SWI2/SNF2 chromatin remodeling gene family in DNA damage response and recombination. Genetics 173:985–994
Liang L, Flury S, Kalck V, Hohn B, Molinier J (2006) CENTRIN2 interacts with the Arabidopsis homolog of the human XPC protein (AtRAD4) and contributes to efficient synthesis-dependent repair of bulky DNA lesions. Plant Mol Biol 61:345–356
Vonarx E, Tabone E, Osmond M, Anderson H, Kunz B (2006) Arabidopsis homologue of human transcription factor IIH/nucleotide excision repair factor p44 can function intranscription and DNA repair and interacts with AtXPD. Plant J 46:512–521
Gentile A, Ditt R, Dias F, Da Silva M, Dornelas M, Menossi M (2009) Characterization of ScMat1, a putative TFIIH subunit from sugarcane. Plant Cell Rep 28:663–672
Gutierrez C (2009) The Arabidopsis cell division cycle. Arabidopsis Book 7:e0120
Van Leene J, Hollunder J, Eeckhout D, Persiau G, Van De Slijke E, Stals H, Van Isterdael G, Verkest A, Neirynck S, Buffel Y, De Bodt S, Maere S, Laukens K, Pharazyn A, Ferreira P, Eloy N, Renne C, Meyer C, Faure J, Steinbrenner J, Beynon J, Larkin J, Van de Peer Y, Hilson P, Kuiper M, De Veylder L, Van Onckelen H, Inzé D, Witters E, De Jaeger G (2010) Targeted interactomics reveals a complex core cell cycle machinery in Arabidopsis thaliana. Mol Syst Biol 6:397
Eschbach V, Kobbe D (2014) Different replication protein A complexes of Arabidopsis thaliana have different DNA-binding properties as a function of heterotrimer composition. Plant Cell Physiol 55:1460–1472
Aklilu B, Soderquist R, Culligan K (2014) Genetic analysis of the Replication Protein A large subunit family in Arabidopsis reveals unique and overlapping roles in DNA repair, meiosis and DNA replication. Nucleic Acids Res 42:3104–3108
Aklilu B, Culligan K (2016) Molecular evolution and functional diversification of Replication Protein A1 in plants. Front Plant Sci 7:33
Peña-Diaz J, Jiricny J (2012) Mammalian mismatch repair: error-free or error-prone? Trends Biochem Sci 37:206–214
Kunkel T, Erie D (2015) Eukaryotic mismatch repair in relation to DNA replication. Annu Rev Genet 49:291–313
Reyes G, Schmidt T, Kolodner R, Hombauer H (2015) New insights into the mechanism of DNA mismatch repair. Chromosoma 124:443–462
Friedhoff P, Li P, Gotthardt J (2016) Protein-protein interactions in DNA mismatch repair. DNA Repair (Amst) 38:50–57
Kolodner R (2016) A personal historical view of DNA mismatch repair with an emphasis on eukaryotic DNA mismatch repair. DNA Repair (Amst) 38:3–13
Sachadyn P (2010) Conservation and diversity of MutS proteins. Mutat Res 694:20–30
Tian L, Gu L, Li G-M (2009) Distinct nucleotide binding/hydrolysis properties and molar ratio of MutSa and MutSb determine their differential mismatch binding activities. J Biol Chem 284:11557–11562
Warren J, Pohlhaus T, Changela A, Iyer R, Modrich P, Beese L (2007) Structure of the human MutSα DNA lesion recognition complex. Mol Cell 26:579–592
Gupta S, Gellert M, Yang W (2011) Mechanism of mismatch recognition revealed by human MutSβ bound to unpaired DNA loops. Nat Struct Mol Biol 19:72–78
Groothuizen F, Sixma T (2016) The conserved molecular machinery in DNA mismatch repair enzyme structures. DNA Repair (Amst) 38:14–23
Owen B, Lang W, McMurray C (2009) The nucleotide binding dynamics of human MSH2-MSH3 are lesion dependent. Nat Struct Mol Biol 16:550–557
Iyer R, Pluciennik A, Genschel J, Tsai M, Beese L, Modrich P (2010) MutLαand proliferating cell nuclear antigen share binding sites on MutSβ. J Biol Chem 285:11730–11739
Edelbrock M, Kaliyaperumal S, Williams K (2013) Structural, molecular and cellular functions of MSH2 and MSH6 during DNA mismatch repair, damage signaling and other noncanonical activities. Mutat Res 743–744:53–66
Mjelle R, Hegre S, Aas P, Slupphaug G, Drabløs F, Saetrom P, Krokan H (2015) Cell cycle regulation of human DNA repair and chromatin remodeling genes. DNA Repair (Amst) 30:53–67
Kadyrova L, Kadyrov F (2016) Endonuclease activities of MutLα and its homologs in DNA mismatch repair. DNA Repair (Amst) 38:42–49
Guarné A, Charbonnier J (2016) Insights from a decade of biophysical studies on MutL: roles in strand discrimination and mismatch removal. Prog Biophys Mol Biol 117:149–156
Sacho E, Kadyrov F, Modrich P, Kunkel T, Erie D (2008) Direct visualization of asymmetric adenine-nucleotide-induced conformational changes in MutLα. Mol Cell 29:112–121
McNally R, Bowman G, Goedken E, O’Donnell M, Kuriyan J (2010) Analysis of the role of PCNA-DNA contacts during clamp loading. BMC Struct Biol 10:3
Pluciennik A, Dzantiev L, Iyer R, Constantin N, Kadyrov F, Modrich P (2010) PCNA function in the activation and strand direction of MutLαendonuclease in mismatch repair. Proc Natl Acad Sci USA 107:16066–16071
Shao H, Baitinger C, Soderblom E, Burdett V, Modrich P (2014) Hydrolytic function of Exo1 in mammalian mismatch repair. Nucleic Acids Res 42:7104–7112
Genschel J, Modrich P (2009) Functions of MutLalpha, replication protein A (RPA), and HMGB1 in 5′-directed mismatch repair. J Biol Chem 284:21536–21544
Kadyrov F, Genschel J, Fang Y, Penland E, Edelmann W, Modrich P (2009) A possible mechanism for exonuclease 1-independent eukaryotic mismatch repair. Proc Natl Acad Sci USA 106:8495–8500
Goellner E, Putnam C, Kolodner R (2015) Exonuclease 1-dependent and independent mismatch repair. DNA Repair (Amst) 32:24–32
Spampinato C, Gomez R, Galles C, Lario L (2009) From bacteria to plants: a compendium of mismatch repair assays. Mutat Res 682:110–128
Tam S, Samipak S, Britt A, Chetelat R (2009) Characterization and comparative sequence analysis of the DNA mismatch repair MSH2 and MSH7 genes from tomato. Genetica 137:341–354
Galles C, Gomez R, Spampinato C (2011) PMS1 from Arabidopsis thaliana: optimization of protein overexpression in Escherichia coli. Mol Biol Rep 38:1063–1070
Gomez R, Galles C, Spampinato C (2011) High-level production of MSH2 from Arabidopsis thaliana: a DNA mismatch repair system key subunit. Mol Biotechnol 47:120–129
Gomez R, Spampinato C (2013) Mismatch recognition function of Arabidopsis thaliana MutSγ. DNA Repair (Amst) 12:257–264
Galles C, Spampinato C (2013) Yeast mutator phenotype enforced by Arabidopsis PMS1 expression. Mol Biol Rep 40:2107–2114
Tam S, Hays J, Chetelat R (2011) Effects of suppressing the DNA mismatch repair system on homeologous recombination in tomato. Theor Appl Genet 123:1445–1458
Xu J, Li M, Chen L, Wu G, Li H (2012) Rapid generation of rice mutants via the dominant negative suppression of the mismatch repair protein OsPMS1. Theor Appl Genet 125:975–986
Van Marcke I, Angenon G (2013) Genomic stability in Nicotiana plants upon silencing of the mismatch repair gene MSH2. Plant Biotechnol Rep 7:467–480
Lario L, Botta P, Casati P, Spampinato C (2015) Role of AtMSH7 in UV-B-induced DNA damage recognition and recombination. J Exp Bot 66:3019–3026
Lario L, Ramirez-Parra E, Gutierrez C, Casati P, Spampinato C (2011) Regulation of plant MSH2 and MSH6 genes in the UV-B-induced DNA damage response. J Exp Bot 62:2925–2937
Ceccaldi R, Rondinelli B, D’Andrea A (2016) Repair pathway choices and consequences at the double-strand break. Trends Cell Biol 26:52–64
Grabarz A, Barascu A, Guirouilh-Barbat J, Lopez B (2012) Initiation of DNA double strand break repair: signaling and single-stranded resection dictate the choice between homologous recombination, non-homologous end-joining and alternative end-joining. Am J Cancer Res 2:249–268
Kakarougkas A, Jeggo P (2014) DNA DSB repair pathway choice: an orchestrated handover mechanism. Br J Radiol 87:20130685
Escribano-Diaz C, Orthwein A, Fradet-Turcotte A, Xing M, Young J, Tkác J, Cook M, Rosebrock A, Munro M, Canny M, Xu D, Durocher D (2013) A cell cycle-dependent regulatory circuit composed of 53BP1-RIF1 and BRCA1-CtIP controls DNA repair pathway choice. Mol Cell 49:872–883
Karanam K, Kafri R, Loewer A, Lahav G (2012) Quantitative live cell imaging reveals a gradual shift between DNA repair mechanisms and a maximal use of HR in mid S phase. Mol Cell 47:320–329
Jasin M, Rothstein R (2013) Repair of strand breaks by homologous recombination. Cold Spring Harb Perspect Biol 5:a012740
Krejci L, Altmannova V, Spirek M, Zhao X (2012) Homologous recombination and its regulation. Nucleic Acids Res 40:5795–5818
Kowalczykowski S (2015) An overview of the molecular mechanisms of recombinational DNA repair. Cold Spring Harb Perspect Biol 7:a016410
Sartori A, Lukas C, Coates J, Mistrik M, Fu S, Bartek J, Baer R, Lukas J, Jackson S (2007) Human CtIP promotes DNA end resection. Nature 450:509–514
Takeda S, Nakamura K, Taniguchi Y, Paull T (2007) Ctp1/CtIP and the MRN complex collaborate in the initial steps of homologous recombination. Mol Cell 28:351–352
Nimonkar A, Genschel J, Kinoshita E, Polaczek P, Campbell J, Wyman C, Modrich P, Kowalczykowski S (2011) BLM-DNA2-RPA-MRN and EXO1-BLM-RPA-MRN constitute two DNA end resection machineries for human DNA break repair. Genes Dev 25:350–362
Huertas P, Jackson S (2009) Human CtIP mediates cell cycle control of DNA end resection and double strand break repair. J Biol Chem 264:9558–9565
Carreira A, Kowalczykowski S (2011) Two classes of BRC repeats in BRCA2 promote RAD51 nucleoprotein filament function by distinct mechanisms. Proc Natl Acad Sci USA 108:10448–10453
Reuter M, Zelensky A, Smal I, Meijering E, van Cappellen W, de Gruiter H, van Belle G, van Royen M, Houtsmuller A, Essers J, Kanaar R, Wyman C (2014) BRCA2 diffuses as oligomeric clusters with RAD51 and changes mobility after DNA damage in live cells. J Cell Biol 207:599–613
Agarwal S, van Cappellen W, Guénolé A, Eppink B, Linsen S, Meijering E, Houtsmuller A, Kanaar R, Essers J (2011) ATP-dependent and independent functions of Rad54 in genome maintenance. J Cell Biol 192:735–750
Ceballos S, Heyer W (2011) Functions of the Snf2/Swi2 family Rad54 motor protein in homologous recombination. Biochim Biophys Acta 1809:509–523
Suwaki N, Klare K, Tarsounas M (2011) RAD51 paralogs: roles in DNA damage signalling, recombinational repair and tumorigenesis. Semin Cell Dev Biol 22:898–905
Sneeden J, Grossi S, Tappin I, Hurwitz J, Heyer W (2013) Reconstitution of recombination-associated DNA synthesis with human proteins. Nucleic Acids Res 41:4913–4925
Blanco M, Matos J (2015) Hold your horSSEs: controlling structure-selective endonucleases MUS81 and Yen1/GEN1. Front Genet 6:253
Matos J, West S (2014) Holliday junction resolution: regulation in space and time. DNA Repair (Amst) 19:176–181
Guirouilh-Barbat J, Lambert S, Bertrand P, Lopez B (2014) Is homologous recombination really an error-free process? Front Genet 5:175
Bétermier M, Bertrand P, Lopez B (2014) Is non-homologous end-joining really an inherently error-prone process? PLoS Genet 10:e1004086
Chiruvella K, Liang Z, Wilson T (2013) Repair of double-strand breaks by end joining. Cold Spring Harb Perspect Biol 5:a012757
Radhakrishnan S, Jette N, Lees-Miller S (2014) Non-homologous end joining: emerging themes and unanswered questions. DNA Repair (Amst) 17:2–8
Waters C, Strande N, Wyatt D, Pryor J, Ramsden D (2014) Nonhomologous end joining: a good solution for bad ends. DNA Repair (Amst) 17:39–51
Williams G, Hammel M, Radhakrishnan S, Ramsden D, Lees-Miller S, Tainer J (2014) Structural insights into NHEJ: building up an integrated picture of the dynamic DSB repair super complex, one component and interaction at a time. DNA Repair (Amst) 17:110–120
Britton S, Coates J, Jackson S (2013) A new method for high-resolution imaging of Ku foci to decipher mechanisms of DNA double-strand break repair. J Cell Biol 202:579–595
Dobbs T, Tainer J, Lees-Miller S (2010) A structural model for regulation of NHEJ by DNA-PKcs autophosphorylation. DNA Repair (Amst) 9:1307–1314
Davis A, Chen B, Chen D (2014) DNA-PK: a dynamic enzyme in a versatile DSB repair pathway. DNA Repair (Amst) 17:21–29
Fell V, Schild-Poulter C (2015) The Ku heterodimer: function in DNA repair and beyond. Mutat Res Rev Mutat Res 763:15–29
Jette N, Lees-Miller S (2015) The DNA-dependent protein kinase: a multifunctional protein kinase with roles in DNA double strand break repair and mitosis. Prog Biophys Mol Biol 117:194–205
Roberts S, Strande N, Burkhalter M, Strom C, Havener J, Hasty P, Ramsden D (2010) Ku is a 5′-dRP/AP lyase that excises nucleotide damage near broken ends. Nature 464:1214–1217
Strande N, Roberts S, Oh S, Hendrickson E, Ramsden D (2012) Specificity of the dRP/AP lyase of Ku promotes nonhomologous end joining (NHEJ) fidelity at damaged ends. J Biol Chem 287:13686–13693
Pommier Y, Huang S, Gao R, Das B, Murai J, Marchand C (2014) Tyrosyl-DNA-phosphodiesterases (TDP1 and TDP2). DNA Repair (Amst) 19:114–129
Coquelle N, Havali-Shahriari Z, Bernstein N, Green R, Glover J (2011) Structural basis for the phosphatase activity of polynucleotide kinase/phosphatase on single and double-stranded DNA substrates. Proc Natl Acad Sci USA 108:21022–21027
Garces F, Pearl L, Oliver A (2011) The structural basis for substrate recognition by mammalian polynucleotide kinase 3′ phosphatase. Mol Cell 44:385–396
Ochi T, Blackford A, Coates J, Jhujh S, Mehmood S, Tamura N, Travers J, Wu Q, Draviam V, Robinson C, Blundell T, Jackson S (2015) PAXX, a paralog of XRCC4 and XLF, interacts with Ku to promote DNA double-strand break repair. Science 347:185–188
Truong L, Li Y, Shi L, Hwang P, He J, Wang H, Razavian N, Berns M, Wu X (2013) Microhomology-mediated end joining and homologous recombination share the initial end resection step to repair DNA double-strand breaks in mammalian cells. Proc Natl Acad Sci USA 110:7720–7725
Decottignies A (2013) Alternative end-joining mechanisms: a historical perspective. Front Genet 4:48
Frit P, Barboule N, Yuan Y, Gomez D, Calsou P (2014) Alternative end-joining pathway(s): bricolage at DNA breaks. DNA Repair (Amst) 17:81–97
Sfeir A, Symington L (2015) Microhomology-mediated end joining: a back-up survival mechanism or dedicated pathway? Trends Biochem Sci 40:701–714
Cheng Q, Barboule N, Frit P, Gomez D, Bombarde O, Couderc B, Ren G, Salles B, Calsou P (2011) Ku counteracts mobilization of PARP1 and MRN in chromatin damaged with DNA double-strand breaks. Nucleic Acids Res 39:9605–9619
Wang M, Wu W, Wu W, Rosidi B, Zhang L, Wang H, Iliakis G (2006) PARP-1 and Ku compete for repair of DNA double strand breaks by distinct NHEJ pathways. Nucleic Acids Res 34:6170–6182
Muthurajan U, Hepler M, Hieb A, Clark N, Kramer M, Yao T, Luger K (2014) Automodification switches PARP-1 function from chromatin architectural protein to histone chaperone. Proc Natl Acad Sci USA 111:12752–12757
Polo S, Jackson S (2011) Dynamics of DNA damage response proteins at DNA breaks: a focus on protein modifications. Genes Dev 25:409–433
Beck C, Robert I, Reina-San-Martin B, Schreiber V, Dantzer F (2014) Poly(ADP-ribose) polymerases in double-strand break repair: focus on PARP1, PARP2 and PARP3. Exp Cell Res 329:18–25
Ko H, Ren E (2012) Functional aspects of PARP1 in DNA repair and transcription. Biomolecules 2:524–548
Pines A, Mullenders L, van Attikum H, Luijsterburg M (2013) Touching base with PARPs: moonlighting in the repair of UV lesions and double-strand breaks. Trends Biochem Sci 38:321–330
Bryant H, Petermann E, Schultz N, Jemth A, Loseva O, Issaeva N, Johansson F, Fernandez S, McGlynn P, Helleday T (2009) PARP is activated at stalled forks to mediate Mre11-dependent replication restart and recombination. EMBO J 28:2601–2615
Haince J, McDonald D, Rodrigue A, Dery U, Masson J, Hendzel M, Poirier G (2008) PARP1-dependent kinetics of recruitment of MRE11 and NBS1 proteins to multiple DNA damage sites. J Biol Chem 283:1197–1208
Della-Maria J, Zhou Y, Tsai M, Kuhnlein J, Carney J, Paull T, Tomkinson A (2011) Human Mre11/human Rad50/Nbs1 and DNA ligase IIIalpha/XRCC1 protein complexes act together in an alternative nonhomologous end joining pathway. J Biol Chem 286:33845–33853
Simsek D, Brunet E, Wong S-W, Katyal S, Gao Y, McKinnon P, Lou J, Zhang L, Li J, Rebar E, Gregory P, Holmes M, Jasin M (2011) DNA ligase III promotes alternative nonhomologous end-joining during chromosomal translocation formation. PLoS Genet 7:e1002080
Baltes N, Voytas D (2015) Enabling plant synthetic biology through genome engineering. Trends Biotechnol 33:120–131
Knoll A, Fauser F, Puchta H (2014) DNA recombination in somatic plant cells: mechanisms and evolutionary consequences. Chromosome Res 22:191–201
Gaj T, Gersbach C, Cr Barbas (2013) ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 31:397–405
Voytas D (2013) Plant genome engineering with sequence-specific nucleases. Annu Rev Plant Biol 64:327–350
Kumar S, Barone P, Smith M (2016) Gene targeting and transgene stacking using intra genomic homologous recombination in plants. Plant Methods 12:11
Puchta H, Fauser F (2014) Synthetic nucleases for genome engineering in plants: prospects for a bright future. Plant J 78:727–741
Sprink T, Metje J, Hartung F (2015) Plant genome editing by novel tools: tALEN and other sequence specific nucleases. Curr Opin Biotechnol 32:47–53
Waterworth W, Altun C, Armstrong S, Roberts N, Dean P, Young K, Weil C, Bray C, West C (2007) NBS1 is involved in DNA repair and plays a synergistic role with ATM in mediating meiotic homologous recombination in plants. Plant J 52:41–52
Uanschou C, Siwiec T, Pedrosa-Harand A, Kerzendorfer C, Sanchez-Moran E, Novatchkova M, Akimcheva S, Woglar A, Klein F, Schlögelhofer P (2007) A novel plant gene essential for meiosis is related to the human CtIP and the yeast COM1/SAE2 gene. EMBO J 26:5061–5070
Akutsu N, Iijima K, Hinata T, Tauchi H (2007) Characterization of the plant homolog of Nijmegen breakage syndrome 1: involvement in DNA repair and recombination. Biochem Biophys Res Commun 353:394–398
Hartung F, Suer S, Puchta H (2007) Two closely related RecQ helicases have antagonistic roles in homologous recombination and DNA repair in Arabidopsis thaliana. Proc Natl Acad Sci USA 104:18836–18841
Mannuss A, Dukowic-Schulze S, Suer S, Hartung F, Pacher M, Puchta H (2010) RAD5A, RECQ4A, and MUS81 have specific functions in homologous recombination and define different pathways of DNA repair in Arabidopsis thaliana. Plant Cell 22:3318–3330
Schröpfer S, Kobbe D, Hartung F, Knoll A, Puchta H (2014) Defining the roles of the N-terminal region and the helicase activity of RECQ4A in DNA repair and homologous recombination in Arabidopsis. Nucleic Acids Res 42:1684–1697
Jia N, Liu X, Gao H (2016) A DNA2 homolog is required for DNA damage repair, cell cycle regulation, and meristem maintenance in plants. Plant Physiol 171:318–333
Da Ines O, Degroote F, Amiard S, Goubely C, Gallego M, White C (2013) Effects of XRCC2 and RAD51B mutations on somatic and meiotic recombination in Arabidopsis thaliana. Plant J 74:959–970
Roth N, Klimesch J, Dukowic-Schulze S, Pacher M, Mannuss A, Puchta H (2012) The requirement for recombination factors differs considerably between different pathways of homologous double-strand break repair in somatic plant cells. Plant J 72:781–790
Serra H, Da Ines O, Degroote F, Gallego M, White C (2013) Roles of XRCC2, RAD51B and RAD51D in RAD51-independent SSA recombination. PLoS Genet 9:e1003971
Wang Y, Xiao R, Wang H, Cheng Z, Li W, Zhu G, Wang Y, Ma H (2014) The Arabidopsis RAD51 paralogs RAD51B, RAD51D and XRCC2 play partially redundant roles in somatic DNA repair and gene regulation. New Phytol 201:292–304
Yao Y, Bilichak A, Titov V, Golubov A, Kovalchuk I (2013) Genome stability of Arabidopsis atm, ku80 and rad51b mutants: somatic and transgenerational responses to stress. Plant Cell Physiol 54:982–989
Abe K, Osakabe K, Ishikawa Y, Tagiri A, Yamanouchi H, Takyuu T, Yoshioka T, Ito T, Kobayashi M, Shinozaki K, Ichikawa H, Toki S (2009) Inefficient double-strand DNA break repair is associated with increased fasciation in Arabidopsis BRCA2 mutants. J Exp Bot 60:2751–2761
Seeliger K, Dukowic-Schulze S, Wurz-Wildersinn R, Pacher M, Puchta H (2012) BRCA2 is a mediator of RAD51- and DMC1-facilitated homologous recombination in Arabidopsis thaliana. New Phytol 193:364–375
Trapp O, Seeliger K, Puchta H (2011) Homologs of breast cancer genes in plants. Front Plant Sci 2:19
Blanck S, Kobbe D, Hartung F, Fengler K, Focke M, Puchta H (2009) A SRS2 homolog from Arabidopsis thaliana disrupts recombinogenic DNA intermediates and facilitates single strand annealing. Nucleic Acids Res 37:7163–7176
Bauknecht M, Kobbe D (2014) AtGEN1 and AtSEND1, two paralogs in Arabidopsis, possess holliday junction resolvase activity. Plant Physiol 166:202–216
Geuting V, Kobbe D, Hartung F, Dürr J, Focke M, Puchta H (2009) Two distinct MUS81-EME1 complexes from Arabidopsis process Holliday junctions. Plant Physiol 150:1062–1071
Hartung F, Suer S, Knoll A, Wurz-Wildersinn R, Puchta H (2008) Topoisomerase 3alpha and RMI1 suppress somatic crossovers and are essential for resolution of meiotic recombination intermediates in Arabidopsis thaliana. PLoS Genet 4:e1000285
Song J, Keppler B, Wise R, Bent A (2015) PARP2 is the predominant Poly(ADP-Ribose) polymerase in Arabidopsis DNA damage and immune responses. PLoS Genet 11:e1005200
Charbonnel C, Allain E, Gallego M, White C (2011) Kinetic analysis of DNA double-strand break repair pathways in Arabidopsis. DNA Repair (Amst) 10:611–619
Charbonnel C, Gallego M, White C (2010) Xrcc1-dependent and Ku-dependent DNA double-strand break repair kinetics in Arabidopsis plants. Plant J 64:280–290
Jia Q, den Dulk-Ras A, Shen H, Hooykaas P, de Pater S (2013) Poly(ADP-ribose)polymerases are involved in microhomology mediated back-up non-homologous end joining in Arabidopsis thaliana. Plant Mol Biol 82:339–351
Nishizawa-Yokoi A, Nonaka S, Saika H, Kwon Y, Osakabe K, Toki S (2012) Suppression of Ku70/80 or Lig4 leads to decreased stable transformation and enhanced homologous recombination in rice. New Phytol 196:1048–1059
Zhang B, Wang M, Tang D, Li Y, Xu M, Gu M, Cheng Z, Yu H (2015) XRCC3 is essential for proper double-strand break repair and homologous recombination in rice meiosis. J Exp Bot 66:5713–5725
Confalonieri M, Faè M, Balestrazzi A, Donà M, Macovei A, Valassi A, Giraffa G, Carbonera D (2014) Enhanced osmotic stress tolerance in Medicago truncatula plants overexpressing the DNA repair gene MtTdp2a (tyrosyl-DNA phosphodiesterase 2). Plant Cell Tiss Organ Cult 116:187–203
Kwon Y, Abe K, Osakabe K, Endo M, Nishizawa-Yokoi A, Saika H, Shimada H, Toki S (2012) Overexpression of OsRecQl4 and/or OsExo1 enhances DSB-induced homologous recombination in rice. Plant Cell Physiol 53:2142–2152
Yoshiyama K (2016) Recent progress in research on DNA damage responsesin animals and plants. Genes Genet Syst 90:185–186
Acknowledgements
I apologize to those whose work is not mentioned here. I acknowledge research support from Agencia Nacional de Promoción Científica y Tecnológica (PICT 2014-3127). C.P.S. is a member of the Researcher Career of CONICET.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Spampinato, C.P. Protecting DNA from errors and damage: an overview of DNA repair mechanisms in plants compared to mammals. Cell. Mol. Life Sci. 74, 1693–1709 (2017). https://doi.org/10.1007/s00018-016-2436-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-016-2436-2