Abstract
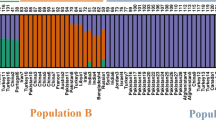
Carthamus tinctorius L. (safflower) is an important oilseed crop that is cultivated in several countries. The present study investigates the genetic diversity and population structure of 531 safflower accessions from 43 countries representing all safflower growing regions of the world. Diversity analysis was performed using ten informative EcoRI/MseI amplified fragment length polymorphism primer pairs that were identified by screening 150 primer combinations. The selected primer pairs generated 381 fragments of which 157 were polymorphic among the analyzed accessions. The genetic diversity indices obtained for the entire collection (I = 0.4536, H = 0.2955) indicated high levels of molecular variability. The distance-based, neighbor-joining method classified the accessions into six clusters with internal subgroupings that were in consonance with 19 clusters obtained using Bayesian model-based BAPS analysis. Clusters obtained through STRUCTURE analysis (at K = 4) could not be correlated with their geographically diverse origins, while BAPS analysis (at K = 19) revealed geographical delineation with low admixture levels among most of the studied accessions. Accessions from Far East and Egypt clustered in distinct groups, indicating conserved nature of their gene pools. The Near East and Iran–Afghanistan regions were collectively found to harbor maximum diversity in accordance with earlier reports. Accessions from the Indian subcontinent showed substantial diversity that was previously undetected. The American accessions showed low molecular variability in contrast to earlier studies. Genetic sub-structuring within gene pools and inter-relationships between accessions belonging to different regional pools was also observed. To the best of our knowledge, this is the first comprehensive study of existing genetic variability in a large collection of safflower germplasm with a global distribution, which provides a more accurate representation of genetic structuring in the crop. This information will facilitate selection of elite genotypes for broadening the genetic base of various breeding programs in safflower.






Similar content being viewed by others
References
Amini F, Saeidi G, Arzani A (2008) Study of genetic diversity in safflower genotypes using agro-morphological traits and RAPD markers. Euphytica 163:21–30
Ashri A (1971a) Evaluation of the world collection of safflower, Carthamus tinctorius L. I. Reaction to several diseases and associations with morphological characters in Israel. Crop Sci 11:253–257
Ashri A (1971b) Evaluation of the world collection of safflower, Carthamus tinctorius L. II. Resistance to the safflower fly, Acanthophilus helianthi R. Euphytica 20:410–415
Ashri A (1975) Evaluation of the germplasm collection of safflower, Carthamus tinctorius L. V. Distribution and regional divergence for morphological characters. Euphytica 24:651–659
Ashri A, Knowles P (1960) Cytogenetics of safflower (Carthamus L.) species and their hybrids. Agron J 52:11–17
Ashri A, Zimmer D, Urie A, Cahaner A, Marani A (1974) Evaluation of the world collection of safflower, Carthamus tinctorius L. IV. Yield and yield components and their relationships. Crop Sci 14:799–802
Bankey P, Billiar T, Wang W, Carlson A, Holman R, Cerra F (1989) Modulation of Kupffer cell membrane phospholipid function by n-3 polyunsaturated fatty acids. J Surg Res 46:439–444
Barati M, Arzani A (2012) Genetic diversity revealed by EST-SSR markers in cultivated and wild safflower. Biochem Syst Ecol 44:117–123
Carlsson AS, Zhu L-H, Andersson M, Hofvander P (2014) Platform crops amenable to genetic engineering—a requirement for successful production of bio-industrial oils through genetic engineering. Biocatal Agric Biotechnol 3:58–64
Carrasco B, Avila P, Perez-Diaz J, Munoz P, García R, Lavandero B, Zurita-Silva A, Retamales JB, Caligari PD (2009) Genetic structure of highland papayas (Vasconcellea pubescens (Lenné et C. Koch) Badillo) cultivated along a geographic gradient in Chile as revealed by inter simple sequence repeats (ISSR). Genet Resour Crop Evol 56:331–337
Chapman MA, Burke JM (2007) DNA sequence diversity and the origin of cultivated safflower (Carthamus tinctorius L.; Asteraceae). BMC Plant Biol 7(1):60
Chapman MA, Hvala J, Strever J, Matvienko M, Kozik A, Michelmore RW, Tang S, Knapp SJ, Burke JM (2009) Development, polymorphism, and cross-taxon utility of EST–SSR markers from safflower (Carthamus tinctorius L.). Theor Appl Genet 120:85–91
Chapman MA, Hvala J, Strever J, Burke JM (2010) Population genetic analysis of safflower (Carthamus tinctorius L.; Asteraceae) reveals a Near Eastern origin and five centers of diversity. Am J Bot 97:831–840
Corander J, Marttinen P (2006) Bayesian identification of admixture events using multilocus molecular markers. Mol Ecol 15:2833–2843
Corander J, Waldmann P, Marttinen P, Sillanpää MJ (2004) BAPS 2: enhanced possibilities for the analysis of genetic population structure. Bioinformatics 20:2363–2369
Dajue L, Mündel H (1996) Safflower (Carthamus tinctorius L.) promoting the conservation and use of underutilized and neglected crops. 7. Inst. Plant Genetic Resources Institute (IPGRI), Rome
Doyle J (1991) DNA protocols for plants—CTAB total DNA isolation. In: Hewitt GM, Johnston A (eds) Molecular techniques in taxonomy. Springer, Berlin, pp 283–293
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
FAO (2012). http://faostat.fao.org/site/567/DesktopDefault.aspx?PageID=567#ancor. Accessed Apr 2014
FAO (2013) Oilseeds market summary. Food outlook June 2013. http://www.fao.org/fileadmin/templates/est/COMM_MARKETS_MONITORING/Oilcrops/Documents/Food_outlook_oilseeds/Food_Outlook_June_13.pdf. Accessed Apr 2014
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Fang J-Y, Chung J-D, Chiang Y-C, Chang C-T, Chen C-Y, Hwang S-Y (2013) Divergent selection and local adaptation in disjunct populations of an endangered conifer, Keteleeria davidiana var. formosana (Pinaceae). PLoS One 8:e70162
Flider FJ (2013) Development and commercialization of GLA safflower oil. Lipid Technol 25:227–229
Hamdan Y, García‐Moreno MJ, Redondo‐Nevado J, Velasco L, Pérez‐Vich B (2011) Development and characterization of genomic microsatellite markers in safflower (Carthamus tinctorius L.). Plant Breed 130:237–241
Hanage WP, Fraser C, Tang J, Connor TR, Corander J (2009) Hyper-recombination, diversity, and antibiotic resistance in Pneumococcus. Science 324:1454–1457
Ilkılıç C, Aydın S, Behcet R, Aydin H (2011) Biodiesel from safflower oil and its application in a diesel engine. Fuel Process Technol 92:356–362
Johnson RC, Kisha T, Evans M (2007) Characterizing safflower germplasm with AFLP molecular markers. Crop Sci 47:1728–1736
Khan MA, von Witzke-Ehbrecht S, Maass BL, Becker HC (2009) Relationships among different geographical groups, agro-morphology, fatty acid composition and RAPD marker diversity in safflower (Carthamus tinctorius L.). Genet Resour Crop Evol 56:19–30
Knowles P (1969) Centers of plant diversity and conservation of crop germ plasm: safflower. Econ Bot 23:324–329
Lee GA, Sung JS, Lee SY, Chung JW, Yi JY, Kim YG, Lee MC (2014) Genetic assessment of safflower (Carthamus tinctorius L.) collection with microsatellite markers acquired via pyrosequencing method. Mol Ecol Resour 14:69–78
Mahasi M, Wachira F, Pathak R, Riungu T (2009) Genetic polymorphism in exotic safflower (Carthamus tinctorious L.) using RAPD markers. J Plant Breed Crop Sci 1:008–012
Marinova E, Riehl S (2009) Carthamus species in the ancient Near East and south-eastern Europe: archaeobotanical evidence for their distribution and use as a source of oil. Veg Hist Archaeobot 18:341–349
McPherson MA, Yang R-C, Good AG, Nielson RL, Hall LM (2009) Potential for seed-mediated gene flow in agroecosystems from transgenic safflower (Carthamus tinctorius L.) intended for plant molecular farming. Transgenic Res 18:281–299
Mokhtari N, Rahimmalek M, Talebi M, Khorrami M (2013) Assessment of genetic diversity among and within Carthamus species using sequence-related amplified polymorphism (SRAP) markers. Plant Syst Evol 299:1285–1294
Nimbkar N (2008) Issues in safflower production in India. In: Proceedings of the Seventh International Safflower Conference, Wagga Wagga.
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539
Pearl SA, Bowers JE, Reyes-Chin-Wo S, Michelmore RW, Burke JM (2014) Genetic analysis of safflower domestication. BMC Plant Biol 14(1):43
Peng S, Feng N, Guo M, Chen Y, Guo Q (2008) Genetic variation of Carthamus tinctorius L. and related species revealed by SRAP analysis. Biochem Syst Ecol 36:531–538
Perrier X, Jacquemoud-Collet J (2006) DARwin software http://darwin.cirad.fr/darwin.
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Ramu P, Billot C, Rami J, Senthilvel S, Upadhyaya H, Reddy LA, Hash C (2013) Assessment of genetic diversity in the sorghum reference set using EST-SSR markers. Theor Appl Genet 126:2051–2064
Rosenberg NA, Pritchard JK, Weber JL, Cann HM, Kidd KK, Zhivotovsky LA, Feldman MW (2002) Genetic structure of human populations. Science 298:2381–2385
Sehgal D, Raina SN (2005) Genotyping safflower (Carthamus tinctorius L.) cultivars by DNA fingerprints. Euphytica 146:67–76
Sehgal D, Rajpal VR, Raina SN, Sasanuma T, Sasakuma T (2009) Assaying polymorphism at DNA level for genetic diversity diagnostics of the safflower (Carthamus tinctorius L.) world germplasm resources. Genetica 135:457–470
Smith JR (1996) History, Chapter 1. In: Safflower. The American Oil Chemists Society Press, Champaign, pp 1–15
Smýkal P, Kenicer G, Flavell AJ, Corander J, Kosterin O, Redden RJ, Ford R, Coyne CJ, Maxted N, Ambrose MJ (2011) Phylogeny, phylogeography and genetic diversity of the Pisum genus. Plant Genet Resour 9:4–18
Sujatha M (2008) Biotechnological interventions for genetic improvement of safflower. In: Proceedings of the Seventh International Safflower Conference, Wagga Wagga, New South Wales pp. 3–6.
Tollefsrud MM, Sønstebø JH, Brochmann C, Johnsen Ø, Skrøppa T, Vendramin GG (2009) Combined analysis of nuclear and mitochondrial markers provide new insight into the genetic structure of North European Picea abies. Heredity 102:549–562
Van Zeist W, Waterbolk-Van Rooijen W (1992) Two interesting floral finds from third millennium BC Tell Hammam et-Turkman, northern Syria. Veg Hist Archaeobot 1(3):157–161
Vavilov NI (1951) The origin, variation, immunity and breeding of cultivated plants. Soil Sci 72:482
Velasco L, Pérez‐Vich B, Fernández‐Martínez J (2005) Identification and genetic characterization of a safflower mutant with a modified tocopherol profile. Plant Breed 124:459–463
Vos P, Hogers R, Bleeker M, Reijans M, van De Lee T, Hornes M, Friters A, Pot J, Paleman J, Kuiper M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–4414
Weiss E (1983) Oilseed crops. In: Safflower. Longman Group Limited, Longman House, London
Yang YX, Wu W, Zheng YL, Chen L, Liu RJ, Huang CY (2007) Genetic diversity and relationships among safflower (Carthamus tinctorius L.) analyzed by inter-simple sequence repeats (ISSRs). Genet Resour Crop Evol 54:1043–1051
Yeh Francis C, Yang R, Boyle Timothy B, Ye Z, Mao Judy X (1999) POPGENE version 1.32, the user-friendly shareware for population genetic analysis. Molecular Biology and Biotechnology Centre, University of Alberta, Canada (http://www.ualbertaca/∼fyeh/).
Acknowledgments
This work was funded by the DST-PURSE grant of the Department of Science and Technology, Government of India, provided to University of Delhi. HA was supported by a research fellowship from University Grants Commission, India. The authors thank Professors R. Geeta and Deepak Pental of University of Delhi for their critical comments and suggestions. The authors would also like to extend their sincere thanks to anonymous reviewers whose critical comments have further improved the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Additional information
S. Kumar and H. Ambreen contributed equally.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(XLSX 23 kb)
Rights and permissions
About this article
Cite this article
Kumar, S., Ambreen, H., Murali, T.V. et al. Assessment of Genetic Diversity and Population Structure in a Global Reference Collection of 531 Accessions of Carthamus tinctorius L. (Safflower) Using AFLP Markers. Plant Mol Biol Rep 33, 1299–1313 (2015). https://doi.org/10.1007/s11105-014-0828-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-014-0828-8